Abstract
Background
Insufficient trophoblast invasion, culminating in suboptimal uterine spiral artery remodeling, is pinpointed as a pivotal contributor to preeclampsia (PE) development. LINC01410 has been documented to be increased in various neoplasms, and is significantly associated with the invasive capabilities of tumor cells. Nonetheless, its function and the mechanisms in the pathogenesis of PE require further investigation.
Methods and results
LINC01410 and methyltransferase-like 3 (METTL3) were ectopically expressed in HTR-8/Svneo cells via lentiviral transduction. Subsequently, the cells’ invasive capabilities and apoptosis rates were evaluated employing Transwell assays and flow cytometry, respectively. The interplay between LINC01410 and METTL3, alongside the m6A methylation of FAS, was probed through RNA immunoprecipitation (RIP). Additionally, the association between FAS and METTL3 was elucidated via Coimmunoprecipitation (Co-IP) assays. The protein level of NF-κB, BAX, and BCL-2 in LINC01410-overexpressing cells was detected by Western blot. Our findings revealed that LINC01410 elevation increased the invasive ability of HTR-8/Svneo cells, directly impacting METTL3 then leading to its reduced expression. Conversely, heightened METTL3 expression mitigated invasiveness while enhancing apoptosis in these cells. Moreover, METTL3’s interaction with FAS led to increased FAS expression, subject to m6A methylation. A surge in LINC01410 markedly decreased both mRNA and protein levels of FAS. Furthermore, LINC01410 overexpression significantly reduced NF-κB and BAX protein levels while augmenting BCL-2.
Conclusions
Upregulation of LINC01410 expression promotes trophoblast cell invasion by inhibiting FAS levels through modified m6A alteration and suppressing the NF-κB pathway. These findings underscore the pivotal role of LINC01410 in regulating trophoblast cell invasion and propose it as a promising therapeutic strategy for preventing or alleviating PE. This offers valuable insights for the clinical treatment of PE, for which definitive targeted therapy methods are currently lacking.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
PE ranks as the third leading etiological factor contributing to maternal and neonatal morbidity and mortality, with it being responsible for over 60,000 maternal deaths annually worldwide [1]. The current medical landscape is notably deficient in efficacious therapeutic or preventive interventions for PE, largely attributed to the enigmatic nature of its pathogenesis [2]. Prominent physiological features of PE encompass augmented apoptosis in trophoblast cells and diminished trophoblast invasion, culminating in suboptimal remodeling of the spiral arteries [3]. Trophoblast cells, which constitute the fetal component of the placenta, are integral to fetal development [4]. Dysfunctional trophoblast cellular processes, manifesting as hindered proliferation, heightened apoptosis, and insufficient invasive capacity, are intricately related to the pathogenetic mechanisms underpinning PE [5]. Accordingly, genes that regulate trophoblast cell behavior are of paramount significance in the progression and pathogenesis of PE [6].
Long non-coding RNAs (lncRNAs), characterized as non-protein coding RNA sequences exceeding 200 nucleotides, are substantiated to exhibit profound associations with a myriad of pathologies, inclusive of neoplastic disorders [7]. Contemporary research has elucidated a nexus between lncRNAs and the pathogenesis and advancement of PE. Multiple lncRNAs have exhibited increased expression in extracellular vesicles derived from plasma of PE patients, suggesting their potential role in the initiation and progression of the condition [8]. Specially, LncRNA INHBA suppresses the invasion and migration of trophoblast cells by inhibiting CENPB expression [9]. lncRNA SH3PXD2A hinders trophoblast invasion and migration during placental development by repressing SH3PXD2A and CCR7 in the transcription level [10].
N6-Methyladenosine (m6A) is a prevalent and dynamic modification found in RNA that exerts significant influence over various biological processes and diseases [11]. Being one of the prevalent chemical alterations in eukaryotic RNAs, m6A holds pivotal roles in influencing RNA stability, localization, translation, splicing, or transport [12]. m6A modification is intricately controlled through its addition, removal, and detection by designated enzymes and proteins. This process involves m6A methyltransferases (METTL3/14/16, WTAP, RBM15/15B, VIRMA, CBLL1, and ZC3H13, collectively known as “writers”), demethylases (FTO and ALKBH5, referred to as “erasers”), and specific m6A-binding proteins (such as YTHDF1/2/3, YTHDC1/2, IGH2BP1/2/3, hnRNPs, eIF3, and FMR1, identified as “readers”) [13]. Moreover, the interplay between m6A and non-coding RNA (ncRNA) significantly contributes to the regulatory mechanisms governing disease progression [14]. The process of m6A modification is known to influence the processing, maturation, and operational capabilities of these ncRNAs, thereby indirectly modulating the expression of oncogenes and tumor suppressor genes [15]. For example, upregulation of the lncRNA BLACAT3 via m6A modification promotes angiogenesis and hematogenous metastasis in bladder cancer [16]. Similarly, increased circMDK levels, facilitated by m6A modification, accelerate cell proliferation, migration and invasion in hepatocellular carcinoma [17].
LINC01410, which encompasses 2877 nucleotides and is situated on chromosome 9 (coordinates: 62,801,461 − 62,813,486), is a novel lncRNA [18]. It has been implicated in facilitating the invasive and migratory capacities of an array of cancers, notably pancreatic, colorectal, gastric, cervical, thyroid, and endometrial carcinomas [19, 20]. Despite this, the exact molecular mechanisms by which LINC01410 influences trophoblast cell invasion remain unclear. This study aims to investigate the role and mechanisms of LINC01410 in trophoblast cells, potentially revealing new therapeutic strategies for the prevention or mitigation of PE.
Materials and methods
HTR-8/Svneo cells culture
Human trophoblast HTR-8/Svneo cells were sourced from the Chinese Academy of Science. These cells were kept in RPMI 1640 medium (Gibco) enriched with 10% FBS (Gibco) and bolstered by 1% penicillin/streptomycin (Gibco). The cells were then maintained in a humidified atmosphere with 5% CO2 at 37˚C.
Vector construction
The lentiviral vector overexpressing LINC01410 was carefully constructed using the full-length human cDNA sequence of the lincRNA LINC01410, acquired from the NCBI GenBank (Gene Bank ID: NR_121647.1, https://www.ncbi.nlm.nih.gov/nuccore/NR_121647.1), by Shanghai GeneChem. The coding sequence of LINC01410 was inserted into the GV367 vector to generate LINC01410-overexpression vector.
The lentiviral vector engineered for METTL3 overexpression was meticulously constructed utilizing human METTL3 genetic sequences sourced from the NCBI GenBank (Gene Bank ID: XM_532627.6, accessible at https://www.ncbi.nlm.nih.gov/nuccore/XM_532627.6) by Shanghai GeneChem. Subsequently, the coding sequence of METTL3 was meticulously inserted into the GV358 vector to yield the METTL3 overexpression vector.
Lentiviral transduction
HTR-8/Svneo cells were cultured in 6-well plates at 37˚C for a duration of 36 h. Upon attaining approximately 70% confluency, the cells were subjected to transfection using Lipofectamine 2000 (Thermo Fisher) for a period of 48 h at 37˚C. Following transfection, each well was uniformly exposed to equivalent volumes (100 µL) of lentiviral particles harboring either METTL3-overexpressing or LINC01410-overexpressing vectors, alongside a negative control (NC; empty vector). After a 48-hour post-transfection interval, the HTR-8/Svneo cells were harvested for subsequent analysis.
Quantitative real-time polymerase chain reaction (qPCR)
Total RNA isolation from HTR-8/Svneo cells was meticulously conducted using the RNA-iSo PluS kit (Takara). Following extraction, the RNA underwent reverse transcription to synthesize complementary DNA (cDNA) with the aid of the cDNA Synthesis SuperMix (Yeasen). Subsequently, the resultant cDNA was subjected to qPCR amplification using the SYBR Green qPCR Mix (Med Chem Express) and analyzed employing the ABI 7500 Real-Time PCR system (Life Technologies, USA). To ensure data comparability, LINC01410 expression levels were normalized to the reference gene GAPDH using the 2−ΔΔCt method [21]. The specific primer sequences employed were: GAPDH (forward: 5’- GCTGTAGCCAAATCGTTGT − 3’, reverse: 5’- CCAGGTGGTCTCCTCTGA − 3’), LINC01410 (forward: 5’- GGCTTACCCATCTGGCAAGT-3’, reverse: 5’-GAGTAGGCAACTGTGCACCT-3’), METTL3 (forward: 5’- GTGATCGTAGCTGAGGTTCGT − 3’, reverse: 5’- GGGTTGCACATTGTGTGGTC − 3’) and FAS (forward: 5’- GACCCTCCTACCTCTGGTTCT − 3’, reverse: 5’- ACCTGGAGGACAGGGCTTAT − 3’).
Western blot
Proteins were meticulously extracted from HTR-8/Svneo cells via RIPA lysis buffer (Med Chem Express) with utmost precision. After extraction, the concentration of the proteins was accurately quantified utilizing the BCA Protein Assay Kit (Thermo Fisher). For the process of gel electrophoresis, an aliquot containing 5 µg of the extracted proteins was combined with 5 × SDS sample buffer and subjected to electrophoretic separation on a 12% SDS-polyacrylamide gel. The proteins were subsequently transferred onto a PVDF membrane. To mitigate non-specific binding, the membranes were blocked with 5% non-fat milk at room temperature for 2 h prior to overnight incubation at 4 °C with the designated primary antibodies: Rabbit anti-METTL3 (1:1000,Abcam), Rabbit anti-FAS (1:1000, Abcam), Rabbit anti- NF-κB (1:1000, Abcam), Rabbit anti- BAX (1:1000, Abcam), Rabbit anti-BCL-2 (1:1000, Proteintech) and Mouse anti-GAPDH (1:5000, Proteintech). Post primary antibody exposure, the membranes underwent an extensive washing procedure prior to being subjected to an incubation period with their respective secondary antibodies (1:5,000, KPL) for a duration of 2 h. Subsequently, the visualization of the targeted protein bands was facilitated through the application of ECL Western blot detection reagents (Thermo Fisher).
Cell viability assay
The assessment of HTR-8/Svneo cell invasion potential post-transfection with LINC01410 overexpressing lentivirus was conducted via the Transwell assay, employing BD Matrigel invasion chambers (BD Biosciences). Furthermore, a Transwell co-culture assay was executed utilizing a 24-well plate configuration. HTR-8/Svneo cells were seeded into the upper chamber of the Transwell with 200 µL of serum-free RPMI 1640 at a density of 7 × 103 cells/well, while the lower chamber was filled with 800 µL of RPMI 1640 supplemented with 10% FBS. Following a 48-hour incubation period at 37˚C, the HTR-8/Svneo cells in the basolateral chamber underwent two PBS washes and stained with 1% crystal violet for 30 min. After another PBS wash, the stained HTR-8/Svneo cells were visualized and imaged using a high-resolution light microscope Olympus cX2 at 100x magnification.
Flow cytometry
Apoptosis in HTR-8/Svneo cells was assessed using the Annexin V-FITC and PI staining kit (Vazyme) in strict accordance with the manufacturer’s instructions. Subsequently, the stained cells underwent analysis via the Beckman DXI800 flow cytometer (Beckman Instruments). Quantification of the apoptotic cell population (PI+) was performed using FlowJo software (version 10.1r5, Tree Star, Inc.). The apoptotic rate (%) was calculated as the combined percentage of early and late apoptotic cells divided by the total cell count.
RIP
The RIP assay utilized the RIP Kit (BersinBio). Post pre-chilled PBS washes, HTR-8/Svneo cells underwent lysis in RIP buffer at 4 °C for a duration of 30 min. Magnetic beads, conjugated with either human anti-METTL3 antibody (Abcam) and normal mouse immunoglobulin G (IgG; Abcam), or anti-m6A antibody (Abcam) and IgG, were employed for RNA capture. Subsequent RNA extraction from the precipitates was facilitated by Trizol reagent (Takara). The expression levels of LINC01410 and FAS were quantified assessed through RT-PCR, utilizing the designated primer sequences for the respective target genes: LINC01410 forward 5′ - GGCTTACCCATCTGGCAAGT − 3′ and reverse 5′ - GAGTAGGCAACTGTGCACCT − 3′; FAS forward 5′ - GACCCTCCTACCTCTGGTTCT − 3′ and reverse 5′ - CTCCTTCCCTTCTTGGCAGG − 3′.
Co-IP assay
Co-IP kit (Thermo Fisher) for Co-IP experiments. Initially, two confluent cultures of HTR-8/Svneo cells were selected. Lysates from these cells were prepared utilizing an IP buffer composed of 150 mM NaCl, 0.1% Triton X-100, 100 mM Tris-HCl, 1 mM EDTA, with the pH adjusted to 7.4. These lysates underwent centrifugation at a speed of 14,000 g for a duration of 20 min. This supernatant was subsequently combined with a Rabbit anti-FAS antibody (Abcam) and incubated at 4℃ overnight. The sample, now bearing the antibody-protein complexes, underwent an additional 4-hour incubation with A/G beads at 4 °C. Following this incubation, the beads were harvested via centrifugation and subjected to three wash steps. The purified beads were subsequently analyzed using SDS-PAGE, and the separated proteins were transferred onto a PVDF membrane. A Rabbit anti-METTL3 antibody (Abcam) was utilized as the probe to detect interactions between METTL3 and FAS proteins. The entire Co-IP procedure was conducted at 4 °C.
Statistical analysis
The quantitative data obtained from the experiments were analyzed using GraphPad Prism software (Version 8.0; California). To ensure the reliability of the results, each experiment was independently replicated thrice. Data presentation was standardized as mean ± standard deviation. For pairwise group comparisons, an unpaired Student’s t-test was employed due to its suitability for comparing the means of two independent groups. In cases involving multiple group comparisons, a one-way ANOVA was utilized to examine the variance among three or more independent groups, followed by a Tukey’s post hoc test to identify specific group differences. This combination of statistical tests was chosen to control for Type I error across multiple comparisons. Statistical significance was established at a threshold of p < 0.05.
Results
LINC01410 promoted the invasion of trophoblast cells
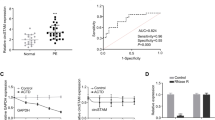
LINC01410 has been implicated in augmenting the invasive and migratory capabilities across various cancer types, including but not limited to pancreatic, colorectal, gastric, cervical, thyroid, and endometrial carcinomas [19, 20]. However, the precise biological function of LINC01410 within trophoblast cells is yet to be elucidated. In this study, we induced overexpression of LINC01410 in HTR-8/Svneo trophoblast cells using lentiviral transfection. Subsequent qPCR confirmed marked increases in mRNA level of LINC01410 in the transfected cells compared to controls (Fig. 1A). High LINC01410 expression notably increased the invasion ability of HTR-8/Svneo cells (Fig. 1B). In addition, elevated LINC01410 led a significant decrease in METTL3 mRNA level (Fig. 1C). LINC01410- METTL3 complexes was detected by RIP, suggesting that LINC01410 can directly interact with METTL3 (Fig. 1D).
Upregulated LINC01410 inhibited the invasion ability of HTR-8/Svneo cells. (A) mRNA level of LINC01410 in HTR-8/Svneo cells was determined by qPCR. (B) The invasion capability of LINC01410-upregulated HTR-8/Svneo cells was determined by Transwell. (C) mRNA level of METTL3 in LINC01410-upregulated HTR-8/Svneo cells was determined by qPCR. (D) The interaction between LINC01410 and METTL3 was detected by RIP. ***p < 0.001. NC: negative control; LINC01410: LINC01410 overexpression
LINC01410 promoted the invasion of trophoblast cells by downregulating METTL3
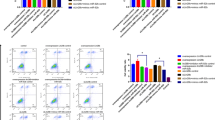
To further investigate the role of METTL3 in trophoblast cells, we upregulated METTL3 in HTR-8/Svneo trophoblast cells via lentiviral transfection. Both mRNA and protein levels of METTL3 markedly increased in the transfected cells compared to controls (Fig. 2AB). Notably, elevated METTL3 expression markedly decreased the invasion ability of HTR-8/Svneo cells (Fig. 2C). Moreover, the upregulation of METTL3 substantially heightened the apoptosis rate of HTR-8/Svneo cells (Fig. 2D). As expected, heightened METTL3 expression countered the facilitative impact of overexpressed LINC01410 on the invasiveness of HTR-8/Svneo cells (Fig. 2E).
Elevated METTL3 promoted the invasion while increased the apoptosis of HTR-8/Svneo cells. (A) mRNA level of METTL3 in HTR-8/Svneo cells was determined by qPCR. (B) Protein level of METTL3 in HTR-8/Svneo cells was determined by Western blot. (C) The invasion capability of METTL3-upregulated HTR-8/Svneo cells was determined by Transwell. (D) Ratio of apoptosis of METTL3-upregulated HTR-8/Svneo cells were examined based on flow cytometric detection. (E) The invasion capability of HTR-8/Svneo cells was determined by Transwell. Data is represented as means ± SD (n = 3). ***p < 0.001. NC: negative control; METTL3: METTL3 overexpression
METTL3 decreased the expression of FAS
FAS has been identified as a putative target gene of LINC01410 [22]. In addition, FAS expression is significantly higher in placenta tissues and may be a susceptibility gene for the development of PE [23, 24]. We further investigated whether FAS is involved in the LINC01410-mediated suppression of METTL3, which promotes the invasion of HTR-8/Svneo cells. Upregulated METTL3 resulted in a significant increase in FAS protein level (Fig. 3A). METTL3-FAS complexes were detected by RIP, suggesting that FAS can directly interact with METTL3 (Fig. 3B). Additionally, it was found that FAS was modified by m6A methylation, as demonstrated by m6A RIP analysis (Fig. 3C). As expected, increased expression of LINC01410 lead to a significant decrease in both the mRNA and protein levels of FAS (Fig. 3D, E).
METTL3 decreased the expression of FAS. (A) Protein level of FAS in METTL3-upregulated HTR-8/Svneo cells was determined by Western blot. (B) The interaction between FAS and METTL3 was detected by Co-IP. (C) m6A modification on FAS was detected by RIP. (D) mRNA level of FAS in LINC01410-upregulated HTR-8/Svneo cells was determined by qPCR. (E) Protein level of FAS in LINC01410-upregulated HTR-8/Svneo cells was determined by Western blot. Grayscale analysis was performed by ImageJ software. Data is represented as means ± SD (n = 3). ***p < 0.001. NC: negative control; METTL3: METTL3 overexpression; LINC01410: LINC01410 overexpression
LINC01410 inhibited the apoptosis of trophoblast cells by suppressing NF-κB signaling cascade
NF-κB has been recognized as a downstream target of FAS/FASL [25]. In this investigation, we identified that heightened expression of LINC01410 notably diminished the protein abundance of NF-κB (Fig. 4A, B). Additionally, LINC01410 suppressed the expression of the pro-apoptotic gene BAX while augmenting the protein expression of the anti-apoptotic gene BCL-2 (Fig. 4A, B). These findings imply that LINC01410 might impede trophoblast cell apoptosis via modulation of the METTL3/FAS/NF-κB pathway.
LINC01410 inhibited the apoptosis of trophoblast cells by suppressing NF-κB signaling cascade. (A) The protein level of NF-κB, BAX and BCL-2 were detected by Western blot. (B) Grayscale analysis was performed by ImageJ software. Data is represented as means ± SD (n = 3). ***p < 0.001. NC: negative control; LINC01410: LINC01410 overexpression
Discussion
PE remains a leading cause of maternal mortality and morbidity [26]. Currently, the only definitive cure for PE is the delivery of the placenta and fetus, which often results in iatrogenic preterm birth [27]. To mitigate this and improve outcomes for mothers, children, and adult offspring, research efforts are increasingly directed not only at treating PE but also at preventing its onset. Recent advances in understanding the pathogenesis of PE, coupled with the necessity to reduce its short- and long-term morbidities, have spurred investigations into novel agents aimed at either preventing or treating the condition [28]. Inadequate trophoblast cell invasion leading to suboptimal uterine spiral artery remodeling is identified as a critical contributing factor to PE development [4, 29]. We identified that the upregulation of LINC01410 significantly enhanced the invasiveness of HTR-8/Svneo cells, suggesting its involvement in PE pathogenesis. Elevated levels of LINC01410 led to the downregulation of METTL3, which, when upregulated, increased FAS expression through changes in m6A modification. Consequently, overexpression of LINC01410 might enhance trophoblast invasiveness by suppressing the METTL3/FAS pathway. This identified LINC01410 as a potential therapeutic approach for preventing or alleviating PE.
LINC01410 has emerged as a significant player in regulating cellular proliferation and invasion across various cancer types, acting through diverse mechanisms involving miRNA interaction and modulation of signaling pathways. Studies have elucidated its role in osteosarcoma, where the LINC01410/miR-122-5p/NDRG3 axis facilitates cell proliferation and migration [30], and in bladder cancer, LINC01410 enhances migration, invasion, and EMT by modulating the miR-4319/Snail1 axis [31]. Furthermore, LINC01410 has been shown to promote osteosarcoma tumorigenesis via the miR-497-5p/HMGA2 axis [32], and its knockdown in cervical cancer leads to suppressed growth and invasion by targeting the miR-2467-3p/VOPP1 axis [33]. Additionally, LINC01410 accelerates invasion and proliferation in osteosarcoma by sponging miR-3128 [34], while in cholangiocarcinoma, it fosters cell proliferation and migration through the miR-124-3p/SMAD5 axis [35]. High expression of LINC01410 has been identified as a potential diagnostic and prognostic marker in colorectal cancer [36]. Its silencing in glioblastoma cells reduces viability but increases apoptosis and sensitivity to temozolomide, acting through the PTEN/AKT pathway by targeting miR-370-3p [20]. Our findings revealed that elevated LINC01410 expression significantly accelerated the invasive potential of HTR-8/Svneo cells, suggesting LINC01410’s role in PE development could be through the promotion of trophoblast cell invasion.
m6A plays an indispensable role in modulating gene expression and cellular functionalities [11]. Within the context of tumorigenesis, m6A exerts a multifaceted influence on RNA stability, translation efficiency, and splicing, thereby functioning as a double-edged sword capable of both promoting and inhibiting tumor proliferation [11, 13]. Significantly, m6A is instrumental in modulating resistance to tumor treatments by impacting the tumor microenvironment and responses to therapeutic interventions, consequently amplifying or mitigating the efficacy of pharmacological agents [37]. While the bulk of research has predominantly concentrated on oncology, emerging studies have commenced exploring the potential implications and mechanisms of m6A modification in obstetric complications, notably PE. Observations have indicated elevated m6A methylation levels and METTL3/14 expression in placentas affected by PE [38]. One study observes no notable alterations in the levels of ALKBH5 and FTO [39], whereas other research indicates a significant increase in ALKBH5 expression in placentas from PE cases, which was associated with a reduction in trophoblast invasion [40, 41]. Contradictory findings on m6A protein expression and function underscore the need for further research into its role in PE pathogenesis. Our findings demonstrated that LINC01410 directly interacted with METTL3, leading to a reduction in its expression. This suggested that m6A modification exerted significant influence over the promotion effect of elevated LINC01410 on trophoblast cell invasion.
METTL3, recognized as the pioneering and pivotal m6A RNA methyltransferase, orchestrates the addition of m6A modifications onto RNA molecules [42]. In majority of tumors, METTL3 functions as an oncogene. For example, in breast cancer, METTL3 enhances m6A modification of PD-L1 mRNA through the IGF2BP3 pathway, reducing tumor immune surveillance and allowing cancer cells to evade immune destruction, thereby promoting their growth and invasion [43]. In hepatocellular carcinoma, the METTL3-m6A-EGFR axis crucially mediates Lenvatinib resistance, highlighting METTL3’s essential role in altering tumor pharmacological responses [44]. Furthermore, METTL3 has been linked to a spectrum of other health conditions, including neurodegenerative diseases and metabolic disorders [45,46,47]. Nevertheless, the functionality and significance of m6A and METTL3 in PE warrant further exploration. In this study, we demonstrated that enhanced METTL3 expression significantly decreased the invasive capabilities of trophoblast cells and concurrently increases their apoptosis rate. Intriguingly, an increase in METTL3 levels mitigated the promotion impact of LINC01410 overexpression on trophoblast cell invasiveness. Our findings indicated that METTL3 acted as a pivotal mediator of LINC01410’s regulatory influence on the invasive properties of trophoblast cells.
The FAS gene, denoted as CD95, encodes the FAS receptor, a critical apoptotic receptor situated on the cellular membrane that precipitates apoptosis through its engagement with the specific ligand, FASLG (FAS ligand) [48]. The dysregulation of FAS/FASLG axis is profoundly implicated in the etiology of a broad spectrum of pathologies, including diverse oncological conditions [49]. For example, the utilization of APG101 to inhibit the FAS/FASLG signaling axis in glioblastoma has demonstrated efficacy in impeding tumor cell invasion and enhancing the therapeutic outcome of radiotherapy [48]. Moreover, the activation of FAS signaling facilitates tumor cell endurance and metastatic propagation [50]. Beyond its regulatory role in apoptosis, FAS also influences the migratory behavior of trophoblast cells. Elevated level of Fas expression facilitates apoptosis, reduces cell viability, and hinders the migratory capacity of human trophoblast cells [51]. Our study demonstrated that METTL3 directly interacted with FAS, leading to increased FAS expression. m6A RIP analysis showed that FAS was modified by m6A methylation. Moreover, LINC01410 elevation significantly reduced FAS mRNA and protein levels. These results suggested that LINC01410 enhanced trophoblast cell invasiveness by suppressing METTL3, which in turn affected m6A modification on FAS and decreased its expression.
NF-κB, a transcription factor ubiquitously present across diverse cellular contexts, orchestrates a pivotal regulatory spectrum encompassing immune modulation, cellular proliferation, apoptosis, and inflammation [52]. The manipulation of NF-κB activity has been shown to influence tumor cell proliferation within hypoxic niches [53] and mitigate the underlying mechanisms of cerebral aneurysm formation [52]. NF-κB in placental tissue upregulates cytokines, triggering immune and inflammatory responses along with placental cell damage, thus hastening PE pathogenesis [54]. NF-κB can be activated by the Fas-related death receptor TNF-R1 [25]. This study revealed that elevated LINC01410 substantially suppressed NF-κB expression. Notably, elevated LINC01410 reduced BAX expression while augmenting BCL-2 expression. These findings suggested that LINC01410 facilitated invasion in trophoblast cells by downregulating FAS, thereby attenuating the NF-κB signaling cascade. These findings indicate that LINC01410 could be a promising therapeutic approach for the prevention or alleviation of PE, a condition for which definitive targeted therapies are currently lacking.
This study is subject to several constraints. Notably, the putative interactions between METTL3 and FAS, as well as between FAS and NF-κB, remain unconfirmed. Furthermore, the investigation was confined to in vitro analyses, underscoring the imperative to develop an animal model to elucidate the function of LINC01410 in PE comprehensively. An additional limitation is the exclusive utilization of a single cell line, HTR-8/Svneo, for the experimental procedures. Consequently, the insights garnered from this research necessitate further corroboration across diverse cell lines and through in vivo studies to fortify their applicability and relevance.
Conclusion
Elevated LINC01410 expression accelerates trophoblast cell invasiveness by downregulating METTL3, leading to decreased FAS levels via altered m6A modification, thereby inhibiting the NF-κB pathway. This highlights the critical role of LINC01410 in controlling trophoblast cell invasion, suggesting that LINC01410 could be an effective therapeutic strategy for PE.
Data availability
No datasets were generated or analysed during the current study.
References
Poon LC, Shennan A, Hyett JA, Kapur A, Hadar E, Divakar H, McAuliffe F, da Silva Costa F, von Dadelszen P, McIntyre HD, Kihara AB, Di Renzo GC, Romero R, D’Alton M, Berghella V, Nicolaides KH, Hod M (2019) The International Federation of Gynecology and Obstetrics (FIGO) initiative on pre-eclampsia: a pragmatic guide for first-trimester screening and prevention. Int J Gynaecol Obstet 145(Suppl 1):1–33. https://doi.org/10.1002/ijgo.12802
Liu YH, Zhang YS, Chen JY, Wang ZJ, Liu YX, Li JQ, Xu XJ, Xie NJ, Lye S, Tan N, Duan CY, Wei YX, He PC (2023) Comparative effectiveness of prophylactic strategies for preeclampsia: a network meta-analysis of randomized controlled trials. Am J Obstet Gynecol 228:535–546. https://doi.org/10.1016/j.ajog.2022.10.014
Miller EC, Wilczek A, Bello NA, Tom S, Wapner R, Suh Y (2022) Pregnancy, preeclampsia and maternal aging: from epidemiology to functional genomics. Ageing Res Rev 73:101535. https://doi.org/10.1016/j.arr.2021.101535
Arutyunyan A, Roberts K, Troule K, Wong FCK, Sheridan MA, Kats I, Garcia-Alonso L, Velten B, Hoo R, Ruiz-Morales ER, Sancho-Serra C, Shilts J, Handfield LF, Marconato L, Tuck E, Gardner L, Mazzeo CI, Li Q, Kelava I, Wright GJ, Prigmore E, Teichmann SA, Bayraktar OA, Moffett A, Stegle O, Turco MY, Vento-Tormo R (2023) Spatial multiomics map of trophoblast development in early pregnancy. Nature 616:143–151. https://doi.org/10.1038/s41586-023-05869-0
Rayburn WF (2020) The placenta: its importance from womb to Tomb. Obstet Gynecol Clin North Am 47:xiii–xiv. https://doi.org/10.1016/j.ogc.2020.01.002
Jie Q, Chen L, Liang J, Yang X, Sun F, Ma Y (2023) Downregulated ETV4 inhibits the proliferation, migration, and invasion of trophoblast cells in preeclampsia. Reproduction 165:373–381. https://doi.org/10.1530/REP-22-0184
Wang H, Feng Y, Zheng X, Xu X (2023) The diagnostic and therapeutic role of snoRNA and lincRNA in bladder Cancer. Cancers (Basel) 15. https://doi.org/10.3390/cancers15041007
Lekva T, Sundaram AYF, Roland MCP, Asheim J, Michelsen AE, Norwitz ER, Aukrust P, Gilfillan GD, Ueland T (2023) Platelet and mitochondrial RNA is decreased in plasma-derived extracellular vesicles in women with preeclampsia-an exploratory study. BMC Med 21:458. https://doi.org/10.1186/s12916-023-03178-x
Jiang S, Chen Q, Liu H, Gao Y, Yang X, Ren Z, Gao Y, Xiao L, Zhong M, Yu Y, Yang X (2020) Preeclampsia-Associated lncRNA INHBA-AS1 regulates the Proliferation, Invasion, and Migration of placental trophoblast cells. Mol Ther Nucleic Acids 22:684–695. https://doi.org/10.1016/j.omtn.2020.09.033
Chen Q, Jiang S, Liu H, Gao Y, Yang X, Ren Z, Gao Y, Xiao L, Hu H, Yu Y, Yang X, Zhong M (2020) Association of lncRNA SH3PXD2A-AS1 with preeclampsia and its function in invasion and migration of placental trophoblast cells. Cell Death Dis 11:583. https://doi.org/10.1038/s41419-020-02796-0
Sendinc E, Shi Y (2023) RNA m6A methylation across the transcriptome. Mol Cell 83:428–441. https://doi.org/10.1016/j.molcel.2023.01.006
Gu J, Cao H, Chen X, Zhang XD, Thorne RF, Liu X (2024) RNA m6A modifications regulate crosstalk between tumor metabolism and immunity. Wiley Interdiscip Rev RNA 15:e1829. https://doi.org/10.1002/wrna.1829
Fang Z, Mei W, Qu C, Lu J, Shang L, Cao F, Li F (2022) Role of m6A writers, erasers and readers in cancer. Exp Hematol Oncol 11:45. https://doi.org/10.1186/s40164-022-00298-7
Jin Y, Fan Z (2024) New insights into the interaction between m6A modification and lncRNA in cancer drug resistance. Cell Prolif 57:e13578. https://doi.org/10.1111/cpr.13578
Chen S, Dong J, Luo X, Nie Z, Lu S, Liu H, Liu J (2022) Interaction between m6A and ncRNAs and its Association with diseases. Cytogenet Genome Res 162:171–187. https://doi.org/10.1159/000526035
Xie J, Zhang H, Wang K, Ni J, Ma X, Khoury CJ, Prifti V, Hoard B, Cerenzia EG, Yin L, Zhang H, Wang R, Zhuo D, Mao W, Peng B (2023) M6A-mediated-upregulation of lncRNA BLACAT3 promotes bladder cancer angiogenesis and hematogenous metastasis through YBX3 nuclear shuttling and enhancing NCF2 transcription. Oncogene 42:2956–2970. https://doi.org/10.1038/s41388-023-02814-3
Du A, Li S, Zhou Y, Disoma C, Liao Y, Zhang Y, Chen Z, Yang Q, Liu P, Liu S, Dong Z, Razzaq A, Tao S, Chen X, Liu Y, Xu L, Zhang Q, Li S, Peng J, Xia Z (2022) M6A-mediated upregulation of circMDK promotes tumorigenesis and acts as a nanotherapeutic target in hepatocellular carcinoma. Mol Cancer 21:109. https://doi.org/10.1186/s12943-022-01575-z
Saleh AA, Elghobashy YA, Kasemy ZA, Hegazy A, AA AL (2024) Impact of Dysregulated LINC01559 and LINC01410 expression on the diagnosis and survival of Non-small Cell Lung Cancer. Biochem Genet. https://doi.org/10.1007/s10528-023-10632-1
Mou L, Wang L, Zhang S, Wang Q (2021) Long noncoding RNA LINC01410 suppresses tumorigenesis and enhances radiosensitivity in Neuroblastoma cells through regulating miR-545-3p/HK2 Axis. Onco Targets Ther 14:3225–3238. https://doi.org/10.2147/OTT.S297969
Fu T, Yang Y, Mu Z, Sun R, Li X, Dong J (2021) Silencing lncRNA LINC01410 suppresses cell viability yet promotes apoptosis and sensitivity to temozolomide in glioblastoma cells by inactivating PTEN/AKT pathway via targeting miR-370-3p. Immunopharmacol Immunotoxicol 43:680–692. https://doi.org/10.1080/08923973.2021.1966031
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Yang Y, Lan RH, Gong HMJ海英 (2021) Clinical significance of lncRNA LINC01410 in preeclampsia and its effect on trophoblast proliferation and invasion. 27:5
Ayala-Ramirez P, Machuca-Acevedo C, Gamez T, Quijano S, Barreto A, Silva JL, Olaya CM, Garcia-Robles R (2021) Assessment of placental extracellular vesicles-Associated Fas ligand and TNF-Related apoptosis-inducing Ligand in pregnancies complicated by early and late Onset Preeclampsia. Front Physiol 12:708824. https://doi.org/10.3389/fphys.2021.708824
Hasan S, Alshaikh B, Yusuf K (2021) Serum levels of soluble Fas and Fas ligand in pregnant women who smoke. Am J Reprod Immunol 85:e13382. https://doi.org/10.1111/aji.13382
Tan S, Liu X, Chen L, Wu X, Tao L, Pan X, Tan S, Liu H, Jiang J, Wu B (2021) Fas/FasL mediates NF-kappaBp65/PUMA-modulated hepatocytes apoptosis via autophagy to drive liver fibrosis. Cell Death Dis 12:474. https://doi.org/10.1038/s41419-021-03749-x
Jung E, Romero R, Yeo L, Gomez-Lopez N, Chaemsaithong P, Jaovisidha A, Gotsch F, Erez O (2022) The etiology of preeclampsia. Am J Obstet Gynecol 226:S844–S866. https://doi.org/10.1016/j.ajog.2021.11.1356
Ma’ayeh M, Costantine MM (2020) Prevention of preeclampsia. Semin Fetal Neonatal Med 25:101123. https://doi.org/10.1016/j.siny.2020.101123
Hauspurg A, Jeyabalan A (2022) Postpartum preeclampsia or eclampsia: defining its place and management among the hypertensive disorders of pregnancy. Am J Obstet Gynecol 226:S1211–S1221. https://doi.org/10.1016/j.ajog.2020.10.027
Jia X, Yang S, Wang X, Ruan J, Huang W (2023) HOXB3 promotes trophoblast cell proliferation, invasion, and migration to alleviate preeclampsia via mediating the Notch/Wnt/beta-catenin pathway. Eur J Pharmacol 960:176015. https://doi.org/10.1016/j.ejphar.2023.176015
Ma W, Zhao X, Xue N, Gao Y, Xu Q (2021) The LINC01410/miR-122-5p/NDRG3 axis is involved in the proliferation and migration of osteosarcoma cells. IUBMB Life 73:705–717. https://doi.org/10.1002/iub.2452
Guo W, Gai Q, Ma Y, Shan Z, Wu J (2021) LINC01410 leads the migration, invasion and EMT of bladder cancer cells by modulating miR-4319 / Snail1. Cancer Cell Int 21:429. https://doi.org/10.1186/s12935-021-02119-z
Ma W, Gao Y, Zhang J, Yao X, Jia L, Xu Q (2021) Long noncoding RNA LINC01410 promotes tumorigenesis of osteosarcoma cells via miR-497-5p/HMGA2 axis. J Biochem Mol Toxicol 35:e22921. https://doi.org/10.1002/jbt.22921
Liu F, Wen C (2020) LINC01410 Knockdown suppresses cervical Cancer Growth and Invasion via Targeting miR-2467-3p/VOPP1 Axis. Cancer Manag Res 12:855–861. https://doi.org/10.2147/CMAR.S236832
Xu Q, He L, Ma L, Fan L, Yan L, Zhao X, Li Y (2020) LINC01410 accelerated the invasion and proliferation of osteosarcoma by sponging miR-3128. Aging 12:24957–24966. https://doi.org/10.18632/aging.103464
Jiang T, Wang C, Zhu Y, Han H (2020) LINC01410 promotes cell proliferation and migration of cholangiocarcinoma through modulating miR-124-3p/SMAD5 axis. J Gene Med 22:e3162. https://doi.org/10.1002/jgm.3162
Xu J, Wang L, Wang Q (2021) High expression of long noncoding RNA 01410 serves as a potential diagnostic and prognostic marker in patients with colorectal Cancer. Clin Lab 67. https://doi.org/10.7754/Clin.Lab.2020.200805
Musso T, Calosso L, Zucca M, Millesimo M, Ravarino D, Giovarelli M, Malavasi F, Ponzi AN, Paus R, Bulfone-Paus S (1999) Human monocytes constitutively express membrane-bound, biologically active, and interferon-gamma-upregulated interleukin-15. Blood 93:3531–3539
Zhang Y, Yang H, Long Y, Zhang Y, Chen R, Shi J, Chen J (2021) circRNA N6-methyladenosine methylation in preeclampsia and the potential role of N6-methyladenosine-modified circPAPPA2 in trophoblast invasion. Sci Rep 11:24357. https://doi.org/10.1038/s41598-021-03662-5
Gu Y, Chu X, Morgan JA, Lewis DF, Wang Y (2021) Upregulation of METTL3 expression and m6A RNA methylation in placental trophoblasts in preeclampsia. Placenta 103:43–49. https://doi.org/10.1016/j.placenta.2020.10.016
Guo Y, Song W, Yang Y (2022) Inhibition of ALKBH5-mediated m(6) a modification of PPARG mRNA alleviates H/R-induced oxidative stress and apoptosis in placenta trophoblast. Environ Toxicol 37:910–924. https://doi.org/10.1002/tox.23454
Li XC, Jin F, Wang BY, Yin XJ, Hong W, Tian FJ (2019) The m6A demethylase ALKBH5 controls trophoblast invasion at the maternal-fetal interface by regulating the stability of CYR61 mRNA. Theranostics 9:3853–3865. https://doi.org/10.7150/thno.31868
Zeng C, Huang W, Li Y, Weng H (2020) Roles of METTL3 in cancer: mechanisms and therapeutic targeting. J Hematol Oncol 13:117. https://doi.org/10.1186/s13045-020-00951-w
Wan W, Ao X, Chen Q, Yu Y, Ao L, Xing W, Guo W, Wu X, Pu C, Hu X, Li Z, Yao M, Luo D, Xu X (2022) METTL3/IGF2BP3 axis inhibits tumor immune surveillance by upregulating N(6)-methyladenosine modification of PD-L1 mRNA in breast cancer. Mol Cancer 21:60. https://doi.org/10.1186/s12943-021-01447-y
Wang L, Yang Q, Zhou Q, Fang F, Lei K, Liu Z, Zheng G, Zhu L, Huo J, Li X, Peng S, Kuang M, Lin S, Huang M, Xu L (2023) METTL3-m(6)A-EGFR-axis drives lenvatinib resistance in hepatocellular carcinoma. Cancer Lett 559:216122. https://doi.org/10.1016/j.canlet.2023.216122
Su X, Qu Y, Mu D (2023) The Regulatory Network of METTL3 in the nervous system: diagnostic biomarkers and therapeutic targets. Biomolecules 13. https://doi.org/10.3390/biom13040664
Zhang B, Jiang H, Dong Z, Sun A, Ge J (2021) The critical roles of m6A modification in metabolic abnormality and cardiovascular diseases. Genes Dis 8:746–758. https://doi.org/10.1016/j.gendis.2020.07.011
Wang Y, Wang Y, Gu J, Su T, Gu X, Feng Y (2022) The role of RNA m6A methylation in lipid metabolism. Front Endocrinol (Lausanne) 13:866116. https://doi.org/10.3389/fendo.2022.866116
Blaes J, Thome CM, Pfenning PN, Rubmann P, Sahm F, Wick A, Bunse T, Schmenger T, Sykora J, von Deimling A, Wiestler B, Merz C, Jugold M, Haberkorn U, Abdollahi A, Debus J, Gieffers C, Kunz C, Bendszus M, Kluge M, Platten M, Fricke H, Wick W, Lemke D (2018) Inhibition of CD95/CD95L (FAS/FASLG) signaling with APG101 prevents Invasion and enhances Radiation Therapy for Glioblastoma. Mol Cancer Res 16:767–776. https://doi.org/10.1158/1541-7786.MCR-17-0563
Magerus A, Bercher-Brayer C, Rieux-Laucat F (2021) The genetic landscape of the FAS pathway deficiencies. Biomed J 44:388–399. https://doi.org/10.1016/j.bj.2021.06.005
Guegan JP, Ginestier C, Charafe-Jauffret E, Ducret T, Quignard JF, Vacher P, Legembre P (2020) CD95/Fas and metastatic disease: what does not kill you makes you stronger. Semin Cancer Biol 60:121–131. https://doi.org/10.1016/j.semcancer.2019.06.004
Lan R, Yang Y, Song J, Wang L, Gong H (2021) Fas regulates the apoptosis and migration of trophoblast cells by targeting NF-kappaB. Exp Ther Med 22:1055. https://doi.org/10.3892/etm.2021.10489
Khan D, Cornelius JF, Muhammad S (2023) The role of NF-kappaB in intracranial aneurysm pathogenesis: a systematic review. Int J Mol Sci 24. https://doi.org/10.3390/ijms241814218
Rastogi S, Aldosary S, Saeedan AS, Ansari MN, Singh M, Kaithwas G (2023) NF-kappaB mediated regulation of tumor cell proliferation in hypoxic microenvironment. Front Pharmacol 14:1108915. https://doi.org/10.3389/fphar.2023.1108915
Zheng L, Shi L, Zhou Z, Chen X, Wang L, Lu Z, Tang R (2018) Placental expression of AChE, alpha7nAChR and NF-kappaB in patients with preeclampsia. Ginekol Pol 89:249–255. https://doi.org/10.5603/GP.a2018.0043
Acknowledgements
Not applicable.
Funding
This study was supported by grants from General Project of the Natural Science Foundation of Hainan Province (821MS123, 821MS124) and Key R&D Projects in Hainan Province (ZDYF2024SHFZ093).
Author information
Authors and Affiliations
Contributions
RL and HG performed the analyses and participated in the study design. HG wrote the manuscript. YY, MC and RL confirm the authenticity of all the raw data. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical approval
This study did not require ethical board approval because it did not perform human or animal trials.
Patient consent for publication
Not applicable for no human trials.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, Y., Chen, M., Lan, R. et al. LINC01410 accelerates the invasion of trophoblast cells by modulating METTL3/Fas. Mol Biol Rep 51, 895 (2024). https://doi.org/10.1007/s11033-024-09834-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11033-024-09834-6