Abstract
The omega-3 polyunsaturated fatty acid, docosahexaenoic acid (DHA) is enriched in neural membranes of the CNS, and recent studies have shown a role of DHA metabolism by 15-lipoxygenase-1 (Alox15) in prefrontal cortex resolvin D1 formation, hippocampo-prefrontal cortical long-term-potentiation, spatial working memory, and anti-nociception/anxiety. In this study, we elucidated epigenetic regulation of Alox15 via histone modifications in neuron-like cells. Treatment of undifferentiated SH-SY5Y human neuroblastoma cells with the histone deacetylase (HDAC) inhibitors trichostatin A (TSA) and sodium butyrate significantly increased Alox15 mRNA expression. Moreover, Alox15 expression was markedly upregulated by Class I HDAC inhibitors, MS-275 and depsipeptide. Co-treatment of undifferentiated SH-SY5Y cells with the p300 histone acetyltransferase (HAT) inhibitor C646 and TSA or sodium butyrate showed that p300 HAT inhibition modulated TSA or sodium butyrate-induced Alox15 upregulation. Differentiation of SH-SY5Y cells with retinoic acid resulted in increased neurite outgrowth and Alox15 mRNA expression, while co-treatment with the p300 HAT inhibitor C646 and retinoic acid modulated the increases, indicating a role of p300 HAT in differentiation-associated Alox15 upregulation. Increasing Alox15 expression was found in primary murine cortical neurons during development from 3 to 10 days-in-vitro, reaching high levels of expression by 10 days-in-vitro—when Alox15 was not further upregulated by HDAC inhibition. Together, results indicate regulation of Alox15 mRNA expression in neuroblastoma cells by histone modifications, and increasing Alox15 expression in differentiating neurons. It is possible that one of the environmental influences on the immature brain that can affect cognition and memory, may take the form of epigenetic effects on Alox15 and metabolites of DHA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
There has been much recent interest in the omega-3 polyunsaturated fatty acid (PUFA) docosahexaenoic acid (DHA) due to its reported anti-inflammatory [1] and neuroprotective roles [2], as well as effects on cognition and behavior [3, 4]. Many studies have also demonstrated beneficial effects of DHA in learning and memory [5,6,7]. Lipoxygenases are enzymes that act on PUFAs including DHA, to generate hydroxyperoxide products that act as signaling molecules or induce structural or physiological changes in the cell. DHA is the main n-3 fatty acid in neuronal membranes [8], and is metabolized by 15-lipoxygenase (Alox15) [9]. The latter produces pro- and anti-inflammatory mediators depending on substrates available. Arachidonic acid released from membrane glycerophospholipids by cytosolic phospholipase A2 (cPLA2) is metabolized by Alox15 to produce 15S-hydroxyeicosatetraenoic acid (15S-HETE) which has pro-inflammatory properties [10]. At the same time 15S-HETE is an endogenous ligand for peroxisome proliferator-activated receptor gamma (PPARγ), which has anti-inflammatory effects [11]. Alox15 can also metabolize DHA to produce resolvin D1 and neuroprotection D1, which have anti-inflammatory actions [12, 13]. Resolvin D1 modulates the infiltration of neutrophils [14] and reduces inflammation by inhibiting production of the pro-inflammatory cytokine, interleukin-1β (IL-1β) [1]. Recently, Alox15 has been found to have effects on synaptic plasticity. Antisense knockdown of the enzyme in the rat prefrontal cortex results in reduced levels of resolvin D1 and deficits in hippocampo-prefrontal cortical long-term-potentiation (LTP) and spatial working memory [15]. Alox15 inhibition in the prefrontal cortex reduces the anti-nociceptive effect of a DHA-rich nutraceutical, in a mouse model of neuropathic orofacial pain, suggesting a role of the enzyme in supraspinal antinociception [16]. At the spinal cord level, resolvin D1 reduces inflammatory pain in mice by normalizing aberrant spinal synaptic plasticity involved in pain hypersensitivity [17].
The 5′-flanking promoter region of the human Alox15 gene contains putative binding sites for transcription factors including signal transducer and activator of transcription 6 (STAT6), activator protein 2 (AP-2), GATA, nuclear factor 1 (NF-1), and P-1 [18]. Transcriptional activation of Alox15 by interleukin-4 (IL-4) has been reported - in monocytes [19], endothelial cells [18] and lung epithelial cells [20]. IL-4 induction of Alox15 expression occurs via the STAT6 pathway [21]. Activation of the cyclic GMP/protein kinase G pathway also induces Alox15 expression in human colon cancer cells [22]. Epigenetic modification of the Alox15 gene by histone deacetylases (HDACs) is an additional mechanism for regulating its expression [23, 24]. HDACs remove acetyl groups from lysine residues on histones, promoting a condensed and transcriptionally-inaccessible form of chromatin (heterochromatin). In contrast, histone acetyltransferases (HATs) add acetyl groups to histone lysine residues to promote a more accessible conformation of chromatin, and facilitate gene transcription.
To date, there is limited data regarding regulation of Alox15 in neuronal cells. In this study, we elucidate epigenetic regulation of Alox15 in SH-SY5Y human neuroblastoma cells using HDAC and HAT inhibitors. Results indicate expression of DHA-metabolizing enzyme Alox15 is regulated by selective histone acetylation in these cells.
Materials and Methods
Chemicals
All reagents were of analytical grade and used as received without further purification. Dimethyl sulfoxide (DMSO), trichostatin A (TSA), tubacin, C646 and retinoic acid (RA) were purchased from Sigma-Aldrich (St. Louis, MO, USA). Sodium butyrate (NaBT), MS-275 and butyrolactone-3 (MB-3) were purchased from Santa Cruz Biotechnology (Santa Cruz, California, USA). Depsipeptide was purchased from ApexBio (ApexBio, Texas, USA). NU9056 was purchased from Tocris Bioscience (Tocris Bioscience, Bristol, UK). TMP 195 was purchased from Axon Medchem (Axon Medchem, Groningen, Netherlands) and PD146176 was obtained from Cayman Chemical (Cayman Chemical, MI, USA). Stock solutions were prepared in DMSO and further diluted in cell culture medium for use. The equivalent concentration of DMSO was used as vehicle controls, with final concentrations of 0.1% or less, in cell culture media.
Cell Cultures
Fully differentiated primary cortical neurons were prepared from E15.5 mouse embryos and cultured in Neurobasal plus B-27 medium (Gibco, NY, USA) containing 2 mM l-glutamine, 10 μM glutamate, 1.6% fetal bovine serum (FBS), 0.4% horse serum and penicillin/streptomycin. On days in vitro (DIV) 3, cells were treated with 1 µM cytosine arabinoside (Ara-C) for 72 h to prevent glial proliferation and maintained in serum-free Neurobasal plus B-27 medium at 37 °C in a humidified 5% CO2 incubator. Experiments were conducted on DIV 10–14 [25].
SH-SY5Y human neuroblastoma cells (CRL-2266) were obtained from American Type Culture Collection (ATCC®). SH-SY5Y cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM) containing 10% heat-inactivated FBS and 1% penicillin/streptomycin (Gibco-Invitrogen, CA, USA). SH-SY5Y cells were maintained in 100 mm dishes (Greiner Bio-One, USA) and incubated under standard conditions of 37 °C and 5% CO2. They were regularly passaged using 25% Trypsin–EDTA (Gibco, NY, USA).
Differentiation of SH-SY5Y cells was carried out by seeding cells in 100 mm culture dishes and cell growth was constantly monitored up to approximately 50–60% confluency. RA was then added at a final concentration of 10 μM in DMEM with 10% FBS.
Effect of HDAC and HAT Inhibitors on Alox15 mRNA Expression in Primary Murine Cortical Neurons
Treatment of Primary Cortical Neurons with Various HAT Inhibitors and TSA
TSA is a pan-HDAC inhibitor that blocks the activity of class I and II HDACs [26]. C646 is a HAT inhibitor that inhibits p300 HAT specifically [27], whereas NU9056 is a Tip60 HAT-specific inhibitor [28]. To examine the effects of HDAC and HAT inhibitors on Alox15 expression, primary murine neurons were co-treated with C646 or NU9056 and 0.5 μM TSA for 24 h. The neurons were treated as follows: (1) DMSO as vehicle control, (2) 0.5 µM TSA, (3) 20 μM C646 and 0.5 µM TSA, or (4) 20 μM NU9056 and 0.5 µM TSA.
Effect of Alox15 Inhibitor, PD146176, on MAP-2, NeuN, α-Tubulin and Alox15 mRNA Expression in Primary Murine Cortical Neurons
Treatment of Primary Cortical Neurons with PD146176
Primary murine neurons were cultured and harvested at DIV 3, 5 and 10, and expression of neuronal markers MAP-2, NeuN and α-tubulin, as well as Alox15 determined at these times. To examine the effects of Alox15 on neuronal markers, primary murine neurons were treated with the specific Alox15 inhibitor, PD146176 (1 µM). Protein expression of MAP-2, NeuN, Alox15, α-tubulin and β-actin were determined by Western blots.
Semi-quantitative RT-PCR Analyses
Total RNA from primary murine neurons (1 μg) was incubated with 200 units of MMLV reverse transcriptase (RTase, Clontech; Palo Alto, CA) in a buffer containing a final concentration of 50 mM Tris–Cl (pH 8.3), 75 mM KCl, 3 mM MgCl2, 20 units of RNase inhibitor, 1 μM polydT oligomer, and 0.5 mM of each dNTP in a final volume of 20 μl. The reaction mixture was incubated at 42 °C for 1 h and then 94 °C for 5 min to inactive the enzyme. A total of 80 μl DEPC treated water was added to the reaction mixture before storage at − 70 °C. 5 μl of the RT reaction solution was used in the PCR reaction. PCR was carried out in a 50 μl final volume containing 200 μM each of dATP, dCTP, dGTP, and dTTP, 5 pmole of each primer, 1.25 units of Tag polymerase (BRL; Gaithersburg, MD), 20 mM Tris–Cl (pH 8.4), 1.5 mM MgCl2, and 50 mM KCl. The mixture was incubated in a thermal cycler for 35 cycles using the following profile: 94 °C for 7 min, then repeat cycles of 94 °C for 45 s; 55 °C for 45 s; 72 °C for 90 s. Samples were then incubated at 72 °C for 7 min and cooled to 4 °C (GeneAmp 2400, PE; Norwalk, CT). PCR products were run on 2% agarose gel for DNA fragment size verification. Primers used were: 5′-CAGGGATCGGAGTACACGTT and 5′-GATTGTGCCATCCTTCCAGT for Alox15 [29]; and 5′-TCCCTCAAGATTGTCAG and 5′-AGATCCACAACGGATAC for GAPDH. Statistical differences were analyzed using one-way ANOVA with post-hoc Fisher’s protected t test, where p < 0.05 was considered significant.
Western Blot Analyses
Western blot analysis of proteins was performed as previously described [30], with mouse monoclonal antibody to Alox15 (1:500, Abcam, New Territories, HK) and antibodies to MAP-2, NeuN or α-tubulin (1:500, Santa Cruz, CA, USA). Protein bands were visualized by an enhanced chemiluminescence system (Merck Millipore, MA, USA). Statistical differences were analyzed using one-way ANOVA with post-hoc Fisher’s protected t test, or Student’s t test, where p < 0.05 was considered significant.
Effect of HDAC and/or HAT Inhibitors on SH-SY5Y Cells
Experiments on Undifferentiated SH-SY5Y Cells
Treatment of Undifferentiated SH-SY5Y Cells with Histone Deacetylase (HDAC) Inhibitors
Undifferentiated SH-SY5Y cells were treated with general HDAC inhibitors TSA (0.5 µM) and sodium butyrate (5 mM), and specific HDAC inhibitors, MS-275 (5 µM) and depsipeptide (10 nM), for 24 h. Sodium butyrate is a general HDAC inhibitor that blocks the activity of class I and II HDACs [31] and is effective in the millimolar range [32]. MS-275 inhibits HDACs 1, 2 and 3 activities, and depsipeptide is an inhibitor of HDACs 1 and 2 [33]. MS-275 inhibits HDAC activity at the micromolar range [34, 35] and depsipeptide is effective at the nanomolar range [24]. TMP 195 is a Class IIa specific HDAC, and is effective in the nanomolar range [36]. Each group consisted of four biological replicates.
Treatment of Undifferentiated SH-SY5Y Cells with HAT Inhibitors and TSA
To investigate the effect of HATs on TSA-induced Alox15 expression, SH-SY5Y cells were treated individually or in combination with 0.5 μM TSA using inhibitors from three different classes of HATs: GCNS5 HAT (MB-3, 20 µM), p300 HAT (C646, 20 µM) and Tip60 HAT (NU9056, 20 µM). The treatments were carried out in four groups: (1) DMSO as vehicle control, (2) 0.5 µM TSA, (3) HAT inhibitor, and (4) HAT inhibitor and 0.5 µM TSA. Cells were co-treated with HAT inhibitors and TSA or vehicle for 24 h.
Treatment of Undifferentiated SH-SY5Y Cells with HAT Inhibitors and Sodium Butyrate
To investigate the effect of HATs on sodium butyrate-induced Alox15 expression, SH-SY5Y cells were treated individually or in combination with sodium butyrate using inhibitors of p300 HAT (C646, 20 µM) or Tip60 HAT (NU9056, 20 µM). The treatments were carried out in four groups: (1) DMSO as vehicle control, (2) 5 mM sodium butyrate, (3) HAT inhibitor, and (4) HAT inhibitor and 5 mM sodium butyrate. Cells were co-treated with HAT inhibitors and sodium butyrate or vehicle for 24 h.
Treatment of Undifferentiated SH-SY5Y Cells with HAT Inhibitors and MS-275
To investigate the effect of HATs on MS-275-induced Alox15 expression, SH-SY5Y cells were treated individually or in combination with MS-275 using inhibitors of p300 HAT (C646, 20 µM) or Tip60 HAT (NU9056, 20 µM). The treatments were carried out in four groups: (1) DMSO as vehicle control, (2) 5 μM MS-275, (3) HAT inhibitor, and (4) HAT inhibitor and 5 μM MS-275. Cells were co-treated with HAT inhibitors and MS-275 or vehicle for 24 h.
Treatment of Undifferentiated SH-SY5Y Cells with HAT Inhibitors and Depsipeptide
To investigate the effect of HATs on depsipeptide-induced Alox15 expression, SH-SY5Y cells were treated individually or in combination with depsipeptide using inhibitors of p300 HAT (C646, 20 µM) or Tip60 HAT (NU9056, 20 µM). The treatments were carried out in four groups: (1) DMSO as vehicle control, (2) 10 nM depsipeptide, (3) HAT inhibitor, and (4) HAT inhibitor and 10 nM depsipeptide. Cells were co-treated with HAT inhibitors and depsipeptide or vehicle for 24 h.
Experiments on Retinoic Acid Treated SH-SY5Y Cells
Treatment of SH-SY5Y Cells with Retinoic Acid and HAT Inhibitors
To examine the effects of HAT inhibitors on Alox15 expression during RA-induced differentiation of SH-SY5Y cells, undifferentiated SH-SY5Y cells were incubated with RA and C646 or NU9056 for 72 h. SH-SY5Y cells were treated as follows: (1) DMSO as vehicle control, (2) 20 μM HAT inhibitor, (3) 10 μM RA, and (4) 10 μM RA and 20 μM HAT inhibitor. The cells were harvested after 72 h and RNA extracted and analyzed by RT-PCR.
Quantification of Neurite Lengths of SH-SY5Y Cells
Images of SH-SY5Y cells were captured using the EVOS FL Cell Imaging System at ×20 magnification, and neurite lengths quantified using the ImageJ Simple Neurite Tracer software plugin, by tracing observable neurites extending from cell bodies in the images [37]. An overall mean neurite length (µm) of 350 cells was obtained for each treatment group. Possible differences between treatment groups were analyzed using two tailed t test and one-way ANOVA with Bonferroni’s multiple comparison post-hoc test, where p < 0.05 was considered significant.
Real-Time RT-PCR
Total RNA from SH-SY5Y cells was extracted with the RNeasy Mini kit (Qiagen, Hamburg, Germany). Reverse transcription of RNA to complementary DNA (cDNA) was performed as previously described [38]. The cDNA obtained was quantified by real-time PCR (RT-PCR) using the TaqMan® Universal PCR Master Mix (Applied Biosystems, CA, USA), with TaqMan® Gene Expression Assay Probes for Alox15 (Hs00609608_m1), NSE (Hs00157360_m1), β-III-tubulin (Hs00801390_s1), and β-actin (#4326315E) (Applied Biosystems, CA, USA). Real-time PCR was performed and the relative fold changes were quantified using the 2−ΔΔCT method as previously described [38]. All reactions were performed in triplicates and the mean and standard error calculated. Statistical differences were analyzed using two tailed t test and one-way ANOVA with Bonferroni’s multiple comparison post-hoc test, where p < 0.05 was considered significant.
Immunocytochemistry
SH-SY5Y cells were plated at a density of 2 × 105 cells on poly-l-lysine-coated coverslips placed in 24-well plates and grown to 80% confluency before administration of treatment. After treatment, the cells were fixed before antigen retrieval and permeabilization. They were then blocked in 1% BSA and incubated overnight at 4 °C with Alox15 specific antibody (ab119774, 1:50 in blocking buffer, Abcam, New Territories, HK), followed by secondary incubation with anti-mouse Alexa Fluor 488 (Applied Biosystems, CA, USA; diluted 1:100 in 1% BSA) for 1 h at room temperature. Nuclei were labelled and coverslips mounted with the Prolong Gold Antifade Mountant DAPI (Invitrogen, USA). Samples were analyzed and images captured using a confocal microscope (Zeiss, OR, USA).
Quantitative Image Analysis
The fluorescence intensity of cells were measured as previously described [38]. The net fluorescence intensity was calculated for each image according to the following formula: Corrected total cell fluorescence (CTCF) = Integrated Density − (Area of selected cell × Mean fluorescence of background readings). The mean and standard error were calculated, and significant differences were analyzed using one-way ANOVA with Bonferroni’s multiple comparison post-hoc test, where p < 0.05 was considered significant.
Lactate Dehydrogenase (LDH) Assay
SH-SY5Y cells were plated at a density of 2 × 105 cells in 24-well plates, and grown to 80% confluency before administration of treatment. The specific Alox15 inhibitor, PD146176, was used for the selective inhibition of Alox15 activity. After 24 h, cell viability was assessed by colorimetric determination of lactate dehydrogenase (LDH) release, using the LDH Cytotoxicity Detection Kit (Roche, Mannheim, Germany). The plate was read at an excitation wavelength of 490 nm on the Tecan Infinite® 200 microplate reader (Tecan Group Ltd., Maennedorf, Switzerland). Percentage cytotoxicity was calculated according to the formula:
The mean cytotoxicity values were then normalized against vehicle-treated controls. The mean and standard errors were calculated, and significant differences analyzed using one-way ANOVA with Bonferroni’s multiple comparison post-hoc test, where p < 0.05 was considered significant.
Results
Basal Alox15 Protein Expression in Primary Murine Cortical Neurons and Undifferentiated SH-SY5Y Cells
Western blot analyses revealed that Alox15 protein was highly expressed in primary murine cortical neurons at day 10 in vitro. In comparison, undifferentiated SH-SY5Y cells expressed very low levels of Alox15 protein (Fig. 1a, b).
Western blot analyses. a Immunoblot of Alox15 protein expression in primary murine cortical neurons and undifferentiated SH-SY5Y neuroblastoma cells at 10 days-in-vitro. b Densitometric analysis of protein band intensity normalized to β-actin. Data were analyzed by Student’s t test. The mean and standard error are indicated. n = 3 in each group. Asterisk indicates significant difference between undifferentiated SH-SY5Y neuroblastoma cells and primary cortical neurons at 10 days-in-vitro, **P < 0.001
Effect of Alox15 on Primary Murine Neurons at 3, 5 and 10 Days-In-Vitro (DIV)
There was an increase in neuronal markers MAP-2, NeuN and α-tubulin with time in culture, consistent with predicted maturation down a neuronal lineage (Fig. 2a). Interestingly, there was also a large increase in Alox15 protein expression with time, consistent with the notion that Alox15 expression increases in mature neurons. Treatment of primary murine neurons with PD146176 did not affect the protein expression of MAP-2, NeuN, α-tubulin or Alox15 itself, indicating the activity of Alox15 does not have a direct effect on maturation of neurons (Fig. 2b).
Effect of Alox15-specific inhibitor PD146176 on primary murine neuronal markers and Alox15 expression during different stages of neuronal development, from DIV3 to DIV10. a Immunoblot of MAP-2, NeuN and Alox15 protein expression in primary cortical neurons of different days in culture. b Densitometric analysis of protein band intensity normalized to β-actin. Data were analyzed by one-way ANOVA with post-hoc Fisher’s protected t test. The mean and standard error are indicated. n = 3 in each group. Asterisk indicates significant differences, ***P < 0.001
Effect of HDAC and HAT Inhibitors on Alox15 mRNA Expression in Primary Murine Cortical Neurons at 10 Days-In-Vitro
No significant differences in Alox15 mRNA and protein expression were found after treatment of primary murine cortical neurons with 0.5 µM TSA nor co-treatment of TSA with 20 µM C646 or NU9056 (data not shown).
Effect of HDAC Inhibitors on Alox15 mRNA Expression in Undifferentiated SH-SY5Y Cells
Significant increases in Alox15 mRNA expression were found after treatment with 0.5 µM TSA (p < 0.001), 5 mM sodium butyrate (p < 0.001), 5 µM MS-275 (p < 0.001) and 10 nM depsipeptide (p < 0.001), compared to vehicle controls (Table 1). In contrast, treatment with TMP 195 showed no increase in Alox15 mRNA expression (Table 1). A small but significant increase in Alox15 mRNA expression (p < 0.05) was detected after treatment with 15 µM tubacin, compared to vehicle controls (Table 1).
Effect of HAT Inhibitors and TSA on Alox15 Expression in Undifferentiated SH-SY5Y Cells
Significant increase in Alox15 mRNA expression was found after treatment with TSA. Co-treatment of 20 μM NU9056 and TSA suppressed the TSA-induced increase in mRNA expression by 67.0% (p < 0.01) (Fig. 3a). Co-treatment with 20 µM C646 suppressed the effect of TSA treatment on Alox15 mRNA expression by 65.7% (p = 0.001) (Fig. 3b), but co-treatment with 20 µM of the GCN5-specific HAT inhibitor MB-3 did not reduce the TSA-induced increase in Alox15 mRNA (data not shown).
Real-time RT-PCR results. a Effect of NU9056 and TSA on Alox15 mRNA expression. b Effect of C646 and TSA on Alox15 mRNA expression. c Effect of MB-3 and TSA on Alox15 mRNA expression. d Effect of NU9056 and sodium butyrate (NaBT) on Alox15 mRNA expression. e Effect of C646 and NaBT on Alox15 mRNA expression. Data were analyzed by one-way ANOVA with Bonferroni’s multiple comparison post-hoc test. The mean and standard error are indicated. n = 4 in each group. Asterisk indicates significant difference compared to vehicle-treated group, ***P < 0.001. Number sign indicates significant difference compared to 0.5 μM TSA, ##P < 0.01, ###P < 0.001
Alox15 immunofluorescence labeling showed localization of Alox15 protein in the cytoplasm. A significant increase in fluorescence intensity was observed after TSA treatment, compared to vehicle controls. Co-treatment of 20 µM NU0956 and 0.5 µM TSA reduced the TSA-induced increase in immunofluorescence (Fig. 4a, b). Co-treatment of 20 µM C646 and 0.5 µM TSA also reduced the TSA-induced increase in immunofluorescence (Fig. 5a, b).
Effect of Tip60-specific HAT inhibitor, NU90546, and TSA. a Quantification of ten high-power microscopic fields from four biological replicates per group, analyzed and computed as average corrected total cell fluorescence (CTCF) values. b Immunofluorescence images of SH-SY5Y cells expressing Alox15 protein after NU9056 and TSA treatment. Scale bar = 20 µm. Data were analyzed by one-way ANOVA with Bonferroni’s multiple comparison post-hoc test. The mean and standard error are indicated. n = 4 in each group. Asterisk indicates significant difference compared to vehicle-treated group, *P < 0.05, **P < 0.01 and ***P < 0.001. Number sign indicates significant difference compared to 0.5 μM TSA, ##P < 0.01, ###P < 0.001
Effect of p300-specific HAT inhibitor, C646, and TSA. a Quantification of ten high-power microscopic fields from four biological replicates per group, analyzed and computed as mean corrected total cell fluorescence (CTCF) values. b Immunofluorescence images of SH-SY5Y cells expressing Alox15 protein after C646 and TSA treatment. Scale bar = 20 µm. Data were analyzed by one-way ANOVA with Bonferroni’s multiple comparison post-hoc test. The mean and standard error are indicated. n = 4 in each group. Asterisk indicates significant difference compared to vehicle-treated group, ***P < 0.001. Number sign indicates significant difference compared to 0.5 μM TSA, ##P < 0.01, ###P < 0.001
Effect of HAT Inhibitors and Sodium Butyrate on Alox15 Expression in Undifferentiated SH-SY5Y Cells
Significant increase in Alox15 mRNA expression was found after treatment with sodium butyrate. Co-treatment of 20 μM NU9056 and sodium butyrate did not reduce the sodium butyrate-induced increase in Alox15 mRNA expression (Fig. 3c). In contrast, co-treatment with 20 μM C646 reduced the sodium butyrate-induced increase in Alox15 mRNA expression by 66.2% (p = 0.004) (Fig. 3d).
Significant increase in Alox15 immunofluorescence intensity was observed in cells after sodium butyrate treatment, compared to vehicle controls. Co-treatment of 20 µM C646 and 5 mM sodium butyrate reduced the sodium butyrate-induced increase in immunofluorescence (p < 0.001) (Fig. 6a, b).
Effect of p300-specific HAT inhibitor C646 and sodium butyrate (NaBT). a Quantification of ten high-power microscopic fields from four biological replicates per group, analyzed and computed as average corrected total cell fluorescence (CTCF) values. b Immunofluorescence images of SH-SY5Y cells expressing Alox15 protein after C646 and sodium butyrate treatment. Scale bar = 20 µm. Data were analyzed by one-way ANOVA with Bonferroni’s multiple comparison post-hoc test. The mean and standard error are indicated. n = 4 in each group. Asterisk indicates significant difference compared to vehicle-treated group, ***P < 0.001. Number sign indicates significant difference compared to 5 mM sodium butyrate, ##P < 0.01, ###P < 0.001
Effect of HAT Inhibitors and MS-275 on Alox15 Expression in Undifferentiated SH-SY5Y Cells
A significant 1127.2-fold increase in Alox15 mRNA expression was found after treatment with 5 μM MS-275 (p < 0.001). Co-treatment of 20 μM NU9056 and MS-275 did not reduce the MS-275 induced increase in Alox15 mRNA expression (Fig. 7a).
Real-time RT-PCR results. a Effect of Tip60-specific HAT inhibitor NU9056 and MS-275 on Alox15 mRNA expression. b Effect of p300-specific HAT inhibitor C646 and MS-275 on Alox15 mRNA expression. c Effect of Tip60-specific HAT inhibitor NU9056 and depsipeptide, on Alox15 mRNA expression. d Effect of p300-specific HAT inhibitor C646 and depsipeptide on Alox15 mRNA expression. Data were analyzed by one-way ANOVA with Bonferroni’s multiple comparison post-hoc test. The mean and standard error are indicated. n = 4 in each group. Asterisk indicates significant difference compared to vehicle-treated group, ***P < 0.001
In another experiment, a significant increase of 577.4-fold in Alox15 mRNA expression was observed after treatment with 5 μM MS-275 (p < 0.001). Co-treatment with 20 μM C646 failed to suppress the MS-275 induced increase in Alox15 mRNA expression (Fig. 7b).
Effect of HAT Inhibitors and Depsipeptide on Alox15 Expression in Undifferentiated SH-SY5Y Cells
A significant increase of 539.3-fold in Alox15 mRNA expression was found after treatment with 10 nM depsipeptide (p < 0.001). Co-treatment with 20 μM NU9056 did not reduce the depsipeptide-induced increase in Alox15 mRNA expression (Fig. 7c).
Likewise, 10 nM depsipeptide significantly increased Alox15 mRNA expression (p < 0.001), and co-treatment with 20 μM C646 failed to suppress the depsipeptide-induced increase in Alox15 mRNA expression (Fig. 7d).
Effect of Alox15 Inhibitor and TSA on Undifferentiated SH-SY5Y Cell Viability
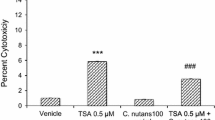
Treatment of SH-SY5Y cells with 0.5 µM TSA caused a significant 10.5-fold in cytotoxicity (p < 0.001). Co-treatment with the specific Alox15 inhibitor PD146176 at a concentration of 1 µM did not significantly reduce the cytotoxicity (Fig. 8).
Effect of Alox15-specific inhibitor PD146176 and TSA on LDH release. Data were analyzed by one-way ANOVA with Bonferroni’s multiple comparison post-hoc test. The mean and standard error are indicated. n = 4 in each group. Asterisk indicates significant difference compared to vehicle-treated group, ***P < 0.001
Effect of RA on SH-SY5Y Cell Morphology, Mean Neurite Length and Neuronal Marker mRNA Expression
Undifferentiated SH-SY5Y cells showed compact morphologies with few neurites from the cell bodies (Fig. 9a). Treatment of these cells with 10 μM RA resulted in increased extension of neurites from cell bodies and reduction in proliferation (Fig. 9b). A significant 2.05-fold increase in neuronal marker NSE mRNA expression was observed after treatment with 10 μM RA, compared with vehicle control (p < 0.001) (Table 2). No significant increase in βIII-tubulin mRNA expression was observed after treatment with 10 μM RA, compared with vehicle control (Table 2).
a, b Effect of RA on SH-SY5Y cell morphology. a Undifferentiated SH-SY5Y cells. b SH-SY5Y cells after treatment with 10 μM RA. Scale bar = 50 μm. c Effect of RA and NU9056 on average neurite length of SH-SY5Y cells. d Effect of RA and C646 on average neurite length of SH-SY5Y cells. e Effect of RA and NU9056 on Alox15 mRNA expression. f Effect of RA and C646 on Alox15 mRNA expression. Data were analyzed by one-way ANOVA with Bonferroni’s multiple comparison post-hoc test. The mean and standard error are indicated. n = 4 in each group. Asterisk indicates significant difference compared to vehicle-treated group, ***P < 0.001. Number sign indicates significant difference compared to group treated with 10 μM RA, ###P < 0.001, #P < 0.05
Significant increase in mean neurite length of SH-SY5Y cells was observed after treatment with 10 μM RA (73.8 ± 1.2 μm) compared to undifferentiated SH-SY5Y cells (22.2 ± 0.2 μm, p < 0.001) (Table 3). Interestingly, a significant 2.9-fold increase in Alox15 mRNA expression was also observed after treatment with 10 μM RA, compared to vehicle control (p < 0.01) (Table 3).
Effect of RA and HAT Inhibitor on Mean Neurite Length and Alox15 mRNA Expression in SH-SY5Y Cells
Significant increase in mean neurite length of SH-SY5Y cells was observed after treatment with 10 μM RA, where mean neurite length was 76.8 ± 1.2 μm, as compared to undifferentiated SH-SY5Y cells where mean neurite length was 20.5 ± 0.1 μm (p < 0.001) (Fig. 9c). Co-treatment with 10 μM RA and 20 μM NU9056 did not result in significant changes in the increase in mean neurite length (Fig. 9c).
In another experiment, significant increase in mean neurite length of SH-SY5Y cells was observed after treatment with 10 μM RA, where mean neurite length was 81.5 ± 1.4 μm, as compared to undifferentiated SH-SY5Y cells where mean neurite length was 23.8 ± 0.2 μm (p < 0.001) (Fig. 9d). Co-treatment with 10 μM RA and 20 μM C646 significantly reduced the mean neurite length by 38.9% to 49.8 ± 0.6 µm (p < 0.001) (Fig. 9d).
Significant increase of 4.7-fold in Alox15 mRNA expression was found after treatment with 10 μM RA (p < 0.001) (Fig. 9e). Co-treatment with 10 μM RA and 20 μM NU9056 did not result in significant reduction in Alox15 mRNA expression (Fig. 9e).
In another experiment, significant increase of 6.1-fold in Alox15 mRNA expression was observed after treatment with 10 μM RA (p < 0.001) (Fig. 9f). Co-treatment with 10 μM RA and 20 μM C646 significantly reduced Alox15 mRNA expression by 45.3% (p < 0.05) (Fig. 9f).
Discussion
The present study was carried out to elucidate the effect of histone acetylation on Alox15 expression in SH-SY5Y cells, which were originally derived from a bone tumor [39]. The current literature provides evidence that undifferentiated SH-SY5Y cells continuously proliferate, express immature neuronal markers, and lack mature neuronal markers [40]. Undifferentiated SH-SY5Y cells are considered to have the properties of immature catecholaminergic neurons. In comparison, fully differentiated SH-SY5Y cells express a variety of neuronal markers such as growth-associated protein (GAP-43), NeuN, synaptophysin (SYN), synaptic vesicle protein II (SV2), NSE and MAP-2; but not glial markers such as glial fibrillary acidic protein (GFAP). In further support that differentiated SH-SY5Y cells have many of the characteristics of neurons, removal of BDNF results in cellular apoptosis. This suggests that survival of differentiated SH-SY5Y cells is dependent on trophic factors, similar to mature neurons [41]. Alox15 mRNA expression was significantly increased after treatment with general HDAC inhibitors TSA and sodium butyrate and the Class I HDAC inhibitors MS-275 and depsipeptide, in undifferentiated SH-SY5Y cells. Results indicate a role of class I HDACs such as HDACs 1, 2, 3 and 8 [42] in regulating Alox15 expression in these cells. In comparison, no significant effect was observed after treatment with the class IIa-specific HDAC inhibitor TMP 195, suggesting that class IIa HDACs are not involved. The class IIb HDAC6 inhibitor tubacin did not upregulate Alox15 mRNA expression, as HDAC6 deacetylates tubulin but not histones [43]. Our observations are consistent with previous studies which showed HDAC inhibitor-mediated transcriptional upregulation of Alox15 in colon [23, 24, 44], pancreatic [45] and breast [46] cancer cells. Global H3 and H4 acetylation [24], as well as demethylation of the CpG islands on the 5′ promoter region [47], have been shown to upregulate Alox15 gene expression in colon cancer cells. HDAC inhibitors may work with transcription factors [48] or synergistically with cytokines [49] to regulate gene expression.
A balance between acetylation and deacetylation of histone proteins is important in regulating gene expression [26], and we therefore elucidated a possible role of HATs in regulating Alox15 expression. Treatment with the p300 HAT inhibitor C646 together with the general HDAC inhibitors TSA or sodium butyrate, showed that p300 HAT is regulating Alox15 expression in SH-SY5Y cells. This is consistent with a previous study, which reported a role of p300 HAT in Alox15 expression in colon cancer cells [24]. p300 HAT and its associated protein CREB-binding protein (CBP) interacts with STAT6 [50], and acetylation of histones by p300 HAT allows IL-4-induced-STAT6 to bind to the promoter, facilitating gene transcription of Alox15 [49]. Treatment with NU9056, a specific Tip60 HAT inhibitor reduced the increase in Alox15 mRNA and protein expression induced by TSA, but NU9056 failed to suppress the increase in Alox15 mRNA expression induced by sodium butyrate. The GCN5 HAT inhibitor MB-3 did not inhibit the increase in Alox15 mRNA expression by TSA, suggesting that the GCN5 HAT is not involved in Alox15 gene regulation (data not shown). Results indicate the importance of p300 HAT, and perhaps Tip60 HAT, in increasing Alox15 expression. Both p300 and Tip60 HAT inhibitors, however, failed to suppress increases in Alox15 mRNA expression that are induced by MS-275 or depsipeptide. The reason for this is unknown, but could be due to histone acetylations that are increased by MS-275 and depsipeptide induced HDAC inhibition, but unaffected by p300 or Tip60 HAT inhibition. Treatment with TSA caused a small amount of cytotoxicity in SH-SY5Y cells as demonstrated by LDH released. This is consistent with previous findings that TSA can be detrimental to neuronal cell viability [51]. Co-treatment with the Alox15 inhibitor PD146176 did not reduce cell death induced by TSA, suggesting that Alox15 is not involved in cell damage under these conditions.
Differentiated SH-SY5Y cells possess neuron-like morphology and express neuronal markers, and are more similar to neurons than undifferentiated cells [52]. Treatment of SH-SY5Y cells with 10 μM RA resulted in upregulation of the neuronal marker NSE, which is consistent with a previous report [53]. Retinoic acid-induced differentiation of SH-SY5Y cells has been shown to lead to changes in acetylation status of histone H3 of genes [54], and in this study, we found significant upregulation of Alox15 mRNA expression after treatment with 10 μM RA. Co-treatment with C646 but not NU9056 suppressed the increase in Alox15 expression, indicating a role of p300 HAT in retinoic-differentiation induced Alox15 upregulation. Increasing Alox15 expression was found in murine primary neurons from 3 days to 10 days-in-vitro, reaching high levels of expression by 10 days-in-vitro; and HDAC inhibitors had no effect on increasing Alox15 expression in these neurons at this time. Inhibition of Alox15 did not affect the expression of MAP-2, NeuN, α-tubulin or Alox15 itself. Together, results indicate regulation of Alox15 mRNA expression in neuroblastoma cells by HDACs and HAT, and increasing levels of Alox15 expression with differentiation.
The prenatal or developing brain is highly sensitive to environmental influences and epigenetic modifications, which can affect gene expression in immature neurons [55]. In contrast, the mature brain is less sensitive to such effects [56]. In view of our recent findings of a role of Alox15 in synaptic plasticity [15] and this study, we postulate that one such environmental influence that can affect learning and memory in the developing brain, may take the form of epigenetic effects on Alox15 and metabolites of DHA. Further work is necessary to elucidate the detailed mechanisms of effect of histone acetylation on Alox15 expression.
References
Hong S, Gronert K, Devchand PR, Moussignac RL, Serhan CN (2003) Novel docosatrienes and 17S-resolvins generated from docosahexaenoic acid in murine brain, human blood, and glial cells. Autacoids in anti-inflammation. J Biol Chem 278(17):14677–14687. https://doi.org/10.1074/jbc.M300218200
Belayev L, Khoutorova L, Atkins KD, Bazan NG (2009) Robust docosahexaenoic acid-mediated neuroprotection in a rat model of transient, focal cerebral ischemia. Stroke 40(9):3121–3126. https://doi.org/10.1161/strokeaha.109.555979
Innis SM (2007) Dietary (n-3) fatty acids and brain development. J Nutr 137(4):855–859
Wurtman RJ (2008) Synapse formation and cognitive brain development: effect of docosahexaenoic acid and other dietary constituents. Metabolism 57(Suppl 2):S6–S10. https://doi.org/10.1016/j.metabol.2008.07.007
Stonehouse W, Conlon CA, Podd J, Hill SR, Minihane AM, Haskell C, Kennedy D (2013) DHA supplementation improved both memory and reaction time in healthy young adults: a randomized controlled trial. Am J Clin Nutr 97(5):1134–1143. https://doi.org/10.3945/ajcn.112.053371
Kuratko CN, Barrett EC, Nelson EB, Salem N Jr (2013) The relationship of docosahexaenoic acid (DHA) with learning and behavior in healthy children: a review. Nutrients 5(7):2777–2810. https://doi.org/10.3390/nu5072777
Zicker SC, Jewell DE, Yamka RM, Milgram NW (2012) Evaluation of cognitive learning, memory, psychomotor, immunologic, and retinal functions in healthy puppies fed foods fortified with docosahexaenoic acid-rich fish oil from 8 to 52 weeks of age. J Am Vet Med Assoc 241(5):583–594. https://doi.org/10.2460/javma.241.5.583
Svennerholm L (1968) Distribution and fatty acid composition of phosphoglycerides in normal human brain. J Lipid Res 9(5):570–579
O’Flaherty JT, Hu Y, Wooten RE, Horita DA, Samuel MP, Thomas MJ, Sun H, Edwards IJ (2012) 15-lipoxygenase metabolites of docosahexaenoic acid inhibit prostate cancer cell proliferation and survival. PLoS ONE 7(9):e45480. https://doi.org/10.1371/journal.pone.0045480
Higgs GA, Salmon JA, Spayne JA (1981) The inflammatory effects of hydroperoxy and hydroxy acid products of arachidonate lipoxygenase in rabbit skin. Br J Pharmacol 74(2):429–433
Sun L, Xu YW, Han J, Liang H, Wang N, Cheng Y (2015) 12/15-Lipoxygenase metabolites of arachidonic acid activate PPARgamma: a possible neuroprotective effect in ischemic brain. J Lipid Res 56(3):502–514. https://doi.org/10.1194/jlr.M053058
Serhan CN, Arita M, Hong S, Gotlinger K (2004) Resolvins, docosatrienes, and neuroprotectins, novel omega-3-derived mediators, and their endogenous aspirin-triggered epimers. Lipids 39(11):1125–1132
Bazan NG (2005) Neuroprotectin D1 (NPD1): a DHA-derived mediator that protects brain and retina against cell injury-induced oxidative stress. Brain Pathol 15(2):159–166
Serhan CN, Hong S, Gronert K, Colgan SP, Devchand PR, Mirick G, Moussignac RL (2002) Resolvins: a family of bioactive products of omega-3 fatty acid transformation circuits initiated by aspirin treatment that counter proinflammation signals. J Exp Med 196(8):1025–1037
Shalini SM, Ho CF, Ng YK, Tong JX, Ong ES, Herr DR, Dawe GS, Ong WY (2017) Distribution of Alox15 in the rat brain and its role in prefrontal cortical resolvin D1 formation and spatial working memory. Mol Neurobiol. https://doi.org/10.1007/s12035-017-0413-x
Shalini S-M, Herr DR, Ong WY (2016) The analgesic and anxiolytic effect of souvenaid, a novel nutraceutical, is mediated by Alox15 activity in the prefrontal cortex. Mol Neurobiol. https://doi.org/10.1007/s12035-016-0138-2
Xu ZZ, Zhang L, Liu T, Park JY, Berta T, Yang R, Serhan CN, Ji RR (2010) Resolvins RvE1 and RvD1 attenuate inflammatory pain via central and peripheral actions. Nat Med 16 (5):592–597. https://doi.org/10.1038/nm.2123
Lee YW, Kuhn H, Kaiser S, Hennig B, Daugherty A, Toborek M (2001) Interleukin 4 induces transcription of the 15-lipoxygenase I gene in human endothelial cells. J Lipid Res 42(5):783–791
Conrad DJ, Kuhn H, Mulkins M, Highland E, Sigal E (1992) Specific inflammatory cytokines regulate the expression of human monocyte 15-lipoxygenase. Proc Natl Acad Sci USA 89(1):217–221
Brinckmann R, Topp MS, Zalan I, Heydeck D, Ludwig P, Kuhn H, Berdel WE, Habenicht JR (1996) Regulation of 15-lipoxygenase expression in lung epithelial cells by interleukin-4. Biochem J 318(Pt 1):305–312
Heydeck D, Thomas L, Schnurr K, Trebus F, Thierfelder WE, Ihle JN, Kuhn H (1998) Interleukin-4 and -13 induce upregulation of the murine macrophage 12/15-lipoxygenase activity: evidence for the involvement of transcription factor STAT6. Blood 92:2503–2510
Deguchi A, Xing SW, Shureiqi I, Yang P, Newman RA, Lippman SM, Feinmark SJ, Oehlen B, Weinstein IB (2005) Activation of protein kinase G up-regulates expression of 15-lipoxygenase-1 in human colon cancer cells. Cancer Res 65(18):8442–8447. https://doi.org/10.1158/0008-5472.can-05-1109
Kamitani H, Taniura S, Ikawa H, Watanabe T, Kelavkar UP, Eling TE (2001) Expression of 15-lipoxygenase-1 is regulated by histone acetylation in human colorectal carcinoma. Carcinogenesis 22(1):187–191
Zuo X, Morris JS, Shureiqi I (2008) Chromatin modification requirements for 15-lipoxygenase-1 transcriptional reactivation in colon cancer cells. J Biol Chem 283(46):31341–31347. https://doi.org/10.1074/jbc.M803729200
Huang CY, Chen JJ, Wu JS, Tsai HD, Lin H, Yan YT, Hsu CY, Ho YS, Lin TN (2015) Novel link of anti-apoptotic ATF3 with pro-apoptotic CTMP in the ischemic brain. Mol Neurobiol 51(2):543–557. https://doi.org/10.1007/s12035-014-8710-0
Ropero S, Esteller M (2007) The role of histone deacetylases (HDACs) in human cancer. Mol Oncol 1(1):19–25. https://doi.org/10.1016/j.molonc.2007.01.001
Bowers EM, Yan G, Mukherjee C, Orry A, Wang L, Holbert MA, Crump NT, Hazzalin CA, Liszczak G, Yuan H, Larocca C, Saldanha SA, Abagyan R, Sun Y, Meyers DJ, Marmorstein R, Mahadevan LC, Alani RM, Cole PA (2010) Virtual ligand screening of the p300/CBP histone acetyltransferase: identification of a selective small molecule inhibitor. Chem Biol 17(5):471–482. https://doi.org/10.1016/j.chembiol.2010.03.006
Coffey K, Blackburn TJ, Cook S, Golding BT, Griffin RJ, Hardcastle IR, Hewitt L, Huberman K, McNeill HV, Newell DR, Roche C, Ryan-Munden CA, Watson A, Robson CN (2012) Characterisation of a Tip60 specific inhibitor, NU9056, in prostate cancer. PLoS ONE 7(10):e45539. https://doi.org/10.1371/journal.pone.0045539
Chen Y, Peng C, Abraham SA, Shan Y, Guo Z, Desouza N, Cheloni G, Li D, Holyoake TL, Li S (2014) Arachidonate 15-lipoxygenase is required for chronic myeloid leukemia stem cell survival. J Clin Investig 124(9):3847–3862. https://doi.org/10.1172/jci66129
Tsai HD, Wu JS, Kao MH, Chen JJ, Sun GY, Ong WY, Lin TN (2016) Clinacanthus nutans protects cortical neurons against hypoxia-induced toxicity by downregulating HDAC1/6. Neuromol Med. https://doi.org/10.1007/s12017-016-8401-2
Candido EP, Reeves R, Davie JR (1978) Sodium butyrate inhibits histone deacetylation in cultured cells. Cell 14(1):105–113
Zhou Q, Dalgard CL, Wynder C, Doughty ML (2011) Histone deacetylase inhibitors SAHA and sodium butyrate block G1-to-S cell cycle progression in neurosphere formation by adult subventricular cells. BMC Neurosci 12:50. https://doi.org/10.1186/1471-2202-12-50
Kim HJ, Bae SC (2011) Histone deacetylase inhibitors: molecular mechanisms of action and clinical trials as anti-cancer drugs. Am J Transl Res 3(2):166–179
Hu E, Dul E, Sung CM, Chen Z, Kirkpatrick R, Zhang GF, Johanson K, Liu R, Lago A, Hofmann G, Macarron R, de los Frailes M, Perez P, Krawiec J, Winkler J, Jaye M (2003) Identification of novel isoform-selective inhibitors within class I histone deacetylases. J Pharmacol Exp Ther 307(2):720–728. https://doi.org/10.1124/jpet.103.055541
Saito A, Yamashita T, Mariko Y, Nosaka Y, Tsuchiya K, Ando T, Suzuki T, Tsuruo T, Nakanishi O (1999) A synthetic inhibitor of histone deacetylase, MS-27–275, with marked in vivo antitumor activity against human tumors. Proc Natl Acad Sci USA 96(8):4592–4597
Lobera M, Madauss KP, Pohlhaus DT, Wright QG, Trocha M, Schmidt DR, Baloglu E, Trump RP, Head MS, Hofmann GA, Murray-Thompson M, Schwartz B, Chakravorty S, Wu Z, Mander PK, Kruidenier L, Reid RA, Burkhart W, Turunen BJ, Rong JX, Wagner C, Moyer MB, Wells C, Hong X, Moore JT, Williams JD, Soler D, Ghosh S, Nolan MA (2013) Selective class IIa histone deacetylase inhibition via a nonchelating zinc-binding group. Nat Chem Biol 9(5):319–325. https://doi.org/10.1038/nchembio.1223
Longair MH, Baker DA, Armstrong JD (2011) Simple Neurite Tracer: open source software for reconstruction, visualization and analysis of neuronal processes. Bioinformatics 27(17):2453–2454. https://doi.org/10.1093/bioinformatics/btr390
Tan CS, Ng YK, Ong WY (2015) Epigenetic Regulation of cytosolic phospholipase A in SH-sy5y human neuroblastoma cells. Mol Neurobiol. https://doi.org/10.1007/s12035-015-9314-z
Kovalevich J, Langford D (2013) Considerations for the use of SH-SY5Y neuroblastoma cells in neurobiology. Methods Mol Biol 1078:9–21. https://doi.org/10.1007/978-1-62703-640-5_2
Pahlman S, Ruusala AI, Abrahamsson L, Mattsson ME, Esscher T (1984) Retinoic acid-induced differentiation of cultured human neuroblastoma cells: a comparison with phorbolester-induced differentiation. Cell Differ 14(2):135–144
Shipley MM, Mangold CA, Szpara ML (2016) Differentiation of the SH-SY5Y human neuroblastoma cell line. J Vis Exp. https://doi.org/10.3791/53193
Delcuve GP, Khan DH, Davie JR (2012) Roles of histone deacetylases in epigenetic regulation: emerging paradigms from studies with inhibitors. Clin Epigenet 4(1):5. https://doi.org/10.1186/1868-7083-4-5
Haggarty SJ, Koeller KM, Wong JC, Grozinger CM, Schreiber SL (2003) Domain-selective small-molecule inhibitor of histone deacetylase 6 (HDAC6)-mediated tubulin deacetylation. Proc Natl Acad Sci USA 100(8):4389–4394. https://doi.org/10.1073/pnas.0430973100
Shureiqi I, Wojno KJ, Poore JA, Reddy RG, Moussalli MJ, Spindler SA, Greenson JK, Normolle D, Hasan AA, Lawrence TS, Brenner DE (1999) Decreased 13-S-hydroxyoctadecadienoic acid levels and 15-lipoxygenase-1 expression in human colon cancers. Carcinogenesis 20(10):1985–1995
Hennig R, Kehl T, Noor S, Ding XZ, Rao SM, Bergmann F, Furstenberger G, Buchler MW, Friess H, Krieg P, Adrian TE (2007) 15-lipoxygenase-1 production is lost in pancreatic cancer and overexpression of the gene inhibits tumor cell growth. Neoplasia 9(11):917–926
Jiang WG, Watkins G, Douglas-Jones A, Mansel RE (2006) Reduction of isoforms of 15-lipoxygenase (15-LOX)-1 and 15-LOX-2 in human breast cancer. Prostaglandins Leukot Essent Fat Acids 74(4):235–245. https://doi.org/10.1016/j.plefa.2006.01.009
Hsi LC, Xi X, Wu Y, Lippman SM (2005) The methyltransferase inhibitor 5-aza-2-deoxycytidine induces apoptosis via induction of 15-lipoxygenase-1 in colorectal cancer cells. Mol Cancer Ther 4(11):1740–1746. https://doi.org/10.1158/1535-7163.mct-05-0218
Shureiqi I, Zuo X, Broaddus R, Wu Y, Guan B, Morris JS, Lippman SM (2007) The transcription factor GATA-6 is overexpressed in vivo and contributes to silencing 15-LOX-1 in vitro in human colon cancer. FASEB J 21(3):743–753. https://doi.org/10.1096/fj.06-6830com
Shankaranarayanan P, Chaitidis P, Kuhn H, Nigam S (2001) Acetylation by histone acetyltransferase CREB-binding protein/p300 of STAT6 is required for transcriptional activation of the 15-lipoxygenase-1 gene. J Biol Chem 276(46):42753–42760. https://doi.org/10.1074/jbc.M102626200
McDonald C, Reich NC (1999) Cooperation of the transcriptional coactivators CBP and p300 with Stat6. J Interferon Cytokine Res 19(7):711–722. https://doi.org/10.1089/107999099313550
Wang Y, Wang X, Liu L, Wang X (2009) HDAC inhibitor trichostatin A-inhibited survival of dopaminergic neuronal cells. Neurosci Lett 467(3):212–216. https://doi.org/10.1016/j.neulet.2009.10.037
Encinas M, Iglesias M, Liu Y, Wang H, Muhaisen A, Cena V, Gallego C, Comella JX (2000) Sequential treatment of SH-SY5Y cells with retinoic acid and brain-derived neurotrophic factor gives rise to fully differentiated, neurotrophic factor-dependent, human neuron-like cells. J Neurochem 75(3):991–1003
Cheung YT, Lau WK, Yu MS, Lai CS, Yeung SC, So KF, Chang RC (2009) Effects of all-trans-retinoic acid on human SH-SY5Y neuroblastoma as in vitro model in neurotoxicity research. Neurotoxicology 30(1):127–135. https://doi.org/10.1016/j.neuro.2008.11.001
Fang HB, Mi Y, Wu NH, Zhang Y, Shen YF (2009) Genome-wide distribution of histone H3 acetylation in all-trans retinoic acid induced neuronal differentiation of SH-SY5Y cells. Chin Sci Bull 54(6):1043–1049. https://doi.org/10.1007/s11434-009-0109-3
Fagiolini M, Jensen CL, Champagne FA (2009) Epigenetic influences on brain development and plasticity. Curr Opin Neurobiol 19(2):207–212. https://doi.org/10.1016/j.conb.2009.05.009
Mychasiuk R (2015) Epigenetics of brain plasticity and behavior. In: Wright JD (ed) International encyclopedia of social and behavioral sciences, vol 7, 2nd edn. Elsevier, Amsterdam
Acknowledgements
This study was supported by a Grant from the National Medical Research Council of Singapore and the National University Health System.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Ethical Approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Ho, C.FY., Bon, C.PE., Ng, YK. et al. Expression of DHA-Metabolizing Enzyme Alox15 is Regulated by Selective Histone Acetylation in Neuroblastoma Cells. Neurochem Res 43, 540–555 (2018). https://doi.org/10.1007/s11064-017-2448-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11064-017-2448-9