Abstract
Genotype-environment interaction is pervasive in forest genetics. Delineation of spatial breeding zones (BZs) is fundamental for accommodating genotype-environment interaction. Here we developed a BZ classification pipeline for the forest tree Eucalyptus globulus in 2 Australian regions based on phenotypic, genomic, and pedigree data, as well on a detailed environmental characterization (“envirotyping”) and spatial mapping of BZs. First, the factor analytic method was used to model additive genetic variance and site–site genetic correlations (rB) in stem volume across 48 trials of 126,467 full-sib progeny from 2 separate breeding programs. Thirty-three trials were envirotyped using 145 environmental variables (EVs), involving soil and landscape (71), climate (73), and management (1) EVs. Next, sparse partial least squares-discriminant analysis was used to identify EVs that were required to predict classification of sites into 5 non-exclusive BZ classes based on rB. Finally, these BZs were spatially mapped across the West Australian and “Green Triangle” commercial estates by enviromic prediction using EVs for 80 locations and 15 sets of observed climate data to represent temporal variation. The factor analytic model explained 85.9% of estimated additive variance. Our environmental classification system produced within-zone mean rB between 0.76 and 0.84, which improves upon the existing values of 0.62 for Western Australia and 0.67 for Green Triangle as regional BZs. The delineation of 5 BZ classes provides a powerful framework for increasing genetic gain by matching genotypes to current and predicted future environments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants are continually responding to a diversity of environmental cues. Genotypes differ in receptivity and reaction to such external stimuli, giving rise to genetically distinct patterns of phenotypes across a range of environments (Arnold et al. 2019; Kang 2002). Such differential “reaction norms” result in genotype by environment interaction (GE) (Costa-Neto and Fritsche-Neto 2021). In the plant breeding context, GE is typically quantified across an experimental network as variance due to a GE interaction component relative to cross-site genetic variance (Shelbourne 1972) or as a genetic correlation (rB) among sites (Burdon 1977).
GE is pervasive in forest genetics, especially for tree growth (Li et al. 2017). In general, GE can be partially explained by static physical attributes such as soil types and a less predictable component of temporal effects. Variance due to unexplained GE decreases genetic gain by reducing overall heritability across sites and compromising the estimation of genetic variance across multiple environments (Li et al. 2017). A common management response is to delineate spatially discrete breeding zones (BZs) within which GE is minimized (e.g., Raymond 2011; Ukrainetz et al. 2018). For species re-planted within their natural range, local populations are generally considered best adapted and safest to use (Ying and Yanchuk 2006) and their BZs are typically based on obvious environmental variables (EVs) such as latitude and temperature (e.g., Chen et al. 2017) or geographically defined regions (e.g., Johnson 1997).
An initial BZ classification may also be made on a regional basis for non-native species. Examples include for Pinus radiata in Australia (Ivković et al. 2015), Pseudotsuga menziesii in New Zealand (Dungey et al. 2012), and for Eucalyptus globulus in Australia (Dutkowski et al. 2015) and Chile (Sanhueza et al. 2002). Such regional classifications are unhelpful when the environmental drivers of GE are highly variable within regions or are more similar between sites in different regions. In these cases, rB estimates from a comprehensive set of trials across diverse sites can be used to produce a zone classification determined by one or more predictive EVs (e.g., based on altitude; Raymond 2011).
Some progress has been made in understanding the EVs that determine GE in forest trees. Mean annual temperature and precipitation (along with altitude, which is highly correlated) appear to be key in explaining GE in P. radiata (Burdon et al. 2017). Latitude and minimum temperature (also highly correlated with each other) explained some variation in GE for P. taeda (Lauer et al. 2021). In a more comprehensive prediction of GE, Ukrainetz et al. (2018) showed that 3 precipitation-related EVs, 4 temperature-related EVs, and 2 EVs related to the frost-free period provided a good zonation outcome for P. contorta across British Columbia. The resulting 4 BZs were precisely mapped across the province in a spatial modeling process that used these 9 significant EVs to predict spatial BZ extent for past and predicted future climates (Ukrainetz et al. 2018).
Despite the advances in the BZ classifications, the quality and resolution of environmental data have constrained the prediction of GE in forest tree species (Dutkowski et al. 2015; Gapare et al. 2015). On the other hand, recent GE studies in agricultural crops have considered vastly more EVs in terms of dimensional feature resolution, such as detailed soil properties and daily weather data (Romay et al. 2010). One key advance connecting envirotyping and crop ecophysiology in crop experiments has been to consider a division of the testing period into discrete stages of plant development or phenological periods (e.g., Costa-Neto et al. 2022; Heslot et al. 2014; Jarquín et al. 2014). The recent abundance of EV data has given rise to “envirotyping,” which involves collating numerous EVs at sites across a network of trials, identifying the most impactful EVs for traits of interest, and using these to define the “envirotype” as an environmental signature of each site (Costa-Neto and Fritsche-Neto 2021; Resende et al. 2021; Xu 2016). Envirotyping can lead to two main “enviromic” (envirotyping-based) outcomes: (1) the “enviromic assembly,” which is a large-scale characterization of the predominant environmental types for a given region or experimental network, in order to compute environmental variance–covariance similarity matrices among sites (Costa-Neto et al. 2021); and (2) the “enviromic prediction” using the enviromic characterization to predict BZs (Heinemann et al. 2022). Both outcomes can enhance a predictive breeding pipeline, such as one based on genomic selection, providing an additional source of variance to complement the genetic relationship matrices in the multiple-environment decision making context (e.g., Costa-Neto et al. 2022, 2021; Jarquín et al. 2014).
An advantage of using highly dimensional environmental data over modeling a linear reaction norm approach for a specific EV is that a combination of many low-signal EVs may be required to characterize the underlying environmental pressures determining GE, as opposed to expecting one or two overarching EVs to explain GE responses (as in Calleja-Rodriguez et al. 2019). This could play an important role for developing a broader understanding of the past and future GE and enable prediction of complex changes to genetic expression resulting from global warming (Crossa et al. 2021). In general, it is expected that locally adapted genotypes will be better suited to more poleward and higher elevation sites given a warmer future climate (Aitken and Bemmels 2016; Sáenz-Romero et al. 2020). For this reason, climate mapping and modeling can produce detailed maps of current and expected future BZs (e.g., Thomson et al. 2010). However, the BZ predictions under climate change scenarios are most reliable when the environmental determinants of GE are known precisely (e.g., Ukrainetz et al. 2018).
Many statistical approaches are available for characterizing GE (Van Eeuwijk et al. 2016), including most notably additive main effects and multiplicative interaction (AMMI), which combines ANOVA with principal component analysis (PCA) techniques (Crossa et al. 1990), factorial regression with geographic coordinates (Costa-Neto et al. 2020), and random regression using an EV such as temperature (Arnold et al. 2019), an environmental index of productivity (e.g., Alves et al. 2020; de Souza et al. 2020), or a latent environmental variable computed by factor analytic (FA) analysis (Meyer 2009; Smith et al. 2001). The FA method provides a complete rB matrix among trial sites while accommodating poor connectivity between sites that is typical of tree breeding experiments (Cullis et al. 2014). It has therefore been extensively used and recommended for analysis of GE in forest tree populations (Calleja-Rodriguez et al. 2019; Chen et al. 2017; Cullis et al. 2014; Gezan et al. 2017; Hardner et al. 2010; Ivković et al. 2015; Ogut et al. 2014; Shalizi and Isik 2019; Ukrainetz et al. 2018). Nevertheless, interpretation of the resultant rB matrix is dependent on the quality of supporting environmental information and is improved by detailed enviromic data (Tolhurst et al. 2022).
Handling large enviromic datasets is complicated by their high multidimensionality and multicollinearity. Partial least squares analysis (PLS), which is related to PCA and multiple linear regression, provides a way to analyze such data with strongly collinear, correlated, noisy and numerous variables (Wold et al. 2001). The method relates one or more response variables Y to a p variables × n samples matrix (X), achieving dimensionality reduction by selecting a subset of p that contributes most to explaining variation in Y. PLS has previously been applied to the problem of selecting EVs to explain crop GE most parsimoniously (e.g., Crossa et al. 1999; Monteverde et al. 2019; Porker et al. 2020). Additionally, PLS could be used to trace environmental signatures across previously characterized breeding zones (Costa-Neto et al. 2022).
Eucalyptus globulus is a commercially important plantation species, especially for pulp and paper production. GE in E. globulus has generally been quantified in very general terms, without revealing the nature of potentially significant spatial patterns. For example, Costa e Silva et al. (2009) calculated a uniform rB of 0.83 for diameter at breast height (DBH) among eight typical Portuguese sites, which is similar to Callister et al.’s (2011) average of 0.76 (0.52 to 1.00) among seven within-series trial pairs in Western Australia (WA). Similarly, Li et al. (2007) fitted uniform rB to subrace, additive, and dominance effects using full-sib families across 10 Australian sites, and reported estimates of 1.02 and 0.71 for DBH and 0.00 and 0.72 for tree height (HT), for sub-race and additive levels, respectively. In a closer examination of GE patterns in E. globulus, Dutkowski et al. (2015) reported that fitting uniform rB values within each of five Australian regions and between each pair of regions produced realized rB estimates ranging from 0.00 to 0.94 between regions (mean 0.57) and from 0.20 to 1.00 within regions (mean 0.69). Notably, uniform rB within WA was 0.59, within the "Green Triangle" region (GT) was 0.66, and between WA and GT was 0.58 (Dutkowski et al. 2015). Using long-term site climate attributes, Dutkowski et al. (2015) found that minimum monthly evaporation in three classes produced uniform within-zone rB values of 0.64, 0.76, and 0.63, and between-zone of 0.52, 0.38, and 0.60.
Approximately 460,000 hectares of E. globulus plantations is managed in Australia (Downham and Gavran 2019) and an improved understanding of GE in the species could contribute substantially to realized gain from genetic improvement programs. Progeny data from two Australian E. globulus improvement programs were recently combined in a unified single-step genomic BLUP analysis (“HBLUP”), using three broad regions as BZs (Callister et al. 2021). Based on the previous, the primary goal of the present study is to use these same full-sib family data and characterize progeny trial sites by environmental attributes to produce a classification for BZs in WA and GT using envirotyping. A second goal of this study is to examine the temporal and spatial variability in BZ distributions across the WA and GT commercial areas. To achieve these goals, we applied an integrative approach combining factor analytic models, enviromics, and spatial modeling.
Materials and methods
Experimental populations
This study was conducted using a subset of the data described by Callister et al. (2021) from two multi-generational breeding populations of E. globulus in southern mainland Australia, EG1 (Australian Bluegum Plantations) and EG2 (HVP Plantations). Forty-eight field trials were established in WA, GT, and Gippsland between 1998 and 2015, representing a total of 126,467 full-sib progeny in 1973 families produced by controlled crossing (see Table 1 for details by population and region). A relatively small number of half-sib families and checks were included in these trials but are not the focus of the present study.
The EG1 population of full-sib families was founded on 112 base-population trees, and the EG2 population was founded on 83 base-population trees in native forests or international landraces (Table SM1). All E. globulus races apart from Tidal River were represented in the combined pedigree (Fig. 1 and Table SM1). A total of 347 EG1 parents and 107 EG2 parents were represented by progeny included in the present analyses.
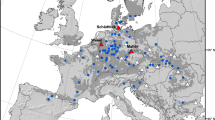
Map of southern Australia with natural distribution and races of E. globulus in brown (after Freeman et al. 2007) and the 33 trial sites used for envirotyping represented by blue diamonds within Albany and Esperance districts (Western Australia region) and Green Triangle region. Gippsland sites used in the FA analysis are located within the natural distribution of Strzelecki Ranges and South Gippsland races (trial sites not shown)
Trials were primarily established in randomized incomplete-block designs with four to eight replicates of each family established in four- to five-tree row plots. Two EG1 trials were single-tree plot designs. Trial layouts were generally contiguous, allowing for the resolution of spatial trend effects within each trial. Planted stocking generally ranged from 800 to 1087 stems/ha, with one site at 556 stem/ha (Fig. SM1). Mean survival was 90% and mean tree height was 12.2 m at assessment (Fig. SM1).
Phenotyping and genotyping
Diameter at breast height (DBH) was measured with a tape for each tree at between 3 and 8 years of age. Tree height (HT) was measured with a hypsometer for each tree in most trials, although a subset of 3–17% (mean 10%) of trees were assessed in 11 of the EG2 trials. In these cases, unmeasured HT data were predicted from the DBH-HT relationship established among measured trees for each trial. Stem volume (VOL) was calculated for each tree using DBH, HT (measured or predicted), and the region-specific volume function provided by each program.
Genotypic relationships among parents in the two populations were used to merge pedigrees for a combined analysis (see Callister et al. 2021 for details). There was little connectivity in the known pedigree, as only four founders were common to both populations. Pedigree-derived relationship coefficients between the two parent cohorts were rare, with two relationships of 0.25, 152 relationships of 0.125, and one expected relationship coefficient of 0.0625.
Leaves were sampled from 164 parents in the EG1 program and from 93 parents in the EG2 program. DNA extraction and genotyping were performed at Gondwana Genomics Pty Ltd, Canberra (Thavamanikumar et al. 2020). The marker panel consisted of 2579 single nucleotide polymorphism (SNP) and small biallelic insertion/deletion (INDEL) markers within previously identified candidate genes (Southerton et al. 2011). The filtered marker matrix included 2444 markers after removing those with minor allele frequency less than 0.05.
Experimental and genetic analysis
Relationship matrix calculation
Joint-program relationship matrices were formed among parents, tested progeny, and their ancestors using preGSf90, which is a module of the Fortran-based BLUPF90 suite (Misztal et al. 2014). The default settings of preGSf90 were used for quality control and calculation of the relationship matrices. The genomic relationship matrix (G) was calculated following the first method of VanRaden (2008). G was rescaled to make it compatible with the comparable section of A representing genotyped individuals (A22) following Christensen et al. (2012). The rescaled matrix Ga was then weighted with 0.05A to avoid difficulties with inversion (Aguilar et al. 2010) and used to calculate H, with inverse (Christensen and Lund 2010; Legarra et al. 2009):
H and A were compared in a preliminary step to highlight identity errors and mistakes in the documented pedigree. These errors were rectified in the pedigree and the process of creating the hybrid relationship matrix was then repeated with the correct pedigree information.
Single-site analyses
All genetic analyses were conducted using ASReml version 4.1 (Gilmour et al. 2015). Single-site, univariate animal models (pedigree BLUP) of VOL were first fitted to data from each trial following Henderson (1984) using the general linear mixed model framework:
Where y is the vector of phenotypic values, X is the incidence matrix for fixed effects, \({\varvec{\tau}}\) is the vector of fixed effects, Z is the incidence matrix for random effects, u is the vector of random effects with E(u) = 0, and e is the vector of residual effects expected to be independently normally distributed with E(e) = 0. Races and landraces were included as fixed genetic groups within the pedigree (Westell et al. 1988). Other fixed effects were the overall mean, replicates, and checklots. Random effects were additive and family-specific genetic effects of full-sib families, incomplete blocks, plots, and half-sib families. The dispersion matrices contained elements A \({\widehat{\sigma }}_{a}^{2}\), I \({\widehat{\sigma }}_{f}^{2}\), I \({\widehat{\sigma }}_{b}^{2}\)¸ I \({\widehat{\sigma }}_{p}^{2}\), I \({\widehat{\sigma }}_{h}^{2}\), and I \({\widehat{\sigma }}_{e}^{2}\), where \({\widehat{\sigma }}_{a}^{2}\) is the estimated additive genetic variance fitted to full-sib families, \({\widehat{\sigma }}_{f}^{2}\) is the estimated non-additive full-sib family-specific variance, \({\widehat{\sigma }}_{b}^{2}\) is the estimated incomplete block variance, \({\widehat{\sigma }}_{p}^{2}\) is the plot variance, \({\widehat{\sigma }}_{h}^{2}\) is half-sib family variance, \({\widehat{\sigma }}_{e}^{2}\) is the residual variance, A was the verified pedigree-based numerator relationship matrix, and I is the identity matrix. Additional within-plot error was fitted for checklots. A two-dimensional autoregressive structure was fitted to model the spatial variability effects within each trial (Dutkowski et al. 2002), which was retained for all but one site after significance testing by two-tailed likelihood ratio tests (Gilmour et al. 2015) and visual inspection of two-dimensional smoothness.
Narrow-sense heritability was estimated for each site: \(\widehat{{h}^{2}}=\frac{{\widehat{\sigma }}_{a}^{2}}{{\widehat{\sigma }}_{a}^{2}+{\widehat{\sigma }}_{f}^{2}+{\widehat{\sigma }}_{b}^{2}+{\widehat{\sigma }}_{p}^{2}+{\widehat{\sigma }}_{e}^{2}}\). VOL data were adjusted for spatial effects (following Ye and Jayawickrama 2008) and standardized by site to have a zero mean and an additive genetic standard deviation of one prior to cross-site analysis (VOLstd).
Factor analytic HBLUP analysis of genotype x environment interaction
VOLstd was analyzed across all 48 sites using a parent model for additive genetic variation and family-specific effects:
where \(\mathbf{y}\) is now the combined vector of VOLstd data for n full-sib progeny across t sites; \({\varvec{\uptau}}\) is the vector of fixed effects of trials with associated design matrix \(\mathbf{X}\); \({{\varvec{Z}}}_{{{\varvec{m}}}_{{\varvec{p}}}}\) and \({{\varvec{Z}}}_{{{\varvec{p}}}_{{\varvec{p}}}}\) are indicator matrices to assign each progeny with a maternal parent and a paternal parent, respectively; \({{\varvec{u}}}_{{{\varvec{a}}}_{{\varvec{p}}}}\) is the vector of additive genetic effects for the parents; \({{\varvec{u}}}_{{\varvec{f}}}\) is the vector of family-specific genetic effects across sites with associated design matrix \({{\varvec{Z}}}_{{\varvec{f}}}\); \({{\varvec{u}}}_{{\varvec{s}}{\varvec{i}}{\varvec{t}}{\varvec{e}}:{\varvec{f}}}\) is the vector of site × family interaction effects with associated design matrix \({{\varvec{Z}}}_{{\varvec{s}}{\varvec{i}}{\varvec{t}}{\varvec{e}}:{\varvec{f}}}\); and \({\varvec{e}}\) is the vector of residuals, which includes Mendelian sampling effects of progeny. \({{\varvec{u}}}_{{\varvec{f}}}\), \({{\varvec{u}}}_{{\varvec{s}}{\varvec{i}}{\varvec{t}}{\varvec{e}}:{\varvec{f}}}\), and \({\varvec{e}}\) were assumed to be normally distributed with mean 0.
\({{\varvec{u}}}_{{{\varvec{a}}}_{{\varvec{p}}}}\) was modeled with a factor analytic (FA) structure across sites. The general form of additive genetic effect i at site j in a FA model with order k (FAk) can be written as
which is a sum of k multiplicative terms and \({\delta }_{{a}_{ij}}\), which represents the lack of fit of the regression model. Each multiplicative term is a product of an environmental effect, or “loading” for the jth site (\({\lambda }_{{a}_{{k}_{j}}}\)) and a genetic value for the ith individual in the relationship matrix (\({f}_{{a}_{{k}_{i}}}\)). The FA model can be represented in matrix notation as
where \({{\varvec{\Lambda}}}_{{\varvec{a}}}\) is the t (site) × k matrix of environmental loadings, \({{\varvec{f}}}_{{\varvec{a}}}\) is the qk × 1 vector of additive genetic scores, and \({{\varvec{\delta}}}_{{\varvec{a}}}\) is the qt × 1 vector of genetic regression residuals, where q is the number of additive effects. We assume that \({{\varvec{f}}}_{{\varvec{a}}}\) and \({{\varvec{\delta}}}_{{\varvec{a}}}\) are mutually independent and distributed as multivariate normal with zero mean and variances given by var(\({{\varvec{f}}}_{{\varvec{a}}}\)) \(=\mathbf{H}\otimes {\mathbf{I}}_{{\varvec{k}}}\) and var(\({{\varvec{\delta}}}_{{\varvec{a}}}\)) \(=\mathbf{H}\otimes {{\varvec{\Psi}}}_{{\varvec{a}}}\), where \(\mathbf{H}\) is the q × q relationship matrix for parents and their ancestors and \({{\varvec{\Psi}}}_{{\varvec{a}}}\) is a t × t diagonal matrix with a variance for each site. The variance of additive genetic effects is var(\({{\varvec{u}}}_{{{\varvec{a}}}_{{\varvec{p}}}}\)) \(=\mathbf{H}\otimes {({{\varvec{\Lambda}}}_{{\varvec{a}}}{{\varvec{\Lambda}}}_{{\varvec{a}}}^{{\varvec{T}}}+{\varvec{\Psi}}}_{{\varvec{a}}})\) so that the additive genetic correlation matrix among sites, \({{\varvec{\rho}}}_{{\varvec{a}}}\), is \({{\varvec{\rho}}}_{{\varvec{a}}}={({{\varvec{\Lambda}}}_{{\varvec{a}}}{{\varvec{\Lambda}}}_{{\varvec{a}}}^{{\varvec{T}}}+{\varvec{\Psi}}}_{{\varvec{a}}})\).
FA model performance was evaluated by calculating the percentage of additive genetic variance accounted for by the k multiplicative terms at each site (Cullis et al. 2014):
and an overall (across-site) percentage accounted for
Factor analytic models were fitted to the 48-trial data first with one, then two k factors, and finally with three factors (FA3) to adequately explain the additive genetic variance across sites.
Selection of sites and definition of BZ classes for model training
The full set of 48 sites was reduced to 33 well-characterized sites for envirotyping (see the “Results” section). All 6 sites in Gippsland were excluded due to lack of directly observed soils information for the region. Six EG1 progeny trials from an earlier generation were also excluded as these sites displayed good rB between each other but were not well correlated to any other site. Three further sites were excluded as their \({v}_{{a}_{j}}\) values were less than 50. The retained trials were established between 2001 and 2015 and were assessed for volume at age 3 years (3 sites), 4 years (6 sites), or 5 years (24 sites). Eleven of the retained trials were in GT (4 from EG1 and 7 from EG2) and 22 in WA (20 from EG1 and 2 from EG2). Information about the level of genetic connection between sites selected for envirotyping is presented in Figs. SM2 and SM3.
Next, a classification was performed with Euclidean hierarchical clustering to assign each of the 33 sites to one of 5 zone classes based on the rB estimates produced by the FA3 models. However, initial efforts using PLS to identify the EVs contributing to membership of unique zone classes did not produce models with satisfactory prediction capacity. Thus, we re-examined the heatmap of rB values and realized that some sites could be assigned to more than one class based on rB. The hypothesis was considered that class-defining EV combinations might afford sites with non-exclusive class membership. The Euclidean hierarchical clusters were considered “core” site membership and then additional “supplemental” sites were added to each class one by one if the mean rB between the candidate site and the existing sites in the class (“marginal rB”) was greater than 0.70 and if the mean rB within the resulting class remained greater than 0.80.
Environmental data collection and envirotyping assembly
A total of 145 EVs were included in the envirotyping study: 71 soil and landscape EVs, 73 climate EVs, and 1 management-associated EV (Table 2). Sources of soil and landscape data were directly observed soil pits or drills, laboratory analyses of soil samples from top 10cm, and the Soil and Landscape Grid of Australia (SLGA), which is a fine resolution (3 arcsecond grid (~ 90 m × 90 m)), public dataset of soil, landscape, and regolith data predictions known to influence crop growth (Grundy et al. 2015). Daily weather data for each site were downloaded from the online SILO platform (Jeffrey et al. 2001) from planting until measurement age or the first 5 years of each trial’s history, whichever was shorter (https://www.longpaddock.qld.gov.au/silo/point-data/, accessed July, 2022).
Soil and landscape attributes were summarized by 71 EVs (Table 2, with details in Table SM2).
Precipitation, minimum temperature, and maximum temperature data were processed into average values for each month across the first 3-year period. These monthly averages were then used to calculate the 19 climate metrics used for species distribution prediction in the BIOCLIM framework (Booth et al. 2014) using the “dismo” R package (see Table SM3 for details).
Studies to predict GE in agricultural crops make good use of well-defined phenological stages such as tillering, jointing, booting, heading, etc., to distinguish periods of weather that affect different physiological processes. For a long-lived perennial such as E. globulus, alternating periods of growth and of survival could be considered phenological stages. Growth periods occur when water and temperature balances allow net anabolism, and survival periods occur when resources must be expended to adapt to unfavorable environmental conditions. For established E. globulus on each of the trial locations studied here, the annual survival stage would be prompted by water limitation and perhaps high temperature, rather than by freezing stress. We considered the first year of a E. globulus plantation to include two unique phenological stages. Both growth and survival are important in the first 4–6 months of establishment, due to the risk of frost death in a period when water and temperature are otherwise favorable for growth. The first summer after growth can also be considered a unique growth period because temperatures are highly suitable for anabolism, while stored soil water is usually capable of meeting a plantation’s water requirements.
Daily weather data from SILO were used to translate the growth stage concept into practice at each of the trial sites as follows. Climate wetness index over a 90-day period (CWI90) was defined as the ratio of total precipitation to total evaporation in the preceding 90 days for each day commencing 91 days after planting. The establishment phase (Estab) was defined from planting until CWI90 reduced to a threshold of 0.5 (Table 3). In the absence of a proven system of phenology for E. globulus, this value was used to mark the commencement of the first-year dry period. The critical threshold of 0.5 for CWI90 was used in each subsequent year to define the end of growth periods and the start of dry periods. Each dry period was ended once 120 mm of rainfall had accumulated since the start of the respective dry period (Table 3).
In addition to the BIOCLIM variables, up to 54 climate EVs were calculated for each site (Table 2 with details in Table SM4). Cardinal values used for calculation of temperature effects on growth were [3, 14, 28, 35], relating to base, minimum optimum, maximum optimum, and maximum cardinal temperatures in degrees C (Smethurst et al. 2022). T.opt.hours was defined as the number of hours between 14 and 28 °C, T.low.hours as the number of hours less than 3 °C, and T.high.hours as the number of hours above 35 °C.
Twenty-five of the 33 envirotyped sites were first-rotation plantations, with either pasture or cropping before establishment. The remaining eight sites were second rotation established after a first rotation of E. globulus. This first- versus second-rotation distinction was the only management-associated EV considered in the envirotyping process.
Partial least squares analysis to discover the major envirotypes for each breeding zone
The next task was to produce a parsimonious characterization of the 5 BZ classes from the curated enviromic dataset of 145 EVs. Each BZ was represented by a training dataset that simply classified sites as included or excluded, based on rB estimates. Sparse PLS-discriminant analysis (sPLS-DA) (Chung and Keles 2010; Lê Cao et al. 2011) was implemented in the R package “Mixomics” (Rohart et al. 2017) to identify which EVs were required to predict classification of sites into membership and non-membership classes for each BZ (a PLS primer is provided in supplemental materials). All EV input data were centered and scaled and the NIPALS algorithm was implemented within Mixomics to input missing values.
For each BZ, the modeling process commenced with specification of 10 components (latent variables) and 20 variables per component, to gauge a maximum classification accuracy possible with the correlation-based training data. The ROC-AUC or area under the receiver-operator characteristic curve (Brown and Davis 2006) was used as an index of classification accuracy. This initial step identified three outlier sites, which were removed from certain training datasets: site no. 16 was removed from BZs 2, 3, and 4 and site no. 31 was removed from BZ1 due to poor EV data quality in both cases, while site no. 21 was removed from BZ2 due to a history of poor weed control.
Parsimony in sPLS-DA model prescription was then achieved for each BZ model by iteratively reducing the number of components and EVs per component, while estimating classification error rates and ROC-AUC statistics. The goal was to achieve classification using 1 or 2 components and as few EVs as possible. Maximum, Euclidean, and Mahalanobis distance metrics were compared for classification on a two-dimensional plane involving two-component solutions. However, visually identified lines separating member sites from non-member sites produced fewer classification errors than these algorithms in each case. EVs used in classification and their respective coefficient values were output for each component for mapping purposes.
Spatial mapping of breeding zones
The goal of this exercise was to produce regional maps of BZs calculated for a sample of 15 planting years in the recent past. A network of 80 prediction locations was created; 40 across the WA estate and 40 across the eastern estate (see Fig. SM4). Priority was given to providing a wide spread of positions across each commercial estate. Locations were also generally chosen where three pre-existing soil pits and three sample collections for soil chemistry could be located within close proximity. Soil EVs for the prediction points were then calculated as the mean of values from these nearby datasets. Notable exceptions were eight sites at the western extent of the WA prediction area for which published chemical information was referenced.
Daily weather data was obtained from SILO for each of these 80 prediction points corresponding to the first 5 years commencing on July 1 on each year from 2002 to 2017 (15 years). Climate EVs that were determinants of BZ class assignment were calculated. The 2010 planting year in GT was excluded as it featured extremely heavy summer rains in the first year. None of the observed progeny trials were planted in 2010 in GT and the heavy summer rain created an atypical environment for the Y1dry phenological stage. BZ class prediction was then performed for each planting year at each of the 80 prediction locations, using the custom distance functions discovered with sPLS-DA. The proportion of planting years (2002–2017) for which each prediction point was classified to each BZ was summarized, and mapped in four quartile classes: 0–25%, 25–50%, 50–75%, and 75–100%. The expected area of each BZ was calculated as the total of products between each quartile mid-class value and its respective area, represented as percentage of area in each region.
Results
Additive genetic correlations across breeding zones
A FA3 model was first used to represent additive genetic variance and site–site correlations (rB) in stem volume across 48 trials of 126,467 full-sib progeny from two separate breeding programs that were connected by HBLUP. This FA3 model explained 85.9% of additive genetic variation across all 48 sites. The percentage of additive genetic variance accounted for by the FA3 model (\({v}_{{a}_{j}})\) ranged from 79.6 to 100 for the 33 envirotyped trials, with a mean of 94.5 (Table SM5 and Fig. SM5). The resultant rB matrix (\({{\varvec{\rho}}}_{{\varvec{a}}}\)) displayed six clear site groups, two of which partially overlapped (see lower left corner of Fig. 2). “Core” constituency was assigned to sites for each of 5 BZs, with BZ2 including core site members from two branches of the dissimilarity dendrogram (Fig. 2).
Heatmap of \({{\varvec{\rho}}}_{{\varvec{a}}}\) matrix of inferred additive genetic correlations among 48 trial sites from the FA3 model. Breeding zone classes used for training the sPLS-DA models are shown below the diagonal: BZ1 (green perimeters), BZ2 (blue perimeters), BZ3 (red perimeter), BZ4 (gold perimeters), and BZ5 (purple perimeters). The numbers in red indicate “core” constituents of each breeding zone (see the “Methods” section). The hierarchical tree uses average clustering. Diagonal cells with \({{\varvec{\rho}}}_{{\varvec{a}}}=1\) are shown in gray
Supplementary sites were added to four of the five BZs (see process outlined in Table SM6). This resulted in BZ1 with three supplementary sites along with four core sites, BZ2 with seven supplementary sites along with eight core sites, BZ3 with only the nine core sites, BZ4 with three supplementary sites along with seven core sites, and BZ5 with four supplementary sites along with five core sites (Fig. 2).
Characteristics of the major environmental signatures for each breeding zone
A subset of 33 trials with good quality soil data and explained variance in the FA3 analysis was envirotyped using 145 environmental variables (EVs). Sparse PLS-discriminant analysis was used to identify EVs that were required to predict classification of sites into five non-exclusive BZ classes on the basis of rB. The results are described below for each breeding zone (from BZ1 to BZ5). Four BZs were defined with two PLS components each (one to five EVs contributing to respective components), and one BZ was defined with a single PLS component (eight EVs). In each case, the goal was to minimize classification errors that may occur by classifying a site that was not assigned to a BV’s training set (i.e., false positive) or failing to classify a site that was assigned to a BV’s training set (i.e., false negative).
Site classification to BZ1 was predicted with one false positive and one false negative (ROC-AUC 0.96) using two PLS components (Table 4, Fig. 3A). The first component was composed of two significantly correlated (r = 0.94, p < 0.005) EVs named Y1.5_optH and Y1.5_GDD (Table 5). These EVs describe the number of optimum growing hours and growing degree days, respectively, in the combined establishment phase, first-year dry season, and growth phases of the first 5 years. The second component was composed of two EVs: most significantly SLT100 (PLS coefficient 0.98), which is the silt content at 60–100 cm, and C_N, which is the C:N ratio (PLS coefficient − 0.22; Table 5). Taken together, these EVs describe BZ1 as an envirotype characterized by optimum growing temperatures and siltier soils with greater nitrogen levels.
Site classification to BZ2 was predicted with no false assignments (ROC-AUC 0.98) using two PLS components (Table 4, Fig. 3B). The first component was composed of five EVs. Two were soil EVs COL_K (Colwell available K) and C_N (C:N ratio) and three were climate EVs estab_hotH (hours with temp > 35 °C in the establishment phase), Y3_precip_tot (total precipitation in year 3, which is strongly correlated to precipitation in growing periods across the first 3 years), and Y1.3dry_days (total length of dry seasons in the first 3 years; Table 5). The second component was composed of 3 EVs: most significantly two correlated (r = 0.84) EVs Y1.5dry_cumVPD and Y1.5_precip_deficit, which are cumulative vapor pressure deficit and difference between cumulative rainfall and evaporation, respectively, in dry seasons across years 1–5 (Table 5). The third EV contributing to the second component of BZ2 was Y1.3_BIO10; mean temperature of the warmest quarter. Taken together, these EVs describe BZ2 as an envirotype characterized by longer, warmer, and drier summers with better-quality soils (higher K and N) and fewer hot days during establishment. Note that with the classified sites for BZ2 in the lower-right of Fig. 3B, more positive values of the first PLS component and more negative values of the second PLS component will support assignment of an envirotype to BZ2 (relevant for interpreting Table 5).
Site classification to BZ3 was predicted with one false positive and one false negative (ROC-AUC 0.96) using 2 PLS components (Table 4, Fig. 3C). The first component was composed of four EVs. The most significant EV contributing to the first component was 1R_BIN, which is the binary coding differentiating first rotation and second rotation sites (PLS coefficient − 0.95, Table 5). This management EV separates the two clear site groups along the X-axis of Fig. 3C, with second-rotation sites to the right. The three lesser EVs contributing to the first component were the correlated (r = − 0.81) EVs SLT60 and SND60 (silt content and sand content, respectively, at 30–60 cm) and Y1.3_radn_tot (total radiation in the combined establishment phase, first-year dry season, and growth phases of the first 3 years; Table 5). The second component was composed of four climate EVs: estab_precip_tot (total precipitation during the establishment phase), Y1dry_Tmax_ave (average daily maximum temperature in the first dry season, highly correlated with dry-season maximum temperatures in the first 3 years), Y1dry_days (length of first dry season), and Y1.3_radn_tot (total radiation in the combined establishment phase, first-year dry season, and growth phases of the first 3 years; Table 5). Taken together, these EVs describe BZ3 as an envirotype characterized by predominantly second rotation, somewhat sandier sites, less establishment-period rainfall, and higher temperatures in summer.
Site classification to BZ4 was predicted with one false positive and two false negatives (ROC-AUC 0.96) using 2 PLS components (Table 4, Fig. 3D). The first component was composed of a single EV: Y2.3_precip_cool6mo (precipitation in the coolest 6 months of years 2–3; Table 5). This EV is significantly correlated with other total precipitation EVs (Table SM4), including total precipitation in the first 3 years (r = 0.92). The second component was composed of four climate EVs: estab_hotH (number of hours with temperature > 35 °C during the establishment phase), Y1.3_BIO17 (precipitation of the driest quarter), Y1.3_BIO10 (mean temperature of the driest quarter), and Y1dry_Tmax_ave (average daily maximum temperature in the first dry season, highly correlated with dry-season maximum temperatures in the first 3 years; Table 5). Taken together, these EVs describe BZ4 as an envirotype characterized by drier winters (and lower overall annual rainfall), cooler at establishment, and somewhat wetter and cooler in summer.
Site classification to BZ5 was predicted with perfect accuracy (ROC-AUC 1.0) using a single PLS component (Table 4). The PLS component was composed of eight EVs, seven of which were related to soils: C_N (C:N ratio), pH_H2O (pH in water), TOT_N (total N), SLT100 (silt content at 60–100 cm), estab_hotH (number of hours with temperature > 35 °C during the establishment phase), SND > 200 (sand content deeper than 200 cm), CLY > 200 (clay content deeper than 200 cm), and S&R_FC > 200 (soil water at field content deeper than 200 cm; Table 5). The last three of these EVs are highly inter-related. Taken together, these EVs describe BZ5 as an envirotype characterized by low nitrogen, higher pH, and somewhat sandier soils and higher established-period temperatures.
Envirotypes at trial sites
Six progeny trial sites qualified as BZ1 following the sPLS-DA classification (see Table SM7 for details). Four of these were concentrated inland of Albany (Fig. 4A), where the milder temperatures and better-quality soil characteristic of this zone may be expected. One BZ1 site was in the East Albany district and shared BZ classification with zones 2 and 3 (Fig. 4A), and one was inland of Esperance (Fig. 4B). BZ2 was the breeding zone most assigned to progeny trials (15 of 33) and was allocated in all three districts (Albany, Esperance, and GT; Fig. 4). It was the sole breeding zone found in the central part of the GT Region (Fig. 4C). BZs 3, 4, and 5 were assigned to nine trials each. BZ3 was distributed through the drier part of the Albany Region and near the Victoria/South Australia border in the GT Region (Fig. 4A and 4C). Trials in the eastern part of Esperance region were characterized predominantly as BZ5 (Fig. 4B) and trials in the eastern part of the GT Region were characterized as BZ4 (Fig. 4C).
Maps of progeny trial sites classified into breeding zones using the sPLS-DA classification in A Albany district, WA, B Esperance district, WA, and C Green Triangle region. See Fig. 1 for geographical context
One site in the East Albany district was classified to BZs 1, 2, and 3 concurrently (see intersection in Fig. 5) and 13 trials were classified to two breeding zones concurrently (Fig. 5). Zones 2 and 3 had four sites in common, as did zones 2 and 4 (Fig. 5).
The envirotype classification system produced within-zone mean rB between 0.76 and 0.84 (Table 6). In contrast, the previous region-based classification system produced within-zone mean rB of 0.62 for WA and 0.67 for GT (Table 6). The distinctions in rB between within- and across-class were also greater for BZs in the envirotype classification system (Table 6).
Spatial mapping of breeding zones
Mapping of envirotypes showed that BZ1 was commonly found in the high-yielding near-coastal areas west of Albany and less frequently inland and east of Albany (Fig. 6). The occurrence of BZ1 was very uncommon in GT (Fig. 7). BZ2 is the most represented envirotype across both regions. Based on recent past climate, 74% of the estate area in WA and 51% of area in GT are expected to represent BZ2 based on climate between 2002 and 2022 (Fig. 7). In WA the areas consistently representing BZ2 are generally in the drier part of the estate (inland), whereas in GT the BZ2 envirotype was widely distributed (Fig. 6).
BZ3 is an envirotype that is relatively rare in WA under recent past climate on 1R sites (expected 11% by area, Fig. 6), but it is more common in WA on 2R sites (expected 42% by area; Fig. 6). In GT, 14% of area is expected to be in the BZ3 envirotype on 1R sites and 62% is expected by area on 2R sites (Fig. 6), making it the most represented envirotype for 2R sites in GT (Fig. 7).
The BZ4 is an unexpected envirotype in WA, but it is important in GT, especially in the more eastern part of the commercial area (Fig. 6). The total expected area of BZ4 in GT is 38% based on past climate (Fig. 7). BZ5 is relatively rare across WA and GT (up to 20% by expected area; Fig. 7). In WA, it was allocated to areas in the south-west that were also consistently allocated to BZ2 (Fig. 6). In GT, BZ5 was distributed most in the western part of the estate, where it overlapped with BZ3 for 2R sites (Fig. 6).
Discussion
This study proposed a detailed environmental classification of breeding zones for E. globulus in two regions of mainland Australia. Previous work with these E. globulus populations has demonstrated the potential of genomic technology to predict genetic values of untested genotypes (Callister et al. 2021). The present results will help to expand our prediction abilities from untested genotypes to untested environments (Crossa et al. 2021). For E. globulus in Australia, the untested environment of greatest interest is the future environment within the existing commercial regions (Pinkard et al. 2015). However, the first hurdle to predicting future climate responses (Fradgley et al. 2023) was to characterize eco-physiologically relevant aspects of the environments in which progeny have already been tested.
Envirotyping of breeding zones helps to understand eco-physiological aspects of adaptation
Challenges to accurate envirotyping for trees compared with agricultural crops include the far deeper soil environment affecting trees and the poorly understood progression of phenological development. Lack of trial-specific EV data with suitable precision has been cited as a limitation to similar studies by Ivković et al. (2015), Gapare et al. (2015), and Dutkowski et al. (2015). The use of direct soil observations wherever possible was an important requirement for accurately identifying the role of soil and landscape EVs in characterizing envirotypes, especially BZ1, BZ2, and BZ5. This highlights the importance of collecting good quality soil data at every genetic trial location, and of carefully positioning trials to avoid crossing boundaries in soil types. The phenological system we applied appears to have been useful, considering the predominance of climate EVs utilizing this system and the generally high levels of BZ classification accuracy produced by the sPLS-DA procedure. Nevertheless, a precise system for defining phenology of E. globulus based on dendrometer and physiological measurements is needed to improve the clarity of climate EVs in future enviromic studies.
Many previous enviromic studies have developed prediction models accounting for GE on continuous variation in EVs (i.e., reaction norms), most notably for annual crops (e.g., Costa-Neto et al. 2022, 2020; Li et al. 2021; Tolhurst et al. 2022). This approach facilitates extrapolation of genetic predictions at unobserved values of EVs and it may offer higher resolution genetic predictions rather than sharply defined boundaries between classes. On the other hand, more advanced treatments of GE in forest trees have classified areas into discrete zones based on EVs (Gapare et al. 2015; Ukrainetz et al. 2018; Yu et al. 2022). It is possible that genetic interactions with environmental attributes are more complex for forest trees than for annual crops, necessitating the inclusion of more EVs than can reasonably be included in a reaction norm model. Another explanation is that annual crop datasets are often much larger than those for forest trees, providing sufficient EV data to model the resolution required for reaction norms.
We took the unusual approach of allowing each site to be classified to multiple zones for training the sPLS-DA model to rB data. Previous GE studies that assigned each site to a single zone based on hierarchical clustering have published rB matrix heatmaps also showing that certain sites were highly correlated to groups outside of their assigned cluster, including for P. radiata (Cullis et al. 2014) and P. contorta (Ukrainetz et al. 2018). The phenomenon of multiple zone membership for a subset of sites may therefore be common to other species and contexts as well as ours, and it makes biological sense that environmental determinants of genetic expression need not be mutually exclusive. For example, if we examine the overlapping site membership between BZ2 and BZ3 we find four sites that meet the requirements of drier summers (BZ2) and lower establishment-period precipitation (BZ3); higher available potassium with lower C:N ratio (BZ2) and second-rotation management (BZ3). For overlapping site membership between BZ2 and BZ4, we find four sites that meet the requirements of drier summers (BZ2) and lower precipitation in the coolest 6 months (BZ4), in other words, drier throughout the year. On the other hand, certain combinations of zones were incompatible, such as BZ5 with BZ1 or BZ2. BZ5 has a positive PLS coefficient for C:N ratio, whereas BZ1 and BZ2 have negative PLS coefficients for C:N (see Table 5). No progeny trial sites were in common between BZ5 and either BZ1 or BZ2 (see Fig. 5), although the far western part of the WA estate did associate strongly with BZ5 and BZ2 due to alignment of other EVs in the classification scheme.
Eucalyptus globulus GE has previously been shown to be influenced by summer precipitation deficit, evaporative demand, and maximum temperature at the subrace level (Costa e Silva et al. 2006), family level (Dutkowski et al. 2015), and in association with genomic markers (Butler et al. 2022). Discrimination along a continuum of summer evaporative demand loosely translates to classification of sites according to BZ2 (drier summers) in our classification system. Dutkowski et al. (2015) also identified minimum month evaporation as an alternative discriminatory EV, and Butler et al. (2022) found isothermality and precipitation to be informative of local adaptation. EVs related to precipitation featured strongly in our zonation, and minimum month evaporation would be related to BIO6 (minimum temperature), which in turn was strongly correlated to optimum growing hours and GDD, both significant to the BZ1 classification. On the other hand, isothermality (BIO3) was not selected by sPLS-DA as a significant EV for classification.
The GE we have observed in these populations could be influenced by interactions with insect herbivores and diseases as well as direct physiological responses to the abiotic environment. The most impactful pests during the first 5 years of these trials are likely to have been Eucalypt weevil (Gonipterus spp.) and autumn gum moth (Mnesampela privata), while the principal disease would have been Teratosphaeria leaf disease (TLD; Teratosphaeria nubilosa and Teratosphaeria cryptica). Higher mean temperature of the warmest quarter (BIO10; characteristic of BZ2) is expected to favor both insect pests, while higher summer rainfall (BIO17; characteristic of BZ4) will favor TLD spore production and growth (Pinkard et al. 2017). It is therefore possible that differential tolerance of these biotic impacts is at least partially responsible for the GE in our E. globulus populations, especially with respect to BZ2 and BZ4.
Technical impacts of BZ mapping for E. globulus breeding
We discovered that BZ3 is more likely to be assigned to a site in second rotation (2R) than in first rotation (1R), which follows Raymond's (2011) finding that prior land use impacts on GE in P. radiata. It is a conclusion of operational significance, as all the Australian E. globulus improvement programs include a large proportion of 1R trials in breeding value estimation for seed that will be predominantly deployed to 2R sites. Nevertheless, we found that 2R status does not classify a site as solely BZ3. Four 2R sites classified as BZ2 as well as BZ3, one 2R site classified as BZ5 as well as BZ3, and one 2R site classified as BZ1. The 2R management factor could be considered a proxy for EVs relating to initial soil water and nitrogen, both of which are generally reduced in 2R relative to 1R (Mendham et al. 2011).
The range of mean within-envirotype rB from 0.76 and 0.84 reported here (see Table 6) compares favorably with published within-class rB from similar studies: 0.60 to 0.81 by Dutkowski et al. (2015) and 0.67 to 0.83 by Ukrainetz et al. (2018). We are confident that the envirotype classification presented here will produce greater within-zone estimates of genetic gain than the previous regional-based classification. Our approach also achieved a reconciliation of data from trials from a district that is no longer part of the commercial estate (Esperance) with the current commercial sites. It is notable that the four eastern-most Esperance sites qualified as BZ5, which is a relatively minor envirotype for ongoing breeding and deployment, so these trials will no longer contribute as significantly to future estimates of breeding value in the EG1 or EG2 programs.
Limitations and future improvements in envirotyping for E. globulus
There are some important limitations to our approach and suggested improvements for implementation subject to future data availability. Envirotype definitions created from the available data excluded explicit influences of pests and diseases, most notably TLD (Andjic et al. 2010), as well as the influence of ground water especially in the Wattle Range (South Australia). We have possibly overlooked a whole phenological stage corresponding to growth in the warmest 3 months after 2 years that would be triggered by rare summer storms. This study also ignored any effect of age on GE, although all phenotypes used in envirotyping were measured within a relatively narrow range of 3 to 5 years post-establishment, during which age-age genetic correlations are expected to be close to 1.0 (Salas et al. 2014; Stackpole et al. 2010). Although the rB matrix resulting from FA analysis is a logical starting point for defining breeding zones, there is a concern that its elements are estimated with varying precision, depending on the degree of genetic connection between pairs of sites. Future studies could apply variable weights to rB to account for precision as part of the BZ definition procedure. This could be done in the context of a critical examination of the non-exclusive BZ concept, specifically to identify a less subjective basis for including “supplemental” sites to “core” groups of sites than we have applied. Lastly, we have ignored GE due to differential levels of genetic expression between environments (Li et al. 2017), which is also an important consideration when structuring breeding programs and planning deployment. These avenues of exploration are all open for future review and will be assisted by the acquisition of new EV data and new sites.
Expectations have increased substantially since Barnes et al. (1984) wrote: “It is probably only realistic to expect to detect, explain and use (GE) when a single environmental factor affects an economically important trait in a predictable manner.” The results of our study show conclusively that a multitude of EVs affect GE in E. globulus in a complex, yet predictable manner. These results will be used to immediately reclassify breeding zones for cross-program analyses of these EG1 and EG2 breeding populations, thereby mitigating the loss of realized gains due to deploying families into less suitable environments. The BZ classification system described here is also being used in conjunction with climate change forecasts to predict which envirotypes will be more prominent in the future, to guide selection and breeding strategy.
Data availability
Pedigree, phenotypic, genomic, and environmental data are available in the repository: https://figshare.com/articles/dataset/E_globulus_enviromic/14402093. This archive contains (A) complete pedigree and phenotypes for 48 trials across both breeding programs, (B) SNP data for the genotyped individuals, and (C) environmental variables and other metadata for 33 envirotyped sites.
References
Aguilar I, Misztal I, Johnson DL, Legarra A, Tsuruta S, Lawlor TJ (2010) A unified approach to utilize phenotypic, full pedigree, and genomic information for genetic evaluation of Holstein final score. J Dairy Sci 93:743–752. https://doi.org/10.3168/jds.2009-2730
Aitken SN, Bemmels JB (2016) Time to get moving: assisted gene flow of forest trees. Evol Appl 9:271–290. https://doi.org/10.1111/eva.12293
Alves RS, de Resende MDV, Azevedo CF, Rocha JRdASdC, Nunes ACP, Carneiro APS, dos Santos GA (2020) Optimization of Eucalyptus breeding through random regression models allowing for reaction norms in response to environmental gradients. Tree Genet Genom 16:1–8. https://doi.org/10.1007/s11295-020-01431-5
Andjic V, Pegg GS, Carnegie AJ, Callister A, Hardy GES, Burgess TI (2010) Teratosphaeria pseudoeucalypti, new cryptic species responsible for leaf blight of Eucalyptus in subtropical and tropical Australia. Plant Pathol 59:900–912. https://doi.org/10.1111/j.1365-3059.2010.02308.x
Arnold PA, Kruuk LE, Nicotra AB (2019) How to analyse plant phenotypic plasticity in response to a changing climate. New Phytol 222:1235–1241. https://doi.org/10.1111/nph.15656
Barnes R, Burley J, Gibson G, Garcia de Leon J (1984) Genotype-environment interactions in tropical pines and their effects on the structure of breeding populations. Silvae Genet 33:186–198
Booth TH, Nix HA, Busby JR, Hutchinson MF (2014) BIOCLIM: the first species distribution modelling package, its early applications and relevance to most current MAXENT studies. Divers Distrib 20:1–9. https://doi.org/10.1111/ddi.12144
Brown CD, Davis HT (2006) Receiver operating characteristics curves and related decision measures: A tutorial. Chemom Intell Lab Syst 80:24–38
Burdon RD (1977) Genetic correlation as a concept for studying genotype-environment interaction in forest tree breeding. Silvae Genet 26:168–175
Burdon RD, Li Y, Suontama M, Dungey HS (2017) Genotype × site × silviculture interactions in radiata pine: knowledge, working hypotheses and pointers for research§. N Z J for Sc 47:1–7. https://doi.org/10.1186/s40490-017-0087-1
Butler JB, Harrison PA, Vaillancourt RE, Steane DA, Tibbits JF, Potts BM (2022) Climate adaptation, drought susceptibility, and genomic-informed predictions of future climate refugia for the Australian forest tree Eucalyptus globulus. Forests 13:575. https://doi.org/10.3390/f13040575
Calleja-Rodriguez A, Andersson Gull B, Wu HX, Mullin TJ, Persson T (2019) Genotype-by-environment interactions and the dynamic relationship between tree vitality and height in northern Pinus sylvestris. Tree Genet Genom 15:1–15. https://doi.org/10.1007/s11295-019-1343-8
Callister AN, England N, Collins S (2011) Genetic analysis of Eucalyptus globulus diameter, straightness, branch size, and forking in Western Australia. Can J for Res 41:1333–1343. https://doi.org/10.1139/x11-036
Callister AN, Bradshaw BP, Elms S, Gillies RW, Sasse JM, Brawner JT (2021) Single-step genomic BLUP enables joint analysis of disconnected breeding programs: an example with Eucalyptus globulus Labill. G3 Genes|Genomes|Genetics 11 https://doi.org/10.1093/g3journal/jkab253
Chen Z-Q, Karlsson B, Wu HX (2017) Patterns of additive genotype-by-environment interaction in tree height of Norway spruce in southern and central Sweden. Tree Genet Genom 13:1–14. https://doi.org/10.1007/s11295-017-1103-6
Christensen OF, Lund MS (2010) Genomic prediction when some animals are not genotyped. Genet Sel Evol 42:2. https://doi.org/10.1186/1297-9686-42-2
Christensen OF, Madsen P, Nielsen B, Ostersen T, Su G (2012) Single-step methods for genomic evaluation in pigs. Animal 6:1565–1571. https://doi.org/10.1017/S1751731112000742
Chung D, Keles S (2010) Sparse partial least squares classification for high dimensional data. Stat Appl Genet Mol Biol 9(1) https://doi.org/10.2202/1544-6115.1492
Costa e Silva J, Potts BM, Dutkowski GW (2006) Genotype by environment interaction for growth of Eucalyptus globulus in Australia. Tree Genet Genom 2:61–75. https://doi.org/10.1007/s11295-005-0025-x
Costa e Silva J, Borralho NMG, Araujo JA, Vaillancourt RE, Potts BM (2009) Genetic parameters for growth, wood density and pulp yield in Eucalyptus globulus. Tree Genet Genom 5:291–305. https://doi.org/10.1007/s11295-008-0174-9
Costa-Neto GMF, Morais Júnior OP, Heinemann AB, de Castro AP, Duarte JB (2020) A novel GIS-based tool to reveal spatial trends in reaction norm: upland rice case study. Euphytica 216:1–16. https://doi.org/10.1007/s10681-020-2573-4
Costa-Neto G, Fritsche-Neto R, Crossa J (2021) Nonlinear kernels, dominance, and envirotyping data increase the accuracy of genome-based prediction in multi-environment trials. Heredity 126:92–106. https://doi.org/10.1038/s41437-020-00353-1
Costa-Neto G, Fritsche-Neto R (2021) Enviromics: bridging different sources of data, building one framework. Crop Breeding and Applied Biotechnology 21 https://doi.org/10.1590/1984-70332021v21Sa25
Costa-Neto G, Crespo-Herrera L, Fradgley N, Gardner K, Bentley AR, Dreisigacker S, Fritsche-Neto R, Montesinos-López OA, Crossa J (2022) Envirome-wide associations enhance multi-year genome-based prediction of historical wheat breeding data. G3 Genes|Genomes|Genetics https://doi.org/10.1093/g3journal/jkac313
Crossa J, Gauch H Jr, Zobel RW (1990) Additive main effects and multiplicative interaction analysis of two international maize cultivar trials. Crop Sci 30:493–500. https://doi.org/10.2135/cropsci1990.0011183X003000030003x
Crossa J, Vargas M, Van Eeuwijk F, Jiang C, Edmeades G, Hoisington D (1999) Interpreting genotype× environment interaction in tropical maize using linked molecular markers and environmental covariables. Theor Appl Genet 99:611–625. https://doi.org/10.1007/s001220051276
Crossa J, Fritsche-Neto R, Montesinos-Lopez OA, Costa-Neto G, Dreisigacker S, Montesinos-Lopez A, Bentley AR (2021) The modern plant breeding triangle: optimizing the use of genomics, phenomics, and enviromics data. Front Plant Sci 12 https://doi.org/10.3389/fpls.2021.651480
Cullis BR, Jefferson P, Thompson R, Smith AB (2014) Factor analytic and reduced animal models for the investigation of additive genotype-by-environment interaction in outcrossing plant species with application to a Pinus radiata breeding programme. Theor Appl Genet 217:2193–2210. https://doi.org/10.1007/s00122-014-2373-0
de Souza MH, Pereira Júnior JD, Steckling SDM, Mencalha J, Dias FdS, Rocha JRdASdC, Carneiro PCS, Carneiro JEdS (2020) Adaptability and stability analyses of plants using random regression models. PLoS ONE 15:e0233200. https://doi.org/10.1371/journal.pone.0233200
Downham R, Gavran M (2019) Australian plantation statistics 2019 update, ABARES technical report 19.2. Canberra https://doi.org/10.25814/5cc65ae71465f
Dungey H, Low C, Lee J, Miller M, Fleet K, Yanchuk A (2012) Developing breeding and deployment options for Douglas-fir in New Zealand: breeding for future forest conditions. Silvae Genet 61:104. https://doi.org/10.1515/sg-2012-0013
Dutkowski GW, Silva JCE, Gilmour AR, Lopez GA (2002) Spatial analysis methods for forest genetic trials. Can J for Res 32:2201–2214. https://doi.org/10.1139/x02-111
Dutkowski GW, Potts BM, Pilbeam D (2015) Genotype x environment interaction of Eucalyptus globulus in Australia has similar patterns at the provenance and additive levels. IUFRO Eucalypt Conference Zhanjiang City, Guangdong, China
Fradgley NS, Bacon J, Bentley AR, Costa-Neto G, Cottrell A, Crossa J, Cuevas J, Kerton M, Pope E, Swarbreck SM (2023) Prediction of near-term climate change impacts on UK wheat quality and the potential for adaptation through plant breeding. Glob Change Biol 29:1296–1313. https://doi.org/10.1111/gcb.16552
Freeman JS, Marques CMP, Carocha V, Borralho N, Potts BM, Vaillancourt RE (2007) Origins and diversity of the Portuguese Landrace of Eucalyptus globulus. Ann for Sci 64:639–647
Gapare WJ, Ivković M, Liepe KJ, Hamann A, Low CB (2015) Drivers of genotype by environment interaction in radiata pine as indicated by multivariate regression trees. For Ecol Man 353:21–29. https://doi.org/10.1016/j.foreco.2015.05.027
Gezan SA, de Carvalho MP, Sherrill J (2017) Statistical methods to explore genotype-by-environment interaction for loblolly pine clonal trials. Tree Genet Genom 13:1–11. https://doi.org/10.1007/s11295-016-1081-0
Gilmour AR, Gogel BJ, Cullis BR, Welham SJ, Thompson R (2015) ASReml User Guide Release 4.1 Functional Specification. VSN International Ltd, Hemel Hempstead, HP1 1ES, UK
Grundy M, Rossel RV, Searle R, Wilson P, Chen C, Gregory L (2015) Soil and landscape grid of Australia. Soil Research 53:835–844. https://doi.org/10.1071/SR15191
Hardner CM, Dieters M, Dale G, DeLacy I, Basford K (2010) Patterns of genotype-by-environment interaction in diameter at breasts height at age 3 for eucalypt hybrid clones grown for reafforestation of lands affected by salinity. Tree Genet Genom 6 https://doi.org/10.1007/s11295-010-0295-9
Heinemann AB, Costa-Neto G, Fritsche-Neto R, da Matta DH, Fernandes IK (2022) Enviromic prediction is useful to define the limits of climate adaptation: a case study of common bean in Brazil. Field Crop Res 286:108628. https://doi.org/10.1016/j.fcr.2022.108628
Henderson CR (1984) Applications of linear models in animal breeding. University of Guelph, Guelph, Ontario
Heslot N, Akdemir D, Sorrells ME, Jannink J-L (2014) Integrating environmental covariates and crop modeling into the genomic selection framework to predict genotype by environment interactions. Theor Appl Genet 127:463–480. https://doi.org/10.1007/s00122-013-2231-5
Ivković M, Gapare W, Yang H, Dutkowski G, Buxton P, Wu H (2015) Pattern of genotype by environment interaction for radiata pine in southern Australia. Ann for Sci 72:391–401. https://doi.org/10.1007/s13595-014-0437-6
Jarquín D, Crossa J, Lacaze X, Du Cheyron P, Daucourt J, Lorgeou J, Piraux F, Guerreiro L, Pérez P, Calus M, Burgueño J, de los Campos G, (2014) A reaction norm model for genomic selection using high-dimensional genomic and environmental data. Theor Appl Genet 127:595–607. https://doi.org/10.1007/s00122-013-2243-1
Jeffrey SJ, Carter JO, Moodie KM, Beswick AR (2001) Using spatial interpolation to construct a comprehensive archive of Australian climate data. Environ Model Softw 16:309–330. https://doi.org/10.1016/S1364-8152(01)00008-1
Johnson G (1997) Site-to-site genetic correlations and their implications on breeding zone size and optimum number of progeny test sites for coastal Douglas-fir. Silvae Genetica 46:280–285
Kang MS (2002) Genotype–environment interaction: progress and prospects. In: Kang MS (ed) Quantitative genetics, genomics, and plant breeding. Cabi Publishing, 221–243 https://doi.org/10.1079/9780851996011.0221
Lauer E, Sims A, McKeand S, Isik F (2021) Genetic parameters and genotype-by-environment interactions in regional progeny tests of Pinus taeda L. in the southern USA. For Sci 67:60–71. https://doi.org/10.1093/forsci/fxaa035
Lê Cao K-A, Boitard S, Besse P (2011) Sparse PLS discriminant analysis: biologically relevant feature selection and graphical displays for multiclass problems. BMC Bioinformatics 12:1–17. https://doi.org/10.1186/1471-2105-12-253
Legarra A, Aguilar I, Misztal I (2009) A relationshp matrix including full pedigree and genomic information. J Dairy Sci 92:4656–4663. https://doi.org/10.3168/jds.2009-2061
Li Y, Dutkowski GW, Apiolaza LA, Pilbeam DJ, Costa e Silva J, Potts BM (2007) The genetic architecture of a Eucalyptus globulus full-sib breeding population in Australia. For Genet 12:167–179
Li Y, Suontama M, Burdon RD, Dungey HS (2017) Genotype by environment interactions in forest tree breeding: review of methodology and perspectives on research and application. Tree Genet Genom 13:60. https://doi.org/10.1007/s11295-017-1144-x
Li X, Guo T, Wang J, Bekele WA, Sukumaran S, Vanous AE, McNellie JP, Tibbs-Cortes LE, Lopes MS, Lamkey KR (2021) An integrated framework reinstating the environmental dimension for GWAS and genomic selection in crops. Mol Plant 14:874–887. https://doi.org/10.1016/j.molp.2021.03.010
Mendham DS, White DA, Battaglia M, McGrath JF, Short TM, Ogden GM, Kinal J (2011) Soil water depletion and replenishment during first- and early second-rotation Eucalyptus globulus plantations with deep soil profiles. Agric for Meteorol 151:1568–1579. https://doi.org/10.1016/j.agrformet.2011.06.014
Meyer K (2009) Factor-analytic models for genotype × environment type problems and structured covariance matrices. Genet Sel Evol 41:1–11. https://doi.org/10.1186/1297-9686-41-21
Misztal I, Tsuruta S, Lourenco DAL, Masuda Y, Aguilar I, Legarra A, Vitezica Z (2014) Manual for BLUPF90 family of programs. University of Georgia, Athens, USA
Monteverde E, Gutierrez L, Blanco P, Pérez de Vida F, Rosas JE, Bonnecarrère V, Quero G, McCouch S (2019) Integrating molecular markers and environmental covariates to interpret genotype by environment interaction in rice (Oryza sativa L.) grown in subtropical areas. G3-Genes Genom Genet 9:1519–1531. https://doi.org/10.1534/g3.119.400064
Ogut F, Maltecca C, Whetten R, McKeand S, Isik F (2014) Genetic analysis of diallel progeny test data using factor analytic linear mixed models. For Sci 60:119–127. https://doi.org/10.5849/forsci.12-108
Pinkard E, Battaglia M, Bruce J, Matthews S, Callister AN, Hetherington S, Last I, Mathieson S, Mitchell C, Mohammed C, Musk R, Ravenwood I, Rombouts J, Stone C, Wardlaw T (2015) A history of forestry management responses to climatic variability and their current relevance for developing climate change adaptation strategies. Forestry 88:155–171. https://doi.org/10.1093/forestry/cpu040
Pinkard E, Wardlaw T, Kriticos D, Ireland K, Bruce J (2017) Climate change and pest risk in temperate eucalypt and radiata pine plantations: a review. Aust for 80:228–241. https://doi.org/10.1080/00049158.2017.1359753
Porker K, Coventry S, Fettell N, Cozzolino D, Eglinton J (2020) Using a novel PLS approach for envirotyping of barley phenology and adaptation. Field Crop Res 246:107697. https://doi.org/10.1016/j.fcr.2019.107697
Raymond CA (2011) Genotype by environment interactions for Pinus radiata in New South Wales, Australia. Tree Genet Genom 7:819–833. https://doi.org/10.1007/s11295-011-0376-4
Resende RT, Piepho HP, Rosa GJM, e Silva-Junior OB, Silva FF, de Resende MDV, Grattapaglia D (2021) Enviromics in breeding: applications and perspectives on envirotypic-assisted selection. Theor Appl Genet 134:95–112. https://doi.org/10.1007/s00122-020-03684-z
Rohart F, Gautier B, Singh A, Lê Cao K-A (2017) mixOmics: an R package for ‘omics feature selection and multiple data integration. PLoS Comput Biol 13(11):e1005752. https://doi.org/10.1371/journal.pcbi.1005752
Romay MC, Malvar RA, Campo L, Alvarez A, Moreno-González J, Ordás A, Revilla P (2010) Climatic and genotypic effects for grain yield in maize under stress conditions. Crop Sci 50:51–58. https://doi.org/10.2135/cropsci2008.12.0695
Sáenz-Romero C, O’Neill G, Aitken SN, Lindig-Cisneros R (2020) Assisted migration field tests in Canada and Mexico: lessons, limitations, and challenges. Forests 12(1):9. https://doi.org/10.3390/f12010009
Salas M, Nieto V, Perafán L, Sánchez A, Borralho NMG (2014) Genetic parameters and comparison between native and local landraces of Eucalyptus globulus Labill. ssp. globulus growing in the central highlands of Colombia. Ann for Sci 71:405–414. https://doi.org/10.1007/s13595-013-0342-4
Sanhueza R, White T, Huber D, Griffin A (2002) Genetic parameters estimates, selection indices and predicted genetic gains from selection of Eucalyptus globulus in Chile. For Genet 9:19–29
Shalizi MN, Isik F (2019) Genetic parameter estimates and GxE interaction in a large cloned population of Pinus taeda L. Tree Genet Genom 15:1–13. https://doi.org/10.1007/s11295-019-1352-7
Shelbourne C (1972) Genotype-environment interactions: its study and its implications in forest tree improvement. IUFRO genetics-SABRAO joint symposia B-1. Tokyo 1972:1–28
Smethurst PJ, McVicar TR, Huth NI, Bradshaw BP, Stewart SB, Baker TG, Benyon RG, McGrath JF, Van Niel TG (2022) Nitrate uptake from an aquifer by two plantation forests: plausibility strengthened by process-based modelling. Forests 13:184. https://doi.org/10.3390/f13020184
Smith A, Cullis B, Thompson R (2001) Analyzing variety by environment data using multiplicative mixed models and adjustments for spatial field trend. Biometrics 57:1138–1147. https://doi.org/10.1111/j.0006-341X.2001.01138.x
Southerton S, Dillon S, Thumma B (2011) Identification of genes and alleles influencing wood development in Eucalyptus. BMC Proc 5(suppl. 7):I5. https://doi.org/10.1186/1753-6561-5-S7-I5
Stackpole DJ, Vaillancourt RE, de Aguiar M, Potts BM (2010) Age trends in genetic parameters for growth and wood density in Eucalyptus globulus. Tree Genet Genom 6:179–193
Thavamanikumar S, Arnold RJ, Luo J, Thumma BR (2020) Genomic studies reveal substantial dominant effects and improved genomic predictions in an open-pollinated breeding population of Eucalyptus pellita. G3-Genes Genom Genet 10:3751–3763. https://doi.org/10.1534/g3.120.401601
Thomson AM, Crowe KA, Parker WH (2010) Optimal white spruce breeding zones for Ontario under current and future climates. Can J for Res 40:1576–1587. https://doi.org/10.1139/X10-112
Tolhurst DJ, Gaynor RC, Gardunia B, Hickey JM, Gorjanc G (2022) Genomic selection using random regressions on known and latent environmental covariates. Theor Appl Genet 135:3393–3415. https://doi.org/10.1007/s00122-022-04186-w
Ukrainetz NK, Yanchuk AD, Mansfield SD (2018) Climatic drivers of genotype–environment interactions in lodgepole pine based on multi-environment trial data and a factor analytic model of additive covariance. Can J for Res 48:835–854. https://doi.org/10.1139/cjfr-2017-0367
Van Eeuwijk FA, Bustos-Korts DV, Malosetti M (2016) What should students in plant breeding know about the statistical aspects of genotype × environment interactions? Crop Sci 56:2119–2140. https://doi.org/10.2135/cropsci2015.06.0375
VanRaden PM (2008) Efficient methods to compute genomic predictions. J Dairy Sci 91:4414–4423. https://doi.org/10.3168/jds.2007-0980
Westell RA, Quaas RL, van Vleck LD (1988) Genetic groups in animal models. J Dairy Sci 71:1310–1318
Wold S, Sjöström M, Eriksson L (2001) PLS-regression: a basic tool of chemometrics. Chemom Intell Lab Syst 58:109–130. https://doi.org/10.1016/S0169-7439(01)00155-1
Xu Y (2016) Envirotyping for deciphering environmental impacts on crop plants. Theor Appl Genet 129:653–673. https://doi.org/10.1007/s00122-016-2691-5
Ye TZ, Jayawickrama KJS (2008) Efficiency of using spatial analysis in first-generation coastal Douglas-fir progeny tests in the US Pacific Northwest. Tree Genet Genom 4:677–692. https://doi.org/10.1007/s11295-008-0142-4
Ying CC, Yanchuk AD (2006) The development of British Columbia’s tree seed transfer guidelines: purpose, concept, methodology, and implementation. For Ecol Man 227:1–13. https://doi.org/10.1016/j.foreco.2006.02.028
Yu Y, Aitken SN, Rieseberg LH, Wang T (2022) Using landscape genomics to delineate seed and breeding zones for lodgepole pine. New Phytol 235:1653–1664. https://doi.org/10.1111/nph.18223
Acknowledgements
The authors are grateful to Rowland Burdon and the anonymous reviewers for helpful comments and suggestions. We acknowledge Australian Bluegum Plantations and Hancock Victorian Plantations for allowing publication of this work. Andrew Callister has received research support from Australian Bluegum Plantations and Hancock Victorian Plantations, including for the completion of this work.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions
Author information
Authors and Affiliations
Contributions
The project was conceptualized by Andrew Callister and Ben Bradshaw. Statistical analyses and manuscript writing were conducted by Andrew Callister. Technical advice and manuscript revision provided by Germano Costa-Neto, Jose Crossa, and Jeremy Brawner. Trial establishment, maintenance, measurement, sampling, and data preparation were conducted by Ben Bradshaw and Stephen Elms.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Communicated by C. Kulheim
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Callister, A.N., Costa-Neto, G., Bradshaw, B.P. et al. Enviromic prediction enables the characterization and mapping of Eucalyptus globulus Labill breeding zones. Tree Genetics & Genomes 20, 3 (2024). https://doi.org/10.1007/s11295-023-01636-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-023-01636-4