Abstract
Feline leukemia virus (FeLV) is responsible for feline leukemia syndrome in domestic cats. The prevention and control of disease caused by FeLV are primarily based on vaccination and identification and isolation of infected subjects. Antigen diagnostic methods, which are the most widely used in clinical practices, can be associated to molecular tests to characterize the FeLV detected. In this study, a quantitative SYBR Green Real-Time PCR (qPCR) assay was used to detect FeLV proviral DNA in blood samples from antigen positive cats referred to a veterinary teaching hospital in Northern Italy in 2018–2021. To genetically characterize the identified viruses, a portion of the viral envelope (env) gene was amplified using six different end-point PCRs and sequenced. Twenty-two of 26 (84.6%) cats included in the study tested positive by qPCR assay. This suggests a high performance of the qPCR adopted but further studies are required to investigate the cause of discordant results between the antigen test and qPCR in four cats. From env gene analysis, 15/22 qPCR-positive cats were infected by FeLV subtype A and 5/15 shown coinfection with subtype B.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Feline leukemia virus (FeLV) is an enveloped RNA virus belonging to the genus Gammaretrovirus occurring worldwide in domestic and small wild cats. In Italy, a recent study estimated a prevalence of FeLV infection of 5.7% (Studer et al. 2019). FeLV genome can integrate into the host cell genome as provirus and may affects the cellular replication, leading to neoplastic transformation and immunosuppression. Endogenous FeLV (enFeLV), in cats’ genome, have been correlated with resistance against exogenous virus (exFeLV) infections and play a crucial role in arising FeLV recombinant subtypes (Stewart et al. 1986; Roy-Burman 1996). The longest known FeLV subtypes are FeLV-A, FeLV-B, and FeLV-C. More recently, FeLV subtypes D, T and TG35 have been identified (Chiu et al. 2018). FeLV-A is the most widespread since it is the only subtype transmissible from cat to cat and it is present in every infected cat.
The clinical outcome in cats is determined by a combination of viral and host factors such as the FeLV subtypes involved, the age of the cat at the time of infection and its immune response. According to this, FeLV infection has four possible outcomes: progressive, regressive, abortive and rarely focal or atypical infection (Little et al. 2020).
The understanding of the infectious status of cats is important before vaccination or to identify and isolate infected subjects within naive multi-cat environments. Point-of-care (PoC) tests based on the detection of p27 antigen of FeLV are widely used to identify infected cats. PoC tests are very performant to detect cats with progressive and focal (atypical) infection, but they show limitations in case of the regressive infections for which the use of molecular methods like PCR assay to detect proviral DNA is recommended (Hofmann-Lehmann and Hartmann 2020). Moreover, the diagnosis of FeLV is very difficult because of the complex pathophysiology and evolution that the infection can have, for these reasons it is not possible to use a single test to determine the status of FeLV infection (Beall et al. 2021; Giselbrecht et al. 2023).
The aims of this study were to detect FeLV provirus in whole blood samples of antigen-positive cats from northern Italy and to genetically characterize the identified viruses.
Materials and methods
Cats tested positive for the presence of FeLV specific antigens were included in the study. All samples were collected between January 2018 and September 2021 at the Veterinary University Hospital (VUH) of the University of Bologna. Plasma, serum, or whole blood samples were tested for FeLV p27 antigen and Feline immunodeficiency virus (FIV) antibodies using a commercial PoC enzyme linked immunosorbent assay (ELISA) based test (SNAP FIV/FeLV Combo Plus test, IDEXX, USA). PoC tests were carried out within one hour from the sampling and the samples were stored at − 20 °C after examination.
Signalment data, clinical signs and clinicopathological findings of enrolled cats were retrieved from medical records. The enrolled cats were grouped as: (i) “symptomatic cats” with clinical signs or clinicopathological abnormalities referable to FeLV infection (SC group) and (ii) “asymptomatic cats” without clinical signs or clinicopathological abnormalities referable to FeLV infection (AC group).
Genomic DNA extraction from K3EDTA blood samples was carried out starting from 200 µl of sample and the DNA was eluited in 100 µl of elution buffer, using a commercially available kit (NucleoSpin Tissue Kit, Macherey-Nagel, Germany).
The presence of FeLV DNA was screened by using a quantitative SYBR Green Real-Time PCR (qPCR) assay (Tandon et al. 2005) (Online Resource 1). The reaction was performed using the PowerUp SYBR Green master mix (Thermo Fisher Scientific, USA) following manufacturer’s instructions. FeLV DNA copies number determination was carried out by absolute quantification using tenfold dilutions of a DNA standard plasmid containing 468 nucleotides sequence from the U3 region (Online Resource 1), cloned into the TOPO TA Cloning vector (Invitrogen, USA). In each reaction, seven tenfold dilutions of the DNA standard plasmid and samples were amplified in duplicates together with a no template control.
Six previously published end-point PCR assays were used to amplify fragments of different length of the FeLV env gene from the qPCR-positive cats. End-point PCRs 1 to 4 were reported to target FeLV A, B and C while PCRs 5 and 6 were reported to be specific only for FeLV B (Erbeck et al. 2021; Watanabe et al. 2013). Reactions were carried out with the proofreading Phusion Hot Start II High-Fidelity DNA Polymerase (Thermo Fisher Scientific, USA), according to the manufacturer’s instructions.
The env gene PCR products were sequenced by Sanger method using the forward and reverse primers adopted for the amplification plus primers designed for sequencing (Online Resource 1). The sequences were assembled with CodonCode Aligner software, aligned with 34 reference sequences retrieved from GenBank database (Online Resource 2) and translated into amino acid sequences using the ClustalW method implemented in the BioEdit 7.0.5 software. Nucleotide and amino acid similarity of obtained sequences with reference sequences and potential recombination breakpoints were evaluated to determine which FeLV subtype the identified viruses belonged to. The assembled nucleotide sequences were analyzed using the BLAST web interface (https://blast.ncbi.nlm.nih.gov/Blast.cgi). Phylogeny was carried out on env gene nucleotide sequence alignment using MEGA 11 version 11.0.10 (Tamura et al. 2021).
Data were evaluated using standard descriptive statistics and reported as median and range. Categorical data such as year of sampling, sex, breed and antibodies test results were analyzed using the Fisher’s exact P-value test or Pearson’s Chi-squared test, while continuous data (age) were analyzed by the Mann-Whitney U test. Statistical significance was set at P < 0.05. Statistical analysis was carried out using the MedCalc Statistical Software version 16.8.4 (MedCalc Software bvba).
Results
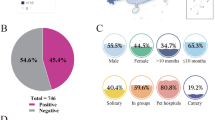
Twenty-six FeLV antigen-positive cats were included in the study. Signalment data, clinical signs, clinicopathological findings and FIV antibodies PoC test results are reported in Tables 1 and 2. Eighteen of 26 (69.2%) cats were included in the SC group and 8/26 (30.8%) were included in the AC group. No statistical association was found regarding the presence of clinical signs or clinicopathological abnormalities associated with FeLV infection and signalment data.
Twenty-two of 26 (84.6%) cats tested positive by qPCR (Table 2). The quantity of FeLV DNA varied between 2.8 and 3 × 106 copies of target amplicon/µL of extract, that correspond to a threshold cycle range from 36.5 to 15.4. Four cats tested negative by qPCR showing a discordance with the PoC test result (Lab IDs: 1128/19, 1126/20, 732/21, 868/21).
The 22 qPCR positive in U3 (LTR) cats showed at least one positive result in one of the six end-point PCR assays based on env gene. In particular, 19/22 cats tested positive by the end-point PCRs 1 to 4 and 15/22 tested positive by the end-point PCRs 5 and 6. Nucleotide sequences of the FeLV env gene amplified by end-point PCRs 1 to 4 were obtained from 15/22 qPCR-positive cats (Table 2), they were of about 1800 nucleotides in length. All these sequences showed an overall nucleotide identity of 92.1–99.6% between them and 89.9–99.9% with FeLV-A reference sequences. For 15/22 qPCR-positive cats, env gene nucleotide sequences were also obtained from amplicons of end-point PCRs 5 and 6 (Table 2), they were of about 1235 nucleotides in length. Ten of this 15 sequences were highly similar between them sharing a nucleotide identity of 95.5–100% with the reference sequences of the endogenous viruses. The remaining 5/15 sequences showed higher nucleotide variability (78–97%) and potentially recombination events between FeLV-A and enFeLV, compatible with FeLV-B viruses.
Phylogenetic analysis identified different clades, indicative of clusterization in FeLV subtype A, endogenous FeLV and FeLV recombinant subtype B, respectively (Fig. 1). Fifteen sequences obtained by end-point PCRs 1 to 4 clustered with the FeLV-A reference sequences from Europe, Japan and the USA. The ten highly similar sequences obtained by end-point PCRs 5 and 6 clustered with the enFeLV reference sequences. Four other sequences obtained by end-point PCRs 5 and 6 clustered with FeLV subtype B reference sequences in an intermediate position between FeLV-A and enFeLV clades while the one from cat 324/2021 grouped with but distant from other FeLV-A sequences obtained in this study.
Phylogenetic tree on the nucleotide sequences of env gene of FeLV. Phylogeny was carried out on FeLV sequences obtained in this study and 34 reference strains (Online Resource 2) using MEGA 11 version 11.0.10. Phylogenetic tree was constructed using Neighbor-Joining method and the Tamura-Nei model with gamma distribution. Statistical support was provided by bootstrapping with 1,000 replicates and values reported on respective branch. The scale bars indicate the estimated numbers of nucleotide substitutions. Identification of the sequences undergoes the following nomenclature: GenBank accession number, strain (only for sequences obtained in this study), country (CH: Switzerland, CN: China, IT: Italy, JP: Japan, UK: United Kingdom, US: United States of America), collection date (or date of database submission), and subtype. Marked in white: sequences obtained in this study belonging to FeLV subtype A. Marked in grey: sequences obtained in this study belonging to endogenous FeLV. Marked in black: sequences obtained in this study belonging to FeLV subtype B
Discussion
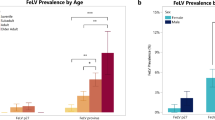
In this study, 26 FeLV antigen-positive cats were tested by molecular assays to confirm the state of infection and the molecular characterization of identified FeLV was performed determining the viral subtypes circulating in Northern Italy. Signalment and history of the cats included in the study partially reflect the risk categories reported in literature: indeed only 57.7% of cats were male (Hofmann-Lehmann et al. 2018). Our data showed higher FeLV prevalence in adult cats as already reported (Levy et al. 2006; Stavisky et al. 2017), confirming that infection should be expected in adult cats as a consequence of reactivation of a previous infection and not only in kittens (Studer et al. 2019). The majority of cats included in the study had clinical signs or clinicopathological abnormalities potentially associated with FeLV infection, probably depending on the investigated cat population: sick cats visiting the VUH (Burling et al. 2017). In our study four cats resulted also positive for FIV antibodies by the PoC test: three were grouped with AC cats while one had clinical manifestations referable to retroviral infection. Considering the Italian epidemiological situation reported in the literature (Battilani et al. 2022), a lower percentage of co-infected cats was expected. On the other hand, the number of cats included in our study and the inclusion criteria adopted, do not allow to draw epidemiological conclusions.
The performance of the qPCR assay adopted in this study was not affected by the modification of the original protocol in which a probe was used instead of the SYBR green (Tandon et al. 2005), in fact FeLV provirus was detected in 84.6% of blood samples examined. Whereas, four cats tested negative, possibly as consequence of a failure of the molecular test, in case of an atypical/focal infection (Giselbrecht et al. 2023) or false positive results of the rapid PoC test adopted. These four cats had anemia or precursor-targeted immune-mediated anemia which may lead to false positive PoC antigen test results (Izquierdo Robert et al. 2023). The cause of these false positive results has not yet been definitively ascertained, although a recent study reported discordant FeLV p27 immunoassay and PCR test results in 21 cats with hematologic disorders emphasizing the importance of follow-up PCR testing in particular clinical situation like blood dyscrasia (Kornya et al. 2023). To better investigate this discrepancy, other tests could be performed, such as a second type of PoC antigen test, a laboratory-based ELISA, or a reverse-transcriptase PCR (Hofmann-Lehmann and Hartmann 2020).
The env gene end-point PCR assays followed by Sanger sequencing allowed to detect FeLV-A in 15 cats. In five of these cats also FeLV-B was detected, a slightly lower prevalence than reported in literature (Phipps et al. 2000). Since FeLV-B arises de novo after multiple recombination events of enFeLV sequences with FeLV-A, the sequence variability of this subtype depends both on recombination site involved and FeLV-A sequence diversity, thus affecting the sensitivity of diagnostic methods. This may also be one of the reasons for the low specificity of the PCRs 5 and 6 (Watanabe et al. 2013), which were expected to amplify only FeLV-B, while most of the sequenced amplicons clustered with enFeLV. Genomic variability of enFeLV coupled with its significant nucleotidic identity with exogenous FeLV makes differentiation by PCR of these two forms challenging (Chiu et al. 2018). Phylogeny of the identified FeLV-B viruses was consistent with several recombination sites at a variety of positions within the 5’ region of the env gene. Indeed, four of the FeLV-B viruses were intermediate between FeLV-A and enFeLV, while the one from cat 324/2021 grouped with FeLV-A, probably because the recombination break point was very close to the N-terminal of the env gene sequence obtained. The common but varied sites of recombination in FeLV-B variants emerged in our study support findings of other studies (Watanabe et al. 2013; Erbeck et al. 2021). This result, together with the failure to obtain env gene nucleotide sequences from some viruses detected by qPCR, is consistent with extensive genetic variation in the surface glycoprotein (Cano-Ortiz et al. 2022; Erbeck et al. 2021; Ortega et al. 2020; Watanabe et al. 2013).
Conclusion
The detection of FeLV proviral DNA in 84.6% antigen-positive cats support the usefulness of qPCR assays in doubtful clinical cases. Validation of the qPCR assay on p27 antigen negative cats would be useful to evaluate its use in the diagnosis of regressive infections, particularly in blood donor cats (Nesina et al. 2015). Molecular investigation allowed to characterize 15 FeLV-A and five FeLV-B viruses, providing new information on subtypes circulating in northern Italy. With regard to the sequence variability of subtype B, it would be useful to develop a molecular tool to correctly identify this virus and proceed with further investigations to correlate this subtype with neoplasms.
Declarations.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files. The nucleotide sequences generated and analyzed during the current study are available in the International Nucleotide Sequence Database Collaboration repository (INSDC, http://www.insdc.org/) with the IDs: OR227248-OR227265, OR227267-OR227273, OR227275-OR227278 and OR227280.
References
Battilani M, Kaehler E, Tirolo A, Balboni A, Dondi F (2022) Clinicopathological findings in cats tested for feline immunodeficiency virus (FIV) and feline leukaemia virus (FeLV). Acta Veterinaria Beograd., 72 419–432. ISSN: 0567–8315, E-ISSN: 1820–7448. https://doi.org/10.2478/acve-2022-0034
Beall MJ, Buch J, Clark G, Estrada M, Rakitin A, Hamman NT, Frenden MK, Jefferson EP, Amirian ES, Levy JK (2021) Feline leukemia virus p27 Antigen Concentration and Proviral DNA load are Associated with Survival in naturally infected cats. Viruses 13302. https://doi.org/10.3390/v13020302
Burling AN, Levy JK, Scott HM, Crandall MM, Tucker SJ, Wood EG, Foster JD (2017) Seroprevalences of feline leukemia virus and feline immunodeficiency virus infection in cats in the United States and Canada and risk factors for seropositivity. J Am Vet Med Assoc 251:187–194. https://doi.org/10.2460/javma.251.2.187
Cano-Ortiz L, Tochetto C, Roehe PM, Franco AC, Junqueira DM (2022) Could phylogenetic analysis be used for Feline Leukemia Virus (FeLV) classification? Viruses. 14:249. https://doi.org/10.3390/v14020249
Chiu ES, Hooverm EA, VandeWoude S (2018) A retrospective examination of feline leukemia subgroup characterization: viral interference assays to deep sequencing. Viruses 10:29. https://doi.org/10.3390/v10010029
Erbeck K, Gagne RB, Kraberger S, Chiu ES, Roelke-Parker M, VandeWoude S (2021) Feline leukemia virus (FeLV) endogenous and exogenous recombination events result in multiple FeLV-B subtypes during natural infection. J Virol 95. https://doi.org/10.1128/jvi.00353-21
Giselbrecht J, Jähne S, Bergmann M, Meli ML, Pineroli B, Boenzli E, Teichmann-Knorrn S, Zablotski Y, Pennisi M-G, Layachi N, Serra R, Bo S, Hofmann-Lehmann R, Hartmann K (2023) Prevalence of different courses of feline leukaemia virus infection in four European countries. Viruses 15(8):1718. https://doi.org/10.3390/v15081718
Hofmann-Lehmann R, Hartmann K (2020) Feline leukaemia virus infection: a practical approach to diagnosis. J Feline Med Surg 22:831–846. https://doi.org/10.1177/1098612x20941785
Hofmann-Lehmann R, Gonczi, Riond B, Meli M, Willi B, Howard J, Schaarschmidt-Kiener D, Regli W, Gilli U, Boretti F (2018) Feline leukemia virus infection: importance and current situation in Switzerland. Schweiz Arch Tierheilkd 160:95–105. https://doi.org/10.17236/sat00146
Izquierdo Robert L, Puig J, Tumbarello M, Farigola M, Seth M, Mesa I, Bernabe LF (2023) A retrospective review of cats with suspected false positive results in point-of-care feline leukemia virus tests and concurrent immune-mediated anemia. J Am Vet Med Assoc 30:1–7. https://doi.org/10.2460/javma.23.02.0059
Kornya M, Bienzle D, Beeler-Marfisi J (2023) Discordant FeLV p27 immunoassay and PCR test results in 21 cats with hematologic disorders. J Feline Med Surg 25:1098612X231183297. https://doi.org/10.1177/1098612x231183297
Levy JK, Scott HM, Lachtara JL, Crawford PC (2006) Seroprevalence of feline leukemia virus and feline immunodeficiency virus infection among cats in North America and risk factors for seropositivity. J Am Vet Med Assoc 228:371–376. https://doi.org/10.2460/javma.228.3.371
Little S, Levy J, Hartmann K, Hofmann-Lehmann R, Hosie M, Olah G, Denis KS (2020) 2020 AAFP feline retrovirus testing and management guidelines. J Feline Med Surg 22:5–30. https://doi.org/10.1177/1098612x19895940
Nesina S, Katrin Helfer-Hungerbuehler A, Riond B, Boretti FS, Willi B, Meli ML, Grest P, Hofmann-Lehmann R (2015) Retroviral DNA-the silent winner: blood transfusion containing latent feline leukemia provirus causes infection and disease in naïve recipient cats. Retrovirology 12:105. https://doi.org/10.1186/s12977-015-0231-z
Ortega C, Valencia AC, Duque-Valencia J, Ruiz-Saenz J (2020) Prevalence and genomic diversity of Feline Leukemia Virus in privately owned and shelter cats in Aburrá Valley. Colombia Viruses 12:464. https://doi.org/10.3390/v12040464
Phipps AJ, Hayes KA, Al-dubaib M, Roy-Burman P, Mathes LE (2000) Inhibition of feline leukemia virus subgroup a infection by coinoculation with subgroup B. Virology 277:40–47. https://doi.org/10.1006/viro.2000.0606
Roy-Burman P (1996) Endogenous env elements: partners in generation of pathogenic feline leukemia viruses. Virus Genes 11:147–161
Stavisky J, Dean RS, Molloy MH (2017) Prevalence of and risk factors for FIV and FeLV infection in two shelters in the United Kingdom (2011–2012). Vet Rec 181:451. https://doi.org/10.1136/vr.103857
Stewart MA, Warnock M, Wheeler A, Wilkie N, Mullins JI, Onions DE, Neil JC (1986) Nucleotide sequences of a feline leukemia virus subgroup a envelope gene and long terminal repeat and evidence for the recombinational origin of subgroup B viruses. J Virol 58:825–834
Studer N, Lutz H, Saegerman C, Gönczi E, Meli ML, Boo G, Hartmann K, Hosie MJ, Moestl K, Tasker S, Belák S, Lloret A, Boucraut-Baralon C, Egberink HF, Pennisi MG, Truyen U, Frymus T, Thiry E, Marsilio F, Addie D, Hochleithner M, Tkalec F, Vizi Z, Brunetti A, Georgiev B, Ludwig-Begall LF, Tschuor F, Mooney CT, Eliasson C, Orro J, Johansen H, Juuti K, Krampl I, Kovalenko K, Šengaut J, Sobral C, Borska P, Kovaříková S, Hofmann-Lehmann R (2019) Pan-european study on the prevalence of the Feline Leukaemia Virus infection - reported by the European Advisory Board on Cat diseases (ABCD Europe). Viruses 11:993. https://doi.org/10.3390/v11110993
Tamura K, Stecher G, Kumar S (2021) MEGA11: Molecular Evolutionary Genetics Analysis version 11. Mol Biol Evol 38:3022–3027. https://doi.org/10.1093/molbev/msab120
Tandon R, Cattori V, Gomes-Keller MA, Meli ML, Golder MC, Lutz H, Hofmann-Lehmann R (2005) Quantitation of feline leukaemia virus viral and proviral loads by TaqMan real-time polymerase chain reaction. J Virol Methods 130:124–132. https://doi.org/10.1016/j.jviromet.2005.06.017
Watanabe S, Kawamura M, Odahara Y, Anai Y, Ochi H, Nakagawa S, Endo Y, Tsujimoto H, Nishigaki K (2013) Phylogenetic and structural diversity in the feline leukemia virus env gene. PLoS ONE 8. https://doi.org/10.1371/journal.pone.0061009
Funding
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Open access funding provided by Alma Mater Studiorum - Università di Bologna within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Contributions
L.G., V.F. and A. B. analyzed the results and wrote the paper. M.C.S., K.V. collected the samples and data. L.G., V.F., N.R., A.T., L.U. and M.M. performed the analysis. F.D. collected the samples and data. L. G., A.B., M.B. Conceived and designed the analysis. F. D: and M.B. Supervised the study. All authors reviewed and approved the article.
Corresponding author
Ethics declarations
Ethics approval
The animal study protocol was approved by the Animal Welfare Committee of the University of Bologna (ID 4475, 03/03/2023) that confirm that the research described in this manuscript does not fall within Directive 63/2010 of the European Parliament and of the Council on the protection of animals used for scientific purposes (transposed into Italian law by Legislative Decree 26/2014) and thus doesn’t require any authorization from the national competent Authorities. The study was not carried out on experimental animals. Only surplus material derived from blood samples taken by clinicians for diagnostic purposes following owner’s informed consent were used.
Informed consent
Informed consent was obtained from all owners of the animals involved in the study.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gallina, L., Facile, V., Roda, N. et al. Molecular investigation and genetic characterization of feline leukemia virus (FeLV) in cats referred to a veterinary teaching hospital in Northern Italy. Vet Res Commun 48, 2683–2689 (2024). https://doi.org/10.1007/s11259-024-10380-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11259-024-10380-6