Abstract
Buckwheat characterises with high susceptibility to in vitro tissue culture conditions, which have been researched extensively to study a plethora of processes. F. tataricum morphogenic callus (MC) is characterised by its capacity for morphogenesis for up to ten years of culture, displaying an extraordinary level of genome stability, and comprises of proembryogenic cell complexes (PECC),which are the structures resembling somatic embryos arrested on the pre-globular stage. The non-morphogenic callus (NC) that appears on the surface of MC after approximately two years of culture due to endoreduplication cycles, is characterised by aneuploidy, rapid growth rate and high level of oxidative stress. F. esculentum embryogenic callus (EC) has different morphological and histological features, remains stable for up to three years of culture, has a dense, globular structure, and is capable of forming embryoids from the masses of embryogenic cells, but does not produce a non-embryogenic clone. In this work, immunocytochemical analyses revealed dynamic epigenetic changes in Fagopyrum calli. We demonstrated that; decreased level of H3K4me2 seems to be associated with pluripotency acquisition in F. esculentum EC and F. tataricum MC; DNA hypomethylation appears to be connected with the acquisition of the embryogenic potential and PECC reinitiation in F. tataricum MC. Moreover, we observed that H4K16ac and H4K5ac exhibited the highest variability during the course of passage in NC. Elevated levels of these modifications on day zero and day six for H4K16ac and H4K5ac, respectively, seem to be connected with endoreplication peaks, the processes which are characteristic of this callus.
Key message
Epigenetic changes accompany the dedifferentiation and re-differentiation processes in long-term callus cultures of Fagopyrum with different capacity for morphogenesis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The buckwheat genus comprises 23 species, and the two most cultivated are Fagopyrum esculentum Moench (common buckwheat) and Fagopyrum tataricum Gaertn. (Tartary buckwheat) (Tomasiak et al. 2022). Buckwheat is highly susceptible to in vitro conditions which have been researched extensively to study the induction of callus, plantlet regeneration from a variety of explants, organogenesis, somatic embryogenesis (SE) and synthesis of secondary metabolites (Fei et al. 2019; Rumyantseva et al. 2005; Rumyantseva et al. 2003; Srejovic and Neskovic 1981; Takahata and Jumonji 1985; Yamane 1985). F. tataricum in vitro tissue culture of the morphogenic callus (MC) characterised by low chromosome variability and the capacity for morphogenesis for up to ten years of culture displaying an extraordinary level of stability (Betekhtin et al. 2017; Rumyantseva et al. 2003). On the contrary, the non-morphogenic callus (NC) of F. tataricum appears on the surface of MC after approximately two years of culture due to endoreduplication cycles (Betekhtin et al. 2019, 2017). NC is characterised by aneuploidy, rapid growth rate and high content of hydrogen peroxide, an indication that NC is in a constant state of oxidative stress (Betekhtin et al. 2017; Kamalova et al. 2009). F. esculentum embryogenic callus (EC) displays different morphological and histological features, remains stable for up to three years of culture, has a dense, globular structure, and is capable of forming embryoids (Rumyantseva et al. 2005) but does not produce a non-embryogenic clone.
Sessile organisms, such as plants, are subjected to the ever-changing environment and have developed a high level of phenotypic plasticity, which to a vast degree, is controlled by the post-translational modifications (PTMs) (Bennett et al. 2021; Ghosh et al. 2021; Miguel and Marum 2011). Under certain circumstances, differentiated plant cells have the ability to return to a previous developmental state (dedifferentiate) and regain pluri- or totipotency, resulting in organogenesis and SE, processes largely involving PTMs (Birnbaum and Sanchez Alvarado 2008; Simsek Geyik et al. 2022; Xu and Huang 2014). The process of dedifferentiation and re-entry into the cell cycle was found to be associated with the global reorganisation of chromatin and changes in gene expression (Avivi et al. 2004; Williams et al. 2003; Zhao et al. 2001). In vitro tissue culture is a good system for studying these processes since they can be easily induced, influenced and modified under controlled conditions. According to the available evidence, it is believed that in vitro conditions destabilise the genetic and epigenetic programs of the plant tissue and can result in alteration of chromosome and DNA sequence, transposon activation or repression, generation of somaclonal variants and recalcitrance (Ckurshumova and Berleth 2015; Neelakandan and Wang 2012).
DNA methylation, one of the most researched modifications, is associated with a variety of molecular mechanisms, including inactivation of the chromatin, gene regulation and plant cell differentiation, among others (Ghosh et al. 2021; Kaeppler et al. 2000; Park et al. 2008; Springer and Schmitz 2017). Highly dynamic mechanisms of global and local DNA methylation changes were found to occur during cell dedifferentiation and re-differentiation processes in callus formation (Horstman et al. 2017; Li et al. 2017b). Recent studies conducted in peach (Prunus persica) demonstrated the beneficial influence of DNA hypomethylation during the transition from leaf to callus (Zheng et al. 2022), and subsequently that genes involved in DNA and histone methylation are up regulated in five- year old callus when compared to newly formed callus (Gao et al. 2023) showing dynamic changes in these epigenetic marks during the callus culture. In Glycine max and Zea mays on the other hand, losses in DNA methylation were observed after many years of consecutive culture which led to the conclusion that tissue culture results in diminishing ability to maintain DNA methylation (Han et al. 2018; Ji et al. 2019). Studies on Eleuterococcus senticosus embryogenic and non-embryogenic calli demonstrated significantly lower global DNA methylation rates in embryogenic calli (Chakrabarty et al. 2003). Similar results were obtained for Agave furcroydes, where hypermethylation on a high but stable level was present throughout the culture in non-embryogenic callus (Monja-Mio et al. 2018). DNA methylation has also been found to be engaged in SE (Chakrabarty et al. 2003; Fraga et al. 2012; Nic-Can et al. 2013; Park et al. 2008). Results obtained for Daucus carota showed that DNA hypomethylation suppressed the formation of embryogenic cells (Yamamoto et al. 2005). DNA methylation contributes to a plethora of processes; however, its role in the cell differentiation/dedifferentiation switch and SE and related processes seem to be species-dependent.
Callus formation and its developmental processes were found to involve global changes in global epigenetic make-up (Bednarek and Orłowska 2019; Miguel and Marum 2011; Us-Camas et al. 2014). Marks involved in open chromatin state such as acetylation on histone H3 and H4, as well as methylation H3K4me3 and H3K36me3 have a tendency to be enriched in the callus cells when compared to somatic cells, while marks responsible for heterochromatin such as H3K9me2/3 and H3K27me2/3 are decreased in dedifferentiated cells (Hemenway and Gehring 2023; Lee and Seo 2018). Histone methylation is an indispensable player in the epigenetic regulation of the gene expression orchestrated by environmental and developmental cues in plants (Cheng et al. 2020; Hu and Du 2022; Yu et al. 2009). Recent research demonstrated that reduced levels of H3K4me2 allow the activation of genes essential for the acquisition of regenerative competency during the callus culture of Arabidopsis (Ishihara et al. 2019). Another study on Arabidopsis reported the reduced accumulation of H3K4me3 during early stages of callus formation (Lee et al. 2019). Similarly, in Coffea canephora global levels of H3K4me were decreased at the beginning of SE induction (Nic-Can et al. 2013). Another modification associated with active transcription H3K36me3 was found to be accompanying the transition processes from differentiated somatic cells to dedifferentiated pluripotent stem cells during the formation of callus in Arabidopsis, leading to the conclusion that this PTM promotes early stages of cellular dedifferentiation by creating an open chromatin structure and maintaining transcriptionally competent state (Lee et al. 2017; Ma et al. 2022; Zhang et al. 2015). Histone acetylation is associated with relaxed euchromatin and transcriptional activation (Kouzarides 2007). It has been reported that reduced levels of histone acetylation due to the compromised function of histone acetyltransferases and deacetylases had a negative effect on morphogenic responses such as callus formation (Kim et al. 2018; Lee et al. 2016; Rymen et al. 2019; Zhang et al. 2020). Indirect evidence of the correlation between dynamic epigenetic changes during SE induction was also demonstrated in Hevea brasiliensis, Peaonia ostii and Arabidopsis, where patterns of expression of the histone acetyltransferases (HAT) and histone deacetylases (HDAC) genes fluctuated during the SE induction (Ci et al. 2022; Li et al. 2017a; Wickramasuriya and Dunwell 2015). In Hevea brasiliensis, reduction in histone deacetylases was observed in the early stages of callus differentiation (Li et al. 2020) Study on Arabidopsis demonstrated that histone acetylation is essential for in vitro acquisition of pluripotency by promoting transcriptional activation of several root-meristem gene loci (Kim et al. 2018). There is limited evidence available in literature decoding the exact function of the histone acetylation, however, the research implicates that acetylation is an essential factor for the appropriate regulation of transcription (Birnbaum and Roudier 2017; Us-Camas et al. 2014).
This study was conducted in order to decipher how the epigenetic modifications influence cell fate transition in long-term cultivated Fagopyrum callus lines with different capacity for morphogenesis. Depicting levels of DNA and histone methylation as well as histone acetylation during the transition from non-embryogenic to embryogenic state will help to comprehend what role the PTMs play in the acquisition of embryogenic potential, thereby expanding the limited evidence available on this topic.
Materials and methods
Plant material
The seeds of F. tataricum, sample k-17 were acquired from the collection of the N.I. Vavilov Institute of Plant Genetic Resources, Saint Petersburg, Russia (plants were grown in field conditions during the period from May to September). For F. esculentum commercially available seeds of the Panda cultivar (the Malopolska Plant Breeding, Poland) were used. EC of F. esculentum and F. tataricum MC were induced from immature zygotic embryos. NC of F. tataricum appears on the surface of MC after approximately two years of culture (Betekhtin et al. 2017). For the present study, EC’s age was approximately one year old and MC’s eight years old, and NC’s five years old. Calli were cultivated in the dark in an incubator at 25 °C ± 1 on an RX medium, according to Betekhtin et al. (2019). It composed of Gamborg B5 including vitamins (Duchefa, Netherlands) (Gamborg et al. 1968), 2 g L−1 N-Z amine A (Sigma-Aldrich, USA), 2 mg L−1 2,4-dichlorophenoxyacetic acid (2,4-D, Sigma-Aldrich), 0.2 mg L−1 kinetin (KIN, Sigma-Aldrich, USA), 0.5 mg L−1 3-indoleacetic acid (IAA, Sigma-Aldrich, USA), 0.5 mg L−1 1-naphthaleneacetic acid (NAA, Sigma-Aldrich, USA), 25 g L−1 sucrose (Chempur, Poland) and 7 g L−1 phyto agar (Duchefa, Netherlands). EC of F. esculentum, MC and NC of F. tataricum were subcultured every four and two weeks, respectively.
Histological and immunostaining procedures
Proembryogenic cell complexes (PECC) from F. tataricum MC were carefully selected in sterile conditions under the stereoscopic Nikon microscope (Japan) and transferred onto a fresh medium for each time point, i.e. day zero, two, six and eleven of the passage. For immunocytochemical analysis NC of F. tataricum and EC of F. esculentum pieces of calli were put onto fresh medium for each time point. The same day, material from one petri dish of F. tataricum MC and NC and F. esculentum EC were fixed in 4% paraformaldehyde (Sigma-Aldrich, USA) in 1 × phosphate buffered saline (PBS), pH 7.3 and placed in the vacuum desiccator for three hours with subsequent incubation in 4 °C overnight. The next day, the fixative was carefully replaced with 1 × PBS and followed by dehydration in a graded ethanol series diluted in 1xPBS solution twice for 15 min in each concentration (10%, 30%, 50%, 70% and 90%) and 99,8% twice for 30 min each. The fixation process was repeated on day two, six and eleven.
The embedding procedure was performed according to Wolny et al. (2014). The de-embedding process involved placing the slides in 99,8% ethanol three times for 10 min, followed by rehydration in ethanol/ 1xPBS solutions: 90%, 70%, 50%, 30% v/v and 1xPBS for 10 min each. The immunostaining and histological processes then commenced. For the histology, slides were stained with 0.05% aqueous solution of Toluidine blue (TBO, Sigma-Aldrich, USA) for 5 min and mounted with distilled water. Observations and photographs were performed with an Olympus BX43F microscope equipped with a Olympus XC50 digital camera.
The immunostaining method used in this experiment was previously described by Braszewska-Zalewska et al. (2010); Braszewska-Zalewska et al. (2012); Braszewska-Zalewska et al. (2013). De-embedded slides were incubated with 5% bovine serum albumin (BSA, Sigma-Aldrich, USA) in 1xPBS for 1 h in the humid chamber at room temperature and subsequently, the primary antibody in 1% BSA in 1xPBS (1:100) was applied and incubated at 4o C overnight. For 5mC primary antibody incubation was preceded with 2N HCl (Sigma-Aldrich, USA) digestion for 45 min. Next, the slides were rinsed by three washes in 1xPBS and secondary antibody in 1% BSA in 1xPBS (1:100) was applied. Samples were incubated at 37 o C in the humid chamber in the dark for one hour. After the incubation, three washes in 1xPBS were performed and nuclei were counterstained with 4',6-diamidyno-2-fenyloindol (DAPI, 2.5 g/ml in Vectashield antifade buffer, Vector Laboratories, USA). The negative control was performed for each of the used modification by conducting the immunostaining procedure omitting the addition of the primary antibody. The antibodies used in the immunocytochemical analysis are listed in Table 1.
Fluorescence intensity measurements and statistical analysis
Images were obtained using Olympus FV1000 confocal system (Olympus, Poland) equipped with an Olympus IX81 inverted microscope. Fluorescence of DAPI (excitation 405 nm, emission 425–475 nm) and Alexa488 (excitation 488 nm, emission 500–600 nm) was acquired from 60 × Plan Apo oil-immersion objective lens (NA 1.35), a 50 mW 405 nm diode laser and a 100 mW multi-line argon ion laser (Melles Griot BV, the Netherlands). Axial series of two-dimensional fluorescence images of the optical sections through the nuclei (z-stacks) was collected with the use of two separate photomultipliers (R6357, Hamamatsu, Japan) set to work in the integration mode at 4 µs pixel dwell time and 12-bit signal digitization (4096 intensity levels). Alexa488 and DAPI fluorescence intensity levels were measured in the ImageJ version 1.53 s software (Wayne Rasband, National Institutes of Health, USA). Images were converted to eight-bits and segmented with the threshold value parameter.
The fluorescence intensity of Alexa488 and DAPI was calculated as mean values from the Integrated Density (IntDen) parameter per one nucleus, which depicted the sum of all of the pixels within the region of interest. Results are presented in relative units. 500 nuclei were analysed for each callus type, four time points, for seven histone modifications, DNA methylation and two controls (unmodified histone H3 and H4). Numerical data is included in Supplementary Materials Fig. S1. Fluorescence data analysis and plotting was done using R, a software environment for statistical computing and graphics in R Studio 2022.12.0 Build 353 (Script included in Supplementary Materials Fig. S3), an integrated development environment for R (Team RC 2022; Team 2022b). An ANOVA’s test with the R Stats package (Team 2022a) followed by Tukey’s HSD test at the significance level p ≤ 0.05. Standard errors were also calculated via the stats package. The package agricolae (Statistical Procedures for Agricultural Research) was used to find significant differences between means (Mendiburu 2021). Subsequently, data was plotted with packages dedicated to data visualization; ggplot2 (Wickham 2022) and ggpubr (Kassambara 2022). Letters on the graphs indicate statistically significant differences between samples.
Results and discussion
In general, callus tissue is of remarkable properties, it contains cells of increased developmental potency and can be formed through a variety of initial pathways which include the same gene regulatory network responsible for stress, hormonal and developmental cues (Feher 2019; Hemenway and Gehring 2023). These processes have also been linked with the chromatin accessibility indicating they are closely intertwined (Bednarek and Orłowska 2019; Birnbaum and Roudier 2017). F. tataricum MC characterises with a stable regeneration potential, the ability to undergo SE and organogenesis while maintaining low chromosome number (2n = 16) throughout approximately ten years of culture (Betekhtin et al. 2017; Kamalova et al. 2009; Rumyantseva et al. 1989). MC’s PECCs have a distinctive structure typical for embryogenic cultures, of the somatic embryos arrested at the pre-globular stage due to the presence of auxin in the culture media as demonstrated previously by Rumyantseva et al. (2003), Souter and Lindsey (2000). PECCs are known to be able to regenerate through organogenesis as well as SE, which makes it a perfect system for studying its response to hormonal cues, whereas an abnormal line, NC appears on the surface of MC due to endoreduplication cycles and is subdued to constant oxidative stress (Betekhtin et al. 2017), making it suitable for research on epigenetic mechanisms involved in stress response pathways. Since they originate from the same species, but are so extraordinary different in structure and responses to tissue culture conditions, comparative analyses can be performed to decipher the role of epigenetic mechanisms behind their development. Since, the closely related species F. esculentum produces an EC typical for that type of callus, able to regenerate through the means of SE, and is not as genetically stable as F. tataricum MC callus, it was significant to research the levels of the same epigenetic modifications to shed some light on similarities in mechanisms regulating responses to stress, hormonal and developmental signals.
Morphology and histology of Fagopyrum esculentum EC, Fagopyrum tataricum MC, NC
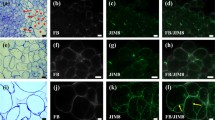
Calli were examined on days zero, two, six and eleven of the passage (Fig. 1). Images of the morphology were acquired with the Keyence VHX-970F digital microscope (Japan) equipped with an ultra-small high-performance zoom lens VH-Z20R/Z20T and wide-area illumination adapter OP-87298. It provided real-time depth composition of images with the 3D scanning technique. The microscope produced images of extraordinary clarity and texture, due to a high resolution dynamic range. In this experiment, day zero is the beginning of a new passage; a time point at which callus is transferred onto fresh RX medium. Most F. esculentum callus induction was obtained from hypocotyls, (Adachi et al. 1989; Hao et al. 1998; Jin et al. 2002; Kwon et al. 2013; Yamane 1985), however this could result in low embryogenic potential and elevated chromosome number. Callus induced from immature zygotic embryos characterised by the presence of meristemoids, was able to produce high amounts of somatic embryos suggesting that the genotype is an important factor for SE (Berbec and Doroszewska 1999; Nesković et al. 1987). EC of F. esculentum has a dense, globular structure, milky glittering surface and is comprised of embryogenic masses (Fig. 1 a–d). Upon transfer to the fresh medium, the secretion is triggered which is marked with red arrows (Fig. 1 a,b). EC of F. esculentum (Fig. 2a) contains embryogenic cells (Fig. 2a, rectangle, a1 arrowheads, a2 arrow, a3 arrow), which are poorly vacuolised, have dense cytoplasm and centrally located nucleus (Fig. 2, a3, arrow). They are surrounded by highly vacuolated parenchymatous cells (Fig. 2a, a1, a2 asterisks). Nucleus has a near-cell wall position. Phenolic containing (PC) cells that accumulate phenolics in their vacuoles are present (Fig. 2a, arrow). EC of F. esculentum also contains cells exhibiting epidermal phenotype (Fig. 2a, a2 open head arrows) which can form embryoids (Rumyantseva et al. 2005).
a Histological section of the EC of F. esculentum; area in rectangle indicates embryogenic cells; open arrows- cells exhibiting epidermal phenotype; asterisks—parenchymatous cells; outlined areas—embryogenic masses; arrow—PCs; scale bar 100 µm (a1) magnification, arrowheads—embryogenic masses; asterisk- parenchymatous cells; scale bar 20 µm (a2) magnification, arrow – embryogenic cells, open arrows—cells exhibiting epidermal phenotype; asterisk- parenchymatous cells; scale bar 20 µm (a3) magnification, arrow—embryogenic cells; scale bar 10 µm b Histological section of the proembryogenic complex of F. tataricum MC; scale bar 50 µm (b1) magnification of area marked on b, arrows—PC, double arrows—meristematic cells; open arrows—parenchymatous cells with starch grains; scale bar 20 µm c Histological section of F. tataricum NC; arrow—nucleus; inset: open arrows – nucleoli; scale bar 10 µm
Upon transfer on fresh medium, in MC of F. tataricum PECC, which has a dense structure and light colour (Fig. 1e) is triggered to disintegrate, release secretion (Fig. 1 e, red arrows) giving rise to ‘soft’ callus cells (SCCs). This is evident on day two and day six (Fig. 1 f, g) where the number of SCCs rises and they become prominent brownish, with loose structure and more secretion (Fig. 1f, red arrows). SCCs may act as support tissue, providing nutrients such as proteins and sugars as well as other conditioning factors, which are secreted into the medium during the disintegration of the PECCs. SCCs do not divide but are metabolically active which postpones death through senescence (Betekhtin et al. 2017). New PECC blowout (white asterisks) supported by SCCs (black asterisks) is noticeable on day 11 (Fig. 1h). Previous study conducted on F. tataricum revealed two cycles of PECCs reinitiation during the course of the passage (Betekhtin et al. 2019, 2017; Rumyantseva et al. 2004). First one occurs approximately on the eleventh day, and the second one around day twenty-second, both supported by the spikes of the mitotic activity and release of the extracellular polymers into the culture medium (Rumyantseva et al. 2004). In this study one cycle was studied, since the events of the PECCs disintegration and reinitiation undergo the exact same processes after day eleven. PECC’s surface (Fig. 2b) is covered by the PC cells (Fig. 2b, rectangle, b1 arrows). Below the PC cell layer, lay poorly vacuolised meristematic cells (Fig. 2b1, double arrows) with dense cytoplasm. Round nuclei and centrally located nucleolus—the source of embryogenically determined cells, from which embryoids and new PECCs can form. Parenchymatous, storage cells (Fig. 2b1, open arrows) containing large starch grains in their plastids and wall-adjacent large nucleolus are located below and constitute the largest part of the PECC. F. tataricum NC (Fig. 1i-l) has unorganised, frail structure, is characterised by very fast growth rate and is devoid of the ability for morphogenesis (Betekhtin et al. 2017). F. tataricum NC (Fig. 2c) comprises exclusively of highly vacuolated parenchymatous cells with irregularly shaped wall-adjacent nucleus (Fig. 2c, arrows) that frequently contains multiple nucleoli (Betekhtin et al. 2017; Rumyantseva et al. 2003) (Fig. 2c, inset, open arrows) PCs are components of the antioxidant defence system in tissue cultures and take part in the regulation of morphogenesis processes (Debeaujon et al. 2003; Sakar and Naik 2000). It has been previously demonstrated that MC accumulates much more phenolic compounds than the NC (Akulov et al. 2018). Moreover, NC cells poorly synthesize chlorophyll as a result of anomalies in the plastid development and lack the capacity to perform photosynthesis. Transformation of MC to the NC seems to be a result of the mutational or epigenetic changes in the DNA and subsequent silencing of genes involved in the PC biosynthesis (Akulov et al. 2018).
Epigenetic modifications in Fagopyrum esculentum EC
The analysis of the level of epigenetic modifications in F. esculentum EC were preceded with the measurement of the unmodified histones H3 and H4 as controls where no significant differences were displayed across the examined days of the passage (Fig. S2 a,b). Levels of DNA methylation across the examined days of the passage displayed significant differences (Fig. 3a). The content of 5mC decreased from day zero to day two which subsequently increased on day six and eleven. A similar result was obtained for Beta vulgaris, where hypomethylation was observed during callus formation (Zakrzewski et al. 2017). Similarly, in Oryza sativa callus, DNA hypermethylation was associated with dedifferentiated state (Stroud et al. 2013). In Triticosecale global DNA demethylation took place during tissue culture regeneration (Machczyńska et al. 2014). H3K36me3 levels (Fig. 3b) fluctuated from high on days zero and six to low on days two and eleven, whereas H3K4me (Fig. 3c, d) PTMs showed the lowest levels on day six and eleven. The deposition of H3K4me3 and H3K36me3 results in an open chromatin and transcriptional activation. H3K4me3 is typically enriched at the transcription start site and is associated with its initiation whereas deposition of H3K36me3 is more widespread and associated with transcriptional elongation (Roudier et al. 2011). H3K4me and H3K36me3 are linked to a variety of developmental processes and responses to environmental cues (Roudier et al. 2011). In Arabidopsis, a mechanism responsible for gene priming involving LYSINE-SPECIFIC DEMETHYLASE 1-LIKE 3 (LDL3) eliminates H3K4me2 during the callus formation and pluripotency acquisition (Ishihara et al. 2019). H3K4me3 increase was observed in Oryza sativa callus and was found to be mainly involved in DNA replication (Zhao et al. 2020) hence suggesting that dynamic cell differentiation in F. esculentum EC takes place during the first days of the passage. H3K18ac was on an overall high level and decreased with the passage progression (Fig. 4a), a pattern displayed also by the levels of the acetylation of H4K12 (Fig. 4b) correlating with the results obtained for histone H3K4me2 methylation. H3K18ac is also mainly found in regions adjacent to the transcription start site and correlated with transcription enhancers. Likewise is H4K12ac which is located on active coding genome regions where it creates binding sites for regulatory factors and promotes transcription (Ayyappan et al. 2015; Inacio et al. 2018). In Brassica napus high levels of H3Ac and H4Ac were detected in the vacuolated microspore, which is a totipotent cell capable of reprogramming and re-entry into the cell cycle upon induction (Rodriguez-Sanz et al. 2014). H4K16ac (Fig. 4c) levels were the lowest on day zero and kept increasing with the passage progression. H4K5ac (Fig. 4d) presented a reverse pattern, decreasing across the passage, with the exception of an increase on day eleven. Both modifications are associated with active transcription suggesting that on day eleven gene transcription might occur in higher frequency and might be related with the acquisition of the embryogenic potential (Moronczyk et al. 2022). It has been previously reported that an increase in histone H4 acetylation was associated with the progression through the S-phase of the cell cycle (Desvoyes et al. 2014). In Vicia faba, Hordeum vulgare and Arabidopsis the intensity of histone H4 acetylation at lysine 5 and 12 was correlated with the cell cycle and evident ‘deposition-related’ acetylation at eu- and heterochromatin during DNA replication and subsequent deacetylation at heterochromatin towards mitosis. On the contrary, significant acetylation of H4K16ac within the chromocenters of endopolyploidy nuclei suggested delayed deacetylation after the endoreduplication (Jasencakova et al. 2001, 2000, 2003).
Epigenetic modifications in F. tataricum NC and MC
Unmodified histone H3 and H4 levels in F. tataricum NC and MC were measured and showed no substantial differences across the examined days of the passage (Fig. S2 c-f). Immunocytochemical analyses revealed dynamic epigenetic changes in F. tataricum calli with different morphogenic potential. Differences in calli structure on the cellular level development might be a contributor to such immense changes in epigenetic modifications. NC is characterised by aneuploidy (Betekhtin et al. 2017; Kamalova et al. 2009). Increased cell ploidy has been related to an extended depth of the nuclear lobes as well as the nucleus area. The occurrence of high-level polyploid cells seemingly justifies the presence of lobed nuclei in the NC cells (Betekhtin et al. 2017). NC exhibited relatively stable but high level of DNA methylation (Fig. 3i), and histone H3K18, H4K12 acetylation (Fig. 4i, j) probably due to the rapid growth rate and high content of hydrogen peroxide, malonic dialdehyde, low catalyse activity and high superoxide dismutase activity indicating that NC is in a constant state of oxidative stress (Kamalova et al. 2009). Additionally, NC has a high level of DNA damage across the passage, which is a normal physiological state for the NC and its cells appear to be adapted to it (Betekhtin et al. 2017). This can be correlated with the high levels of DNA methylation (Fig. 3i) and significantly reduced and unstable levels of the acetylation of H4K16 and H4K5 (Fig. 4k, l) during the course of the passage.
The most noticeable and prominent pattern of the epigenetic modifications across the course of the passage in F. tataricum MC was DNA methylation (Fig. 3e). On day zero and eleven the levels of 5mC were significantly decreased when compared to day two and six. This might be due to the fact that PECCs on day zero have the highest morphogenic capacity, and upon disintegration and the loss of the capacity the 5mC content rises. Subsequent reinitiating of PECCs on day eleven results in regaining the morphogenic capacity and a decrease in the DNA methylation level. The levels of 5mC in NC (Fig. 3i), on the other hand were on a much higher level when compared to MC, however except for day six stayed on a stable level across the course of the passage. Betekhtin et al. (2019) examined the expression of DNA methyltransferases in F. tataricum calli and demonstrated significantly higher expression of MET1 and MET2 in the NC, while demethylases (DME1, DME2, DME3) expression was on the higher level in MC. Similar results were obtained for Siberian ginseng where significantly lower global DNA methylation rates in embryogenic calli were present in contrast to non-embryogenic calli (Chakrabarty et al. 2003). Additionally, in Agave furcroydes, hypermethylation on high but stable level was present throughout the culture in non-embryogenic callus and in the embryogenic clone the hypomethylation was observed in the beginning and towards the end of the passage (Monja-Mio et al. 2018), which correlates with the results obtained for MC of F. tataricum. H3K36me3 (Fig. 3f, j) was on the stable level with no vast fluctuations throughout the days of the passage in MC and in NC. H3K4me marks (Fig. 3g, h) were present in decreased levels on days zero and two in MC, similarly to Coffea canephora global levels of H3K4me which were decreased at the beginning of SE induction (Nic-Can et al. 2013). As mentioned earlier, elimination of H3K4me2 was observed in Arabidopsis during the pluripotency acquisition (Ishihara et al. 2019), therefore the differentiation/dedifferentiation switch might be initiated during the first days of the culture in MC. In NC, the level of H3K4me2 (Fig. 3k) was on the stable high level throughout the culture, whereas in H3K4me3 (Fig. 3l) it was decreased on days zero and two, comparable to MC. H3K18ac levels in MC and NC (Fig. 4e, i) were on an overall high level. However, an interesting pattern was prevalent in NC, where the lowest level presented on day zero and eleven, elevating on day two and six. H4K12ac is the mark associated with euchromatin and active transcription and in MC and NC (Fig. 4f, j) the pattern is opposing, namely in MC the H4K12ac level was the highest on day zero and eleven, whereas in NC those days levels were the lowest. This might be due to active gene expression correlated with PECCs reinitiating in the MC. An increase in the acetylation of histone H4 and acetylation of H3K18 has been associated with the progression through the S-phase of the cell cycle (Jasencakova et al. 2001, 2003). Therefore, the elevated levels of H4K12ac (Fig. 4j) in NC on days two and six as well as H4K5ac (Fig. 4l) on day zero and H4K16ac (Fig. 4k) on day six might be a result of the endoreplication (Betekhtin et al. 2017). The levels of H4K16ac in MC (Fig. 4g) presented a pattern of lower levels on day zero and eleven and elevated levels on day two and six, which seem to correlate with intense cell division and an increase in the mitotic activity (Rumyantseva et al. 2003). H4K5ac in MC (Fig. 4h) presented a gradual increase from day zero to day eleven, while in NC the significant reduction of this modification was observed on day zero two and eleven. Elevated level of this modification on day six may suggest that upon this day the transcription is activated and rapid transcription takes place. Betekhtin et al. (2019) examined the expression of genes connected with ethylene biosynthesis, a factor with a significant role in the senescence processes, 1-AMINOCYCLOPROPANE-1-CARBOXYLIC SYNTHASE (ACS2 and ACS6), 1-AMINOCYCLOPROPANE-1-CARBOXYLIC ACID OXIDASE(ACO1) and ETHYLENE RESPONSE FACTOR (ERF), in MC and NC. Their research showed that, ACS2 and ACS6 expression was higher in the NC, which led to the conclusion that such vast differences in the expression of these genes can result in the ethylene overproduction and the induction of fast senescence processes in the NC. Additionally, an increased expression of the ERF1 in NC in comparison to MC seem to be related to the high levels of oxidative stress present throughout the culture. It was demonstrated, that ERF1 is upregulated in the response to abiotic stress triggers such as salt and cold treatments or water deficit (Lestari et al. 2018; Makhloufi et al. 2014).
Conclusions
This study investigated the influence of the epigenetic modifications on the cell fate transition in long-term cultivated in vitro Fagopyrum callus lines with different capacity for morphogenesis. It was exhibited that; decreased level of H3K4me2 seems to be connected with pluripotency acquisition in F. esculentum EC and F. tataricum MC; DNA hypomethylation appears to be correlated with the acquisition of the embryogenic potential and PECC reinitiation in F. tataricum MC; out of all examined PTMs, H4K16ac and H4K5ac revealed the highest variability during the course of passage in NC. Elevated levels of these modifications on day zero and day six for H4K16ac and H4K5ac respectively, seem to be correlated with endoreplication peaks, the processes which are distinctive for this callus. To sum up, while F. esculentum EC displays the characteristics typical to irregular, disorganised callus tissue, F. tataricum MC behaves in a completely different way. The cyclical development, disintegration and reinitiation of PECCs allowed to examine epigenetic changes and associate them with these processes. Comprehending how the chromatin reprogramming underlies the changes in the cell fate can provide a way for future manipulation towards the crop improvement, since once established during the callus formation, those modifications can be inherited through the epigenetic allele’s transmission and become strategies for the crop manipulation in the future (Hemenway and Gehring 2023; Lee and Seo 2018). In the future study, annotation of the genes involved in the process of dedifferentiation/re-differentiation switch, their expression and their epigenetic status in connection of the PTMs should be considered.
Data availability
All data generated or analysed during this study are included in this published article.
Abbreviations
- 5mC:
-
5-Methylcytosine
- H3K18ac:
-
Acetylation of Histone H3 on lysine 18
- H4K16ac:
-
Acetylation of Histone H4 on lysine 16
- H4K12ac:
-
Acetylation of Histone H4 on lysine 12
- H4K5ac:
-
Acetylation of Histone H4 on lysine 5
- H3K4me2:
-
Dimethylation of Histone H3 on lysine 4
- H3K4me3:
-
Trimethylation of Histone H3 on lysine 4
- H3K36me3:
-
Trimethylation of Histone H3 on lysine 36
- PTMs:
-
Post-Translational Modifications
- SE:
-
Somatic embryogenesis
- TBO:
-
Toluidine Blue O
References
Adachi T, Yamaguchi A, Miike Y, Hoffmann F (1989) Plant regeneration from protoplasts of common buckwheat (Fagopyrum esculentum). Plant Cell Rep 8:247–250
Akulov AN, Gumerova EA & Rumyantseva NI (2018) Cell cultures of Fagopyrum tataricum as a source of biologically active phenolic compounds. Buckwheat Germplasm in the World. 259–270
Avivi Y, Morad V, Ben-Meir H, Zhao J, Kashkush K, Tzfira T, Citovsky V, Grafi G (2004) Reorganization of specific chromosomal domains and activation of silent genes in plant cells acquiring pluripotentiality. Dev Dyn 230(1):12–22. https://doi.org/10.1002/dvdy.20006
Ayyappan V, Kalavacharla V, Thimmapuram J, Bhide KP, Sripathi VR, Smolinski TG, Manoharan M, Thurston Y, Todd A, Kingham B (2015) Genome-wide profiling of histone modifications (H3K9me2 and H4K12ac) and gene expression in rust (Uromyces appendiculatus) inoculated common bean (Phaseolus vulgaris L.). PLoS One 10(7):e0132176. https://doi.org/10.1371/journal.pone.0132176
Bednarek PT, Orłowska R (2019) Plant tissue culture environment as a switch-key of (epi)genetic changes. Plant Cell Tiss Org 140(2):245–257. https://doi.org/10.1007/s11240-019-01724-1
Bennett M, Cleaves K, Hewezi T (2021) Expression patterns of DNA methylation and demethylation genes during plant development and in response to phytohormones. Int J Mo Sci 22(18):9681. https://doi.org/10.3390/ijms22189681
Berbec A, Doroszewska T (1999) Regeneration in vitro of three cultivars of buckwheat (Fagopyrum esculentum Moench.) as affected by medium compositron. Fagopyrum 16:49–52
Betekhtin A, Rojek M, Jaskowiak J, Milewska-Hendel A, Kwasniewska J, Kostyukova Y, Kurczynska E, Rumyantseva N, Hasterok R (2017) Nuclear genome stability in long-term cultivated callus lines of Fagopyrum tataricum (L.) Gaertn. PLoS One 12(3):e0173537. https://doi.org/10.1371/journal.pone.0173537
Betekhtin A, Pinski A, Milewska-Hendel A, Kurczynska E, Hasterok R (2019) Stability and instability processes in the calli of Fagopyrum tataricum that have different morphogenic potentials. Plant Cell Tiss Org 137(2):343–357. https://doi.org/10.1007/s11240-019-01575-w
Birnbaum KD, Roudier F (2017) Epigenetic memory and cell fate reprogramming in plants. Regeneration 4(1):15–20. https://doi.org/10.1002/reg2.73
Birnbaum KD, Sanchez Alvarado A (2008) Slicing across kingdoms: regeneration in plants and animals. Cell 132(4):697–710. https://doi.org/10.1016/j.cell.2008.01.040
Braszewska-Zalewska A, Bernas T, Maluszynska J (2010) Epigenetic chromatin modifications in Brassica genomes. Genome 53(3):203–210. https://doi.org/10.1139/g09-088
Braszewska-Zalewska A, Dziurlikowska A, Maluszynska J (2012) Histone H3 methylation patterns in Brassica nigra, Brassica juncea, and Brassica carinata species. Genome 55(1):68–74. https://doi.org/10.1139/g11-076
Braszewska-Zalewska AJ, Wolny EA, Smialek L, Hasterok R (2013) Tissue-specific epigenetic modifications in root apical meristem cells of Hordeum vulgare. PLoS One 8(7):e69204. https://doi.org/10.1371/journal.pone.0069204
Chakrabarty D, Yu KW, Paek KY (2003) Detection of DNA methylation changes during somatic embryogenesis of Siberian ginseng (Eleuterococcus senticosus). Plant Sci 165(1):61–68. https://doi.org/10.1016/s0168-9452(03)00127-4
Cheng K, Xu Y, Yang C, Ouellette L, Niu L, Zhou X, Chu L, Zhuang F, Liu J, Wu H, Charron JB, Luo M (2020) Histone tales: lysine methylation, a protagonist in Arabidopsis development. J Exp Bot 71(3):793–807. https://doi.org/10.1093/jxb/erz435
Ci H, Li C, Aung TT, Wang S, Yun C, Wang F, Ren X, Zhang X (2022) A Comparative transcriptome analysis reveals the molecular mechanisms that underlie somatic embryogenesis in Peaonia ostii “Fengdan.” Int J Mol Sci 23(18):10595. https://doi.org/10.3390/ijms231810595
Ckurshumova W, Berleth T (2015) Overcoming recalcitrance - Auxin response factor functions in plant regeneration. Plant Signal Behav 10(7):e993293. https://doi.org/10.4161/15592324.2014.993293
Debeaujon I, Nesi N, Perez P, Devic M, Grandjean O, Caboche M, Lepiniec L (2003) Proanthocyanidin-accumulating cells in Arabidopsis testa: regulation of differentiation and role in seed development. Plant Cell 15(11):2514–2531. https://doi.org/10.1105/tpc.014043
Desvoyes B, Fernandez-Marcos M, Sequeira-Mendes J, Otero S, Vergara Z, Gutierrez C (2014) Looking at plant cell cycle from the chromatin window. Front Plant Sci 5:369. https://doi.org/10.3389/fpls.2014.00369
Feher A (2019) Callus, dedifferentiation, totipotency, somatic embryogenesis: what these terms mean in the era of molecular plant biology? Front Plant Sci 10:536. https://doi.org/10.3389/fpls.2019.00536
Fei Y, Wang LX, Fang ZW, Liu ZX (2019) Somatic embryogenesis and plant regeneration from cotyledon and hypocotyl explants of Fagopyrum esculentum Moench lpls mutant. Agronomy 9(11):768. https://doi.org/10.3390/agronomy9110768
Fraga HP, Vieira LN, Caprestano CA, Steinmacher DA, Micke GA, Spudeit DA, Pescador R, Guerra MP (2012) 5-Azacytidine combined with 2,4-D improves somatic embryogenesis of Acca sellowiana (O. Berg) Burret by means of changes in global DNA methylation levels. Plant Cell Rep 31(12):2165–76. https://doi.org/10.1007/s00299-012-1327-8
Gamborg OL, Miller RA, Ojima K (1968) Nutrient requirements of suspension cultures of soybean root cells. Exp Cell Res 50(1):151–158. https://doi.org/10.1016/0014-4827(68)90403-5
Gao L, Liu J, Liao L, Gao A, Njuguna BN, Zhao C, Zheng B, Han Y (2023) Callus induction and adventitious root regeneration of cotyledon explants in peach trees. Horticulturae 9(8):850. https://doi.org/10.3390/horticulturae9080850
Ghosh A, Igamberdiev AU, Debnath SC (2021) Tissue culture-induced DNA methylation in crop plants: a review. Mol Biol Rep 48(1):823–841. https://doi.org/10.1007/s11033-020-06062-6
Han Z, Crisp PA, Stelpflug S, Kaeppler SM, Li Q, Springer NM (2018) Heritable epigenomic changes to the maize methylome resulting from tissue culture. Genetics 209(4):983–995. https://doi.org/10.1534/genetics.118.300987
Hao J, Pei Y, Qu Y & Zheng C Study on callus differentiation conditions of common buckwheat. In: 7th International Symposium on Buckwheat, Winnipeg, MB, Canada,, 12–14 August 1998. p 12–14
Hemenway EA, Gehring M (2023) Epigenetic regulation during plant development and the capacity for epigenetic memory. Annu Rev Plant Biol 74:87–109. https://doi.org/10.1146/annurev-arplant-070122-025047
Horstman A, Li M, Heidmann I, Weemen M, Chen B, Muino JM, Angenent GC, Boutilier K (2017) The BABY BOOM transcription factor activates the LEC1-ABI3-FUS3-LEC2 network to induce somatic embryogenesis. Plant Physiol 175(2):848–857. https://doi.org/10.1104/pp.17.00232
Hu H, Du J (2022) Structure and mechanism of histone methylation dynamics in Arabidopsis. Curr Opin Plant Biol 67:102211. https://doi.org/10.1016/j.pbi.2022.102211
Inacio V, Martins MT, Graca J, Morais-Cecilio L (2018) Cork oak young and traumatic periderms show PCD typical chromatin patterns but different chromatin-modifying genes expression. Front Plant Sci 9:1194. https://doi.org/10.3389/fpls.2018.01194
Ishihara H, Sugimoto K, Tarr PT, Temman H, Kadokura S, Inui Y, Sakamoto T, Sasaki T, Aida M, Suzuki T, Inagaki S, Morohashi K, Seki M, Kakutani T, Meyerowitz EM, Matsunaga S (2019) Primed histone demethylation regulates shoot regenerative competency. Nat Commun 10(1):1786. https://doi.org/10.1038/s41467-019-09386-5
Jasencakova Z, Meister A, Walter J, Turner BM, Schubert I (2000) Histone H4 acetylation of euchromatin and heterochromatin is cell cycle dependent and correlated with replication rather than with transcription. Plant Cell 12:2087–2100
Jasencakova Z, Meister A, Schubert I (2001) Chromatin organization and its relation to replication and histone acetylation during the cell cycle in barley. Chromosoma 110(2):83–92. https://doi.org/10.1007/s004120100132
Jasencakova Z, Soppe WJ, Meister A, Gernand D, Turner BM, Schubert I (2003) Histone modifications in Arabidopsis- high methylation of H3 lysine 9 is dispensable for constitutive heterochromatin. Plant J 33(3):471–480. https://doi.org/10.1046/j.1365-313x.2003.01638.x
Ji L, Mathioni SM, Johnson S, Tucker D, Bewick AJ, Do Kim K, Daron J, Slotkin RK, Jackson SA, Parrott WA, Meyers BC, Schmitz RJ (2019) Genome-wide reinforcement of DNA methylation occurs during somatic embryogenesis in soybean. Plant Cell 31(10):2315–2331. https://doi.org/10.1105/tpc.19.00255
Jiang D, Kong NC, Gu X, Li Z, He Y (2011) Arabidopsis COMPASS-like complexes mediate histone H3 lysine-4 trimethylation to control floral transition and plant development. PLoS Genet 7(3):e1001330. https://doi.org/10.1371/journal.pgen.1001330
Jin H, Jia JF, Hao JG (2002) Efficient plant regeneration in vitro in buckwheat. Plant Cell Tiss Org 69:293–295
Kaeppler SM, Keppler HF, Rhee Y (2000) Epigenetic aspects of somaclonal variation in plants. Plant Mol Biol 43:179–188
Kamalova GV, Akulov AN, Rumyantseva NI (2009) Comparison of redox state of cells of tatar buckwheat morphogenic calluses and non-morphogenic calluses obtained from them. Biochemistry (mosc) 74(6):686–694. https://doi.org/10.1134/s0006297909060145
Kassambara A (2022) 'ggplot2' Based Publication Ready Plots, R package version 0.4.0, <https://CRAN.R-project.org/package=ggpubr>. R package version 0.4.0 edn
Kim JY, Yang W, Forner J, Lohmann JU, Noh B, Noh YS (2018) Epigenetic reprogramming by histone acetyltransferase HAG1/AtGCN5 is required for pluripotency acquisition in Arabidopsis. EMBO J. https://doi.org/10.15252/embj.201798726
Kouzarides T (2007) Chromatin modifications and their function. Cell 128(4):693–705. https://doi.org/10.1016/j.cell.2007.02.005
Kwon SJ, Han MH, Huh YS, Roy SK, Lee CW, Woo SH (2013) Plantlet regeneration via somatic embryogenesis from hypocotyls of common buckwheat (Fagopyrum esculentum Moench.). Korean J Crop Sci 58(4):331–335. https://doi.org/10.7740/kjcs.2013.58.4.331
Lee K, Seo PJ (2018) Dynamic epigenetic changes during plant regeneration. Trends Plant Sci 23(3):235–247. https://doi.org/10.1016/j.tplants.2017.11.009
Lee K, Park OS, Jung SJ, Seo PJ (2016) Histone deacetylation-mediated cellular dedifferentiation in Arabidopsis. J Plant Physiol 191:95–100. https://doi.org/10.1016/j.jplph.2015.12.006
Lee KP, Park O-S, Seo PJ (2017) Arabidopsis ATXR2 deposits H3K36me3 at the promoters of LBD genes to facilitate cellular dedifferentiation. Sic Signal 10:eaan0316
Lee K, Park OS, Choi CY, Seo PJ (2019) ARABIDOPSIS TRITHORAX 4 facilitates shoot identity establishment during the plant regeneration process. Plant Cell Physiol 60(4):826–834. https://doi.org/10.1093/pcp/pcy248
Leljak-Levanic D, Bauer N, Mihaljevic S, Jelaska S (2004) Changes in DNA methylation during somatic embryogenesis in Cucurbita pepo L. Plant Cell Rep 23(3):120–127. https://doi.org/10.1007/s00299-004-0819-6
Lestari R, Rio M, Martin F, Leclercq J, Woraathasin N, Roques S, Dessailly F, Clement-Vidal A, Sanier C, Fabre D, Melliti S, Suharsono S, Montoro P (2018) Overexpression of Hevea brasiliensis ethylene response factor HbERF-IXc5 enhances growth and tolerance to abiotic stress and affects laticifer differentiation. Plant Biotechnol J 16(1):322–336. https://doi.org/10.1111/pbi.12774
Li HL, Guo D, Zhu JH, Wang Y, Peng SQ (2017) Identification and expression analysis of genes involved in histone acetylation in Hevea brasiliensis. Tree Genet Genomes. https://doi.org/10.1007/s11295-017-1178-0
Li X, Han JD, Fang YH, Bai SN, Rao GY (2017) Expression analyses of embryogenesis-associated genes during somatic embryogenesis of Adiantum capillus-veneris L. in vitro: new insights into the evolution of reproductive organs in land plants. Front Plant Sci 8:658. https://doi.org/10.3389/fpls.2017.00658
Li HL, Guo D, Zhu JH, Wang Y, Peng SQ (2020) Identification of histone methylation modifiers and their expression patterns during somatic embryogenesis in Hevea brasiliensis. Genet Mol Biol 43(1):e20180141. https://doi.org/10.1590/1678-4685-GMB-2018-0141
Ma J, Li Q, Zhang L, Cai S, Liu Y, Lin J, Huang R, Yu Y, Wen M, Xu T (2022) High auxin stimulates callus through SDG8-mediated histone H3K36 methylation in Arabidopsis. J Integr Plant Biol 64(12):2425–2437. https://doi.org/10.1111/jipb.13387
Machczyńska J, Orłowska R, Mańkowski DR, Zimny J, Bednarek PT (2014) DNA methylation changes in triticale due to in vitro culture plant regeneration and consecutive reproduction. Plant Cell Tiss Org 119(2):289–299. https://doi.org/10.1007/s11240-014-0533-1
Makhloufi E, Yousfi FE, Marande W, Mila I, Hanana M, Berges H, Mzid R, Bouzayen M (2014) Isolation and molecular characterization of ERF1, an ethylene response factor gene from durum wheat (Triticum turgidum L. subsp. durum), potentially involved in salt-stress responses. J Exp Bot 65(22):6359–71. https://doi.org/10.1093/jxb/eru352
Mendiburu F (2021) Statistical procedures for agricultural research, R package version 1.3–5, <https://CRAN.R-project.org/package=agricolae>. In. https://CRAN.R-project.org/package=agricolae
Miguel C, Marum L (2011) An epigenetic view of plant cells cultured in vitro: somaclonal variation and beyond. J Exp Bot 62(11):3713–3725. https://doi.org/10.1093/jxb/err155
Monja-Mio KM, Quiroz-Moreno A, Herrera-Herrera G, Montero-Muñoz JL, Sánchez-Teyer F, Robert ML (2018) Analysis of two clonal lines (embryogenic and non-embryogenic) of Agave fourcroydes using AFLP and MSAP. A J Plant Sci 9(4):745–762. https://doi.org/10.4236/ajps.2018.94059
Moronczyk J, Braszewska A, Wojcikowska B, Chwialkowska K, Nowak K, Wojcik AM, Kwasniewski M, Gaj MD (2022) Insights into the histone acetylation-mediated regulation of the transcription factor genes that control the embryogenic transition in the somatic cells of Arabidopsis. Cells 11(5):863. https://doi.org/10.3390/cells11050863
Neelakandan AK, Wang K (2012) Recent progress in the understanding of tissue culture-induced genome level changes in plants and potential applications. Plant Cell Rep 31(4):597–620. https://doi.org/10.1007/s00299-011-1202-z
Nesković M, Vujicić R, Budimir S (1987) Somatic embryogenesis and bud formation from immature embryos of buckwheat (Fagopyrum esculentum Moench.). Plant Cell Rep 6:423–426
Nic-Can GI, Lopez-Torres A, Barredo-Pool F, Wrobel K, Loyola-Vargas VM, Rojas-Herrera R, De-la-Pena C (2013) New insights into somatic embryogenesis: LEAFY COTYLEDON1, BABY BOOM1 and WUSCHEL-RELATED HOMEOBOX4 are epigenetically regulated in Coffea canephora. PLoS One 8(8):e72160. https://doi.org/10.1371/journal.pone.0072160
Park SY, Murthy HN, Chakrabarthy D, Paek KY (2008) Detection of epigenetic variation in tissue-culture-derived plants of Doritaenopsis by methylation-sensitive amplification polymorphism (MSAP) analysis. In Vitro Cell Dev Biol Plant 45(1):104–108. https://doi.org/10.1007/s11627-008-9166-6
Rodriguez-Sanz H, Moreno-Romero J, Solis MT, Kohler C, Risueno MC, Testillano PS (2014) Changes in histone methylation and acetylation during microspore reprogramming to embryogenesis occur concomitantly with Bn HKMT and Bn HAT expression and are associated with cell totipotency, proliferation, and differentiation in Brassica napus. Cytogenet Genome Res 143(1–3):209–218. https://doi.org/10.1159/000365261
Roudier F, Ahmed I, Berard C, Sarazin A, Mary-Huard T, Cortijo S, Bouyer D, Caillieux E, Duvernois-Berthet E, Al-Shikhley L, Giraut L, Despres B, Drevensek S, Barneche F, Derozier S, Brunaud V, Aubourg S, Schnittger A, Bowler C, Martin-Magniette ML, Robin S, Caboche M, Colot V (2011) Integrative epigenomic mapping defines four main chromatin states in Arabidopsis. EMBO J 30(10):1928–1938. https://doi.org/10.1038/emboj.2011.103
Rumyantseva NI, Sergejewa NB, Khakimova LE, Salnikov BB, Gumerova EA, Lozowaja BB (1989) Organogenesis and somatic embryogenesis in tissue culture of two buckwheat species. Russ J Plant Physl 36:187–194
Rumyantseva NI, Samaj J, Ensikat H-J, Sal’nikov VV, Kostyukova YA, BaluSka F, Volkmann D (2003) Changes in the extracellular matrix surface network during cyclic reproduction of proembryonic cell complexes in the Fagopyrum tataricum (L.) Gaertn callus. Dokl Biol Sci 391:375–378
Rumyantseva NI, Akulov AN, Mukhitov AR (2004) Extracellular polymers in callus cultures of Fagopyrum tataricum (L.) Gaertn. with different morphogenic activities: time courses during the culture cycle. Appl Biochem Microbiol 40(5):494–500
Rumyantseva NI, Sal’nikov VV, Lebedeva VV (2005) Structural changes of cell surface in callus of Fagopyrum esculentum Moench. during induction of morphogenesis. Russ J Plant Physiol 52(3):381–387
Rymen B, Kawamura A, Lambolez A, Inagaki S, Takebayashi A, Iwase A, Sakamoto Y, Sako K, Favero DS, Ikeuchi M, Suzuki T, Seki M, Kakutani T, Roudier F, Sugimoto K (2019) Histone acetylation orchestrates wound-induced transcriptional activation and cellular reprogramming in Arabidopsis. Commun Biol 2:404. https://doi.org/10.1038/s42003-019-0646-5
Sakar D, Naik PS (2000) Phloroglucinol enhances growth and rate of axillary shoot proliferation in potato shoot tip cultures in vitro. Plant Cell Tiss Org 60(2):139–149
Simsek Geyik M, Yazicilar B, Bezirganoglu İ (2022) Microscopic and physiological analysis of somatic embryos under in vitro culture in Triticale. Icontech Int J 6(1):73–80. https://doi.org/10.46291/ICONTECHvol6iss1pp73-80
Souter M, Lindsey K (2000) Polarity and signalling in plant embryogenesis. J Exp Bot 51(347):971–983. https://doi.org/10.1093/jexbot/51.347.971
Springer NM, Schmitz RJ (2017) Exploiting induced and natural epigenetic variation for crop improvement. Nat Rev Gen 18(9):563–575. https://doi.org/10.1038/nrg.2017.45
Srejovic V, Neskovic M (1981) Regeneration of plants from cotyledon fragments of buckwheat (Fagopyrum esculentum Moench.). Z Pflanzenphysiol 104:37–42
Stroud H, Ding B, Simon SA, Feng S, Bellizzi M, Pellegrini M, Wang GL, Meyers BC, Jacobsen SE (2013) Plants regenerated from tissue culture contain stable epigenome changes in rice. Elife 2:e00354. https://doi.org/10.7554/eLife.00354
Takahata YJ, Jumonji E (1985) Plant regeneration from hypocotyl section and callus in buckwheat (Fagopyrum esculentum Moench.). Ann Rep Fac Educ 45:137–142
Team RC (2022a) R: A Language and environment for statistical computing, Vienna, Austria: R foundation for statistical computing. URL: https://www.R-project.org/. In. https://www.R-project.org/ Accessed 1.02.2023
Team RS (2022b) RStudio: Integrated development environment for R; RStudio, PBC, Boston, MA. URL: http://www.rstudio.com/. In. http://www.rstudio.com/ Accessed 1.02.2023
Tomasiak A, Zhou M, Betekhtin A (2022) Buckwheat in tissue culture research: current status and future perspectives. Int J Mol Sci 23(4):2298. https://doi.org/10.3390/ijms23042298
Us-Camas R, Rivera-Solís G, Duarte-Aké F, De-la-Peña C (2014) In vitro culture: an epigenetic challenge for plants. Plant Cell Tiss Org 118(2):187–201. https://doi.org/10.1007/s11240-014-0482-8
Viejo M, Rodriguez R, Valledor L, Perez M, Canal MJ, Hasbun R (2010) DNA methylation during sexual embryogenesis and implications on the induction of somatic embryogenesis in Castanea sativa Miller. Sex Plant Reprod 23(4):315–323. https://doi.org/10.1007/s00497-010-0145-9
Wickham H (2022) Easily install and load the 'Tidyverse' R package version 1.2.0, <https://CRAN.R-project.org/package=tidyverse>. In. https://cran.r-project.org/package=tidyverse
Wickramasuriya AM, Dunwell JM (2015) Global scale transcriptome analysis of Arabidopsis embryogenesis in vitro. BMC Genomics 16(1):301. https://doi.org/10.1186/s12864-015-1504-6
Williams L, Zhao J, Morozova N, Li Y, Avivi Y, Grafi G (2003) Chromatin reorganization accompanying cellular dedifferentiation is associated with modifications of histone H3, redistribution of HP1, and activation of E2F-target genes. Dev Dyn 228(1):113–120. https://doi.org/10.1002/dvdy.10348
Wolny E, Braszewska-Zalewska A, Hasterok R (2014) Spatial distribution of epigenetic modifications in Brachypodium distachyon embryos during seed maturation and germination. PLoS One 9(7):e101246. https://doi.org/10.1371/journal.pone.0101246
Xu L, Huang H (2014) Genetic and epigenetic controls of plant regeneration. Curr Top Dev Biol 108:1–33. https://doi.org/10.1016/B978-0-12-391498-9.00009-7
Yamamoto N, Kobayashi H, Togashi T, Mori Y, Kikuchi K, Kuriyama K, Tokuji Y (2005) Formation of embryogenic cell clumps from carrot epidermal cells is suppressed by 5-azacytidine, a DNA methylation inhibitor. J Plant Physiol 162(1):47–54. https://doi.org/10.1016/j.jplph.2004.05.013
Yamane Y (1985) Induced differentiation of buckwheat plants from subcultured calluses in vitro. Jpn J Genet 49:139–146
Yao X, Feng H, Yu Y, Dong A, Shen WH (2013) SDG2-mediated H3K4 methylation is required for proper Arabidopsis root growth and development. PLoS One 8(2):e56537. https://doi.org/10.1371/journal.pone.0056537
Yu Y, Bu Z, Shen W-H, Dong A (2009) An update on histone lysine methylation in plants. Prog Nat Sci 19(4):407–413. https://doi.org/10.1016/j.pnsc.2008.07.015
Zakrzewski F, Schmidt M, Van Lijsebettens M, Schmidt T (2017) DNA methylation of retrotransposons, DNA transposons and genes in sugar beet (Beta vulgaris L.). Plant J 90(6):1156–1175. https://doi.org/10.1111/tpj.13526
Zhang T, Cooper S, Brockdorff N (2015) The interplay of histone modifications-writers that read. EMBO Rep 16(11):1467–1481. https://doi.org/10.15252/embr.201540945
Zhang H, Guo F, Qi P, Huang Y, Xie Y, Xu L, Han N, Xu L, Bian H (2020) OsHDA710-mediated histone deacetylation regulates callus formation of rice mature embryo. Plant Cell Physiol 61(9):1646–1660. https://doi.org/10.1093/pcp/pcaa086
Zhao J, Morozova N, Williams L, Libs L, Avivi Y, Grafi G (2001) Two phases of chromatin decondensation during dedifferentiation of plant cells: distinction between competence for cell fate switch and a commitment for S phase. J Biol Chem 276(25):22772–22778. https://doi.org/10.1074/jbc.M101756200
Zhao N, Zhang K, Wang C, Yan H, Liu Y, Xu W, Su Z (2020) Systematic analysis of differential H3K27me3 and H3K4me3 deposition in callus and seedling reveals the epigenetic regulatory mechanisms involved in callus formation in rice. Front Genet 11:766. https://doi.org/10.3389/fgene.2020.00766
Zheng B, Liu J, Gao A, Chen X, Gao L, Liao L, Luo B, Ogutu CO, Han Y (2022) Epigenetic reprogramming of H3K27me3 and DNA methylation during leaf-to-callus transition in peach. Hortic Res 9:uhac132. https://doi.org/10.1093/hr/uhac132
Acknowledgements
This research was funded by the National Science Centre Poland, Project OPUS19 grant number 2020/37/B/NZ9/01499.
Author information
Authors and Affiliations
Contributions
AB acquired funding, AB and ABr, conceived, designed the study and supervised its execution, AT carried out the experiments with supervision from KSC and ABr, LSB prepared the statistical analysis, AT, AB, ABr analysed the obtained results, AT drafted the manuscript with contributions from all of the authors, AB, ABr performed review and editing. All of the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The use of all plant materials in this study complies with relevant institutional, national, and international guidelines and legislation. Seeds of F. tataricum (sample k-17) are from the collections of the N. I. Vavilov Institute of Plant Genetic Resources, Saint Petersburg, Russia. Obtained seeds were multiplied in Plant Cytogenetic and Molecular Biology Group Institute of Biology, Biotechnology and Environmental Protection, Faculty of Natural Sciences, University of Silesia in Katowice, Poland. F. tataricum sample k-17 is a common cultivated landrace of F. tataricum and seeds are available on request from authors of the publication. Seeds of F. esculentum genotype Panda are commercially available and were purchased from the Malopolska Plant Breeding Company.
Additional information
Communicated by Danny Geelen.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tomasiak, A., Sala-Cholewa, K., Berg, L.S. et al. Global epigenetic analysis revealed dynamic fluctuations in levels of DNA methylation and histone modifications in the calli of Fagopyrum with different capacity for morphogenesis. Plant Cell Tiss Organ Cult 155, 743–757 (2023). https://doi.org/10.1007/s11240-023-02595-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-023-02595-3