Abstract
Background
Breast cancer (BC) is one of the most prevalent cancers that contribute to mortality among women worldwide. Despite contradictory findings, considerable evidence suggests that single nucleotide polymorphisms (SNPs) in the FSCN1 and HOTAIR genes may have a causative impact on the development of BC. This case–control study was conducted to evaluate the association of genotype frequency in FSCN1 rs852479, rs1640233, and HOTAIR rs920778 with susceptibility and prognosis of BC, as well as the impact of clinical stages and hormonal features.
Methods and results
FSCN1 (rs852479, rs1640233) and HOTAIR (rs920778) were genotyped using TaqMan real-time PCR assay in 200 BC patients and 200 cancer-free controls, all representing Egyptian women. Genotypic analyses in association with clinicopathological factors and disease risk were assessed. As a result, a significant association with BC risk was observed for CC genotype frequency of FSCN1 rs852479 A > C (OR = 0.395, 95% CI 0.204–0.76, p-value = 0.005). However, no significant correlation was detected between the FSCN1 rs1640233 C > T and HOTAIR rs920778 C > T polymorphic variants and susceptibility to BC. Interestingly, CC genotype of FSCN1 rs1640233 was more likely to progress tumor size and lymph node invasion in BC cases (p-value = 0.04 and 0.02, respectively). Moreover, it was revealed that there was a non-significant correlation between the haplotype distributions of FSCN1 rs852479 and rs1640233 and the probability of BC.
Conclusions
Based on the sample size and genetic characteristics of the subjects involved in the present study, our findings indicated that FSCN1 rs852479 may contribute to BC susceptibility in a sample of the Egyptian population.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer (BC) is the most prevalent malignancy in women worldwide. According to its death rate, it is considered the second most frequent cause of cancer mortality among women [1]. Globally, there were 2.3 million women diagnosed with BC and 685,000 deaths from this disease in 2020 [2]. The incidence and mortality rates of BC vary according to the region [3]. An estimated 313,510 new instances of invasive BC in women and 611,720 cancer-related deaths will occur in the United States in 2024 [4]. In Egypt, more than 22,000 new cases of cancer are diagnosed each year, making it the main cause of cancer-related mortality among Egyptian women [5]. The ratio of mortality/incidence rate of BC cases in Egypt was approximately double the ratio (41%) when compared with developed countries (23%) [6]. Many variables are associated with the risk of BC, including age, environmental, gynecological, and genetic factors [7].
Polymorphisms of a DNA sequence caused by a single nucleotide variation in humans are known as single-nucleotide polymorphisms (SNPs), which are the most prevalent types of genetic variations in the human genome. SNPs in genes can potentially alter the protein structure or affect the expression level of the gene product [8, 9], which in turn changes disease susceptibility, affecting tumorigenesis and cancer progression as well as drug resistance [10, 11]. Certain genetic polymorphisms can predict an individual’s susceptibility to BC and also influence disease management and progression [12].
Fascin-1 (FSCN1) is a 55-kDa actin-bundling protein coded by a gene located on chromosome 7p22.1 with about 13.84 kb in length and includes five exons. Human FSCN1 is thought to be involved in the assembly of actin filament bundles found in lamellipodia, filopodia, microspikes, and stress fibers [13, 14]. FSCN1 is abundantly expressed in many types of normal cells, including neurons, endothelial cells, glial cells, mesenchymal, and antigen-presenting dendritic cells, and is low or absent in normal epithelial cells [15]. Based on the occurrence of FSCN1 in different organs, it is predictably participating in more biological functions in the human body [16]. In contrast to normal tissues, increased FSCN1 expression has been associated with several types of malignancies, including lung, colon, breast, ovary, and oral squamous cell carcinoma [17,18,19,20,21]. As an oncogene, FSCN1 can influence mitochondrial remodeling in cancerous cells, in addition to promoting invasion, tumor migration, metastatic colonization, cancer cell self-renewal, and drug resistance. In BC, FSCN1 is crucial for predicting aggressive tumor behavior, especially in advanced stages [22]. Recent evidence suggests that aberrant STAT3 signaling accelerates the growth of breast tumors by downregulating the expression of downstream target genes that regulate angiogenesis, such as hypoxia-inducible factor-1 (HIF-1) and nuclear factor-kappaB (NF-κB), and by binding to the promoter of the FSCN1 gene, triggering its expression [23]. Upregulation of FSCN1 enhances the severity and prognosis of human BC and can serve as a diagnostic marker to differentiate triple-negative subtypes of BC from other types of the disease [14]. Interestingly, different SNPs have been reported in FSCN1 to modulate the risk of BC development [24].
HOX transcript antisense RNA (HOTAIR) is a transcript that originates from the antisense strand of the HOXC gene cluster with an approximate length of 2.2 kb. The human HOTAIR gene is found between HOXC11 and HOXC12 genes on the long arm of chromosome 12q13.13 [25]. It is an example of an oncogenic long noncoding RNA (lncRNA), which has emerged as a master regulator of cancer [12]. The HOTAIR gene controls several cellular and biochemical processes to promote the proliferation, invasion, survival, drug resistance, and prognosis of various tumors. Some reports indicate that polymorphisms of the HOTAIR gene are associated with a variety of cancers, including breast [25], pancreatic [26], gastric [27], thyroid [28], and colorectal cancers [29]. HOTAIR gene expression in BC cells is modulated by numerous epigenetic and transcriptional mechanisms [25]. Several SNPs, located in the intronic region of the HOTAIR gene, have been reported to regulate its expression level [30,31,32]. These SNPs are expected to be related to the occurrence, progression, recurrence, and metastasis of BC and serve as a novel therapeutic target for the disease [33, 34].
Recently, the relationship of FSCN1 and HOTAIR polymorphisms with breast tumor development has been investigated [34,35,36]. However, some conclusions are still controversial and require further analysis to fully understand the relationship between these genes’ polymorphisms and BC risk. Therefore, this study was conducted to elucidate the association between FSCN1 rs852479, rs1640233, and HOTAIR (rs920778) with the risk or prognosis of BC concerning several clinicopathological variables in the Egyptian population.
Subjects and methods
Study subjects
This study enrolled 200 Egyptian women with BC (cases) and 200 healthy women without BC (controls), matched by age and comparable socioeconomic factors. All participants were recruited from Beni-Suef University Hospital in the period between 2021 and 2023.
The study protocol was approved by the Ethics Committee of the Faculty of Pharmacy (Girls), Al-Azhar University (REC number: 436), and all study procedures were conducted in accordance with the Declaration of Helsinki. All study participants provided fully informed written consent at the time of study entry.
All samples underwent genotyping for three SNPs: FSCN1 rs852479, rs1640233, and HOTAIR rs920778 to evaluate the association between gene polymorphisms and BC risk. Clinical examinations were detected to assess the impact of polymorphisms on BC patients based on menopausal status, tumor size, lymph node invasion, and histological grade. Additionally, BC prognostic biomarkers, including estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor 2 (HER2), were investigated.
DNA extraction
A peripheral blood sample (3 ml) was withdrawn from all study participants under complete aseptic conditions. Genomic DNA was extracted from blood samples using the salting‐out method [37]. The concentration and quality of the DNA were checked by measuring the absorbance at 260 and 280 nm using a UV spectrophotometer, NanoDrop 2000 (Thermo-Fisher Scientific, Wilmington, USA). Pure preparations of DNA have OD260/OD280 values of 1.7–2.0. The extracted DNA concentration ranged from 50 to 100 ng DNA/μl. The extracted DNA samples were maintained at a temperature of − 20 °C until the genotyping procedure.
Polymorphisms genotyping
Genotyping of FSCN1 rs852479, rs1640233, and HOTAIR rs920778 was done by TaqMan real‐time PCR method using the pre-designed assays for allelic discrimination, containing specific TaqMan probes with fluorescent dyes for each allele. The total PCR volume was 20 µl, containing 5 μl DNA, 10 μl TaqMan Universal PCR Master Mix, 0.05 μl (40×) Assay Mix, and 4.5 μl RNase‐free water. The PCR reaction conditions were the same for the three SNPs, with a pre-denaturation cycle at 95 °C for 10 min, followed by 45 cycles of 95 °C denaturation for 10 s, 60 °C annealing for 30 s, and final extension at 72 °C for 30 s. For genotyping quality control, deionized water was used to replace template DNA as a negative control. The PCR results (changing fluorescence level) were analyzed using the provided software.
Sample size and statistical analysis
The sample size was calculated using G*Power software version 3.1.9.7 for power analysis and sample size [38]. A total sample size of 400 was required, 200 in each group, with a power of 80% and a significance level of 5%. SPSS 22.0 software package (SPSS Inc., Chicago, IL, USA) was utilized for statistical analysis. Categorical variables are expressed as frequencies and percentages, while continuous variables are given as mean ± standard deviation (SD). Differences in clinical characteristics were compared between patients and healthy control groups using independent-sample t-tests (continuous variables) and chi-square tests (categorical variables). Hardy–Weinberg equilibrium (HWE) analysis was performed for each SNP assay. The chi-square (χ2) test was used to test differences between the two groups for each SNP genotype and allele. Allele frequencies were calculated with the gene counting method. The most common genotypes were selected as the reference. Odds ratios (OR) were calculated with a 95% confidence interval (CI) to estimate the degree of the association between genotypes and the risk of BC. SNPStats (https://www.snpstats.net/) was used to perform haplotype analysis test for linkage disequilibrium (LD) [39]. The significance level was set at a p-value < 0.05.
Results
General demographic and clinicopathological characteristics of the studied groups
The demographic and clinicopathological features of BC patients and controls in this study are shown in Table 1. The mean age of the controls at the time of enrollment was not significantly different from that of the BC cases (48.39 vs. 49.5 years, respectively, p-value = 0.394). Regarding BC cases, 47% were premenopausal, and 53% were postmenopausal. In respect to ER, 126 cases (63%) tested positive, and 125 cases (62.5%) tested positive for PR, while 45 (22.5%) of the cases were positive for HER2. Referring to the Nottingham prognostic index (NPI), the percentage of BC patients with T 1, 2, 3, and 4 was 9, 37.5, 28.5, and 25%, respectively, and N 0, 1, 2, and 3 emerged in 28, 43.5, 19, and 9.5% of patients, respectively. Concerning the histology grades of BC, 9 patients were classified as grade I (4.5%), 141 as grade II (70.5%), and 50 as grade III (25%).
Distribution frequencies of genotypes and alleles in BC patients and controls
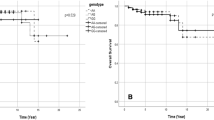
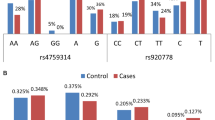
The distribution patterns of FSCN1 rs852479 and rs1640233, and HOTAIR rs920778 genotypes for all subjects are shown in Table 2 and Fig. 1. In the healthy controls and cases groups, all genotypic frequencies were in HWE (p-value > 0.05). Genotype analysis of FSCN1 polymorphism in both controls and cases revealed that most of those with rs852479 SNP were homozygous for the AA genotype, while with rs1640233 SNP, most of them were homozygous for CC genotype. Likewise, for HOTAIR rs920778 SNP, most of the controls and cases were homozygous for CC genotype (Table 2).
According to the logistic regression analysis for each genetic polymorphism in BC patients and controls as given in Table 2, we observed that women with CC genotype frequency of FSCN1 rs852479 A > C have a significantly high incidence of developing BC when compared with AC genotype (CC vs. AC, OR = 0.395; 95% CI 0.204–0.76, p-value = 0.005; OR = 1.053, 95% CI 0.67–1.64, p-value = 0.82, respectively). In addition, those with the C allele of the FSCN1 rs852479 polymorphism were more likely than those with the A allele to develop BC (C allele OR = 0.63; 95% CI 0.46–0.88, p-value = 0.01) (Table 2; Fig. 1A). Regarding FSCN1 rs1640233 C > T and HOTAIR rs920778 C > T polymorphism, differences in all genotypes were not significant for BC patients compared with healthy controls (Table 2; Fig. 1B, C).
The association between genotypes and clinicopathological features of BC patients
In this study, the possible relationship between some clinicopathological parameters of patients with BC and the distribution of SNP genotypes was explored (Table 3). Regarding the clinical characteristics, only the FSCN1 rs1640233 polymorphism of the CC genotype was significantly associated with developing tumor size and lymph node involvement among BC cases (p-value = 0.04 and 0.02, respectively). Otherwise, no significant differences were found in the frequencies of FSCN1 rs852479 and HOTAIR rs920778 genotypes in the patients’ group based on all evaluated features (p-value > 0.05).
The association of FSCN1 haplotype frequencies with BC in the studied groups
Association analysis between the risk of BC and haplotypes of FSCN1 rs852479 and rs1640233 among BC cases and controls is summarized in Table 4. LD was estimated for the two SNPs (r2 = 0.65, D′ = 0.88), which is expected under linkage disequilibrium. The haplotypes’ distribution showed that the AC haplotype was the most frequent in both cases and controls (69.9 and 77.2%, respectively), while AT haplotype showed the lowest frequency among both groups. Overall, none of the considered haplotypes were significantly associated with the development of BC (p-value > 0.05).
Discussion
BC is a complicated and heterogeneous disease with a multifaceted etiology caused by a combination of genetic and lifestyle-related factors. Various studies suggest that SNP genotyping may contribute to risk assessment and guide BC management. In the present case–control study, we evaluated the frequency distributions of the FSCN1 (rs852479, rs1640233) and HOTAIR (rs920778) SNPs and their associations with BC susceptibility in Egyptian women.
Regarding FSCN1, our findings revealed that women with CC genotype frequency of rs852479 C > A are significantly associated with a high risk of developing BC when compared with AC genotype (CC vs. AC, OR = 0.395, 95% CI 0.204–0.76, p-value = 0.005; OR = 1.053, 95% CI 0.67–1.64, p-value = 0.82; respectively). Furthermore, the FSCN1 rs852479 C allele polymorphism is attributed to increased BC risk when compared with the frequency of A allele (C allele OR = 0.63; 95% CI 0.46–0.88, p-value = 0.01). In contrast, the rs1640233 SNP polymorphism of patients and controls did not differ significantly across all genotypes (p-value > 0.05).
Wang et al. [24] investigated the relationship between six SNPs of the FSCN1 gene in a cohort of Han Chinese women. There were no significant variations detected in the genotypes’ frequency of the rs8772, rs3801004, rs2966447, rs852479, and rs1640233 polymorphisms between BC patients and the healthy control group [24]. Nevertheless, another study revealed that Egyptian females with the FSCN1 rs3801004 C > G polymorphisms had a significantly higher risk of BC [36]. Liu et al. [40] suggest that there might be an association between FSCN1 and the development of BC. They further confirmed the possible functional relevance of FSCN1 expression in the development of Triple-Negative Breast Cancer (TNBC) because it was substantially higher in TNBC than in the non-TNBC subtype. Consequently, these results assist in elucidating the functional significance of FSCN1 in the pathogenesis of TNBC and may provide perspectives on the mechanisms behind cancerous progression [40].
Concerning HOTAIR rs920778 C > T polymorphism, we reported no significant difference between the cases and control group (p-value > 0.05) for all genotypes and alleles. Our results are consistent with a recent study on the Egyptian population, which discovered that the rs920778 C > T polymorphism was not significantly related to BC progression [35]. According to prior research, the allelic frequencies of the HOTAIR gene (rs12826786, rs1899663, and rs4759314) were not statistically different between BC patients and cancer-free controls and were not likely to develop BC [41].
Contrary to our findings, it has been observed that there was a significant relationship between the rs920778 polymorphism and a high incidence of BC in women from Turkey [42], Iran [43], India [44], and China [30, 45]. Based on a meta-analysis of 4 studies with 4936 cases and 5214 healthy controls investigating the association of four HOTAIR SNPs with BC vulnerability, it was found that rs920778 polymorphism significantly lowered the risk of BC under heterozygous, homozygous, and recessive models among the West Asians, and increased BC risk under dominant and allele models within the East Asian population [34]. Furthermore, some reports have indicated that HOTAIR SNP rs920778 exhibits variable results in the same population but in distinct cancer types such as gastric [46] and breast [42], which suggests that there are variations in the polymorphism throughout different malignancies.
These disparities in the results could be caused by genetic diversity among ethnic populations resulting from different gene–gene and gene–environment interactions, or they could be the result of additional constraints associated with the number of cases and sampling techniques. As elucidation, the HapMap data (https://www.ncbi.nlm.nih.gov/snp/rs920778) indicates that there are notable variations in the allele frequency of the HOTAIR rs920778 polymorphism between various ethnic communities. Additionally, the assessment of HOTAIR expression in tumor samples could help in better recognition of the role of these polymorphisms in cancer progression, which ought to be investigated further [32].
Numerous investigations have been conducted on the relationship between gene polymorphisms involved in different cellular processes and the risk and clinicopathological aspects of BC. When we analyzed the clinical aspects of rs852479 and rs1640233 FSCN1 and rs920778 HOTAIR genotypic frequencies among BC patients, we found that CC genotype of FSCN1 rs1640233 was significantly associated with developed tumor size and lymph node invasion (p-value = 0.04 and 0.02; respectively). Besides, no statistically significant differences were identified in the frequencies of FSCN1 rs852479 and HOTAIR rs920778 genotypes concerning all evaluated parameters (p-value > 0.05). Within the same context, other investigations suggested that there was no significant correlation between the clinicopathological aspects of BC patients and the HOTAIR rs920778 polymorphism [30]. Interestingly, Hassanzarei et al. [43] discovered that the frequencies of different HOTAIR genotypes in the Iranian population weren’t associated with any clinicopathological features except for rs920778, which was significantly related to ER status. Conversely, Bayram et al. [42] found that the CC genotype of HOTAIR rs920778 polymorphism was associated with advanced TNM classification, larger tumor size, poor histological grade, and the presence of distant metastasis in BC patients but was not related to other clinic-laboratory or hormonal parameters.
In a comparison of clinic-pathological aspects with FSCN1 genotypes, Wang et al. [24] discovered that BC patients with the FSCN1 rs852479 and rs1640233 were not statistically correlated to any clinical status of the tumor. Using immunohistochemistry, Min et al. [47] investigated FSCN1 expression in a microarray of 194 samples from patients with invasive breast cancer. Findings suggested a strong correlation between the expression of FSCN1 and some clinicopathological characteristics, such as high histological grade, tumor necrosis, and status of ER- and PR-negativity. They further found that FSCN1 expression was significantly associated with BC survival, especially in patients with advanced-stage BC [47]. Moreover, in Chinese and African-American women, FSCN1 expression is suggested to be associated with TNBC and also linked to more severe clinical aspects and negative hormone receptors [48, 49]. Haplotype analyses may provide evidence about the genetic involvement in disease incidence [50]. We examined the impacts of different haplotype combinations of two FSCN1 SNPs rs852479 and rs1640233 upon the risk of BC, and no significant relation between haplotypes and BC susceptibility was detected. Overall, as related to other functional polymorphisms in other genes, the effect of genetic polymorphisms of FSCN1 and HOTAIR on predisposition to BC would be affected by additional factors in these genes or perhaps other genes, and the assessment should be customized on a population-specific criterion.
Conclusions
The findings of this study suggest that FSCN1 rs852479 C > A polymorphism is implicated in BC risk and development among Egyptian women. Furthermore, CC variant of FSCN1 rs1640233 C > T has been found to be significantly associated with some BC prognostic factors, potentially worsening the prognosis for those carrying the polymorphism. Otherwise, no significant relationship between the HOTAIR rs920778 C > T polymorphism and BC risk in our patients was detected. To our knowledge, this is the first study regarding FSCN1 rs852479 and rs1640233 polymorphisms and their association with BC susceptibility in Egyptian women. Further studies are needed to be conducted in larger patient cohorts to explore specific clinical and pathological characteristics as well as in patients from different populations.
Data availability
The data sets generated and/or analyzed over the course of the study are not publicly available but are available from the corresponding author upon reasonable request.
Abbreviations
- BC:
-
Breast cancer
- SNPs:
-
Single nucleotide polymorphisms
- FSCN1 :
-
Fascin-1
- HOTAIR :
-
HOX Transcript antisense RNA
- TNBC:
-
Triple-negative breast cancer
References
Ahmed H, Hussein M (2022) Impacts of genetic polymorphisms on breast cancer. Int J Oncol Res 5:037. https://doi.org/10.23937/2643-4563/1710037
Sung H, Ferlay J, Siegel RL et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71(3):209–249. https://doi.org/10.3322/CAAC.21660
Lima SM, Kehm RD, Terry MB (2021) Global breast cancer incidence and mortality trends by region, age-groups, and fertility patterns. EClinicalMedicine 38:100985. https://doi.org/10.1016/j.eclinm.2021.100985
Siegel RL, Giaquinto AN, Jemal A (2024) Cancer statistics, 2024. CA Cancer J Clin 74(1):12–49. https://doi.org/10.3322/caac.21820
Abdelaziz A, Shawki M, Shaaban A et al (2020) Breast cancer awareness among Egyptian women and the impact of caring for patients with breast cancer on family caregivers’ knowledge and behaviour. Res Oncol 17(1):1–8. https://doi.org/10.21608/RESONCOL.2020.42340.1114
Azim HA, Elghazawy H, Ghazy RM et al (2023) Clinicopathologic features of breast cancer in Egypt—contemporary profile and future needs: a systematic review and meta-analysis. JCO Glob Oncol 9:e2200387. https://doi.org/10.1200/go.22.00387
Osman Y, Elsharkawy T, Hashim TM et al (2022) Study of single nucleotide polymorphisms associated with breast cancer patients among Arab ancestries. Int J Breast Cancer 2022:2442109. https://doi.org/10.1155/2022/2442109
Robert F, Pelletier J (2018) Exploring the impact of single-nucleotide polymorphisms on translation. Front Genet 9:507. https://doi.org/10.3389/FGENE.2018.00507
Ali HM, Ellakwa DES, Elaraby NM, Zaher AM, Amr KS (2023) Study the association of microRNA polymorphisms (miR-146a, miR-4513) with the risk of coronary heart diseases in Egyptian population. J Biochem Mol Toxicol 37(3):e23284. https://doi.org/10.1002/JBT.23284
Berry NK, Scott RJ, Rowlings P, Enjeti AK (2019) Clinical use of SNP-microarrays for the detection of genome-wide changes in haematological malignancies. Crit Rev Oncol Hematol 142:58–67. https://doi.org/10.1016/J.CRITREVONC.2019.07.016
Deng N, Zhou H, Fan H, Yuan Y (2017) Single nucleotide polymorphisms and cancer susceptibility. Oncotarget 8(66):110635. https://doi.org/10.18632/ONCOTARGET.22372
Van Veen EM, Brentnall AR, Byers H et al (2018) Use of single-nucleotide polymorphisms and mammographic density plus classic risk factors for breast cancer risk prediction. JAMA Oncol 4(4):476–482. https://doi.org/10.1001/JAMAONCOL.2017.4881
Li CH, Chan MH, Liang SM, Chang YC, Hsiao M (2022) Fascin-1: Updated biological functions and therapeutic implications in cancer biology. BBA Adv 2:100052. https://doi.org/10.1016/J.BBADVA.2022.100052
Liu H, Zhang Y, Li L et al (2021) Fascin actin-bundling protein 1 in human cancer: promising biomarker or therapeutic target? Mol Ther Oncolytics 20:240–264. https://doi.org/10.1016/J.OMTO.2020.12.014
Poli G, Ruggiero C, Cantini G et al (2019) Fascin-1 is a novel prognostic biomarker associated with tumor invasiveness in adrenocortical carcinoma. J Clin Endocrinol Metab 104(5):1712–1724. https://doi.org/10.1210/jc.2018-01717
Abbasi A, Noroozinia F, Anvar S, Abbasi MA, Hosseinzadeh S, Seyed Mokhtari A (2020) Fascin overexpression is associated with higher grades of breast cancer. Pol J Pathol 70(4):264–268. https://doi.org/10.5114/PJP.2019.93128
Zhao W, Gao J, Wu J et al (2015) Expression of Fascin-1 on human lung cancer and paracarcinoma tissue and its relation to clinicopathological characteristics in patients with lung cancer. Onco Targets Ther 8:2571. https://doi.org/10.2147/OTT.S81915
Piskor BM, Pryczynicz A, Lubowicka E et al (2018) Immunohistochemical expression of Fascin-1 in colorectal cancer in relation to clinical and pathological parameters. Folia Histochem Cytobiol 1(2):106–112. https://doi.org/10.5603/FHC.A2018.0011
Gupta I, Vranic S, Al-Thawadi H et al (2021) Fascin in gynecological cancers: an update of the literature. Cancers 13(22):5760. https://doi.org/10.3390/CANCERS13225760
Park SH, Song JY, Kim YK et al (2014) Fascin1 expression in high-grade serous ovarian carcinoma is a prognostic marker and knockdown of fascin1 suppresses the proliferation of ovarian cancer cells. Int J Oncol 44(3):637–646. https://doi.org/10.3892/IJO.2013.2232
Natesan SC, Ramakrishnan BP, Krishnapillai R, Thomas P (2019) Immunohistochemical expression of fascin in oral epithelial dysplasia and oral squamous cell carcinoma. World J Dent 10(5):340–345. https://doi.org/10.5005/jp-journals-10015-1658
Sarantelli E, Mourkakis A, Zacharia LC, Stylianou A, Gkretsi V (2023) Fascin-1 in cancer cell metastasis: old target-new insights. Int J Mol Sci 24(14):11253. https://doi.org/10.3390/IJMS241411253
Chang D, Liu XX, Liu R, Sun JW (2023) The role and regulatory mechanism of FSCN1 in breast tumorigenesis and progression. Yi Chuan 45(2):115–127. https://doi.org/10.16288/J.YCZZ.22-346
Wang CQ, Tang CH, Wang Y et al (2017) FSCN1 gene polymorphisms: biomarkers for the development and progression of breast cancer. Sci Rep 7(1):15887. https://doi.org/10.1038/S41598-017-16196-6
Raju GSR, Pavitra E, Bandaru SS et al (2023) HOTAIR: a potential metastatic, drug-resistant and prognostic regulator of breast cancer. Mol Cancer 22(1):1–15. https://doi.org/10.1186/S12943-023-01765-3
Jiang D, Xu L, Ni J, Zhang J, Cai M, Shen L (2019) Functional polymorphisms in LncRNA HOTAIR contribute to susceptibility of pancreatic cancer. Cancer Cell Int 19(1):1–8. https://doi.org/10.1186/S12935-019-0761-X
Pan W, Liu L, Wei J, Ge Y, Zhang J, Chen H et al (2015) A functional lncrna hotair genetic variant contributes to gastric cancer susceptibility. Mol Carcinog 55(1):90–96. https://doi.org/10.1002/mc.22261
Madadi Rad R, Pouladi N, Bosharani NN, Alizadeh A (2022) Association between HOTAIR rs1899663 G>T gene polymorphism and thyroid cancer susceptibility. J Babol Univ Med Sci 24(1):95–102
Kim JO, Jun HH, Kim EJ et al (2020) Genetic variants of HOTAIR associated with colorectal cancer susceptibility and mortality. Front Oncol 10:72. https://doi.org/10.3389/FONC.2020.00072
Lv Z, Kou C, Chen N et al (2021) Single nucleotide polymorphisms in HOTAIR are related to breast cancer risk and prognosis in the Northeastern Chinese population. Front Oncol 11:706428. https://doi.org/10.3389/FONC.2021.706428
Qi Q, Wang J, Huang B et al (2016) Association of HOTAIR polymorphisms rs4759314 and rs920778 with cancer susceptibility on the basis of ethnicity and cancer type. Oncotarget 7(25):38775–38784. https://doi.org/10.18632/ONCOTARGET.9608
Tian T, Li C, Xiao J et al (2016) Quantitative assessment of the polymorphisms in the HOTAIR lncRNA and cancer risk: a meta-analysis of 8 case-control studies. PLoS ONE 11(3):e0152296. https://doi.org/10.1371/JOURNAL.PONE.0152296
Hajjari M, Rahnama S (2019) Association between SNPs of long non-coding RNA HOTAIR and risk of different cancers. Front Genet 10:113. https://doi.org/10.3389/fgene.2019.00113
Wang B, Yuan F, Zhang F, Miao Z, Jiang D (2023) A systematic review and meta-analysis of the association between HOTAIR polymorphisms and susceptibility to breast cancer. Arch Med Sci 19(1):128–137. https://doi.org/10.5114/AOMS.2019.87537
Anber N, Tarabay MM, Elmougy R, Abdel-Dayem MA, Elbendary EY (2023) Association of HOTAIR gene rs920778 (C > T) and rs4759314 (A > G) polymorphism with breast cancer in Egyptian women. Mol Biol Rep 50(11):9153. https://doi.org/10.1007/S11033-023-08725-6
Ibrahim NE, Hamed RMR, Refaat A, Mosaad YO, Mekawy DM (2023) Genetic polymorphism in FSCN1 rs3801004 C/G and CD44 rs353639 A/C, as prognostic factor in Egyptian breast cancer patients. Asian Pac J Cancer Prev 24(10):3517–3523. https://doi.org/10.31557/APJCP.2023.24.10.3517
Miller SA, Dykes DD, Polesky HF (1988) A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res 16(3):1215–1215. https://doi.org/10.1093/NAR/16.3.1215
Kang H (2021) Sample size determination and power analysis using the G*Power software. J Educ Eval Health Prof 18:17. https://doi.org/10.3352/jeehp.2021.18.17
Solé X, Guinó E, Valls J, Iniesta R, Moreno V (2006) SNPStats: a web tool for the analysis of association studies. Bioinformatics 22(15):1928–1929. https://doi.org/10.1093/BIOINFORMATICS/BTL268
Liu Z, Ning G, Xu R, Cao Y, Meng A, Wang Q (2016) Fscn1 is required for the trafficking of TGF-β family type I receptors during endoderm formation. Nat Commun 7:12603. https://doi.org/10.1038/NCOMMS12603
Khorshidi HR, Taheri M, Noroozi R, Soudyab M, Sayad A, Ghafouri-Fard S (2017) Investigation of the association of HOTAIR single nucleotide polymorphisms and risk of breast cancer in an Iranian population. Int J Cancer Manag 10(5):7498. https://doi.org/10.5812/IJCM.7498
Bayram S, Sümbül AT, Batmacı CY, Genç A (2015) Effect of HOTAIR rs920778 polymorphism on breast cancer susceptibility and clinicopathologic features in a Turkish population. Tumour Biol 36(5):3863–3870. https://doi.org/10.1007/S13277-014-3028-0
Hassanzarei S, Hashemi M, Sattarifard H, Hashemi SM, Bahari G, Ghavami S (2017) Genetic polymorphisms of HOTAIR gene are associated with the risk of breast cancer in a sample of southeast Iranian population. Tumor Biol 39(10):1–8. https://doi.org/10.1177/1010428317727539
Rajagopal T, Seshachalam A, Akshaya RL et al (2020) Association of HOTAIR (rs920778 and rs1899663) and NME1 (rs16949649 and rs2302254) gene polymorphisms with breast cancer risk in India. Gene 762:145033. https://doi.org/10.1016/J.GENE.2020.145033
Yan R, Cao J, Song C et al (2015) Polymorphisms in lncRNA HOTAIR and susceptibility to breast cancer in a Chinese population. Cancer Epidemiol 39(6):978–985. https://doi.org/10.1016/J.CANEP.2015.10.025
Bayram S, Ülger Y, Sümbül AT, Kaya BY, Rencüzoğulları A, Genç A, Sevgiler Y, Bozkurt O, Rencüzoğulları E (2015) A functional HOTAIR rs920778 polymorphism does not contributes to gastric cancer in a Turkish population: a case-control study. Fam Cancer 14(4):561–567. https://doi.org/10.1007/s10689-015-9813-0
Min KW, Chae SW, Kim DH et al (2015) Fascin expression predicts an aggressive clinical course in patients with advanced breast cancer. Oncol Lett 10(1):121. https://doi.org/10.3892/OL.2015.3191
Wang CQ, Li Y, Huang BF et al (2017) EGFR conjunct FSCN1 as a novel therapeutic strategy in triple-negative breast cancer. Sci Rep 7(1):15654. https://doi.org/10.1038/S41598-017-15939-9
Esnakula AK, Ricks-Santi L, Kwagyan J et al (2014) Strong association of fascin expression with triple negative breast cancer and basal-like phenotype in African-American women. J Clin Pathol 67(2):153–160. https://doi.org/10.1136/JCLINPATH-2013-201698
D’Amelio AM, Monroy C, El-Zein R, Etzel CJ (2012) Using haplotype analysis to elucidate significant associations between genes and Hodgkin lymphoma. Leuk Res 36(11):1359. https://doi.org/10.1016/J.LEUKRES.2012.07.014
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Contributions
NHE, ERG, LKA, and YE: performed the molecular experiments, collected the DNA samples, and contributed to validation and interpretation of the data; ERG and NHE: made statistics and analysis of data; SEPS and DAA: participated in the management of the study subjects, examined the patients, collected biological samples, and performed clinical-pathological analysis; ERG: wrote the first draft of the manuscript, reviewed the manuscript before submission, and corresponded with the journal. All authors have read, revised, and approved the final form of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflicts of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
This study has been approved by the Ethics Committee of the Faculty of Pharmacy (Girls), Al-Azhar University, Cairo, Egypt (REC number: 436).
Informed consent
Written informed consent was obtained from all participants following a full explanation of the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Galal, E.R., Abdelhakam, D.A., Ahmed, L.K. et al. The association of FSCN1 (rs852479, rs1640233) and HOTAIR (rs920778) polymorphisms with the risk of breast cancer in Egyptian women. Mol Biol Rep 51, 495 (2024). https://doi.org/10.1007/s11033-024-09459-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11033-024-09459-9