Abstract
In vitro plant regeneration was established in Echinacea pallida, a plant that is commonly used as a folk medicine to treat the common cold, fevers, inflammation and so on. Conditions for callus induction, lateral root and shoot regeneration were determined. Subsequently, two vectors pCHS and pOSAG78, carrying different selection marker genes resistant to kanamycin and hygromycin, respectively, were independently used to transform leaf explants of E. pallida using an Agrobacterium-mediated method. Genomic PCR analysis confirmed the presence of the transgene and selection marker gene in obtained transgenic lines. Southern hybridization indicated that the T-DNA insertion in some transgenic E. pallida was single copy. Among them, transformants carrying Petunia chalcone synthase (CHS) were selected for further study. CHS is a key enzyme in the biosynthesis of diverse flavonoids including anthocyanin pigmentation. Here, we analyzed the roles and compared the gene expression of two clusters of CHSs, EpaCHS-A and EpaCHS-B (EpaCHS-B1 and EpaCHS-B2), isolated from E. pallida. Two of the genes, EpaCHS-A and EpaCHS-B1, were abundantly expressed in petals, whereas EpaCHS-B2 was expressed at high levels in leaves. The expression of EpaCHSs remained constant in leaves and roots of Petunia CHS transformants, while EpaCHS-B2 expression was changed in flowers of transgenic plants. The biosynthesis of caffeic acid derivatives, cichoric acid and caftaric acid, was increased in leaves and roots of CHS transformants, respectively, while the amount of echinacoside in roots of transgenic plants was decreased. This is the first report on genetic engineering of E. pallida. The information contained herein can be used as a tool for further study of the biological pathways and secondary metabolism of specific compounds from medicinal Echinacea species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Echinacea species have been used to treat the common cold, headaches, fevers, inflammation, and used against infections and for stimulation of various immune cells for hundreds of years in North America and Europe (Barrett 2003). Among the dozen or so species that make up the Echinacea genus, three species, namely Echinacea purpurea (also known as purple coneflower), Echinacea pallida (also known as pale purple coneflower) and E. angustifolia (also known as narrow-leaf coneflower), are commonly used as medicinal plants. These three species share similar morphological characteristics and pharmacological use. Products made from E. purpurea, E. pallida and E. angustifolia are sold commercially as herbal supplements; however, there are significant differences in the chemical composition of these species (Binns et al. 2002; Laasonen et al. 2002; Barrett 2003; Barnes et al. 2005; Mistríková and Vaverková 2007). Many years are required to breed inbred lines and some species are self-incompatible (Ison et al. 2014). Clonal micropropagation is an approach that is capable of producing large numbers of genetically similar, disease-free individual plants with superior characteristics in a short period of time and limited space. This approach also has potential for the development of techniques to facilitate phytochemical production of high quality and study new methods of biosynthesis (Abbasi et al. 2007a, b). Previous reports have described hairy root culture and genetic engineering of E. purpurea (Koroch et al. 2002; Liu et al. 2006), but there have been few reports on strategies about genetic engineering of E. pallida.
Conventional propagation of Echinacea includes seed germination and protocols for optimum growth (Zheng et al. 2006; Maria et al. 2012; Qu and Widrlechner 2012), whereas in vitro tissue culture of Echinacea provides rapid multiplication and genetic modification (Harbage 2001; Wang and To 2004). Regeneration of Echinacea can occur by organogenesis and/or somatic embryogenesis. Plant growth regulators (PGRs) play a key role in adjusting tissue differentiation in vitro (Koroch et al. 2003; Jones et al. 2007). In most studies, the presence of 6-benzyladenine (BA) was essential for shoot organogenesis in Echinacea species. Murashige and Skoog (MS; Murashige and Skoog 1962) medium with different combinations of BA, kinetin and indole-3-butyric acid (IBA) was effective in inducing shoot organogenesis from intact leaves and the medium containing 3,6-dichloro-o-anisic acid (DC) or 2,4-dichlorophenoxyacetic acid (2,4-D) affected somatic embryogenesis from hypocotyls of E. pallida (Lakshmanan et al. 2002). The BA and α-naphthaleneacetic acid (NAA) combination was used for regeneration of plantlets via indirect shoot organogenesis from the leaf explants of Echinacea (Koroch et al. 2003; Sauve et al. 2004; Wang and To 2004). MS supplemented with BA (26.6 μM) and NAA (0.11 μM) achieved the optimum shoot regeneration frequency (63%) and was associated with a high number of shoots per explant of E. pallida (Koroch et al. 2003).
All species of Echinacea are grown as medicinal and ornamental plants (McKeown 1999; Walck et al. 2002). Flavonoids are a class of plant secondary metabolites and flavones and flavonols are used to change color hues. Chalcone synthase (CHS) is the first committed step of flavonoid biosynthesis. In E. purpurea, biosynthesis of caffeic acid derivatives (CADs) and anthocyanins, which are derived from flavonoids were increased in hairy root cultures grown in the light (Abbasi et al. 2007b). Regulation of CHS genes was thus designed to change the flavonoid biosynthesis of E. pallida. In this study, two vectors pCHS (carrying expression cassettes for selection marker nptII conferring kanamycin resistance and Petunia CHS) and pOSAG78 (carrying expression cassettes for OSAG78 from Oncidium orchid, selection marker hptII conferring hygromycin resistance and GUS reporter gene) were employed independently to transform E. pallida using an Agrobacterium-mediated method. The vector pCHS bearing the Petunia CHS gene, which is capable of increasing the expression of exogenous CHS, was introduced into E. pallida and the interactions of the three Echinacea CHS genes were analyzed. OSAG78 in another vector pOSAG78 encodes a patatin-like protein isolated from senescent flowers in Oncidium Grower Ramsey (Lin et al. 2011). Patatin-like proteins have lipase and phospholipase A activities. Previously, transgenic Arabidopsis plants overexpressing OSAG78 showed higher lipase activity and altered phenotypes including smaller leaves, rounder flowers and later flowering (Lin et al. 2011). Here pOSAG78 was chosen to investigate (1) whether hygromycin can be used as a selection agent in E. pallida; and (2) whether GUS expression in vector pOSAG78 can be visualized in transgenic E. pallida plants. We did not focus on the function and effect of overexpressing OSAG78 on transgenic E. pallida plants. The results of this study may provide an alternative method for the study of the biological pathways and secondary metabolites in Echinacea. The transformation system of E. pallida established provides the possibility of investigating the effect of expression of various genes, and may contribute to the successful modification of medicinal plants for higher yield of natural products.
Materials and methods
Plant materials and culture conditions
Seeds of E. pallida were purchased from Johnny’s Selected Seeds (Maine, USA), surface sterilized according to a previously published procedure (Wang and To 2004) and transferred to a basal nutrient medium (MS) containing MS salts (Murashige and Skoog 1962), 3% sucrose and 0.7% agar. The pH value of the medium was adjusted to 5.7. Seeds were incubated at 25 °C with a 16-h illumination (100 µmol m−2 s−1) for germination. Leaves from in vitro germinated seedlings were used as the source of explants to generate the shoot multiplication and in vitro explants were used to establish plant regeneration and the Agrobacterium-mediated transformation system.
Plant regeneration from in vitro leaf explants
For callus induction and shoot initiation, the leaves were cut into small pieces (0.5 × 0.5 cm2) onto a callus induction Ref medium (MS salts; 30 g 1−1 sucrose; 0.4 mg l−1 thiamine; 100 mg l−1 myo-inositol; 0.02 mg l−1 NAA; 6 mg l−1 BA; 0.8% Bacto-agar; pH 5.7) in the dark for 3 weeks and then transferred to sub-culture shoot induction N0 medium (MS salts; 30 g 1−1 sucrose; 0.004 mg l−1 NAA; 0.8 mg l−1 BA; 0.8% Bacto-agar; pH 5.7) in a 22 °C growth chamber with a photoperiod of 8-h light and 16-h darkness. Shoots about 4–5 cm long were transferred to MS medium supplemented with a low concentration of IBA and GA3, alone or in combination, for 8 weeks. The numbers of roots per shoot were recorded after 16 weeks of culture. For acclimatization, the plantlets were transferred to pots in a mist chamber for 2 weeks and then moved to the greenhouse at Academia Sinica, Taipei, Taiwan.
Plasmid construction
The plasmid pCHS contains a full-length cDNA encoding Petunia chalcone synthase (PhCHS; 1170 bp) gene and a neomycin phosphotransferase II (nptII; 795 bp) gene driven by the cauliflower mosaic virus (CaMV) 35S promoter and a nopaline synthase (nos) promoter, respectively (Wang and To 2004). In addition, pOSAG78 was constructed by carrying a full-length cDNA of Oncidium senescence-associated gene (OSAG78; 1224 bp) (Lin et al. 2011) alone downstream of the CaMV 35S promoter and then cloned into binary vector pCAMBIA1301 which contains a hygromycin phosphotransferase II (hptII) gene driven by the CaMV 35S promoter. The pCHS confers kanamycin resistance whereas the pOSAG78 confers hygromycin resistance (Supplementary Fig. S1). These vectors were transformed individually into the Agrobacterium tumefaciens strain LBA4404 by electroporation (Gene Pulser II, Bio-Rad) and bacteria were prepared for transformation as previously described in AB/kanamycin medium (Wang and To 2004).
Agrobacterium-mediated transformation
Leaves from in vitro explants were cultured in Ref medium for 2 days. The 2-day pre-cultured leaves were transferred into the Agrobacterium suspension (OD600 = 0.8–1.0) containing 100 µM acetosyringone (AS; Fluka Chemika, Switzerland), and soaked for 1 h, and then blotted dry on sterile filter paper to remove excess bacterial suspension. The infected explants were incubated Ref-co medium (50 g 1−1 glucose in Ref medium was substituted by 30 g 1−1 sucrose and supplemented with 100 µM AS) for 5 days in the dark. After co-cultivation, explants were washed by shaking in a Ref solution containing 200 mg 1−1 timentin (Duchefa, The Netherlands) for inhibition of Agrobacterium. The explants were blotted dry and transferred onto selection medium containing Ref medium or N0 medium supplemented with 200 mg 1−1 timentin and 50 mg 1−1 kanamycin (for pCHS vector) or 10 mg 1−1 hygromycin (for pOSAG78 vector) for approximately 2 weeks. For the pre-selection phase, after co-cultivation, leaf explants were cultured on Ref medium supplemented with 200 mg l−1 timentin for 2 weeks, and then selection was carried out as detailed above. Afterwards, putative transformants were sub-cultured in N1 medium (MS salts; 30 g 1−1 sucrose; 0.004 mg l−1 NAA; 0.2 mg l−1 BA; 0.8% Bacto-agar; pH 5.7; antibiotics), transferred to rooting medium and then grown in pots.
Molecular analysis of putative transformants
DNA was extracted from the leaf using the CTAB method (Michiels et al. 2003) and PCR-amplification was carried out using primer pairs: Kana-5′/Kana-3′, GUS-5′/GUS-3′, and Hygro-5′/Hygro-3′ (Supplementary Table S1) to detect the kanamycin resistance gene, the GUS gene, and the hygromycin resistance gene, respectively. GUS staining of the leaves and roots of the transformants OSAG78 was carried out as previously described (Wang et al. 2015).
Southern blotting
For transformant analysis, genomic DNA (15 μg) was digested with EcoRI (New England BioLabs, MA, USA) at 37 °C overnight. After electrophoresis on a 0.8% agarose gel in TAE buffer, the gels were denatured and blotted onto Hybond-N+ nylon membrane (GE Healthcare, UK). The non-radioactive DIG-labelled 35S-pro-CHS probe was prepared by PCR with 35S pro-5′/CHS-R1 (Supplementary Table S1) as primers and plasmid pCHS as template; the non-radioactive DIG-labelled Hygro probe was prepared by PCR with Hygro-5′/Hygro-3′ (Supplementary Table S1) as primers and plasmid pOSAG78 as template. For endogenous CHS gene identification in E. pallida, 15 μg genomic DNA of wild-type was digested by restriction enzymes BscGI, SpeI, NdeI, AlfII, BanI, NcoI, NsiI, EcoRV, HindIII or ScaI (New England BioLabs, MA, USA), and hybridization with the probes CHS-A and CHS-B1 was carried out using primer pairs comprising CHSA-5′/CHSA-3′ and CHSB1-5′/CHS1-3′ (Supplementary Table S1) based on information from EpaCHS-A and EpaCHS-B1 (Supplementary Fig. S2), respectively. Conditions for probe preparation, membrane hybridization and detection were as previously described (Wang and To 2004).
Total RNA isolation, reverse transcription (RT)-PCR and real-time PCR
RNA was extracted using the Trizol method (Invitrogen, http://www.invitrogen.com) according to the manufacturer’s protocol and the concentration was determined by the A260 measured using a NanoDrop ND1000 spectrophotometer (NanoDrop Technologies). RT reaction was performed using the OneStep RT-PCR Kit (Qiagen, Hilden, Germany). Each reaction consisted of 150 ng RNA amplified for 35 cycles. E. pallida actin gene served as an internal control. Specific primer sets for amplification of the full-length Petunia CHS cDNA and Oncidium OSAG78 cDNA are listed in Supplementary Table S1. cDNA synthesis was performed by the M-MLV Reverse Transcriptase and RNaseOUT Recombinant Ribonuclease Inhibitor (Invitrogen, Carlsbad, CA, USA). The real-time PCR reaction was carried out using KAPK SYBR FAST qPCR Kit (KAPABIOSYSTEMS, Boston, USA) and the reaction was performed on an Applied Biosystems 7500 Real-Time PCR System (Applied Biosystems). Three biological replicates were used for quantification, and data analysis was performed on an ABI PRISM 7500 thermocycler. The ratios relative to the lowest value, fold-change = \({2^{ - \Delta (\Delta {C_{\text{T}}})}}\), are shown as the relative expression level. Based on the isolated sequences of EpaCHS genes, the PCR fragments for detecting EpaCHS-A, EpaCHS-B1 and EpaCHS-B2 were in positions 855–1027, 1–228, and 296–486 of Supplementary Fig. S2, respectively.
Cloning of CHS genes in E. pallida and phylogenetic analysis
To investigate CHS genes in E. pallida, degenerate primers were designed based on homologous sequences from the National Center for Biotechnology Information (NCBI). The candidate sequences were amplified using the SMART RACE cDNA amplification kit (Clontech, Mountain View, CA) and sequences were assembled by ContigExpress of Vector NTI Suite 9.0 (Invitrogen, Carlsbad, CA) to obtain full-length cDNA encoding E. pallida chalcone synthase. The expected size from start codon and stop codon was gel-purified (Qiagen, Valencia, CA), cloned into the pGEM-T Easy vector (Promega, WI, USA), and transformed into E. coli JM109 competent cells. Plasmid DNA was isolated and then sequenced. Nucleotide sequences encoding chalcone synthase in E. pallida were submitted into GenBank with the following accession numbers: EpaCHSA, KY081676; EpaCHSB1, KY094647; EpaCHSB2, KY094648. For phylogenetic analysis, protein sequences of selected chalcone synthases were obtained from GenBank. The accession numbers are: Petunia CHSA, AAF60297; Petunia CHSB, CAA32732; Petunia CHSD, CAA32733; Petunia CHSG, P22927.2; Petunia CHSJ, CAA32737; Antirrhinum CHS, CAA27338; Arabidopsis CHS, NP_196897; Dahlia CHS1, BAK08887; Dahlia CHS2, BAK08888; Gerbera CHS1, CAA86218; Gerbera CHS2, CAA86219; Gerbera CHS3, CAA86220; Ipomoea CHSA, AAB62591; Ipomoea CHSB, AAC49030; Pea CHS1, CAA44933; Pea CHS2, CAA44934; Pea CHS3, CAA44935. A neighbor-joining phylogenetic tree based on the alignment of the polypeptides was constructed with the ClustalW program (Vector NTI Suite 9.0; Invitrogen, Carlsbad, CA) and calculated with MEGA software 6.0 by maximum likelihood with bootstrap resampling of 1000 replicates (Tamura et al. 2011).
High-performance liquid chromatography and mass spectrometry analyses
Phenolic acids (caftaric acid, cichoric acid, and echinacoside) in extracts of E. pallida were prepared with 60:40 methanol/water (Brown et al. 2010). Tissues (100 mg) were extracted with 1 ml of extraction solvent, and each sample was mixed with 1 ml of 10:90 DMSO/methanol containing 3 μg luteolin-7-o-glucoside (Extrasynthese, Genay Cedex, France) as an internal standard. High-performance liquid chromatography (HPLC) analysis was conducted using an Agilent 1200 Chemstation HPLC system and a C18 reverse-phase column (YMC-Pack Polymer C18, YMC, Japan) as described (Wang et al. 2015). Phenolic compounds were monitored at 332 nm, and the mobile phase consisted of 0.1% formic acid in H2O (A) and acetonnitrile (B), and separations were performed using the following solvent gradient: 0 (10% B), 2 (15% B), 12 (30%B), 20 (65% B), 27 (80% B), 32 (90% B), and 50 min (10% B) (Pellati et al. 2012). The injection volume was 20 μl. The fractions of caftaric acid, cichoric acid and echinacoside were collected using HPLC and further analyzed by LC–MS (Pellati et al. 2012; Thomsen et al. 2012).
Results and discussion
In vitro plantlet regeneration
At present, even though there are several micropropagation strategies available to Echinacea manufacturers (Harbage 2001; Lakshmanan et al. 2002; Sauve et al. 2004), only a few integrated regeneration programs are successful in E. pallida (Lakshmanan et al. 2002; Koroch et al. 2003). In the initial shoot regeneration experiment, leaves were excited from in vitro-grown plantlets, cut into small pieces, and cultured on MS medium supplemented with 0.02 mg l−1 NAA and 6 mg l−1 BA in a growth chamber at 25 °C under a photoperiod of 16-h illumination (100 μmol m−2 s−1) and 8-h darkness for 1 month. As a result, frequency of 60% leaf discs producing shoots was scored; and it is consistent with a high number of shoots per explant as previously described (Koroch et al. 2003). The efficiency of root induction resolves the same problem in whole-plant regeneration of E. pallida micropropagation (Harbage 2001; Lakshmanan et al. 2002; Koroch et al. 2003). Of the several combinations tested, MS medium supplemented with MS vitamins, 0.2 mg l−1 IBA and 3 mg l−1 GA3 was the most effective for root regeneration from explants. Roots were developed and callus production at the basal cut end of shoots was inhibited (Table 1). Moreover, the percentage of rooting reached 78–89% of all the explants and at least 3 roots per plantlet were observed after 16 weeks of rooting treatment. Generally, the average number of roots per plantlet was over five (Fig. 1) and more than 90% of the plantlets propagated through tissue culture survived acclimation. Auxins induce plant rooting and regulate root growth (Overvoorde et al. 2010; Saini et al. 2013; Perianez-Rodriguez et al. 2014; Chandler and Werr 2015), but auxin is not necessary for root organogenesis in in vitro regeneration of Echinacea species (Harbage 2001). In addition, even a very low concentration of auxins inhibits Echinacea root induction. Presence of IBA above 1 μM caused considerable callus production at the basal cut end of shoots, and it is likely that the auxin-induced callus at the stem base might have inhibited root production (Lakshmanan et al. 2002). However, it was reported that medium without regulator as well as medium with IBA at concentrations from 0.49 to 9.8 μM resulted in no significant difference in the efficiency of rooting in E. pallida (Koroch et al. 2003).
Rooting and whole-plant regeneration of E. pallida. a Photograph from the bottom of tubes showing induction and growth of multiple roots from single shoot explant. b In vitro healthy plantlet. c Five plantlets were used for root length analysis under medium treatments at 16 weeks after transplantation
The physiological functions of GA in root growth have been discussed for Arabidopsis (Tanimoto 2012; Mauriat et al. 2014) and crosstalk between indole-3-acetic acid (IAA) and GA has been demonstrated in the signaling system (Fu and Harberd 2003; Gou et al. 2010). Supplementation of GA3 was reported to induce increases in secondary metabolite production and also affected cell viability and root morphogenesis of E. purpurea in hairy root cultures (Abbasi et al. 2012). E. purpura with a fibrous root system was easily rooted without auxins in vitro; however, rooting response of both E. pallida and E. angustifolia in taproot systems in the field was poor (Harbage 2001; Koroch et al. 2003).
Agrobacterium-mediated transformation
Transformation via leaf organogenesis has been described in many species (Dayal et al. 2003; Husaini 2010; Van et al. 2010; Liu et al. 2011; Verma and Mathur 2011). Using leaf tissue as an explant source in transformation for further secondary metabolism investigation provides a convenient approach for non-destructive selection and avoids the outcrossing reproductive nature of the genus from seed or seedling explants (Koroch et al. 2003). In our transformation experiment, we found that E. pallida leaf explant is very sensitive to Agrobacterium infection and pre-culture was found to improve the survival rate of regenerated shoots under selection medium. Moreover, selection agents interfered with regeneration of E. pallida transformants (Table 2; Fig. 2a, b), and there was a significant effect on percentage of survival shoots, causing an increase of 11–23% of leaf discs regenerating shoots with a 14-day pre-selection phase, while the explants rarely regenerated shoots when subjected to selection pressure immediately after co-cultivation. When the inoculated leaves were pre-selected on shoot regeneration medium, they showed tiny brown calli (Fig. 2c, d), and some calli and shoots became necrotic on different selective media, while putative transgenic shoots further elongated (Fig. 2e–h). These shoots regenerated into plantlets on rooting medium and were then grown in pots (Fig. 2i, j). Pre-incubation before co-cultivation and pre-selection of explants effectively minimized such inhibition which was not observed in the transformation of E. purpurea (Wang and To 2004) and tomato (Van et al. 2010). Increases in the efficiencies of transformation using pre- and/or post-agroinfection have been reported in strawberry (Husaini 2010), petunia (Liu et al. 2011), orchid (Chai et al. 2002), Catharanthus roseus (Verma and Mathur 2011), and Brassica napus (Ovesná et al. 1993; Cardoza and Stewart 2003). In this study, the percentage of shoot regeneration under kanamycin-containing medium from E. pallida was 24% (Table 2) which is lower than for E. purpurea (47%; Wang and To 2004).
Agrobacterium-mediated transformation of the medicinal E. pallida. A typical no pre-selection plate (a) and a pre-selection plate (b). Formation of tiny calli (c) and shoot buds (d), brown in color. e, f Putative transformants (red arrow) and non-transformants (blue arrow) showed green shoot buds and shoot necrosis, respectively, in selection medium containing antibiotics. g An enlarged photo showing transformants with healthy development of shoots and leaves. h An enlarged photo showing non-transformants with brown necrotic shoots that finally died. Transgenic plantlets grown in vitro (i) and transferred to a pot (j)
Selectable marker genes have been pivotal in identifying and selecting transformed progeny. Plant transformation technologies require the use of different marker genes to monitor the multi-genic traits of transgenic plants by the multi-step transformation. In our studies, we tested the most commonly used antibiotics, kanamycin and hygromycin (Miki and McHugh 2004). It is commonly found that some selection systems are more effective in certain plant species and regeneration systems than others. An example is the lower efficiency of kanamycin resistance as a selection system in cereals than in dicots (Twyman et al. 2002). E. pallida has lower shoot regeneration frequency with hygromycin (Table 2), and transgenic lines OSAG78-4 and OSAG78-6 had retarded shoot regeneration.
Characterization of transgenic plants
PCR amplifications of DNAs from CHS transformants carrying kanamycin-resistant nptII gene were considered to be transgenic positive plants (Fig. 3a). In parallel, Southern blot analysis was carried out to estimate transgenic copy number and demonstrated that all transgenic plants were independent integration events (Fig. 3b). Transgenic CHS-3, CHS-4 and CHS-5 were suggested to be single copy while CHS-2 and CHS-7 had multiple copies. However, only nptII gene conferring kanamycin resistance could be detected in CHS-1 in which Petunia CHS was lost as revealed by Southern blot analysis. Thus, in this study, the transformation efficiency from E. pallida was calculated as 4% (5/121) which is similar to E. purpurea (3.4%; Wang and To 2004). Genome-walking was further used to characterize the plant/T-DNA insertion sites in the transgenic CHS-3, CHS-4 and CHS-5. The genomic sequence in the T-DNA integration site in transformants had no similarity to nucleotide databases and EST databases (Supplementary Fig. S3). We found that 160 bases were deleted in a flanking sequence in CHS-3, and a reverse recombination was detected in the left border of CHS-5. In addition, the flanking sequences which were isolated from CHS-4 could not be confirmed by wild-type plants (data not shown). It seems complicated by this transformation process.
Characterization of CHS transgenic plants. a Genomic PCR analysis of transgenic plants. The kanamycin resistance gene (nptII) was detected by PCR with primer set Kana-5′/Kana-3′. Vector construction and primer information are presented in Fig. S1 and Table S1, respectively. One microgram of DNA isolated from the extended leaves was used for genomic PCR. b Southern blot of six individual transgenic plants. Eighteen micrograms of DNA was digested with EcoRI and probed with DIG-labeled PCR product containing 35S promoter and Petunia CHS cDNA. c Expression patterns of PhCHS in the E. pallida leaf and root. Primer information is presented in Table S1
For transformant OSAG78, the reporter GUS gene and the hygromycin-resistant hptII gene were detected in genomic PCR analysis (Fig. 4a), and Southern blot analysis was also observed. A single hybridization band was revealed in OSAG78-1 and OSAG78-2; three hybridization bands were detected in OSAG78-5 (Fig. 4b). However, no GUS product of PCR amplification was detected in OSAG78-1 (Fig. 4c). Moreover, the large deletion of a transforming vector has been reported from transgenic plants (Wu et al. 2006; Wang et al. 2015). Truncation of T-DNA results in loss of transgene function, as was the case in our study for CHS-1 and OSAG78-1, in which only the selection marker was detected (Figs. 3, 4a). In addition, herb, floricultural plant, wheat, triticale, orchid and barley transgenic plants have been shown to carry incomplete T-DNAs after Agrobacterium-mediated transformation (Bartlett et al. 2008; Hensel et al. 2012; Tsai et al. 2012; Wang et al. 2012, 2015).
Characterization of OSAG78 transgenic plants. a Genomic PCR analysis of transgenic plants. The GUS gene and hygomycin resistance gene (hptII) were detected by PCR with primer sets GUS-5′/GUS-3′ and Hygo-5′/Hygo-3′, respectively. b Southern blot of three individual transgenic plants. Eighteen micrograms of DNA was digested with EcoRI and probed with DIG-labeled PCR product of Oncidium OSAG78 cDNA. c GUS stains of leaves and roots. d Expression patterns of Onc. OSAG78 in E. pallida
Histochemical staining for GUS activity produced blue coloration in the tissues of OSAG78-2 and OSAG78-5 but not OSAG78-1 (Fig. 4c). Unfortunately, transformants CHS-6, OSAG78-3, OSAG78-4 and OSAG78-6 did not survive due to pathogen infection.
Expression of foreign genes in transgenic plants was analyzed by RT-PCR (Figs. 3c, 4d). Petunia CHS was highly expressed in the leaves and roots of all transgenic lines except CHS-1 in which Petunia CHS was lost (Fig. 3b, c). The expression of OSAG78 was only observed in OSAG78-2; its expression was not detectable in transgenic lines OSAG78-1 and OSAG78-5 (Fig. 4d). The expression of OSAG78 was not detected, but GUS was highly expressed in OSAG78-5 (Fig. 4c, d).
Isolation and comparison of E. pallida CHS genes
In this study, we cloned CHS-like genes by conserved regions from the NCBI. Three copies of the EpaCHSs genes were found: EpaCHS-A, EpaCHS-B1, and EpaCHS-B2 (Supplementary Fig. S2) and the presence of multiple CHS genes in the E. pallida genome was analyzed by Southern blotting (Fig. 5). The CHS multigene family has been divided into two subgroups according to the degree of nucleotide identity in the open reading frames (Durbin et al. 2000; Tuteja et al. 2004; Wan et al. 2011). Based on the homology sequences, one gene forming the original CHSA, and three DNA fragments producing the second duplicate subgroup were obtained by the digestion of different restriction enzymes (Fig. 5). EpaCHS-A with 389 amino acids was classified as cluster A, and a total of 1366 base pairs (bp), including 1170 bp of exon and 196 bp of the single intron, were sequenced. The other two cDNA fragments EpaCHS-B1 and EpaCHS-B2 encoded 398 amino acids, with 86.7% nucleotide similarity belonging to the CHSB subgroup. They exhibited 88.8% protein sequence identity. The two exons of the EpaCHS-B1 gene are 188 and 1009 bp in length, respectively. They are separated by a 1070 bp intron. The EpaCHS-B1 and EpaCHS-B2 sequences were 68.6 and 66.9% identical to that of EpaCHS-A, respectively. Nucleotide sequences have been submitted into GenBank with accession numbers as follows: EpaCHS-A, KY081676; EpaCHS-B1, KY094647; EpaCHS-B2, KY094648. The sequence analysis suggested that more copies of CHS exist in the E. pallida genome.
The protein sequences of EpaCHS-B1 and EpaCHS-A shared 78.4% identity, whereas the EpaCHS-B2 and EpaCHS-A protein sequences shared 74.5% identity (Supplementary Fig. S4). The phylogenetic tree analysis (Supplementary Fig. S5) based on the chalcone synthase amino acid sequences revealed two notable characteristics. First, Echinacea CHS falls into Dahlia CHS of the Helianthodae subtribe. Second, the protein sequences of EpaCHSA and petunia CHSA encoded 389 amino acids, with 83.1% identity, and were therefore more similar than the CHSB subgroup (Supplementary Figs. S4, S5). The EpaCHSB1 and EpaCHSB2 protein sequences shared 81.5 and 77.3% identity to petunia CHSA, respectively. We used quantitative real-time PCR to confirm EpaCHS expression in various tissues, and the results indicated that EpaCHS-A and EpaCHS-B1 are abundantly expressed in petals, whereas expression of EpaCHS-B2 was strongly detected in leaves (Fig. 6).
The CHS family with a single intron is commonly found in the CHS genes of most angiosperms except for Antirrhinus majus (Sommer and Saedler 1986). The CHS genes of E. pallida contain an intron which splits a cysteine codon at a conserved position, and have identical amino acids at four chemically reactive residues (Cys164, Phe215, His303, Ans336 from Medicago sativa) of a well-known CHS-related enzyme (Wan et al. 2011). The CHS enzyme is encoded by a small multigene family in plants and most of the duplicated genes may have sub-functionalization and tissue-specificity (Durbin et al. 2000; Ohno et al. 2011; Wan et al. 2011). There was 78.4% identity between CHSA1 and CHSB1 in E. pallida, and the CHSB subgroup exhibited 88.8% identity (Supplementary Fig. S4). However, EpaCHS-A1 and EpaCHS-B1 are abundantly expressed in petals, and EpaCHS-B1 and EpaCHS-B2 have been shown to be regulated in different tissue (Fig. 6). In addition, EpaCHS-B2 had Ile inserted before The194, but Ile254 was lost as an active site residue in CHS2 of M. sativa (Supplementary Fig. S4). These conservative changes may change the structure and chemical properties of the protein, and differentially regulate anthocyanin and flavone synthesis (Wang et al. 2007; Ma et al. 2009; Ohno et al. 2011). The different expression patterns of two homologous EpaCHSBs in tissues of E. pallida imply these two chalcone synthase paralogs possibly play distinct roles in different tissues.
EpaCHS expression and metabolite analysis in CHS transgenic plants
Transgenic plants were grown to maturity and no phenotypic changes were observed. In the current study, a change in the distribution of secondary metabolite main products was observed in the leaves and roots of transgenic CHS-3, CHS-5 and CHS-7 by HPLC (Fig. 7), and there were no significant differences in expressions of EpaCHS in these tissues by quantitative real-time PCR (Supplementary Fig. S6). Furthermore, real-time PCR showed that EpaCHS-B2 which was expressed in leaves was present at dramatically increased levels in the flower of CHS-5, and Petunia CHS was reduced (Fig. 8). The expression pattern in which EpaCHS-B2 was increased and Petunia CHS decreased was also revealed in transgenic CHS-7 (Fig. 8). Therefore, EpaCHS-B2 and Petunia CHS may interact with each other in CHS-5 and CHS-7. The caffeic acid derivatives (CADs) are responsible for the flavonoid biosynthesis in secondary metabolism of E. pallida. To study the effects of the CHS transgene on CAD biosynthesis by Petunia CHS gene transformation, we analyzed the content of leaf and root caftaric acid, echinacoside and cichoric acid by HPLC. Biosynthesis of CADs including cichoric acid and caftaric acid was increased in leaves and roots of CHS transformants, respectively, while the amount of echinacoside in roots of transgenic plants was decreased in comparison to that in the WT (Fig. 7). The flavonoid biosynthesis of 4-coumaroyl-CoA and CADs biosynthesis of caffeic acid begins with coumaric acid precursors via the generation of the phenylalanine and tyrosine pathway (Brenda 2001; Jia et al. 2015). However, the amounts of cichoric acid and caftaric acid were increased in Petunia CHS transformants. These peaks in cichoric acid and caftaric acid purity from transgenic CHS plants were further determined by HPLC–ESI-MS (Table 3), and were confirmed to be a single compound. The interaction between the flavonoid pathway and CAD biosynthesis in Echinacea species has not been elucidated. These results may indicate that CADs are responsible for the flavonoid biosynthesis in secondary metabolism of E. pallida by exogenous CHS gene transformation.
Expression patterns of CHS in the flower of E. pallida. a Expression patterns of PhCHS in the E. pallida flower. b Expression levels of EpaCHS in the flower of wild-type and CHS lines were analyzed by real-time PCR. Data were normalized against the expression levels of the actin gene, and the ratios of reactions from the flower of wild-type plant (wt flower) were designated a value of one to determine the relative ratios of other reactions
Conclusion
In this study, conditions for callus induction and plant regeneration from leaf explants of in vitro-grown E. pallida were determined. Subsequently, two vectors pCHS and pOSAG78, carrying different selection marker genes, were independently used to transform leaf explants of E. pallida using an Agrobacterium-mediated method. The transgenic plants obtained were confirmed and characterized. Among them, transformants carrying Petunia CHS were further selected to study the function of foreign CHS in E. pallida, which contains at least three copies of endogenous CHS. Furthermore, HPLC analysis coupled with mass spectrometry identification revealed that two CADs, cichoric acid and caftaric acid, were increased in leaves and roots of CHS transformants, respectively, whilst the amount of echinacoside in roots of transgenic plants was decreased. There are relatively few studies of genetically engineered Echinacea (Abbasi et al. 2007a). To our best knowledge, this is the first report regarding metabolic engineering in this medicinal plant. This study successfully produced transgenic E. pallida plants with monitored plant secondary metabolites. This procedure may improve genetic engineering of Echinacea species.
Abbreviations
- 2,4-D:
-
2,4-Dichlorophenoxyacetic acid
- BA:
-
6-Benzyladenine
- CADs:
-
Caffeic acid derivatives
- CaMV:
-
Cauliflower mosaic virus
- CHS:
-
Chalcone synthase
- DC:
-
3,6-Dichloro-o-anisic acid
- Epa:
-
Echinacea pallida
- GA:
-
Gibberellin
- GA3 :
-
Gibberellin A3
- GUS:
-
β-Glucuronidase
- HPLC:
-
High-performance liquid chromatography
- hptII :
-
Hygromycin phosphotransferase II
- IBA:
-
Indole-3-butyric acid
- IAA:
-
Indole-3-acetic acid
- MS:
-
Murashige and Skoog (1962)
- NAA:
-
α-Naphthaleneacetic acid
- nos :
-
Nopaline synthase
- nptII :
-
Neomycin phosphotransferase II
- OSAG:
-
Oncidium senescence-associated gene
References
Abbasi BH, Saxena PK, Murch SJ, Liu CZ (2007a) Echinacea biotechnology: challenges and opportunities. In Vitro Cell Dev Biol 43:481–492
Abbasi BH, Tian CL, Murch SJ, Saxena PK, Liu CZ (2007b) Light-enhanced caffeic acid derivatives biosynthesis in hairy root cultures of Echinacea purpurea. Plant Cell Rep 26:1367–1372
Abbasi BH, Stiles AR, Saxena PK, Liu CZ (2012) Gibberellic acid increases secondary metabolite production in Echinacea purpurea hairy roots. Appl Biochem Biotechnol 168:2057–2066
Barnes J, Anderson LA, Gibbons S, Phillipson JD (2005) Echinacea species (Echinacea angustifolia (DC.) Hell., Echinacea pallida (Nutt.) Nutt., Echinacea purpurea (L.) Moench): a review of their chemistry, pharmacology and clinical properties. J Pharm Pharmacol 57:929–954
Barrett B (2003) Medicinal properties of Echinacea: a critical review. Phytomedicine 10:66–86
Bartlett JG, Alves SC, Smedley M, Snape JW, Harwood WA (2008) High-throughput Agrobacterium-mediated barley transformation. Plant Methods 4:22–33
Binns SE, Livesey JF, Arnason JT, Baum BR (2002) Phytochemical variation in Echinacea from roots and flowerheads of wild and cultivated populations. J Agric Food Chem 50:3673–3687
Brenda WS (2001) Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol 126:485–493
Brown PN, Chan M, Betz JM (2010) Optimization and single-laboratory validation study of a high-performance liquid chromatography (HPLC) method for the determination of phenolic Echinacea constituents. Anal Bioanal Chem 397:1883–1892
Cardoza V, Stewart CN (2003) Increased Agrobacterium-mediated transformation and rooting efficiencies in canola (Brassica napus L.) from hypocotyl segment explants. Plant Cell Rep 21:599–604
Chai ML, Xu CJ, Senthil KK, Kim JY, Kim DH (2002) Stable transformation of protocorm-like bodies in Phalaenopsis orchid mediated by Agrobacterium tumefaciens. Sci Hortic 96:213–224
Chandler JW, Werr W (2015) Cytokinin-auxin crosstalk in cell type specification. Trends Plant Sci 20:292–300
Dayal S, Lavanya M, Devi P, Sharma KK (2003) An efficient protocol for shoot regeneration and genetic transformation of pigeonpea [Cajanus cajan (L.) Millsp.] using leaf explants. Plant Cell Rep 21:1072–1079
Durbin ML, McCaig B, Clegg MT (2000) Molecular evolution of the chalcone synthase multigene family in the morning glory genome. Plant Mol Biol 42:79–92
Fu X, Harberd NP (2003) Auxin promotes Arabidopsis root growth by modulating gibberellin response. Nature 421:740–743
Gou J, Strauss SH, Tsai CJ, Fang K, Chen Y, Jiang X, Busova VB (2010) Gibberellins regulate lateral root formation in populus through interactions with auxin and other hormones. Plant Cell 22:623–639
Harbage JF (2001) Micropropagation of Echinacea angustifolia, E. pallida, and E purpurea from stem and seed explants. HortScience 36:360–364
Hensel G, Oleszczuk S, Daghma DES, Zimny J, Melzer M, Kumlehn J (2012) Analysis of T-DNA integration and generative segregation in transgenic winter triticale (x Triticosecale Wittmack). BMC Plant Biol 12:171–181
Husaini AM (2010) Pre- and post-agroinfection strategies for efficient leaf disk transformation and regeneration of transgenic strawberry plants. Plant Cell Rep 29:97–110
Ison JL, Wagenius S, Reitz D, Ashley MV (2014) Mating between Echinacea angustifolia (Asteraceae) individuals increases with flowering synchrony and spatial proximity. Am J Bot 101:180–189
Jia J, Zhang F, Li Z, Qin X, Zhang L (2015) Comparison of fruits of Forsythia suspensa at two different maturation stages by NMR-based metabolomics. Molecules 20:10065–10081
Jones MPA, Yi Z, Murch SJ, Saxena PK (2007) Thidiazuron-induced regeneration of Echinacea purpurea L.: micropropagation in solid and liquid culture systems. Plant Cell Rep 26:13–19
Koroch AR, Kapteyn J, Juliani HR, Simon JE (2002) In vitro regeneration and Agrobacterium transformation of Echinacea purpurea leaf explants. In: Janick J, Whipkey A (eds) Trends in new crops and new uses. ASHS Press, Alexandria, pp 522–526
Koroch AR, Kapteyn J, Juliani HR, Simon JE (2003) In vitro regeneration of Echinacea pallida from leaf explants. In Vitro Cell Dev Biol 39:415–418
Laasonen M, Wennberg T, Harmia-Pulkkinen T, Vuorela H (2002) Simultaneous analysis of alkamides and caffeic acid derivatives for the identification of Echinacea purpurea, Echinacea angustifolia, Echinacea pallid and Parthenium integrifolium roots. Planta Med 68:572–574
Lakshmanan P, Danesh M, Taji A (2002) Production of four commercially cultivated Echinacea species by different methods of in vitro regeneration. J Hortic Sci Biotechnol 77:158–163
Lin CC, Chu CF, Liu PH, Lin HH, Liang SC, Hsu WE, Lin JS, Wang HM, Chang LL, Chien CT, Jeng ST (2011) Expression of an Oncidium gene encoding a patatin-like protein delays flowering in Arabidopsis by reducing gibberellin synthesis. Plant Cell Physiol 52:421–435
Liu CZ, Abbasi BH, Gao M, Murch SJ, Saxena PK (2006) Caffeic acid derivatives production by hairy root cultures of Echinacea purpurea. J Agric Food Chem 54:8456–8460
Liu Y, Lou Q, Xu W, Xin Y, Bassett C, Wang Y (2011) Characterization of a chalcone synthase (CHS) flower-specific promoter from Lilium orential ‘Sorbonne’. Plant Cell Rep 30:2187–2194
Ma LQ, Pang XB, Shen HY, Pu GB, Wang HH, Lei CY, Wang H, Li GF, Liu BY, Ye HC (2009) A novel type III polyketide synthase encoded by a three-intron gene from Polygonum cuspidatum. Planta 229:457–469
Maria O, Thomsen MO, Fretté XC, Christensen KB, Christensen LP, Grevsen K (2012) Seasonal variations in the concentrations of lipophilic compounds and phenolic acids in the roots of Echinacea purpurea and Echinacea pallid. J Agric Food Chem 60:12131–12141
Mauriat M, Petterle A, Bellini C, Moritz T (2014) Gibberellins inhibit adventitious rooting in hybrid aspen and Arabidopsis by affecting auxin transport. Plant J 78:372–384
McKeown KA (1999) A review of the taxonomy of the genus Echinacea. In: Janick J (ed) Perspectives on new crops and new uses. ASHS Press, Alexandria, pp 482–489
Michiels A, Ende WV, Tucker M, Riet LV, Laere AV (2003) Extraction of high-quality genomic DNA from latex-containing plants. Anal Biochem 315:85–89
Miki B, McHugh S (2004) Selectable marker genes in transgenic plants: applications, alternatives and biosafety. J Biotechnol 107:193–232
Mistríková I, Vaverková S (2007) Morphology and anatomy of Echinacea purpurea, E. angustifolia, E. pallida and Parthenium integrifolium. Biologia 62:2–5
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Ohno S, Hosokawa M, Kojima M, Kitamura Y, Hoshino A, Tatsuzawa F, Doi M, Yazawa S (2011) Simultaneous post-transcriptional gene silencing of two different chalcone synthase genes resulting in pure white flowers in the octoploid dahlia. Planta 234:945–958
Overvoorde P, Fukaki H, Beeckman T (2010) Auxin control of root development. Cold Spring Harb Perspect Biol 2:a001537
Ovesná J, Ptáček L, Opatrný Z (1993) Factors influencing the regeneration capacity of oilseed rape and cauliflower in transformation experiments. Biol Plant 35:107–112
Pellati F, Orlandini G, Benvenuti S (2012) Simultaneous metabolite fingerprinting of hydrophilic and lipophilic compounds in Echinacea pallida by high-performance liquid chromatography with diode array and electrospray ionization-mass spectrometry detection. J Chromatogr A 1242:43–58
Perianez-Rodriguez J, Manzano C, Moreno-Risueno MA (2014) Post-embryonic organogenesis and plant regeneration from tissues: two sides of the same coin? Front Plant Sci 5:727–737
Qu L, Widrlechner MP (2012) Reduction of seed dormancy in Echinacea pallida (Nutt.) Nutt. by in-dark seed selection and breeding. Ind Crops Prod 36:88–93
Saini S, Sharma I, Kaur N, Pati PK (2013) Auxin: a master regulator in plant root development. Plant Cell Rep 32:741–757
Sauve RJ, Mmbaga MT, Zhou S (2004) In vitro regeneration of the Tennessee coneflower (Echinacea tennesseensis). In Vitro Cell Dev Biol 40:325–328
Sommer H, Saedler H (1986) Structure of the chalcone synthase gene of Antirrhinum majus. Mol Genet Genomics 202:429–434
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 2:2731–2739
Tanimoto E (2012) Tall or short? Slender or thick? A plant strategy for regulating elongation growth of roots by low concentrations of gibberellin. Ann Bot 110:373–381
Thomsen MO, Fretté XC, Christensen KB, Christensen LP, Grevsen K (2012) Seasonal variations in the concentrations of lipophilic compounds and phenolic acids in the roots of Echinacea purpurea and Echinacea pallida. J Agric Food Chem 60:12131–12141
Tsai YT, Chen PY, To KY (2012) Plant regeneration and stable transformation in the floricultural plant Cleome spinosa, a C3 plant closely related to the C4 plant C. gynandra. Plant Cell Rep 31:1189–1198
Tuteja JH, Clough SJ, Chan WC, Vodkin OV (2004) Tissue-specific gene silencing mediated by a naturally occurring chalcone synthase gene cluster in Glycine max. Plant Cell 16:819–835
Twyman RM, Stöger E, Kohli A, Capell T, Christou P (2002) Selectable and screenable markers for rice transformation. In: Jackson JF, Linskens HF, Inman RB (eds) Testing for genetic manipulation in plants. Springer, Berlin, pp 1–17
Van DT, Ferro N, Jacobsen HJ (2010) Development of a simple and effective protocol for Agrobacterium tumefaciens mediated leaf disc transformation of commercial tomato cultivars. GM Crops 1:312–321
Verma P, Mathur AK (2011) Agrobacterium tumefaciens-mediated transgenic plant production via direct shoot bud organogenesis from pre-plasmolyzed leaf explants of Catharanthus roseus. Biotechnol Lett 33:1053–1060
Walck JL, Hemmerly TE, Hidayati SN (2002) The endangered Tennessee purple coneflower, Echinacea tennesseensis (Asteraceae): its ecology and conservation. Native Plants J 3:54–64
Wan D, Wang A, Zhang X, Wang Z, Li Z (2011) Gene duplication and adaptive evolution of the CHS-like genes within the genus Rheum (Polygonaceae). Biochem Syst Ecol 39:651–659
Wang HM, To KY (2004) Agrobacterium-mediated transformation in the high-value medicinal plant Echinacea purpurea. Plant Sci 166:1087–1096
Wang WK, Schaal BA, Chiou YM, Murakami N, Ge XJ, Huang CC, Chiang TY (2007) Diverse selective modes among orthologs/paralogs of the chalcone synthase (Chs) gene family of Arabidopsis thaliana and its relative A. halleri ssp. gemmifera. Mol Phylogenet Evol 44:503–520
Wang CK, Hsu SY, Chen PY, To KY (2012) Transformation and characterization of transgenic Bidens pilosa L. Plant Cell Tissue Org 109:457–464
Wang HM, To KY, Lai HM, Jeng ST (2015) Modification of flower color by suppressing β-ring carotene hydroxylase genes in Oncidium. Plant Biol 18:220–229
Wu H, Sparks CA, Jones HD (2006) Characterisation of T-DNA loci and vector backbone sequences in transgenic wheat produced by Agrobacterium-mediated transformation. Mol Breed 18:195–208
Zheng Y, Dixon MA, Saxena PK (2006) Growing environment and nutrient availability affect the content of some phenolic compounds in Echinacea purpurea and Echinacea angustifolia. Planta Med 72:1407–1414
Acknowledgements
The authors would like to express thanks to Miranda Loney for critical reading and editing of the manuscript. We also thank Dr. Chin-Yu Lin for LC-MS identification and the Metabolomics Core Facility in Academia Sinica for technical assistance. This work was supported by Academia Sinica of Taiwan to Dr. Kin-Ying To.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.

11240_2017_1208_MOESM1_ESM.jpg
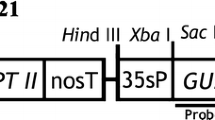
Supplementary Fig. S1 pCHS and pOSAG78 construct. The PhCHS driven by cauliflower mosaic virus 35S promoter (P35S), and the kanamycin resistance gene (nptII) with nopaline synthase promoter (NosP) in binary vector pBI221 (Wang and To 2004); the OSAG78, hygromycin resistance gene (hptII) and β-glucuronidase gene with catalase intron (gusA) under the control of cauliflower mosaic virus 35S promote (P35S) in pCAMBIA1301. CaMV, Endo, and Nos are transcription terminators (LB: left border; RB: right border) (JPG 525 KB)

11240_2017_1208_MOESM2_ESM.jpg
Supplementary Fig. S2 Comparison of CHS-A, CHS-B1, and CHS-B2 encoding chalcone synthase (CHS) in E. pallida. The sequences begin at the start codon and end at the stop codon. Identical positions are represented as white font on a black background, and the conserved positions are indicated as white over gray (JPG 9880 KB)

11240_2017_1208_MOESM3_ESM.jpg
Supplementary Fig. S3 T-DNA insertion position of transgenes CHS-3 and CHS-5 using genome-walking. The T-DNA insertion site is indicated as white font on a black background, the left flanking sequence is black over gray and the right is in black font. The reversion region is indicated as black over dark gray in CHS-5 (JPG 3406 KB)


11240_2017_1208_MOESM5_ESM.jpg
Supplementary Fig. S5 Phylogenic tree of plant chalcone synthase. The bootstrap values of 1000 retrials are indicated on each branch, and the scale shows 0.05 amino acid substitutions per site. The alignment of the amino acid sequences is presented in Supplementary Fig. S4, and the accession numbers of proteins are listed in Materials and Methods (JPG 1181 KB)

11240_2017_1208_MOESM6_ESM.jpg
Supplementary Fig. S6 Expression levels of EpaCHS in the leaf and root of wild-type and CHS lines were analyzed by real-time PCR. Data were normalized against the expression levels of the actin gene, and the ratios of reactions from the leaf of a wild-type plant (wt leaf) were designated the value of one to determine the relative ratios of other reactions (JPG 877 KB)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Wang, HM., Jeng, ST. & To, KY. In vitro regeneration, Agrobacterium-mediated transformation, and genetic assay of chalcone synthase in the medicinal plant Echinacea pallida . Plant Cell Tiss Organ Cult 130, 117–130 (2017). https://doi.org/10.1007/s11240-017-1208-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-017-1208-5