Abstract
Key message
Association analysis resulted in the identification of specific StGWD alleles causing either an increase or decrease in starch phosphate content which was verified in diploid and tetraploid potato mapping populations.
Abstract
Potatoes are grown for various purposes like French fries, table potatoes, crisps and for their starch. One of the most important aspects of potato starch is that it contains a high amount of phosphate ester groups which are considered to be important for providing improved functionalization after derivatization processes. Little is known about the variation in phosphate content as such in different potato varieties and thus we studied the genetic diversity for this trait. From other studies it was clear that the phosphate content is controlled by a quantitative trait locus (QTL) underlying the candidate gene α-Glucan Water Dikinase (StGWD) on chromosome 5. We performed direct amplicon sequencing of this gene by Sanger sequencing. Sequences of two StGWD amplicons from a global collection of 398 commercial cultivars and progenitor lines were used to identify 16 different haplotypes. By assigning tag SNPs to these haplotypes, each of the four alleles present in a cultivar could be deduced and linked to a phosphate content. A high value for intra-individual heterozygosity was observed (Ho = 0.765). The average number of different haplotypes per individual (Ai) was 3.1. Pedigree analysis confirmed that the haplotypes are identical-by-descent (IBD) and offered insight in the breeding history of elite potato germplasm. Haplotypes originating from introgression of wild potato accessions carrying resistance genes could be traced. Furthermore, association analysis resulted in the identification of specific StGWD alleles causing either an increase or decrease in starch phosphate content varying from 12 nmol PO4/mg starch to 38 nmol PO4/mg starch. These allele effects were verified in diploid and tetraploid mapping populations and offer possibilities to breed and select for this trait.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Potato is a healthy and nutritious part of the average Western human diet, contributing carbohydrates and important amino acids and vitamins. It is, next to corn and wheat, one of the main sources of starch. Starch and its derivatives are widely employed in the manufacture of paper, textiles and adhesives, and due to their biodegradable and renewable nature they are increasingly being considered as an environmentally-friendly alternative to using synthetic additives in many other products, including plastics, detergents, pharmaceutical tablets, pesticides, cosmetics and even oil-drilling fluids (Kraak 1992). The thermal and rheological properties of potato starch, as well as properties in processing are related to the degree of starch phosphorylation (Veselovsky 1940). The presence of phosphate groups in starch increases the water-binding capacity, viscosity, transparency and freeze–thaw stability of processed potato starch (Craig et al. 1989; Swinkels 1985). Although natural starch from many plant species contains small amounts of covalently-bound phosphate, potato starch is particularly rich in phosphate (Jobling 2004). Phosphate groups are found attached to amylopectin chains at both C-3 and C-6 positions of the glucose residue (Bay-Smidt et al. 1994; Hizukuri et al. 1970; Sonnewald and Kossmann 2013). Several genes involved in starch biosynthesis and breakdown affect the level of starch-bound phosphate (Kötting et al. 2010; Carpenter et al. 2015) and can be proposed as candidate genes for starch phosphorylation. The most promising one is Glucan Water Dikinase (GWD1 or StGWD), a gene first described as the R-locus in potato (Lorberth et al. 1998). This single copy gene is a key enzyme in starch breakdown (EC 2.7.9.4) and catalyzes the transfer of phosphate to the C-6 position of glucosyl residues of the amylopectin fraction (Ritte et al. 2006; Zeeman et al. 2007). Genetic modification of plants overexpressing this gene permits the production of high-phosphate starch (Lorberth et al. 1998; Ritte et al. 2002). Likewise, silencing experiments have shown that potatoes with lower activity of this enzyme have a significantly lower level of starch-bound phosphate (Lorberth et al. 1998; Viksø-Nielsen et al. 2001; Wickramasinghe et al. 2009). The lower amount of starch-bound phosphate decreases starch degradation and sugar accumulation in potatoes during cold storage (Lorberth et al. 1998). Hence, a further reduction of phosphate in potato starch might contribute to potatoes with increased resistance to cold-sweetening. Breeding for a further increase of the phosphate content in potato starch on the other hand is highly desirable because a high natural degree of phosphorylation could lead to a higher substitution degree thus making chemical modification processes more efficient and potentially more environmentally friendly.
In the past two decades of potato research, identification of genes and markers that control the genetic variation of complex quantitative traits like starch phosphate content has mainly been done by linkage analysis in bi-parental segregating populations. These mapping populations are often developed from diploid parents that originate partly or completely from wild species. Such populations sample a maximum of four alleles in a single study and observed gene effects are often not representative of those found in elite tetraploid cultivars (Simko et al. 2004). In contrast to linkage mapping, association analysis samples a much larger number of alleles and usually cultivars with existing phenotypic information are used, representative for an elite genepool. Conceptually, there are two different approaches to identify DNA polymorphisms associated with quantitative trait loci (QTL) within an association analysis framework: a genome wide association analysis (GWAS) and a candidate gene approach. D'Hoop et al. (2008) has conducted an initial study to explore the potential of GWAS in potato by applying a genome-wide set of AFLP and SSR markers. In this study, a germplasm panel of 430 tetraploid potato cultivars was assembled and phenotyped. The panel covered a world-wide set of cultivars and progenitor lines, complemented by breeding lines, covering the entire range of commercial potato with respect to country of origin, year of release and market segment (consumption, frying and starch industry).
In a candidate gene-based association mapping approach, genotyping is targeted to functional and positional candidate genes for the trait under consideration. Background information on the physiology and biochemistry of the trait, together with knowledge on gene function from model organisms may suggest functional involvement of the candidate gene. Additional support may be provided by positional information of QTL from linkage maps or physical maps that locate a gene to the chromosome region suspected of being involved in the trait. In potato, candidate gene-based association mapping has been conducted by re-sequencing (Li et al. 2005; Simko et al. 2004; Wolters et al. 2010), by High Resolution Melting (HRM) analysis (de Koeyer et al. 2009) and by sequence based analysis using micro arrays or SNPs derived from Genotyping by Sequencing (GBS, Carpenter et al. 2015; Klaassen et al. 2019; Prodhomme et al. 2020; Rosyara et al. 2016).
Haase and Plate (1996) calculated a high heritability (h2 = 0.83) for starch-bound phosphate content in potato. Werij et al. (2012) confirmed StGWD as a candidate gene that underlies one of the three starch phosphate QTLs in the backcross diploid C × E mapping population of potato. The QTL analysis showed three major additive QTLs on chromosomes 2, 5 and 9, each explaining approximately 20% of the observed variance. In other studies these different chromosomes were identified to be important as well (Carpenter et al. 2015; van Harsselaar et al. 2017). The QTL on chromosome 5 co-localizes with the StGWD locus. The BC1 structure of the diploid C × E population however only allows the characterization of three GWD alleles in four possible combinations in the offspring. Thus, an unresolved question is how the level of starch phosphorylation is influenced by StGWD alleles in elite potato cultivars in which many more alleles, in different allelic compositions, are expected.
In this paper we investigate whether the collective information of quantitatively scored (i.e. dosage of) SNPs would enable us to deduce the composition of GWD haplotypes in individual tetraploid potato cultivars. We identified 16 distinct and highly diverse haplotypes and assigned tag SNPs to each of them. With this set of tag SNPs we were able to identify the fully informative four-allele GWD configurations for almost all of the nearly 400 sampled potato cultivars. The genetic composition of the cultivars was used to identify genotypic and allelic associations with starch-bound phosphate. We validated the marker-trait associations found in a diploid mapping population and show that one specific allele has a reducing effect on starch phosphate. We also identified a novel allele associated with an increase in starch phosphorylation, ready to be used in marker assisted breeding. We show that with a specific combination of four alleles a threefold higher level of phosphate content in starch could be achieved in tetraploid potato.
Materials and methods
Plant materials and DNA isolation
To aid haplotype identification five monoploid potato reference genotypes were used: 7322 (H7322, or AM79.7322 originally from G. Wenzel, Institut für Genetik, Grünbach, Germany), M47 and M133 (1022M-47 and 1022M-133) (Hoogkamp et al. 2000) and M5 and M38 (851-5 and 851-38) (Uijtewaal 1987). Furthermore, DNA from nine diploid reference genotypes was used: C and E (US-W5337.3 and 77.2102.37) (Hanneman Jr and Peloquin 1967; Jacobsen 1980), 1024-2 and 1029-31 (87.1024/2 and 87.1029/31) (Jacobsen et al. 1989), RH and SH (RH89-039-16 and SH82-93-488) (Rouppe Van Der Voort et al. 1997; Van Os et al. 2006), RH90 and RH88 (RH90-038-21 and RH88-025-50) (Park et al. 2005), and G254 (Olsder and Hermsen 1976). The cultivar collection used consisted of 430 tetraploid potato cultivars and progenitor lines chosen to represent a diverse range of commercial potato cultivars with respect to country of origin, year of release and market segment (D'hoop et al. 2008). Offspring plants of the diploid cross C × E as well as from the tetraploid cross cv. Astarte × cv. Voran and from a selfing population of cv. Sunrise were used for validation experiments. Genomic DNA was extracted from leaf material according to the protocol of van der Beek et al. (1992).
StGWD gene sequence
As no genomic sequence of the potato GWD gene was available at the time we started our research we sequenced a full length genomic allele of the StGWD gene (Genebank JQ388473) from a BAC clone of the diploid RH89-039-16 genotype. The BAC clone (RH033J14, Genbank AC237986) was anchored to the ultra-dense SH × RH genetic map (Van Os et al. 2006) and located StGWD to BIN37 of the upper arm of chromosome 5, between markers GP179 (BIN27) and the centromere (BIN46) and at a 12 cM distance of the marker SPUD237 (BIN20) (De Jong et al. 1997) (data not shown). The 16.5-kb gene contains 34 exons and encodes 1464 amino acids. In the sequenced Solanum phureja DM whole genome assembly (Xu et al. 2011) the StGWD gene is located on superscaffold PGSC0003DMB000000248 of chromosome 5, and formerly annotated as gene PGSC0003DMG400007677 (iTAG transcript ID Sotub05g014130.1.1), recently as Soltu.DM.05G009520 (DM v6.1 annotation).
PCR amplification and sequencing
Amplification and sequencing primers (Supplementary Table 1) were designed based on the consensus sequence of available genomic, mRNA and EST sequences and amplified both coding and non-coding sequence intervals of the StGWD gene. PCR amplicons for sequencing were generated from 50 ng genomic DNA template. Amplifications were performed in 20 μl reactions using 1 u of Taq Polymerase, 1 × reaction buffer, 200 nM dNTP and 250 nM of each primer. Standard cycling conditions were: 4 min initial denaturation at 94 °C, followed by 35 cycles of 1 min denaturation at 94 °C, 30 s annealing at 57 °C and 40 s extension at 72 °C. Reactions were finished by 7 min incubation at 72 °C. PCR products were examined for quality on ethidium bromide-stained agarose gels. PCR products were directly sequenced on ABI377 or ABI3700 sequencers (Biosciences, WUR) using the dideoxy chain-termination method and ABI PRISM Reaction Kit. Forward amplification primers were used as sequencing primers. To obtain phased haplotypes PCR products of eight genotypes of the GWDex7 amplicon and six of the GWD56 amplicon were cloned in pGEM-Teasy vector (Promega) and sequenced. On average twelve cloned PCR products were sequenced for each GWD haplotype to obtain a consensus sequence.
Sequence variant detection and analysis
Alignment and quality scoring was done using the Staden software package (Staden 1996). Sequence variations (SNPs and short Indels) were detected using NovoSNP (Weckx et al. 2005). The allele copy number of SNPs was scored using both the Data Acquisition & Data Analysis software DAx7.1 (Van Mierlo Software Consultancy) and manual scoring. For nucleotide diversity and phylogenetic analysis the consensus haplotype sequences were compared with one another and with S. lycopersicum-derived sequences using MEGA 4 (Tamura et al. 2007) and TREECON (Van de Peer and De Wachter 1994) software. Similarity between each pair of sequences was calculated on the basis of percentage identity and tree construction was performed using the Neighbor-joining method. To estimate gene frequencies a program for the analysis of autotetraploid genotypic data, AUTOTET (Thrall and Young 2000) was used. The following statistics were calculated to describe the levels of genetic diversity: Ai, the average number of alleles per individual at a locus; Ho, the observed heterozygosity; and He, the expected heterozygosity. In order to compare the genotype proportions with those expected under Hardy–Weinberg equilibrium the mean fixation index (F) was calculated and the chi-squared (χ2) test was used to evaluate deviations of F from zero. Pedigree information was collected from the potato pedigree database (Van Berloo et al. 2007) and inspected for abnormalities in StGWD allele transmission using Pajek (De Nooy et al. 2005) and Cytoscape (Shannon et al. 2003).
High resolution melting analysis
Amplicons for HRM genotyping were generated from 15 ng genomic DNA template. PCR amplifications were performed in 10 μl reactions using 2 μl of F-524 Phire™ 5 × reaction buffer (Finnzymes), 0.1 μl Phire™ Hot Start DNA Polymerase (Finnzymes), 1 μl LCGreen™ Plus + (BioChem) and 0.25 μl of 5 mM primers. PCR and heteroduplex formation were performed using the following conditions: 94 °C, 2 min; 40 cycles, 94 °C, 5 s; fragment-dependent Tm, 10 s; 72 °C, 10 s; a denaturation step of 30 s at 94 °C and renaturation by cooling to 30 °C. Amplicons were genotyped using the LightScanner® System (Idaho Technology).
Phenotypic data collection
The tetraploid genotypes of this study were grown in two years and starch was isolated from both years. The phosphate content of starch was however analyzed for only one year and for a subset of 207 genotypes, because the assay is laborious. Starch phosphate measurements of individual samples were repeated in triplicate. For this measurement, approximately 20 mg starch (dry weight) was added to 250 μl 70% HClO4 and heated at 250 °C for 25 min. 50 μl 30% H2O2 was added and the mixture was heated at 250 °C for another 5 min. After cooling down the volume was increased to 2 ml by adding H2O. 100 μl of the sample was pipetted into a 96-well microtiter plate and 200 μl of color reagent (0.75% (NH4)6Mo7O24·4H2O, 3% FeSO4·7H2O and 0.75% SDS dissolved in 0.375 M H2SO4) was added. After incubation for 10 min at room temperature, the absorbance was measured at 750 nm, and compared to the absorption of a calibration curve to determine the sample PO4 concentration in nmol PO4/mg starch.
Association analysis
For the analysis of phenotypic data and marker-trait association SPSS (IBM) was used. The association analysis was performed using a linear mixed model. The multivariate model was arranged to simultaneously assess the significance of all haplotype effects. Copy number of each of the haplotypes were modeled as fixed effects and haplotypes A1 to A5 were modeled as nested factors of grouped haplotype A. Variance components were estimated by the REML method. A general linear model was applied to estimate the explained phenotypic variance of associated haplotypes. For this, copy number of the haplotype within each clone was tested separately.
Results
Sequence diversity and haplotype analysis
A panel of five monoploid and nine diploid potato accessions was selected to gain an initial insight into StGWD nucleotide polymorphism among S. tuberosum clones. Seven PCR amplicons were Sanger sequenced and assessed for single locus amplification, SNPs and Indels. Amplicons derived from the monoploid accessions had sequence chromatogram peaks representing a single haplotype. Amplicons of diploid accessions displayed double chromatogram peaks at discrete nucleotide positions as expected for heterozygous accessions. Of the seven different amplicons, three amplicons showed no indel polymorphisms. Indel polymorphisms can result in undecipherable sequence chromatograms. Two amplicons were selected to identify SNPs and haplotypes in a broader panel of 430 tetraploid potato cultivar and progenitor lines. The GWDex7 amplicon (627 bp) includes a large part of the gene region from exon 8 to exon 9. The GWD56 amplicon (606 bp) covers exon 15 to exon 17 (Fig. 1).
For the tetraploid accessions, high quality sequence chromatograms with an average read length of 523 bp were generated for 398 cultivars. A small number of accessions showed low quality chromatograms in repetitive runs. In the approximately 1 kb of accessible DNA sequence of the two amplicons, 81 polymorphisms were detected and quantitatively scored. Four of these polymorphisms were of multi-allelic (three tri-allelic, one tetra-allelic) nature and 77 of bi-allelic nature. The average number of polymorphic sites—which ignores the fact that polymorphisms co-segregate in haplotype blocks—was 1 polymorphism/12 bp.
Using the sequence information of both monoploid and diploid accessions seven initial haplotypes were inferred (A1, A2, A3, B, C, D, F). Three haplotypes (A1, A2, B) were observed among the five monoploid accessions. Haplotypes of the diploids were deduced by subtracting already identified haplotypes from the sequence chromatograms. Haplotypes of the tetraploid potato germplasm collection could not be directly inferred from the unphased sequence chromatograms due to the highly heterozygous state and high SNP frequency. Putative haplotype models for these accessions were deduced by identifying sets of co-segregating SNPs. For this we calculated the squared correlation coefficient (r2) between the copy numbers of all polymorphisms. Co-segregating polymorphisms were assigned to putative haplotypes and novel haplotypes were identified by sequencing cloned amplicons of a number of corresponding potato accessions. All polymorphisms were assigned to 16 verified haplotypes, and haplotype-specific tag SNPs could be identified (Table 1).
The quantitatively scored tag SNPs were used to assign an allele copy number and genotype composition to each cultivar. In case haplotypes contained multiple tag SNPs tagging the same haplotype the best quantifiable SNP was selected for copy number estimate. When a haplotype contained no unique tag SNP the allele copy number was inferred by subtracting the copy number of already tagged allele(s) from the “tag” SNP shared by the alleles (Table 2). For the GWDex7 amplicon identical-in-state haplotypes A2, A3, A4 and A5 and haplotype D were multi-marker defined. For the GWD56 amplicon haplotypes A1 and A2 were identical-in-state and there were four multi-marker defined haplotypes. The allele copy numbers found for the haplotypes of the GWDex7 amplicon invariably matched the allele copy numbers in the GWD56 amplicon. Using the selected tag SNPs it was possible to assign a four-allele genotype to 384 (96%) of the tetraploid potato cultivars (Supplementary Fig. 1).
To investigate the sequence similarity between the detected haplotypes, a Neighbor-joining dendrogram was constructed using amplicon sequences of S. lycopersicum as out-group (Fig. 2). Over the 1 kb of DNA sequence of the two amplicons, the tomato haplotype was to a high degree similar (95.4–96.5%) to the haplotypes observed in the potato germplasm set. Sequence similarity between the 16 potato haplotypes ranged from 96.8 to 99.9%. Distance between the two most distant potato haplotypes (A1 and E) approached the sequence divergence observed between potato and tomato.

source is given. The tomato (S. lycopersicum cv. Heinz) sequence was used as out-group to root the tree
Dendogram of the 16 GWD haplotypes. The distances were computed using the Jukes-Cantor method and the tree inferred using the Neighbor-joining method. The percentage of replicate trees in which the associated haplotypes clustered together in the bootstrap test (1000 replicates) are shown next to the branches. For each allele the frequency and—when identified—the
The estimated nucleotide diversity between the 16 potato haplotypes was π = 18.5 × 10−3 and translated into an average SNP diversity of ≈ 1 SNP/54 bp (1/π). At the protein level, the analyzed haplotypes included 302 codons of five exons. Of those codons 36 showed polymorphisms, causing 24 non-synonymous changes and 15 synonymous changes. No well-defined dysfunctional mutations such as stop codon, splicing site or frame shift mutations were found. For estimates of nucleotide diversity at the population level, the frequencies at which haplotypes occurred was considered. Six haplotypes had an allele frequency above 5%, and 10 had a frequency below 5% (Table 3). In the sampled population of 398 cultivars, we found a population frequency adjusted nucleotide diversity value of π = 16.2 × 10−3. Between two randomly selected homologues alleles, this translated into ≈ 1 SNP/62 bp.
Pedigree analysis
To verify that the identified haplotypes were identical-by-descent and to identify putative sources of the haplotypes we performed a pedigree analysis. For 218 fully genotyped cultivars at least one parental cultivar had also been genotyped, and for 56 of these both parents were genotyped. For 22 out of the 218 genotyped parents/offspring pairs a mismatch was observed. In 12 occasions the mismatch repeatedly involved the parental genotypes of the cultivars AM 78-3704, Sirtema, Early Rose and Patersons Victoria. For several of the alleles, the putative source of the allele was found (Table 3). Haplotypes A1, H, I, J and K were found to be relatively new in the analyzed genepool. Haplotype G was found only in five heirloom potato cultivars. Other haplotypes were present in both ancient and new potato cultivars.
Genetic diversity
Gallais (2003) proposed the following terms to describe tetraploid genotypes: monogenic (aaaa), digenic-simplex (aaab), digenic-duplex (aabb), trigenic (aabc) and tetragenic (abcd), terms we adopt here. When for a cultivar only one of the GWD amplicons was successfully re-sequenced, the five haplotypes that were identical-in-state in either of the amplicons (A1, A2, A3, A4 and A5) could not always be fully resolved. To strengthen the analysis of genotypic variation, these similar haplotypes were grouped into a single haplotype A group. A monogenic condition was observed in nine cultivars that were monogenic for haplotype A. Four of these contained only the major haplotype A3 and were truly homozygous at the StGWD locus. The five other cultivars contained three copies of the A3 allele and a copy of either the A1 or A2 allele. All other cultivars were heterozygous. We observed 77 tetragenic, 185 trigenic, 76 digenic-simplex and 37 digenic-duplex cultivars. The average number of alleles per individual (Ai) was 2.86 when the haplotypes A were grouped and estimated at 3.08 when using all 16 haplotypes. A total of 111 different genotypic classes were observed. The number of cultivars per class ranged from 1 to 27 (3.5 cultivars per class on average). The most abundant genotypic class was AAAB occurring in 27 cultivars, followed by AABB, AABD and AABF. Observed and expected heterozygosity (Ho = 0.765, He = 0.758) were in close agreement when assuming random chromosome segregation. A χ2 test showed that the mean fixation index (F) was in accordance with Hardy–Weinberg expectations.
Associations with starch phosphate content
Starch phosphate content was measured for 203 of the 398 genotyped cultivars. It ranged from 12.6 to 37.7 nmol PO4/mg starch, with an average of 22.5 ± 4.3 nmol PO4/mg starch. Variation in starch phosphate content within the 80 genotypic classes with measured starch phosphate contents was substantial. Two genotypic classes differed significantly from the other classes. Average starch phosphate content of homozygous class AAAA (14.6 nmol PO4/mg starch, n = 4) was significantly lower than the other classes. The single cultivar representing class BBCH had a significantly higher starch phosphate content (37.7 nmol PO4/mg starch). Linear mixed model analysis, modeling all haplotypes, identified significant independent associations to starch phosphate content for the grouped haplotype A (p-value 0.009) and haplotype H (p-value 0.015). The haplotype A association explained approximately 13.4% of the populations phenotypic variance and showed a negative association with starch phosphate content. Haplotype H showed a positive association and explained around 4.7% of the variance (Fig. 3).
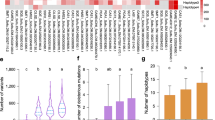
A Association between GWD allele-copy number and starch phosphate content for haplotypes A and H. Cultivars with the same four-allele genotype (80 different four-allele genotypes were found in 203 cultivars with measured starch phosphate content) have the same x-coordinate. Cultivars with different genotypes are ordered according to the increasing average starch phosphate content of the genotype. B Combined bar plot of the association between haplotypes A and H and the starch phosphate content of cultivars. Error bars show the standard error of the mean
Validation in segregating populations
To confirm the association of the haplotype A, the GWD genotypes of 93 plants of the diploid potato C × E mapping population were resolved using HRM. Three distinct GWD haplotypes were observed in the parental genotypes. Haplotype A2 is shared between both parents, haplotype F is unique to the C-parent and haplotype B unique to the E-parent. Similar to the results found for the association analysis, the C × E mapping population plants lacking allele A had significant higher starch phosphate content while the offspring homozygous for allele A had significant lower starch phosphate content (Fig. 4).
Amount of phosphorylated starch in the four genotypic GWD classes of the C × E population. Similar to the results found for the association analysis, the C × E mapping population plants lacking allele A have significantly higher starch phosphate content while the offspring homozygous for allele A have significant lower starch phosphate content. Error bars (dark grey) show the standard error of the mean
To verify the association of the haplotype A and H we also genotyped 76 tetraploid offspring of a cross Astarte (A3A3CI) × Voran (A3A3CC) and 34 tetraploid offspring of a selfing population of Sunrise (BBHH) using HMR. In a study by Noda et al. (2004), who measured starch phosphorous content in six potato cultivars, a low phosphorous content was reported for cultivar Astarte. Starch phosphate content was measured in 34 offspring of the Astarte × Voran cross and in 19 offspring of Sunrise. For the Sunrise offspring we only obtained offspring genotypes with either no, one or two copies of the H allele. For the Astarte × Voran cross we analyzed only those plants that had allele A3 and/or C. Offspring with allele H showed a clear tendency (p value 0.070) towards higher starch phosphate content and offspring homozygous for allele A3 had a significant (p value < 0.001) lower phosphate content (Fig. 5).
Discussion
Nucleotide diversity
Two regions of the StGWD gene of 627 bp and 606 bp were analyzed by direct sequencing of PCR products from monoploid, diploid and tetraploid potato clones. Analysis of the sequence chromatograms, along with verification of a number of haplotypes using cloned PCR products, allowed us to identify a set of 16 haplotypes and their tag SNPs. The tag SNPs were instrumental to fully genotype 384 tetraploid potato cultivars and to study the genetic variation and phenotypic effect of the StGWD gene.
DNA sequence variation in potato is exceptionally abundant. We found an overall frequency of polymorphic sites of one variant every 12 bp for the StGWD gene. For this large panel of cultivars and progenitor lines, the number of polymorphic sites even exceeds the level of one variant per 21–23 bp found in previous studies (Rickert et al. 2003; Simko et al. 2006). The frequency of polymorphic sites and the molecular diversity can however vary widely, depending on how many clones, which regions and which genotypes are being analyzed. The study of Simko et al. (2006) involved 47 samples, including some wild accessions, and was re-sequenced for 66 loci. Comparison of nucleotide diversity between Simko et al. (2006) and our study shows that this statistic is more stable across studies (π = 14.6 × 10−3 and π = 16.2 × 10−3, respectively).
As expected, the average within-potato sequence diversity in our study does not exceed the nucleotide diversity between Solanum species at the StGWD locus. However, the level of sequence homology of the two most distant potato haplotypes approaches the tomato/potato species sequence divergence. Considering tomato/potato divergence time of 7.3 MYA (Wang et al. 2008), most observed haplotypes seem to have diverged long time ago, at least predating domestication in potato. Some haplotypes were observed only in recent cultivars and likely resulted from efforts to introduce late blight and nematode resistance from related species (Bradshaw and Ramsay 2005). Analyzed cultivars harboring haplotype H have a lineage descending from clone USDA 96-56. It is likely that this haplotype has been introduced into the S. tuberosum genepool together with the introduction of the chromosome 5 Phytophthora infestans R1 resistance gene from S. demissum (Sinden and Sanford 1981). For another haplotype, haplotype A1 it is likely that it has been introgressed into the cultivated genepool from the CPC 1673 donor of the chromosome 5 potato cyst nematode (Globodera rostochiensis) resistance allele H1 (Ellenby 1952). And haplotype K seems to be introduced into the genepool by introgression of the chromosome 5 Globodera pallida nematode resistance from clone VTN 62-33-3 (Ross and Hunnius 1986).
Genetic diversity
The haplotype tag SNPs, with a quantitative scoring of allele copy number for two independent amplicons, gave us the possibility to exploit the full genotypic information. We used this genotypic information to evaluate the genetic diversity in the analyzed set of cultivars and accessions. Intra-individual heterozygosity (Ho = 0.77) and the mean observed number of haplotypes per plant (Ai ≈ 3.1) are high at the StGWD locus. Both the number of haplotypes and heterozygosity are markedly higher than those reported in an earlier allozyme study of 13 loci in tetraploid potato cultivars (Oliver and Martínez Zapater 1984), which seems to demonstrate the superior resolution of SNP markers compared to allozyme studies. The number of observed haplotypes was also higher than those reported in more recent potato re-sequencing studies (Li et al. 2005; Sattarzadeh et al. 2006; Simko et al. 2004) and comparable to a multi-locus study employing SSR markers on the same cultivar set (D'hoop et al. 2010). Furthermore, the above average levels of alleles and heterozygosity observed in individual potato cultivars suggest that an underestimation of heterozygosity caused by allele homoplasy should be of a minor magnitude and strengthen our conclusion that the alleles are identical-by-descent. Full allelic resolution of the StGWD locus would however require the complete gene to be re-sequenced, while in this study we only sequenced parts of the gene. Therefore, it cannot be excluded that currently unresolved haplotypes, identical-in-state to the identified haplotypes, remain in the genepool. Near complete re-sequencing of the StGWD gene in 84 potato cultivars and accessions using massively parallel sequencing has however not identified new alleles of the StGWD gene (Uitdewilligen et al. 2013).
Association analysis of StGWD haplotypes with starch phosphate content
In potato there is only a small diversification into subpopulations. This diversification is along cultivars used for fresh consumption, processing (chips, fries), and potatoes used for the starch industry (D'hoop et al. 2010). We did not observe differences in StGWD allele frequencies in these subpopulations and therefore omitted correction for population structure in the association analysis.
Starch phosphate content is hardly influenced by environmental conditions (Haase and Plate 1996; Werij et al. 2012) and can be measured with a small technical error (Noda et al. 2006). A large variation in starch phosphate content within each genotypic class of GWD alleles was however observed in the tetraploid association mapping panel. This large variation indicates that, similar to the diploid C × E mapping population, in tetraploid cultivars multiple QTL with major effects on different genomic locations are associated with the trait. In fact the involvement of GWD2, Soluble Starch Synthase III and Branching Enzyme 1 and 2 has been shown by Carpenter et al. (2015).
Starch phosphate measurements and QTL analysis in the diploid C × E potato mapping population has been described previously (Werij et al. 2012). The QTL analysis showed three major additive QTLs on chromosomes 2, 5 and 9, each explaining approximately 20% of the observed variance. The QTL on chromosome 5 co-localized with the StGWD locus, a key enzyme involved in starch phosphorylation (Zeeman et al. 2007). We re-sequenced the parental genotypes of the C × E population and identified the GWD haplotypes in the C × E offspring using HRM. The reducing effect of haplotype A on starch phosphate content detected by the association analysis, is confirmed in the C × E population. Additionally, we verified the phenotypic effect of haplotype A and haplotype H in two tetraploid cross populations. Whether the association between starch phosphate content and these haplotypes is caused by a difference in RNA expression level or functional variation in the StGWD protein needs to be determined by further study.
Results from the present study indicate that a haplotype association analysis approach is a robust tool for mapping quantitative loci with relatively strong effects in commercially important potato populations, even without considering population structure. The fact that only a small proportion of the explained variance can be captured in this way is clear but also that this has already profound effects on the total phosphate content. Whether the observed effects connected to allele A (decrease) or allele H (increase) are sufficient to merit breeding efforts on starch phosphate content for commercial purposes remains to be evaluated but it is evident that even for these types of quantitative traits in a tetraploid background specific effects of individual alleles can contribute to the final phenotype and can be used as breeding targets.
Availability of data and material
All data is enclosed either in main text or as supplementary data. Other data can be requested from the corresponding author.
Code availability
Not applicable.
References
Bay-Smidt AM, Wischmann B, Olsen CE, Nielsen TH (1994) Starch bound phosphate in potato as studied by a simple method for determination of organic phosphate and 31P-NMR. Starch 46:167–172
Bradshaw JE, Ramsay G (2005) Utilisation of the commonwealth potato collection in potato breeding. Euphytica 146:9–19
Carpenter MA, Joyce NI, Genet RA, Cooper RD, Murray SR, Noble AD, Butler RC, Timmerman-Vaughan GM (2015) Starch phosphorylation in potato tubers is influenced by allelic variation in the genes encoding glucan water dikinase, starch branching enzymes I and II, and starch synthase III. Front Plant Sci 6:143
Craig SAS, Maningat CC, Seib PA, Hoseney RC (1989) Starch paste clarity. Cereal Chem 66:173–182
D’hoop BB, Paulo MJ, Kowitwanich K, Sengers M, Visser RGF, van Eck HJ, van Eeuwijk FA (2010) Population structure and linkage disequilibrium unravelled in tetraploid potato. Theor Appl Genet 121:1151–1170
De Jong W, Forsyth A, Leister D, Gebhardt C, Baulcombe DC (1997) A potato hypersensitive resistance gene against potato virus X maps to a resistance gene cluster on chromosome 5. Theor Appl Genet 95:246–252
De Nooy W, Mrvar A, Batagelj V (2005) Exploratory social network analysis with Pajek. Psychometrika 71(3):605
de Koeyer D, Douglass K, Murphy A, Whitney S, Nolan L, Song Y, de Jong W (2009) Application of high-resolution DNA melting for genotyping and variant scanning of diploid and autotetraploid potato. Mol Breeding 25:67–90
Dhoop BB, Paulo MJ, Mank RA, van Eck HJ, van Eeuwijk FA (2008) Association mapping of quality traits in potato (Solanum tuberosum L.). Euphytica 161:47–60
Ellenby C (1952) Resistance to the potato root eelworm, Heterodera rostochiensis Wollenweber [2]. Nature 170:1016
Gallais A (2003) Quantitative genetics and breeding methods in autopolyploid plants. Institut national de la recherche agronomique, Paris
Haase NU, Plate J (1996) Properties of potato starch in relation to varieties and environmental factors. Starch/staerke 48:167–171
Hanneman RE Jr, Peloquin SJ (1967) Croosability of 24-chromosome potato hybrids with 48-chromosome cultivars. Eur Potato J 10:62–73
Hizukuri S, Tabata S, Kagoshima NZ (1970) Studies on starch phosphate part 1. Estimation of glucose-6-phosphate residues in starch and the presence of other bound phosphate(s). Starch/staerke 22(10):338–343
Hoogkamp TJH, Van Den Ende RGT, Jacobsen E, Visser RGF (2000) Development of amylose-free (amf) monoploid potatoes as new basic material for mutation breeding in vitro. Potato Res 43:179–189
Jacobsen E (1980) Increase of diplandroid formation and seed set in 4x × 2x crosses in potatoes by genetical manipulation of dihaploids and some theoretical consequences. Z Pflanzenzuchtung 85:110–121
Jacobsen E, Hovenkamp-Hermelink JHM, Krijgsheld HT, Nijdam H, Pijnacker LP, Witholt B, Feenstra WJ (1989) Phenotypic and genotypic characterization of an amylose-free starch mutant of the potato. Euphytica 44:43–48
Jobling S (2004) Improving starch for food and industrial applications. Curr Opin Plant Biol 7:210–218
Klaassen MT, Willemsen JH, Vos PG, Visser RGF, van Eck HJ, Maliepaard C, Trindade LM (2019) Genome-wide association analysis in tetraploid potato reveals four QTLs for protein content. Mol Breed 39:151
Kötting O, Kossmann J, Zeeman SC, Lloyd JR (2010) Regulation of starch metabolism: the age of enlightenment? Curr Opin Plant Biol 13:321–329
Kraak A (1992) Industrial applications of potato starch products. Ind Crop Prod 1:107–112
Li L, Strahwald J, Hofferbert HR, Lübeck J, Tacke E, Junghans H, Wunder J, Gebhardt C (2005) DNA variation at the invertase locus invGE/GF is associated with tuber quality traits in populations of potato breeding clones. Genetics 170:813–821
Lorberth R, Ritte G, Willmitzer L, Kossmann J (1998) Inhibition of a starch-granule-bound protein leads to modified starch and repression of cold sweetening. Nat Biotechnol 16:473–477
Noda T, Tsuda S, Mori M, Takigawa S, Matsuura-Endo C, Saito K, Mangalika HAM, Hanaoka A, Suzuki Y, Yamauchi H (2004) The effect of harvest dates on the starch properties of various potato cultivars. Food Chem 86:119–125
Noda T, Tsuda S, Mori M, Takigawa S, Matsuura-Endo C, Kim S-J, Hashimoto N, Yamauchi H (2006) Determination of the phosphorus content in potato starch using an energy-dispersive X-ray fluorescence method. Food Chem 95:632–637
Oliver JL, Martínez Zapater JM (1984) Allozyme variability and phylogenetic relationships in the cultivated potato (Solanum tuberosum) and related species. Plant Syst Evol 148:1–18
Olsder J, Hermsen JGT (1976) Genetics of self-compatibility in dihaploids of Solanum tuberosum L. I. Breeding behaviour of two self-compatible dihaploids. Euphytica 25:597–607
Park TH, Vleeshouwers VGAA, Kim JB, Hutten RCB, Visser RGF (2005) Dissection of foliage and tuber late blight resistance in mapping populations of potato. Euphytica 143:75–83
Prodhomme C, Vos PG, Tammes J, João Paulo M, Visser RGF, Vossen JH, van Eck HJ (2020) Distribution of P1(D1) wart disease resistance in potato germplasm and GWAS identification of haplotype specific SNP markers. Theor Appl Genet 133:1859–1871
Rickert AM, Kim JH, Meyer S, Nagel A, Ballvora A, Oefner PJ, Gebhardt C (2003) First-generation SNP/InDel markers tagging loci for pathogen resistance in the potato genome. Plant Biotechnol J 1:399–410
Ritte G, Lloyd JR, Eckermann N, Rottmann A, Kossmann J, Steup M (2002) The starch-related R1 protein is an α-glucan, water dikinase. Proc Natl Acad Sci USA 99:7166–7171
Ritte G, Heydenreich M, Mahlow S, Haebel S, Kotting O, Steup M (2006) Phosphorylation of C6- and C3-positions of glucosyl residues in starch is catalysed by distinct dikinases. FEBS Lett 580:4872–4876
Ross H, Hunnius W (1986) Potato breeding: problems and perspectives. P. Parey
Rosyara UR, De Jong WS, Douches DS, Endelman JB (2016) Software for genome-wideassociation studies in autopolyploids and its application to potato. Plant Genome 9:2
Rouppe Van Der Voort JNAM, Van Zandvoort P, Van Eck HJ, Folkertsma RT, Hutten RCB, Draaistra J, Gommers FJ, Jacobsen E, Helder J, Bakker J (1997) Use of allele specificity of comigrating AFLP markers to align genetic maps from different potato genotypes. Mol Gen Genet 255:438–447
Sattarzadeh A, Achenbach U, Lubeck J, Strahwald J, Tacke E, Hofferbert HR, Rothsteyn T, Gebhardt C (2006) Single nucleotide polymorphism (SNP) genotyping as basis for developing a PCR-based marker highly diagnostic for potato varieties with high resistance to Globodera pallida pathotype Pa2/3. Mol Breed 18:301–312
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Simko I, Costanzo S, Haynes KG, Christ BJ, Jones RW (2004) Linkage disequilibrium mapping of a Verticillium dahliae resistance quantitative trait locus in tetraploid potato (Solanum tuberosum) through a candidate gene approach. Theor Appl Genet 108:217–224
Simko I, Haynes KG, Jones RW (2006) Assessment of linkage disequilibrium in potato genome with single nucleotide polymorphism markers. Genetics 173:2237–2245
Sinden SL, Sanford LL (1981) Origin and inheritance of solarmarine glycoalkaloids in commercial potato cultivars. Am Potato J 58:305–325
Sonnewald U, Kossmann J (2013) Starches - from current models to genetic engineering. Plant Biotechnol J 11:223–232
Staden R (1996) The Staden sequence analysis package. Mol Biotechnol 5:233–241
Swinkels JJM (1985) Composition and properties of commercial native starches. Starch/stärke 37:1–5
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thrall PH, Young A (2000) Autotet: a program for analysis of autotetraploid genotypic data. J Hered 91:348–349
Uijtewaal BA (1987) Ploidy variability in greenhouse cultured and in vitro propagated potato (Solanum tuberosum) monohaploids (2n=x=12) as determined by flow cytometry. Plant Cell Rep 6:252–255
Uitdewilligen JGAML, Wolters A-MA, D’hoop BB, Borm TJA, Visser RGF, van Eck HJ (2013) A next-generation sequencing method for genotyping by-sequencing of highly heterozygous autotetraploid potato. PLoS ONE 8(5):e62355
Van Berloo R, Hutten RCB, Van Eck HJ, Visser RGF (2007) An online potato pedigree database resource. Potato Res 50:45–57
Van Harsselaar JK, Lorenz J, Senning M, Sonnewald U, Sonnewald S (2017) Genome-wide analysis of starch metabolism genes in potato (Solanum tuberosum L.). BMC Genom 18:37
Van Os H, Andrzejewski S, Bakker E, Barrena I, Bryan GJ, Caromel B, Ghareeb B, Isidore E, De Jong W, Van Koert P, Lefebvre V, Milbourne D, Ritter E, Rouppe Van Der Voort JNAM, Rousselle-Bourgeois F, Van Vliet J, Waugh R, Visser RGF, Bakker J, Van Eck HJ (2006) Construction of a 10,000-marker ultradense genetic recombination map of potato: providing a framework for accelerated gene isolation and a genomewide physical map. Genetics 173:1075–1087
Van de Peer Y, De Wachter R (1994) TREECON for Windows: a software package for the construction and drawing of evolutionary trees for the Microsoft Windows environment. Comput Appl Biosci 10:569–570
van der Beek JG, Verkerk R, Zabel P, Lindhout P (1992) Mapping strategy for resistance genes in tomato based on RFLPs between cultivars: Cf9 (resistance to Cladosporium fulvum) on chromosome 1. Theor Appl Genet 84:106–112
Veselovsky IA (1940) Biochemical and anatomical properties of starch of different varieties of potatoes and their importance for industrial purposes. Am Potato J 17:330–339
Viksø-Nielsen A, Blennow A, Kristensen KH, Jensen A, Møller BL (2001) Structural, physicochemical, and pasting properties of starches from potato plants with repressed r1-gene. Biomacromol 3:836–841
Wang Y, Diehl A, Wu F, Vrebalov J, Giovannoni J, Siepel A, Tanksley SD (2008) Sequencing and comparative analysis of a conserved syntenic segment in the solanaceae. Genetics 180:391–408
Weckx S, Del-Favero J, Rademakers R, Claes L, Cruts M, De Jonghe P, Van Broeckhoven C, De Rijk P (2005) novoSNP, a novel computational tool for sequence variation discovery. Genome Res 15:436–442
Werij JS, Furrer H, van Eck HJ, Visser RGF, Bachem CWB (2012) A limited set of starch related genes explain several interrelated traits in potato. Euphytica 186:501–516
Wickramasinghe HAM, Blennow A, Noda T (2009) Physico-chemical and degradative properties of in-planta re-structured potato starch. Carbohydr Polym 77:118–124
Wolters AMA, Uitdewilligen JGAML, Kloosterman BA, Hutten RCB, Visser RGF, van Eck HJ (2010) Identification of alleles of carotenoid pathway genes important for zeaxanthin accumulation in potato tubers. Plant Mol Biol 73:659–671
Xu X, Pan S, Cheng S, Zhang B, Mu D, Ni P, Zhang G, Yang S, Li R, Wang J, Orjeda G, Guzman F, Torres M, Lozano R, Ponce O, Martinez D, De la Cruz G, Chakrabarti SK, Patil VU, Skryabin KG, Kuznetsov BB, Ravin NV, Kolganova TV, Beletsky AV, Mardanov AV, Di Genova A, Bolser DM, Martin DM, Li G, Yang Y, Kuang H, Hu Q, Xiong X, Bishop GJ, Sagredo B, Mejia N, Zagorski W, Gromadka R, Gawor J, Szczesny P, Huang S, Zhang Z, Liang C, He J, Li Y, He Y, Xu J, Zhang Y, Xie B, Du Y, Qu D, Bonierbale M, Ghislain M, Herrera Mdel R, Giuliano G, Pietrella M, Perrotta G, Facella P, O’Brien K, Feingold SE, Barreiro LE, Massa GA, Diambra L, Whitty BR, Vaillancourt B, Lin H, Massa AN, Geoffroy M, Lundback S, DellaPenna D, Buell CR, Sharma SK, Marshall DF, Waugh R, Bryan GJ, Destefanis M, Nagy I, Milbourne D, Thomson SJ, Fiers M, Jacobs JM, Nielsen KL, Sonderkaer M, Iovene M, Torres GA, Jiang J, Veilleux RE, Bachem CW, de Boer J, Borm T, Kloosterman B, van Eck H, Datema E, Hekkert BL, Goverse A, van Ham RC, Visser RG (2011) Genome sequence and analysis of the tuber crop potato. Nature 475:189–195
Zeeman SC, Delatte T, Messerli G, Umhang M, Stettler M, Mettler T, Streb S, Reinhold H, Kötting O (2007) Starch breakdown: Recent discoveries suggest distinct pathways and novel mechanisms. Funct Plant Biol 34:465–473
Acknowledgements
The authors would like to thank Dr. Nick de Vetten formerly of Averis Seeds, Valthermond, the Netherlands, for producing and phenotyping the two validation crosses. We thank Heleen Furrer (Plant Breeding, WUR) for measuring starch phosphate content and Dirk Jan Huigen and Dr. Ronald Hutten (Plant Breeding, WUR) for help with greenhouse activities. This research was supported by a grant of the Dutch technology foundation STW, project WPB-7926. This manuscript was part of the PhD thesis of the first author J.G.A.M.L. Uitdewilligen entitled: ‘Discovery and genotyping of existing and induced DNA sequence variation in potato’, 166 pages. Thesis, Wageningen University, Wageningen, NL (2012) ISBN 978-94-6173-233-0.
Funding
This research was supported by a grant of the Dutch technology foundation STW, project WPB-7926.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by JGAM Uitdewilligen, AMA Wolters and HJ van Eck. The first draft of the manuscript was written by JGAM Uitdewilligen and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
Not applicable.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Uitdewilligen, J.G.A.M.L., Wolters, A.M.A., van Eck, H.J. et al. Allelic variation for alpha-Glucan Water Dikinase is associated with starch phosphate content in tetraploid potato. Plant Mol Biol 108, 469–480 (2022). https://doi.org/10.1007/s11103-021-01236-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-021-01236-7