Abstract
Mitochondrial dysfunction is a well-known contributor to aging and age-related diseases. The precise mechanisms through which mitochondria impact human lifespan, however, remain unclear. We hypothesize that humans with exceptional longevity harbor rare variants in nuclear-encoded mitochondrial genes (mitonuclear genes) that confer resistance against age-related mitochondrial dysfunction. Here we report an integrated functional genomics study to identify rare functional variants in ~ 660 mitonuclear candidate genes discovered by target capture sequencing analysis of 496 centenarians and 572 controls of Ashkenazi Jewish descent. We identify and prioritize longevity-associated variants, genes, and mitochondrial pathways that are enriched with rare variants. We provide functional gene variants such as those in MTOR (Y2396Lfs*29), CPS1 (T1406N), and MFN2 (G548*) as well as LRPPRC (S1378G) that is predicted to affect mitochondrial translation. Taken together, our results suggest a functional role for specific mitonuclear genes and pathways in human longevity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Geroscience research has been focused on understanding the molecular mechanisms driving the cellular and physiological functional decline that comes with increased age [1, 2]. Understanding the aging process to develop geroprotective interventions that extend health, quality of life, and independence during the final decades of life is one goal of geroscience research [2, 3]. Geroscience research has succeeded in identifying key biological pathways, labeled as “hallmarks or pillars of aging,” that drive the aging process [4, 5]. One of these hallmarks is the age-related dysfunction of mitochondria, found to contribute to biological aging across evolutionarily divergent species [6].
Mitochondrial dysfunction contributes to multiple physiological and functional deficits during aging, such as sarcopenia, frailty, reduced cardiac output, chronic inflammation (“inflammaging”), and cognitive decline, as well as numerous age-related diseases in humans and other animals [4, 5, 7]. Decreased mitochondrial function has been associated with overall age-related functional decline in nearly all tissues, especially those demanding higher energy, such as the heart, muscle, brain, and liver [8, 9]. Preservation of mitochondrial function through genetic or pharmacological measures has been shown to alter lifespan, the progression of aging, and age-related disease pathologies in model organisms [10,11,12,13,14,15,16,17,18]. For example, lifespan is extended in mice when mitochondrial uncoupling protein 2 (UCP2) [19] or mitochondrial catalase [20] is overexpressed. Somewhat paradoxically, lifespan can also be increased upon modest inhibition of mitochondrial electron transport chain (ETC) function, such as knockdown (KD) of the complex IV component SURF1 in mice and in models of reduced ETC function in worms, yeast, and flies [21, 22]. The underlying mechanisms remain unresolved, and strong ETC inhibition uniformly causes severe disease and early death in both mice and humans. Mitochondrial function is also tightly interconnected with conserved longevity-assurance pathways that regulate aging in laboratory animals. These include nutrient sensing and metabolic regulation through the mammalian target of rapamycin (MTOR), insulin/insulin-like growth factor signaling (IIS), AMP-activated protein kinase (AMPK), and sirtuins [10, 23, 24].
Despite the extensive research in model organisms, it is still unclear whether the same genes and pathways in mitochondrial function are important for human longevity. The natural occurrence of genetic variants in humans that affect longevity offers an ideal starting point for addressing the knowledge gap. The best examples of such “natural mutants” are human centenarians, who managed to delay or ward off the diseases that normally begin to plague humans starting at middle age [25]. As demonstrated by multiple studies, this extreme human longevity has a strong genetic component and longevity is likely mediated via heritable resilience to age-associated diseases [26,27,28,29,30,31]. Given that the frequency of centenarians is only ~ 1/5000–7000 individuals [32], it has been hypothesized that genetic variation that contributes to exceptional longevity in centenarians is rare or absent in younger control populations [32, 33].
Identification of potentially protective rare variants in centenarians requires deep sequencing analysis of centenarian genomes [34,35,36]. Alternatively, targeted sequencing of candidate genes involved in the conserved pathways of aging provides an opportunity to test the relevance of candidate genes and pathways in human longevity. Indeed, by a comprehensive resequencing analysis of candidate genes focused on the well-established conserved pathways of aging, we have identified and characterized rare, functional variants in IGF1R [37, 38], NFKBIA [39], CLU [39], and PRKCH [39] that are associated with human longevity. While these variants are enriched in centenarians at a nominal significance (p < 0.05), functional analysis of these variants using cell models demonstrated parallel mechanisms between humans and model organisms. For example, the longevity-associated variants in the IGF1R gene cause reduced signaling through the insulin/insulin-like growth factor 1 signaling (IIS) pathway [37, 40], one of the major conserved pathways of aging [37, 38]. Furthermore, functional analyses of the longevity-associated gene variants in NFKBIA, CLU, and PRKCH suggest reduced signaling through the PKC/NF-κB pathways [39]. Thus, deep sequencing analysis of candidate genes in centenarians has the translational potential to reveal functional relevance of conserved genes and pathways on human longevity.
Here we describe an integrated functional genomics approach to identify rare functional variants in mitonuclear genes discovered by target capture sequencing analysis of centenarians and controls. We identify and prioritize candidate variants, genes, and mitochondrial pathways that are associated with human longevity and provide novel variants worthy of functional studies to investigate underlying mechanisms. Our results suggest that naturally occurring, potentially functional, rare DNA sequence variation in mitonuclear genes may contribute to human longevity and healthy aging by affecting mitochondrial function and homeostasis as has been demonstrated in model organisms.
Methods
Compiling and categorizing mitonuclear candidate genes for sequencing
The list of candidate genes to include for sequencing was compiled by comprehensive literature and database mining. The GeneCards Human Gene Database localization information [41] (https://www.genecards.org/), MitoCarta version 1 accessed 2014–2015 [42], and MitoMiner version 3 accessed 2014–2015 [43] (http://mitominer.mrc-mbu.cam.ac.uk/release-4.0/begin.do) were used to identify nuclear-encoded genes in the mitochondrial proteome. The Online Mendelian Inheritance in Man database was used to gather disease and phenotype information for candidate genes (OMIM®), accessed 2015 [44] (https://www.omim.org/). Human Ageing Genomic Resources (HAGR) [45] (https://genomics.senescence.info/), such as the Ageing Gene Database (GenAge, accessed 2015 and updated January 2019) and the Digital Aging Atlas (DAA) [46] (http://ageing-map.org/, accessed 2015 and updated January 2019), were used to identify and verify candidate genes associated with lifespan, aging, age-related disease, and other related traits in model organisms and humans.
Functional categorization of the candidate genes was done using a combination of detailed gene information from GeneCards gene summaries, Gene Ontology (GO) from the Reactome database [47] (https://www.reactome.org/), and Gene Ontology Resource [48] (http://geneontology.org/). Subcellular compartment localization scores were sourced from the Compartments database, which integrates numerous types of protein localization evidence from other sources and is linked through GeneCards (accessed January 2019, [49]) (https://compartments.jensenlab.org/Search).
Sequencing and post-sequencing data annotation and summary statistics
Target capture next-generation pooled sequencing was conducted using equimolar concentrations of DNA samples from the well-established AJ longevity cohort of 494 cases (synonymous with probands or centenarians) and 572 controls [50]. The samples were combined as previously described [51] in 43 total pools for sequencing, each containing samples from 25 individuals per pool. The 43 pools were combined into 4 captured samples, with 11 pools per the first three captured samples and 10 pools in the fourth captured sample. Target capture probes were designed to enrich for all coding regions and some regulatory regions: exons, exon–intron junctions, 5′-untranslated and 3′-untranslated regions, and 2 kb upstream of the proximal promoter regions of candidate genes. The total target size for the mitonuclear candidate genes spanned ~ 4.6 Mb. The pool sequencing achieved a median coverage per pool of ~ 378.12-fold in total, which is equivalent to ~ 15-fold coverage per sample. Libraries were sequenced on the Illumina HiSeq2000 flow-cell at Axeq Technologies. The resulting data was processed at Einstein’s Genomics Computational Core. P-value was calculated by Fisher’s exact test. Final annotations including mapped gene, variant location, and variant impact were adjusted (unless otherwise stated) for each detected variant by Ensembl Variant Effect Predictor (VEP) (http://grch37.ensembl.org/info/docs/tools/vep/index.html) or by manually checking against the human reference genome (GRCh37/hg19) in the UCSC genome browser (University of California Santa Cruz Genomics Institute (https://genome.ucsc.edu/cgi-bin/hgGateway)) [52]. The variant ID annotations were updated with dbSNP144, accessed April 2016. The Manhattan plot was generated in R and all other summary statistics were calculated in Microsoft Excel. To determine the proportion of variants that are centenarian-enriched and common or centenarian-enriched and rare, the alternative (ALT) and reference (REF) alleles were first adjusted to ensure that ALT is the minor allele (MIN), followed by calculation of both MAF in each group and the cohort minor allele frequency (MAF), both based on observed diploid allele counts.
GWAS analysis
The GWAS catalog from the National Human Genome Research Institute (NHGRI) and European Molecular Biology Laboratory-European Bioinformatics Institute (EMBL-EBI) (NHGRI-EBI) was mined for age-related trait associations with mitonuclear candidate genes (accessed 04/2016, http://www.ebi.ac.uk/gwas). Data was collected and compiled as described previously [53]. Briefly, all data for candidate mitonuclear genes reported in the GWAS catalog were extracted for analysis and categorized based on associated trait(s). Specifically, genes associated with age-related disease at p ≤ 10−8 and “lifespan,” “aging,” and “longevity” at p ≤ 10−6 were included for downstream analysis. All genes associated with diseases of related categories were grouped together (e.g., cardiovascular disease traits include “coronary heart disease,” “myocardial infarction,” “homocysteine levels,” and other disease terms and high confidence risk biomarkers that fit into the larger category of “cardiovascular disease”). If a gene and trait association from the same study (i.e., not a replication study) was reported multiple times, the association with the most significant p-value was retained for this analysis and duplicates were removed. From there, the number of associations and unique genes was quantified and graphed per category.
We used the recently developed R script for Enrichment of Age-Related Diseases in GWAS (EARDiG) https://github.com/biosinodx/EARDiG.
Gene-based SKAT analysis
Gene-based rare variant analysis by Sequence Kernel Association Tests (SKAT) was accomplished through the R package “SKAT” [54]. SKATs were run twice, on sequencing data including and excluding singletons. The SKAT results reported are based on data excluding singletons as this is more stringent and there were few differences between SKAT results including and excluding singletons. Model organism lifespan data was extracted from GenAge, linked through HAGR using orthologs reported by GeneCards, the homologous human gene symbol, or Entrez IDs.
To ensure consistent results for GO enrichment on SKAT results, we compared against different reference backgrounds including the whole human genome (~ 21,000 genes), the mitochondrial proteome (~ 2220 genes), and the mitonuclear candidate list (~ 660 genes). We used the Protein Analysis Through Evolutionary Relationships database (PANTHER, http://pantherdb.org/).
We defined a comprehensive annotation criteria to filter top variants of interest for in silico prediction: (1) single-variant association signal with longevity (p ≤ 0.05); (2) directionality of enrichment, with higher priority for those that are enriched in centenarians compared to controls (i.e., observed MAFcases > observed MAFcontrols); (3) location, with higher priority for nonsynonymous variants (nsVs) over non-coding variants (ncVs); (4) impact, with higher priority for nsVs with non-conserved (NC) biochemical and physiochemical amino acid changes, and damaging or deleterious predictions, and higher priority for ncVs with strong Regulome database scores, that fall within predicted regulatory regions as assessed by manual checking of DNase hypersensitivity, promoter-associated and enhancer-associated histone marks, and TF-binding evidence in the UCSC genome browser ensemble track hubs [55]; (5) validation by genotyping and other population MAF information obtainable for further confidence—global MAF from 1000Genomes, average MAF from GnomAD (~ 15,000 whole genomes and ~ 125,000 exomes) and Exac (~ 60,000 exomes) and MAF from the Geisinger and Regeneron whole-exome sequencing study of the DiscovEHR cohort containing ~ 50,700 individuals [56]; and (6) comprehensive annotations showing a priori biological evidence of phenotypes associated with the SNP- or its respective gene in humans or model organisms including evolutionary conservation at the variant site, protein domains or post-translational modifications mapped at or near the variant site, gene-associated or SNP-associated phenotype information including gene expression changes with age in humans or model organisms and model organism longevity information.
Many sources were used in conjunction with the Genome Reference Consortium Human Build 37 to obtain and corroborate annotations for variant filtering. Ensembl VEP [57] was used as a source of variant position information, global or average control MAF information from the 1000Genomes Project, gnomAD, and ExAC and prediction scores for protein impact of nsVs, including Condel. The Model organism Aggregated Resources for Rare Variant ExpLoration (MARRVEL) was a source of variant site evolutionary conservation in model organisms, CADD protein impact prediction scores, gnomAD and ExAC population allele frequencies, and SNP-associated human phenotype ontology (HPO) data from the sequencing of ~ 10,500 exomes at the University of Washington’s Mendelian Genetics Center via the linked source—Geno2MP (Marrvel.org) [58].
The UCSC genome browser was used with track hubs to annotate gene and SNP associations (Genetic Association Database of Complex Disease and Disorders (GAD), The NHGRI-GWAS Catalog, clinically associated SNPs by Flagged SNPs in dbSNP144, and the Genomenon Mastermind track for literature mining of genetic variants). ENCODE tracks hubs on UCSC genome browser were used to annotate regulatory regions (ENCODE/broad chromatin state segmentation, DNase hypersensitivity cluster, histone modifications, and transcription factor ChIP-seq, and the Genetic Information Research Institute RepeatMasker track for overlapping elements such as LINEs/SINEs) [59]. The UCSC genome browser was also used with tracks to annotate protein information such as domains or structures, modifications, sequences, and alignments (UniProt Protein annotations, and PeptideAtlas MassSpec peptide sequence).
Amino acid biochemical conservation was manually annotated as conservative (‘C), non-conservative (NC), or radical (‘R) based on information about amino acid R-group properties including charge, polarity, hydropathy, and size [60,61,62,63,64] (https://en.wikipedia.org/wiki/Proteinogenic_amino_acid). Amino acid substitutions designated “C” have zero biochemical property differences, while “NC” substitutions have at least 1 biochemical difference, and “R” substitutions have ≥ 2 changes including a change in size which is regarded as more radical change because of the potential for steric hindrance.
Prediction annotations of the likelihood that a polymorphism is neutral or damaging to protein function were determined using two independent, experimentally validated algorithms that automatically provide prediction scores based on multiple lines of evidence for such polymorphisms: the CONsensus DELeteriousness score of missense mutations (Condel) [65] and the Combined Annotation Dependent Depletion scores (CADD, CADDphred) [66, 67]. When the reference and alternative amino acids are “C,” the biochemical and physiochemical characteristics are more neutral and reduce the likelihood of a functional protein change; e.g., the reference and alternative amino acids are of similar size, charge, or polarity so the biochemical change is not severe and the expectation is that Condel and CADD would predict the SNP to be “neutral or benign,” whereas “NC” or “R” substitutions infer that the properties of the two amino acids differ enough to potentially alter protein structure and/or function and the Condel and CADD predictions would be “probably or possible damaging or deleterious.”
Genotyping
High-throughput iPLEX mass array genotyping of 112 variants was conducted at the Einstein Genomics Core using amplified DNA for 1083 AJ cohort individuals. Genotyping analysis and variants were validated based on comparison of allele frequencies between pooled sequencing and genotyping. A total of 100 of the variants were successfully genotyped, while 12 variants failed at primer design. Thirty-four variants failed to validate for technical and quality reasons.
Protein modeling
The 2D protein domain mapping and schematic was created using data and figures modified from feature viewers on UniProt [68] (https://www.uniprot.org/) and PDB [69] (The Protein Data Bank, rcsb.org) for the LRPPRC protein (ID: P42704). The 3D rendering of the partial structure of LRPPRC is a homology-based model generated from a template (5nnr.1.A) based on a protein with similar sequence (i.e., homology model) using the “Build Model” function with the FASTA sequence for the full 130 kDa protein (UniProtKB/Swiss-Prot: P42704.3) on SWISS-MODEL ExPASy [70] (https://swissmodel.expasy.org/). This was the best choice homology model that encompassed the C-terminal region of interest in our protein as no crystal structure for LRPPRC has been resolved yet. The University of California San Francisco’s (UCSF) Resource for Biocomputing, Visualization, and Informatics (RBVI) Chimera tool was used to depict and manipulate the tertiary structure of the homology model, generate graphics, and analyze and calculate distances between nearest neighboring atoms and residues and between potential hydrogen bonds [71].
Results
Selection of candidate aging-related mitonuclear genes
Candidate nuclear-encoded mitochondrial genes (mitonuclear genes) were chosen on the basis of biological evidence for their function in mitochondria, aging processes, age-related diseases, and longevity or lifespan in model organisms or humans (see the “Methods” section). We excluded genes encoded by the mitochondrial genome (mtDNA) from our candidate list in order to focus specifically on mitonuclear gene variants. In total, ~ 680 mitonuclear genes were chosen as candidates for the target capture sequencing-based discovery of longevity-associated variants (LAVs) and genes (LAGs). There are many mitochondrial pathways that are important in aging and age-related pathology; thus, we were interested in how our large list of candidates represented these mitochondrial functions, which could be useful in downstream analysis. We divided all mitonuclear candidates into categories according to their respective functions and pathways within and around the mitochondria, according to GO and detailed gene summaries provided by GeneCards (Fig. 1a and Tables S1, S2). As expected, many of the candidate genes have primary functions in metabolism; however, the mitonuclear candidate genes also represent a diversity of mitochondrial functions in cellular homeostasis. The categories representing the most candidate genes, with > 100 genes per category, are lipid metabolism, oxidative phosphorylation, protein expression, and proteostasis. Although not a requisite, 78% of our candidate genes have high mitochondrial localization scores (≥ 3; with 65% having the highest confidence score of 5) as indicated in GeneCards and Compartments (Fig. 1b).
Mitochondrial functions and localization score of selected mitonuclear candidate genes. a Nuclear-encoded genes with mitochondrial functions placed in respective functional categories. Candidates chosen based on evidence for association with aging or longevity in model organisms or humans. b Localization scores were retrieved from Compartments, linked through GeneCards. Increasing localization score indicates increasing confidence that the gene is localized to the subcellular compartment (0—lowest confidence, 5—highest confidence score for mitochondrial localization)
Characterization of candidate mitonuclear genes by GWAS-detected phenotypes
Common human genetic variation associated with different phenotypes and traits, including disease risk variants and lifespan-associated variants, can be sourced from published GWAS data. Lifespan-associated variants are defined by trait designations including “aging (time to death/event),” “lifespan,” “longevity (85 +),” and “parental longevity.” We utilized GWAS data to further annotate biological information, in the context of human aging and age-related disease, to our mitonuclear candidate genes. To identify the number of age-related disease and lifespan associations with our candidate genes, we extracted the data pertaining to mitonuclear candidate genes and age-related diseases from the GWAS Catalog using a significance threshold of p < 5 × 10−8 for disease associations, and p < 5 × 10−6 for lifespan associations. In total, ~ 16% of the 659 mitonuclear candidates harbor common variants associated with human age-related disease (Fig. S1a) or lifespan in GWAS (Fig. S1b). Metabolic disease contains the greatest number of associations, while inflammatory/immunological diseases are associated with the greatest number of unique mitonuclear genes. The most common candidate genes associating with multiple disease categories were LIPC and FADS1/2, which are lipid metabolism genes, and CPS1, which is important for amino acid metabolism and the urea cycle. Only 6 candidates were associated with lifespan terms: SUCLA2, ECHS1, PARKIN, CYP51A1, PFKM, and TOMM40-APOE. Using our newly developed tool (EARDiG) for determining if genes within a query set are statistically enriched in GWAS of age-related diseases (“Methods” section), it was determined that the mitonuclear candidates are significantly enriched in metabolic diseases (p = 0.018) (Fig. S1c) but not in any other category (Fig. S1d).
Discovery of genetic variants in candidate mitonuclear genes
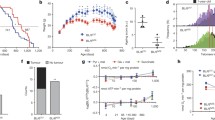
To efficiently sequence all possible variants, we previously established a method of next-generation sequencing on pooled samples with probes designed to capture specific DNA targets [51, 72]. This method of sequencing permits the capture of rare coding and regulatory variants. Samples sequenced are from 496 probands and 572 controls of the AJ Longevity Genes Project [73]. Proband or case samples include centenarians with an average lifespan ≥ 95 years, while control samples come from individuals with no family history of longevity whose parents lived to a maximum of 85 years and are often spouses of probands’ offspring allowing further control for the environment and lifestyle.
In total, 66,419 variants were sequenced and annotated to ~ 659 of our 683 mitonuclear candidate genes. Twenty-four genes were removed after quality control or because no variation was detected in those target regions, suggesting high sequence conservation. Due to the lack of statistical power that often restricts single-variant analysis in rare variant association studies, no variants reached multiple testing correction significance except the most significant common variant, rs689454, located upstream of the NQO1 gene with an association of p = 1 × 10−6 (FDR = 0.050) (Fig. 2). The top 10 variants are located in the following genes (in order of highest to lowest significance): NQO1, NOS1, SUCLG2, DNM1L, PARP1, PI4K2A, EIF4G2, NRF1, ZNF746, and FOXM1.
Significant gene variants discovered by pooled sequencing. Manhattan plot representing all 66,419 variants discovered as individual dots, with association signal significance (-log10 (p-value)) with longevity in the AJ cohort as a function of chromosomal location. Blue line is p = 0.001, red line is p = 0.01, and black line is p = 0.05. Red box highlights the most significant NQO1 variant. The majority of variants are novel (~ 78% as determined by Ensembl VEP and based on lack of variant IDs from dbSNP144)
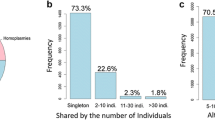
The majority of detected variants are novel (~ 78% as determined by Ensembl VEP and based on lack of variant IDs from dbSNP144); however, nearly half of the total variants are singletons, representing a large portion of the novel variant pool that can be detected by deep sequencing (Fig. 3a). The total number of variants with a nominal association p-value ≤ 0.050 is 854, of which ~ 40% (328) are common (MAF > 0.010) and ~ 60% (526) are rare (MAF ≤ 0.010). As expected, the proportion of non-coding (ncVs) or non-exonic variants exceeds that of coding (cVs) or exonic variants. Among cVs, 67% are nonsynonymous (nsVs), 26% are synonymous (sVs), 4% are stop gain (sg or nonsense), 3% are frameshifts (fs), and < 1% are in-frame insertions or deletions (InDels.) (Fig. 3b). Excluding singletons does not substantially change the proportions of the different types of variants (Fig. 3c).
The distribution and characteristics of mitonuclear gene variants discovered by pooled sequencing. a The relationship between singleton variants, allele frequency, and association significance in sequencing data. Singleton variants account for approximately half of the variants identified by sequencing. b Distribution and percentages of different types of genetic variations as annotated by genic location (exonic/coding, non-exonic/non-coding) and impact. Pie charts express both numbers and the percent of variants in the respective pie. c Distribution of different types of genetic variations excluding singletons as annotated by genic location (exonic/coding, non-exonic/non-coding) and sequence impact (i.e., 1% less ncVs, 1% more cVs: 3% less 3′UTR, 3% more intronic, and 5% less nsVs). Pie charts express both numbers and the percent of variants in the respective pie. d The proportion of centenarian-enriched to centenarian-depleted variants is based on MAF in cases versus MAF in controls and separated based on observed cohort MAF. Common = MAF > 0.010. Rare = MAF ≤ 0.010. Green bars = percent of variants enriched in centenarians, blue bars = percent of variants depleted in centenarians. The analysis was done after excluding singletons
When a variant has a greater MAF in centenarians, as compared to controls, we call the variant “enriched” with regard to centenarians and “depleted” in controls. When considering all variants (regardless of p-value), the proportion of variants enriched and depleted in centenarians remains ~ 50% (Fig. 3d). However, when only significant variants (p ≤ 0.050) are considered, the proportion of rare variants that are enriched in centenarians is far greater. Nearly 80% of rare variants are enriched in centenarians and the remaining 20% are depleted (Fig. 3d). In addition, we wanted to know if centenarians harbor more of a specific type of variant, e.g., do they have more cVs or ncVs compared to controls? The distributions of different types of variants between centenarians and controls do not differ very much (Fig. S2a). Excluding singletons only alters the proportions of some types of variants by no more than 5% (e.g., there are 5% less nsVs in total, and centenarians have 1% more frameshifts compared to controls after excluding singletons) (Fig. S2b and S3a, b).
Identifying longevity-associated mitonuclear genes and pathways by gene-based analysis
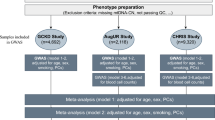
We applied a workflow to variants discovered by target capture sequencing to systematically filter top longevity-associated genes (LAGs), variants (LAVs), and pathways (LAPs). It has been done under the hypothesis that rare centenarian-enriched variants may be protective and functionally causal in longevity compared to common variants and that these causal rare variants may or may not function in aggregate (Fig. 4). Gene-based analyses can help overcome rare variant statistical challenges by treating all variants within a single locus as a group to measure their aggregate longevity-association statistic. Thus, data were filtered to identify LAGs by using three types of Sequence Kernel Association Test (SKAT, SKAT-C, and SKAT-O), a rare variant burden test for joint significance of multiple genetic variants within a single gene [54]. Identifying genes that are significant across all SKATs or in specific SKATs can be informative for downstream interpretation because SKATs analyze different information: SKAT is specific for rare/low-frequency variants, SKAT-C includes common variants as well as rare variants, and SKAT-O gives more weight to variants that are strongly enriched in one group over another. Hence, with SKAT-O, we can assess the significance of overall directionality among cases and controls for all variants within a locus (i.e., whether variants in a given gene are significantly enriched in AJ centenarians).
Workflow to filter, identify, and analyze longevity-associated mitonuclear genes and variants. Gene-based variant aggregate analyses by SKATs were used to identify longevity-associated genes, while individual SNP-based analysis was performed by screening variants meeting certain criteria of interest: mainly nsVs, rare variants (MAF ≤ 0.010), and centenarian-enriched variants were filtered for subsequent downstream analyses, including gene set pathway enrichment, variant effect prediction, and modeling in silico
Gene-based analysis revealed 76 nominally significant (p ≤ 0.050) longevity-associated genes, and we call these 76 genes the “longevity-associated gene set” (Tables S3 and S4). Mitochondrial genes well-known for their associations with aging, age-related disease, and lifespan, including PPARG, TP53, PARP2, and APIP, were found to be significantly associated with AJ longevity. However, none of the genes is significant after multiple test correction (Bonferroni, 660 tests, p = 7.5 × 10−5). Nonetheless, by incorporating biological evidence, we can gather further support for the genes that are more likely to be true positives. Thus, we further analyzed and annotated the longevity-associated gene set in terms of their representative mitochondrial functions and pathways, and their aging biology context, by data and text mining.
A summary schematic of the 21 top (p ≤ 0.010) longevity-associated mitonuclear genes illustrates the mitochondrial pathways represented, the experimental evidence for regulation of lifespan in model organisms, and the overlapping genes found to be significant in multiple SKATs (Fig. 5). For example, LETM1, a mitochondrial inner membrane calcium and proton (Ca2+ /H+) antiporter that also regulates mitochondrial morphology and dynamics, and which is SKAT-O significant (p = 0.0026) in our analysis, is reported to be an anti-longevity gene in Caenorhabditis elegans due to the 21% extension of mean lifespan that is observed after letm-1 RNAi-mediated knockdown [74]. We identified 46 variants annotated to LETM1: 37 as rare, 25 enriched in centenarians, 23 both enriched and rare, and 35 ncVs, with the majority (32) mapping to the 3′UTR of LETM1.
Longevity-associated genes and pathways discovered by gene-based association study. Mitonuclear genes with the greatest significance (p ≤ 0.01) in at least one SKAT are shown in their respective functional pathway(s); genes are not significant after MTC. A pro-longevity gene indicates evidence that overexpression of the gene can increase lifespan or a loss of function can decrease lifespan in the indicated model organism; an anti-longevity gene indicates evidence for overexpression decreasing lifespan or loss of function increasing lifespan in the indicated model organism. ΔΨm = mitochondrial membrane potential; Pyr = pyruvate; AcCoA = acetyl-coA; PPP = pentose phosphate pathway; TCA = the citric acid cycle; Cis.Aco. = cis-aconitase; IsoC. = isocitrate; α-KG = alpha-ketoglutarate; Succ.CoA = succinyl-CoA; Succ. = succinate; Fum. = fumarate; Mal. = malate; Ox.Ac = oxaloacetate; ETC = electron transport chain; Fe-S = iron-sulfur; MAM = mitochondrial ER-associated membrane; ROS = reactive oxygen species; ATP = adenosine triphosphate; KBs = ketone bodies; UAC = uric acid cycle; NH4 + = ammonia; Arg. = arginine; Asp. = aspartate; Arg.Succinate = argininosuccinate; CPS1 = carbamoyl phosphate synthase
Using the PANTHER tool for GO analysis, we assessed the enrichment of mitochondrial pathways in the longevity-associated gene set for different reference backgrounds (Table S5). The fold enrichment of ketone body metabolism in the longevity-associated gene set was significant—passing multiple test correction against all backgrounds except the most stringent mitonuclear candidate list, as expected. There are ~ 12 ketone metabolism genes reported by PANTHER in the human reference list, and half were included in the candidate gene list (AACS, ACAT1, BDH1, HMGCL, HMGCS2, and SLC27A5), representing < 1% of the total mitonuclear candidate genes and all except 1 (ACAT1) are longevity-associated by SKATs (p ≤ 0.050). SLC27A5/FATP5, a liver-specific transporter of bile and fatty acids for lipid uptake and bile-acid synthesis, and HMGCL, the lyase catalyzing the conversion of HMGCoA to the ketone body acetoacetate, are among the top significant longevity-associated ketone metabolism genes, with SLC27A5 showing significance in all SKATs (p ≤ 0.035). However, 2 other top significant genes (ACACA and HMGCS1) are not annotated to ketone metabolism in PANTHER but do play roles in the pathway, suggesting that the enrichment of ketone metabolism is actually under-represented by PANTHER GO analysis. These genes may not be annotated to “ketone metabolism” because ACACA and HMGCS1 have primary roles in fatty-acid beta-oxidation which is an important upstream step to producing AcCoA substrates for hepatic mitochondrial ketone synthesis, and mevalonate substrates for cholesterol synthesis, which can occur as a non-oxidative fate of ketone bodies, respectively [75, 76].
Prioritizing potentially functional longevity-associated mitonuclear variants
To identify functional longevity-associated variants, we further prioritized rare LAVs (Fig. 4) on the basis of their functionality of coding variants predicted by CADD and Condel algorithms for variant deleteriousness impact scores and enrichment in centenarians as compared to controls (see the “Methods” section). As a result, we identified 112 longevity-associated variants in 86 genes with different annotations (Table S6) and identified 59 nsVs (Table S7). The majority of prioritized variants are centenarian-enriched (~ 88%; 90) and rare (63%), 31 and 26 of which fall into regions of high vertebrate conservation and in conserved residues in invertebrates (yeast, worms, or flies), respectively. Among the 59 prioritized nsVs (Table S7), 54 resulted in not conserved amino acid substitutions, 14 in Radial (R) substitutions, and Condel predicted 21 to be deleterious, while 29 are reported possibly—likely damaging by CADD and are therefore very strong candidates. Given the rarity of LAVs (Table S7), we performed genotyping analysis to validate 65 LAVs without any technical and quality issues, of which 30 nsVs were predicted to be functional—18 LAVs by CADD and 12 LAVs by Condel, representing our strongest candidate variants (Table S8).
We also prioritized potentially functional longevity-associated genes (LAGs). Genes containing multiple top nsVs could represent “hotspots” for rare, potentially functional longevity associations. Thus, we note that 5 genes harbor multiple prioritized nsVs: ACACB, ALDOA, ATP5S, MRPL51, and LRPPRC. These genes are involved in various mitochondrial functions: ACACB encodes the key lipid synthesis enzyme that carboxylates acetyl-CoA to malonyl-CoA; ALDOA is the muscle-type isoform of the glycolytic aldolase enzyme that catalyzes the fourth step of glycolysis—the condensation of 6-carbon fructose 1,6-bisphosphate to 3-carbon glyceraldehyde 3-phosphate; ATP5S/DMAC2L encodes a subunit of the membrane and proton channel component (Fo) in the terminal ETC complex mitochondrial ATP synthase (complex V); MRPL51 encodes a protein component of the large 39S mitochondrial ribosomal subunit; and LRPPRC/LRP130 (leucine-rich pentatricopeptide repeat containing 130 kDa protein) encodes an RNA-binding protein that primarily positively regulates mitochondrial mRNA stability and translation. Furthermore, LAVs in the CPS1 (rs1047891; T1406N), MFN2 (G548*), and MTOR (Y2396Lfs*29) have been implicated in longevity, age-related disease, and endophenotypes associated with lifespan determination (see the “Discussion” section).
Modeling and predicting biological impact of LRPPRC variants
Of our highest interests are two missense variants in LRPPRC. These variants are very rare and only detected in centenarians (MAFs < 0.005) but not in controls or any other populations (MAF = 0) (Tables S7 and S8). rs149693840 (S1378G) is predicted to be deleterious (Table S8), while rs146630100 (D1352E) is neutral. Neither of the affected amino acids maps to positions known or predicted to undergo post-translational modification; however, a lysine ubiquitinylation site is just 2 residues upstream of D1352 and downstream of S1378, a tyrosine phosphorylation site is 4 residues away (Fig. 6a). The crystal structure of LRPPRC has never been resolved, and as of yet there is no prediction on its 3D structure.
Longevity-associated variants in a proposed tertiary structure of LRPPRC. a 2D LRPPRC protein schematic showing the location of the N-terminal mitochondrial targeting signal (MTS), pentatricopeptide repeats (PPR), C-terminal RNA-binding domain (RNA-bd), and post-translational modification sites: phosphorylation (Pi), ubiquitinylation (Ub.), acetylation (Ac.), and methylation (Me), and longevity-associated variants D1352E and S1378G. Black box outlines the region magnified. Gray box with dashed lines approximates the portion of LRPPRC modeled by the homology structure in b. b 3D homology model with predicted PPR H–L-H motifs (green and purple) and the locations of amino acids altered by longevity-associated variants (black arrows). Gray box with dashed lines around S1378G approximates the region magnified in c. c Magnification shows loop region with distances between the oxygen atom of Ser1378 R-group with closest neighboring hydrogen atoms from Glu1374, Pro1375, and Glu1377. Distance labeled in blue and shown in angstroms (Å), distances > 5 Å omitted. d Potential hydrogen-bonding interactions between S1378 with the surrounding loop region and terminal PPR (helix-loop-helix motif in purple) motif are shown. Thick blue line—hydrogen bonds from Serine1378; thin orange lines—other hydrogen bonds from indicated residues
Using publicly available protein modeling tools, we built a partial 3D homology model (“Methods” section) to visualize the location of these LAVs and their potential interactions with other residues in the tertiary structure of the C-terminus of LRPPRC (Fig. 6b-d). D1352 is located in an alpha helix, while S1378 is located in a loop that is between two alpha helices (helix-loop-helix (H-L-H) motif) (Fig. 6b, c). The closest residues to S1378 include multiple Pro and Glu residues (Fig. 6c)—P1375, P1376, E1374, E1377, and E1380—suggesting that the homology model accurately predicts loop regions as Pro residues often act as loop stabilizers due to their rigidity [77]. Moreover, loops in folded protein structures are often associated with functional domains at the surface of the protein, such as protein–protein interaction regions, especially highly ordered loops [78, 79]. The negatively charged Glu residues could participate in electrostatic interactions with other molecules and the surrounding solution; this might suggest that this is a very dynamic loop since negative charges from Glu might change the loop angle or shape to interact less with the slightly basic environment of the mitochondrial matrix where LRPPRC action takes place. Serine’s R-group contains a slightly polar carboxyl group with hydrogen-bonding capacity and we detected a potential hydrogen bond from S1378 to the backbone at residue T1322 located in the terminal H-L-H PPR motif that is involved in RNA binding (Fig. 6d). This could be important in altering the angle or position of the RNA-binding PPR or other residues and structures that are nearby in the folded tertiary structure. However, the Gly R-group is a hydrogen atom that lacks the electrostatic potential to participate in hydrogen bonding, suggesting that a substitution from S → G at residue 1378 has the potential to alter the structure and/or biochemistry of the C-terminal RNA-binding domain in LRPPRC. LRPPRC has yet not been directly investigated in the context of aging and longevity, making it a novel target for our future study.
Discussion
By combining a targeted candidate sequencing approach with integrative analysis of potentially functional centenarian-enriched, rare variants, we discovered 35 high-priority functional variants (LAVs) and 75 mitonuclear genes (LAGs) in multiple mitochondrial pathways that are associated with longevity (p < 0.05). Our data support the hypothesis that rare variants are likely functional and enriched in individuals exhibiting the rare phenotype of extreme longevity.
Given the rarity of these variants in the rare population of centenarians, it is highly challenging, if not impossible, to establish the statistical significance of the longevity association. However, focusing on the well-established conserved pathways of aging, we have identified and characterized longevity-associated functional rare coding variants in IGF1R [37, 38] and functional rare non-coding variants in NFKBIA [39], CLU [39], and PRKCH [39] genes by using cell models. Our results demonstrated parallel mechanisms between humans and model organisms, increasing our confidence that longevity-associated genes/pathways found in the conserved pathways may be relevant to human longevity.
We identified nutrient sensing through MTOR, ketone body metabolism, calcium signaling, the urea cycle, 1-carbon metabolism (1C), and mitochondrial translation as potential key targets for the investigation of the mitochondrial mechanisms of human longevity. We targeted two rare variants in LRPPRC gene, rs149693840 (S1378G) and rs146630100 (D1352E) that are located in the C-terminal RNA-binding domain of the mitochondrial translation regulator [80]. These variants are adjacent to residues that undergo post-translational modification, suggesting that altering these amino acids has the potential to impact protein–protein interactions and signaling. The S1378G variant has the potential to alter electrostatic interactions that stabilize a loop structure and the terminal H-L-H PPR motif, involved in the RNA-binding function of LRPPRC. Autosomal recessive missense mutations in LRPPRC cause the French-Canadian Type of Leigh Syndrome (LSFC), a rare mitochondrial disease-causing cytochrome c oxidase (COX, complex IV (CIV)) deficiency, which results in severe neurological, muscular, and kidney pathology, lactic acidosis, and premature death [81,82,83]. Loss of function of LRPPRC in worms and human cell lines disrupts ETC stoichiometry, especially at CIV, leading to mitochondrial protein imbalance, upregulated mtUPR, altered ATP/ADP ratio, and altered mitochondrial dynamics [84, 85]. Given previous enigmatic work on ETC dysfunction in model organism longevity [14, 74, 86, 87], the importance of mitochondrial translation and proteostasis in mediating several mechanisms of longevity and aging in model organisms, and the biochemical importance of COX in regulating the rate of respiration [88], we believe these LAVs could be interesting targets for future investigation.
We identified MFN2 as a LAG in two SKATs and validated a rare enriched stop gain LAV (G548*) in this gene that is predicted to be deleterious by CADD and located at a highly evolutionarily conserved residue. MFN2 is a positive regulator of mitochondrial dynamics and morphology and tethers the mitochondria and ER membranes to form mitochondrial-associated membranes (MAMs) that regulate lipid and Ca2+ exchange at the ER, and regulate ubquinone (UQ) synthesis [89,90,91]. Given that complete knockout (KO) of MFN2 is embryonic lethal in mice and loss of function mutations in humans lead to the severe mitochondrial disease Charcot Marie Tooth Type 2A (CMT2A) [92], the heterozygous carriers of this variant as detected in centenarians may minimally reduce protein abundance or function. Alternatively, the truncated protein only interrupts a short portion of the cytoplasmic domain but does not completely abolish the remaining C-terminal region of the protein which would render it completely non-functional [93]. Moreover, it could be that this MFN2 variant partially alters localization of MFN2 [94], or interactions with other proteins, including PINK1 and PARKIN which, as mentioned above, are important mediators in the etiology of dementia, including Alzheimer’s disease (AD) and PD [95], or the variant may alter UQ biosynthesis or export to extramitochondrial endomembrane systems [89, 96].
A rare frameshift in MTOR (Y2396Lfs*29) enriched in centenarians has the strong potential to impair MTOR function. This MTOR frameshift was the only significant coding variant identified in the MTOR gene and the only significant frameshift variant in the sequencing data, suggesting that this variant is extremely unique. Given the background and established role of MTOR on aging and lifespan determination as well as on mitochondrial biogenesis and metabolism [97,98,99], it is intriguing to speculate that this frameshift produces some functional effect on MTOR protein abundance or signaling. We hypothesize that this variant may reduce mTOR-mediated nutrient sensing and signaling, thereby increasing lifespan and healthspan through multiple downstream mechanisms, including enhanced autophagy, improved mitochondrial function, and altered metabolism, as has been shown in model organisms and cultured human cells [10, 100,101,102,103].
The functional CPS1 variant (rs1047891; T1406N) occurs in the allosteric binding domain of the protein. Studies suggest it confers protection from coronary artery disease (CAD) risk in females specifically and metabolic disease (MD) through control of weight gain, suggesting that this variant could be an example of a sexually dimorphic variant in longevity [104,105,106,107,108,109]. CPS1 is an enzymatic hub in mitochondrial metabolism linked to caloric restriction, SIRT5, the TCA cycle, 1C metabolism, and NO production, and its primary role is in detoxification of NH4+ by the urea cycle which is increased under nutrient deprivation stress when protein catabolism increases [105, 110,111,112].
A functional investigation of these variants, either in vitro or in vivo, may help resolve the biological mechanisms behind complex phenotypes like longevity across species. Future studies of these rare variants with knock-in models in human cell lines or mammalian model organisms are critical to providing the functional evidence that validates these in silico results. In studying the role of mitochondria in human aging and longevity, it is critical to consider how cellular metabolic, redox, and oxygen environments might modify the phenotype or how polymorphic longevity-associated pathways might change in response to different cellular stress conditions.
It is known that complex phenotypes are determined by multiple alleles functioning in combination (i.e., haplotypes), which should be extended to trans-genomic haplotypes and interactions between mtDNA-nDNA in future research [113,114,115]. With future developments in the integration of genomics, epigenomics, transcriptomics, proteomics, and metabolomics (-omics) data, there is greater hope that functional genetic variation in any pathway can be investigated for any phenotype, and in humans, on a more holistic system-biology level [1, 2, 116]. This kind of work will also help researchers plan for focused data-driven experiments aimed at explaining genotype–to–phenotype mechanisms.
Our results suggest new hypotheses and support existing hypotheses that may help guide future experiments aimed at linking important functions of mitonuclear genes to human longevity. Future studies should explore these hypotheses in functional biological models.
Code availability
Not applicable.
References
Partridge L, Deelen J, Slagboom PE, Eline S. Facing up to the global challenges of ageing. Nature. 2018;561(7721):45–56.
Schork NJ, Raghavachari N. Report: NIA workshop on translating genetic variants associated with longevity into drug targets On behalf of Workshop Speakers and Participants. GeroScience. 2018;40:523–38.
Sierra F. The emergence of geroscience as an interdisciplinary approach to the enhancement of health span and life span. Cold Spring Harb Perspect Med. 2016;6(4):a025163–a025163.
Kennedy BK, Berger SL, Brunet A, Campisi J, Cuervo AM, Epel ES, Franceschi C, Lithgow GJ, Morimoto RI, Pessin JE, Rando TA, Richardson A, Schadt EE, Wyss-Coray T, Sierra F. Geroscience: linking aging to chronic disease. Cell. 2014;159(4):709–13.
Lopez-Otin C, Blasco MA, Partridge L, Serrano M, Kroemer G. The hallmarks of aging. Cell. 2013;153(6):1194–217.
Berry BJ, Kaeberlein M. An energetics perspective on geroscience: mitochondrial protonmotive force and aging. GeroScience. 2021;43(4):1591–604.
Barbosa MC, Grosso RA, Fader CM. Hallmarks of aging: an autophagic perspective. Front Endocrinol. 2019;9:790–790.
Short KR, Bigelow ML, Kahl J, Singh R, Coenen-Schimke J, Raghavakaimal S, Nair KS. Decline in skeletal muscle mitochondrial function with aging in humans. Proc Natl Acad Sci U S A. 2005;102(15):5618–23.
Chan DC. Mitochondria: dynamic organelles in disease, aging, and development. Cell. 2006;125(7):1241–52.
Hwang AB, Jeong DE, Lee SJ. Mitochondria and organismal longevity. Curr Genomics. 2012;13(7):519–32.
Dillin A, Hsu AL, Arantes-Oliveira N, Lehrer-Graiwer J, Hsin H, Fraser AG, Kamath RS, Ahringer J, Kenyon C. Rates of behavior and aging specified by mitochondrial function during development. Science. 2002;298(5602):2398–401.
Caldeira Da Silva CC, Cerqueira FM, Barbosa LFLvF, Medeiros MHGG, Kowaltowski AJ. Mild mitochondrial uncoupling in mice affects energy metabolism, redox balance and longevity. Aging Cell. 2008;7(4):552–60.
Copeland JM, Cho J, Lo T, Hur JH, Bahadorani S, Arabyan T, Rabie J, Soh J, Walker DW. Extension of Drosophila life span by RNAi of the mitochondrial respiratory chain. Current biology : CB. 2009;19(19):1591–8.
Houtkooper RH, Mouchiroud L, Ryu D, Moullan N, Katsyuba E, Knott G, Williams RW, Auwerx J. Mitonuclear protein imbalance as a conserved longevity mechanism. Nature. 2013;497(7450):451–7.
Kujoth GC, Hiona A, Pugh TD, Someya S, Panzer K, Wohlgemuth SE, Hofer T, Seo AY, Sullivan R, Jobling WA, Morrow JD, van Remmen H, Sedivy JM, Yamasoba T, Tanokura M, Weindruch R, Leeuwenburgh C, Prolla TA. Mitochondrial DNA mutations, oxidative stress, and apoptosis in mammalian aging. Science (New York, NY). 2005;309(5733):481–4.
Morrow GGV, Battistini S, Zhang P, Tanguay RM. Decreased lifespan in the absence of expression of the mitochondrial small heat shock protein Hsp22 in Drosophila. J Biol Chem. 2004;279(42):43382–5.
Kirchman PA, Kim S, Lai CY, Jazwinski SM. Interorganelle signaling is a determinant of longevity in Saccharomyces cerevisiae. Genetics. 1999;152(1):179–90.
Morrow G, Samson M, Michaud S, Tanguay RM. Overexpression of the small mitochondrial Hsp22 extends Drosophila life span and increases resistance to oxidative stress. FASEB J. 2004;18(3):598–9.
Conti B, Sanchez-Alavez M, Winsky-Sommerer R, Morale MC, Lucero J, Brownell S, Fabre V, Huitron-Resendiz S, Henriksen S, Zorrilla EP, Lecea Ld, Bartfai T. Transgenic mice with a reduced core body temperature have an increased life span. Science. 2006;314(5800):825–8.
Schriner SE, Linford NJ, Martin GM, Treuting P, Ogburn CE, Emond M, Coskun PE, Ladiges W, Wolf N, Van Remmen H, Wallace DC, Rabinovitch PS. Extension of murine life span by overexpression of catalase targeted to mitochondria. Science. 2005;308(5730):1909–11.
Pulliam DA, Bhattacharya A, Van Remmen H. Mitochondrial dysfunction in aging and longevity: a causal or protective role? Antioxid Redox Signal. 2013;19(12):1373–87.
Munkácsy E, Rea SL. The paradox of mitochondrial dysfunction and extended longevity. Exp Gerontol. 2014;56:221–33.
Dai DF, Chen T, Johnson SC, Szeto H, Rabinovitch PS. Cardiac aging: from molecular mechanisms to significance in human health and disease. Antioxid Redox Signal. 2012;16(12):1492–526.
Dai DF, Chiao YA, Marcinek DJ, Szeto HH, Rabinovitch PS. Mitochondrial oxidative stress in aging and healthspan. Longev Healthspan. 2014;3:6.
Zhang ZD, Milman S, Lin J-R, Wierbowski S, Yu H, Barzilai N, Gorbunova V, Ladiges WC, Niedernhofer LJ, Suh Y, Robbins PD, Vijg J. Genetics of extreme human longevity to guide drug discovery for healthy ageing. Nat Metab. 2020;2(8):663–72.
Atzmon G, Schechter C, Greiner W, Davidson D, Rennert G, Barzilai N. Clinical phenotype of families with longevity. J Am Geriatr Soc. 2004;52(2):274–7.
Perls T, Kohler IV, Andersen S, Schoenhofen E, Pennington J, Young R, Terry D, Elo IT. Survival of parents and siblings of supercentenarians. J Gerontol A Biol Sci Med Sci. 2007;62(9):1028–34.
Schoenmaker M, de Craen AJ, de Meijer PH, Beekman M, Blauw GJ, Slagboom PE, Westendorp RG. Evidence of genetic enrichment for exceptional survival using a family approach: the Leiden Longevity Study. Eur J Hum Genet : EJHG. 2006;14(1):79–84.
Westendorp RG, van Heemst D, Rozing MP, Frolich M, Mooijaart SP, Blauw GJ, Beekman M, Heijmans BT, de Craen AJ, Slagboom PE. Nonagenarian siblings and their offspring display lower risk of mortality and morbidity than sporadic nonagenarians: the Leiden Longevity Study. J Am Geriatr Soc. 2009;57(9):1634–7.
Terry DF, Wilcox MA, McCormick MA, Pennington JY, Schoenhofen EA, Andersen SL, Perls TT. Lower all-cause, cardiovascular, and cancer mortality in centenarians’ offspring. J Am Geriatr Soc. 2004;52(12):2074–6.
Perls TT, Wilmoth J, Levenson R, Drinkwater M, Cohen M, Bogan H, Joyce E, Brewster S, Kunkel L, Puca A. Life-long sustained mortality advantage of siblings of centenarians. Proc Natl Acad Sci USA. 2002;99(12):8442–7.
Andersen SL, Sebastiani P, Da Dworkis, Feldman L, Perls TT. Health span approximates life span among many supercentenarians: compression of morbidity at the approximate limit of life span. J Gerontol A Biol Sci Med Sci. 2012;67(4):395–405.
Gorlov IP, Gorlova OY, Sunyaev SR, Spitz MR, Amos CI. Shifting paradigm of association studies: value of rare single-nucleotide polymorphisms. Am J Hum Genet. 2008;82(1):100–12.
Bloss CS, Pawlikowska L, Schork NJ. Contemporary human genetic strategies in aging research. Ageing Res Rev. 2011;10(2):191–200.
Glatt SJ, Chayavichitsilp P, Depp C, Schork NJ, Jeste DV. Successful aging: from phenotype to genotype. Biol Psychiat. 2007;62(4):282–93.
Sebastiani P, Solovieff N, DeWan AT, Walsh KM, Puca A, Hartley SW, Melista E, Andersen S, Dworkis DA, Wilk JB, Myers RH, Steinberg MH, Montano M, Baldwin CT, Hoh J, Perls TT. Genetic signatures of exceptional longevity in humans. PLoS ONE. 2012;7(1):e29848–e29848.
Tazearslan C, Huang J, Barzilai N, Suh Y. Impaired IGF1R signaling in cells expressing longevity-associated human IGF1R alleles. Aging Cell. 2011;10(3):551–4.
Suh Y, Atzmon G, Cho MO, Hwang D, Liu B, Leahy DJ, Barzilai N, Cohen P. Functionally significant insulin-like growth factor I receptor mutations in centenarians. Proc Natl Acad Sci U S A. 2008;105(9):3438–42.
Ryu S, Han J, Norden-Krichmar TM, Zhang Q, Lee S, Zhang Z, Atzmon G, Niedernhofer LJ, Robbins PD, Barzilai N, Schork NJ, Suh Y. Genetic signature of human longevity in PKC and NF-kappaB signaling. Aging Cell. 2021;20(7):e13362.
Suh Y, Atzmon G, Cho M-O, Hwang D, Liu B, Leahy DJ, Barzilai N, Cohen P. Functionally significant insulin-like growth factor I receptor mutations in centenarians. Proc Natl Acad Sci USA. 2008;105(9):3438–42.
Stelzer G, Rosen N, Plaschkes I, Zimmerman S, Twik M, Fishilevich S, Stein TI, Nudel R, Lieder I, Mazor Y, Kaplan S, Dahary D, Warshawsky D, Guan-Golan Y, Kohn A, Rappaport N, Safran M, Lancet D. The GeneCards Suite: from gene data mining to disease genome sequence analyses. Curr. Protoc. Bioinform. 2016;54(1):1.30.1–1.30.33.
Pagliarini DJ, Calvo SE, Chang B, Sheth SA, Vafai SB, Ong SE, Walford GA, Sugiana C, Boneh A, Chen WK, Hill DE, Vidal M, Evans JG, Thorburn DR, Carr SA, Mootha VK. A mitochondrial protein compendium elucidates complex I disease biology. Cell. 2008;134(1):112–23.
Rhee H-W, Zou P, Udeshi ND, Martell JD, Mootha VK, Carr SA, Ting AY. Proteomic mapping of mitochondria in living cells via spatially restricted enzymatic tagging. Science. 2015;339(6125):1328–31.
Amberger JS, Bocchini CA, Ois Schiettecatte F, Scott AF, Hamosh A. OMIM.org: Online Mendelian Inheritance in Man (OMIM R), an online catalog of human genes and genetic disorders. Nucleic Acids Research. 2014;43:789–98.
Tacutu R, Thornton D, Johnson E, Budovsky A, Barardo D, Craig T, Diana E, Lehmann G, Toren D, Wang J, Fraifeld VE, de Magalhães JP. Human Ageing Genomic Resources: new and updated databases. Nucleic Acids Res. 2018;46(D1):D1083–90.
Craig T, Smelick C, Tacutu R, Wuttke D, Wood SH, Stanley H, Janssens G, Savitskaya E, Moskalev a, Arking R, de Magalhaes JP. The Digital Ageing Atlas: integrating the diversity of age-related changes into a unified resource. Nucleic Acids Res. 2014;43(D1):D873–8.
Fabregat A, Jupe S, Matthews L, Sidiropoulos K, Gillespie M, Garapati P, Haw R, Jassal B, Korninger F, May B, Milacic M, Roca CD, Rothfels K, Sevilla C, Shamovsky V, Shorser S, Varusai T, Viteri G, Weiser J, Wu G, Stein L, Hermjakob H, D’Eustachio P. The reactome pathway knowledgebase. Nucleic Acids Res. 2018;46(D1):D649–55.
Consortium TGO. The Gene Ontology Resource: 20 years and still GOing strong. Nucleic Acids Res. 2019;47(D1):D330–8.
Binder JX, Pletscher-Frankild S, Tsafou K, Stolte C, O’Donoghue SI, Schneider R, Jensen LJ. COMPARTMENTS: unification and visualization of protein subcellular localization evidence. Database. 2014;2014(0):bua012–bua012.
Barzilai N, Atzmon G, Schechter C, Schaefer EJ, Cupples AL, Lipton R, Cheng S, Shuldiner AR. Unique lipoprotein phenotype and genotype. J Am Med Assoc (JAMA). 2003;290(15):2030–40.
Ryu S, Han J, Norden-Krichmar TM, Schork NJ, Suh Y. Effective discovery of rare variants by pooled target capture sequencing: a comparative analysis with individually indexed target capture sequencing. Mutat Res. 2018;809:24–31.
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D. The human genome browser at UCSC. Genome Res. 2002;12(6):996–1006.
Johnson SC, Gonzalez B, Zhang Q, Milholland B, Zhang Z, Suh Y. Network analysis of mitonuclear GWAS reveals functional networks and tissue expression profiles of disease-associated genes. Hum Genet. 2017;136(1):55–65.
Wu MC, Lee S, Cai T, Li Y, Boehnke M, Lin X. Rare-variant association testing for sequencing data with the sequence kernel association test. Am J Hum Genet. 2011;89(1):82–93.
Raney BJ, Dreszer TR, Barber GP, Clawson H, Fujita PA, Wang T, Nguyen N, Paten B, Zweig AS, Karolchik D, Kent WJ. Track data hubs enable visualization of user-defined genome-wide annotations on the UCSC Genome Browser. Bioinformatics. 2014;30(7):1003–5.
Dewey FE, Murray MF, Overton JD, Habegger L, Leader JB, Fetterolf SN, O’Dushlaine C, Van Hout CV, Staples J, Gonzaga-Jauregui C, Metpally R, Pendergrass SA, Giovanni MA, Kirchner HL, Balasubramanian S, Abul-Husn NS, Hartzel DN, Lavage DR, Kost KA, Packer JS, Lopez AE, Penn J, Mukherjee S, Gosalia N, Kanagaraj M, Li AH, Mitnaul LJ, Adams LJ, Person TN, Praveen K, Marcketta A, Lebo MS, Austin-Tse CA, Mason-Suares HM, Bruse S, Mellis S, Phillips R, Stahl N, Murphy A, Economides A, Skelding KA, Still CD, Elmore JR, Borecki IB, Yancopoulos GD, Davis FD, Faucett WA, Gottesman O, Ritchie MD, Shuldiner AR, Reid JG, Ledbetter DH, Baras A, Carey DJ. Distribution and clinical impact of functional variants in 50,726 whole-exome sequences from the DiscovEHR study. Science (New York, NY). 2016;354(6319):aaf6814–aaf6814.
Hunt SE, McLaren W, Gil L, Thormann A, Schuilenburg H, Sheppard D, Parton A, Armean IM, Trevanion SJ, Flicek P, Cunningham F. Ensembl variation resources. Database. 2018;2018. https://doi.org/10.1093/database/bay119.
Wang J, Al-Ouran R, Hu Y, Kim S-Y, Wan Y-W, Wangler MF, Yamamoto S, Chao H-T, Comjean A, Mohr SE, Perrimon N, Liu Z, Bellen HJ, Adams CJ, Adams DR, Alejandro ME, Allard P, Ashley EA, Azamian MS, Bacino CA, Balasubramanyam A, Barseghyan H, Beggs AH, Bellen HJ, Bernstein JA, Bican A, Bick DP, Birch CL, Boone BE, Briere LC, Brown DM, Brush M, Burke EA, Burrage LC, Chao KR, Clark GD, Cogan JD, Cooper CM, Craigen WJ, Davids M, Dayal JG, Dell’Angelica EC, Dhar SU, Dipple KM, Donnell-Fink LA, Dorrani N, Dorset DC, Draper DD, Dries AM, Eckstein DJ, Emrick LT, Eng CM, Esteves C, Estwick T, Fisher PG, Frisby TS, Frost K, Gahl WA, Gartner V, Godfrey RA, Goheen M, Golas GA, Goldstein DB, Gordon MG, Gould SE, Gourdine J-PF, Graham BH, Groden CA, Gropman AL, Hackbarth ME, Haendel M, Hamid R, Hanchard NA, Handley LH, Hardee I, Herzog MR, Holm IA, Howerton EM, Jacob HJ, Jain M, Jiang Y-h, Johnston JM, Jones AL, Koehler AE, Koeller DM, Kohane IS, Kohler JN, Krasnewich DM, Krieg EL, Krier JB, Kyle JE, Lalani SR, Latham L, Latour YL, Lau CC, Lazar J, Lee BH, Lee H, Lee PR, Levy SE, Levy DJ, Lewis RA, Liebendorfer AP, Lincoln SA, Loomis CR, Loscalzo J, Maas RL, Macnamara EF, MacRae CA, Maduro VV, Malicdan MCV, Mamounas LA, Manolio TA, Markello TC, Mazur P, McCarty AJ, McConkie-Rosell A, McCray AT, Metz TO, Might M, Moretti PM, Mulvihill JJ, Murphy JL, Muzny DM, Nehrebecky ME, Nelson SF, Newberry JS, Newman JH, Nicholas SK, Novacic D, Orange JS, Pallais JC, Palmer CGS, Papp JC, Pena LDM, Phillips JA, Posey JE, Postlethwait JH, Potocki L, Pusey BN, Ramoni RB, Robertson AK, Rodan LH, Rosenfeld JA, Sadozai S, Schaffer KE, Schoch K, Schroeder MC, Scott DA, Sharma P, Shashi V, Silverman EK, Sinsheimer JS, Soldatos AG, Spillmann RC, Splinter K, Stoler JM, Stong N, Strong KA, Sullivan JA, Sweetser DA, Thomas SP, Tifft CJ, Tolman NJ, Toro C, Tran AA, Valivullah ZM, Vilain E, Waggott DM, Wahl CE, Walley NM, Walsh CA, Wangler MF, Warburton M, Ward PA, Waters KM, Webb-Robertson B-JM, Weech AA, Westerfield M, Wheeler MT, Wise AL, Wolfe LA, Worthey EA, Yamamoto S, Yang Y, Yu G, Zornio PA, Perrimon N, Liu Z, Bellen HJ. MARRVEL: integration of human and model organism genetic resources to facilitate functional annotation of the human genome. Am J Hum Genet. 2017;100(6):843–53.
Rosenbloom KR, Sloan CA, Malladi VS, Dreszer TR, Learned K, Kirkup VM, Wong MC, Maddren M, Fang R, Heitner SG, Lee BT, Barber GP, Harte RA, Diekhans M, Long JC, Wilder SP, Zweig AS, Karolchik D, Kuhn RM, Haussler D, Kent WJ. ENCODE Data in the UCSC Genome Browser: year 5 update. Nucleic Acids Res. 2012;41(D1):D56–63.
Gromiha MM. Protein bioinformatics: from sequence to function. Cambridge: Academic Press; 2010. pp. 320.
Vitkup D, Sander C, Church GM. The amino-acid mutational spectrum of human genetic disease. Genome Biol. 2003;4(11):R72.
Yampolsky LY, Stoltzfus A. The exchangeability of amino acids in proteins. Genetics. 2005;170(4):1459–72.
Zhang J. Rates of conservative and radical nonsynonymous nucleotide substitutions in mammalian nuclear genes. J Mol Evol. 2000;50(1):56–68.
Betts MJ, Russell RB, Amino acid properties and consequences of substitutions, in Bioinformatics for Geneticists. 2003 289–316.
González-Pérez A, Ló P-B. Improving the assessment of the outcome of nonsynonymous SNVs with a consensus deleteriousness score, Condel. Am J Hum Genet. 2011;88:440–9.
Kircher M, Witten DM, Jain P, O’Roak BJ, Cooper GM, Shendure J. A general framework for estimating the relative pathogenicity of human genetic variants. Nat Genet. 2014;46(3):310–5.
Rentzsch P, Witten D, Cooper GM, Shendure J, Kircher M. CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res. 2019;47(D1):D886–94.
Consortium TU. UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res. 2019;47(D1):D506–15.
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE. The Protein Data Bank. Nucleic Acids Res. 2000;28(1):235–42.
Waterhouse A, Bertoni M, Bienert S, Studer G, Tauriello G, Gumienny R, Heer FT, De Beer TAP, Rempfer C, Bordoli L, Lepore R, Schwede T, de Beer TAP, Rempfer C, Bordoli L, Lepore R, Schwede T. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 2018;46(W1):W296–303.
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE. UCSF Chimera-a visualization system for exploratory research and analysis. J Comput Chem. 2004;25:1605–12.
Han J, Ryu S, Moskowitz DM, Rothenberg D, Leahy DJ, Atzmon G, Barzilai N, Suh Y. Discovery of novel non-synonymous SNP variants in 988 candidate genes from 6 centenarians by target capture and next-generation sequencing. Mech Ageing Dev. 2013;134(10):478–85.
Barzilai N, Gabriely I, Gabriely M, Iankowitz N, Sorkin JD. Offspring of centenarians have a favorable lipid profile. J Am Geriatr Soc. 2001;49(1):76–9.
Bennett CF, Vander Wende H, Simko M, Klum S, Barfield S, Choi H, Pineda VV, Kaeberlein M. Activation of the mitochondrial unfolded protein response does not predict longevity in Caenorhabditis elegans. Nat Commun. 2014;5:3483–3483.
Cotter DG, Schugar RC, Crawford PA. Ketone body metabolism and cardiovascular disease. Am J Physiol-Heart Circ Physiol. 2013;304(8):H1060–76.
Puchalska P, Crawford PA. Multi-dimensional roles of ketone bodies in fuel metabolism, signaling, and therapeutics. Cell Metab. 2017;25(2):262–84.
Bajaj K, Madhusudhan MS, Adkar BV, Chakrabarti P, Ramakrishnan C, Sali A, Varadarajan R. Stereochemical criteria for prediction of the effects of proline mutations on protein stability. PLoS Comput Biol. 2007;3(12):e241–e241.
Hu X, Wang H, Ke H, Kuhlman B. High-resolution design of a protein loop. Proc Natl Acad Sci U S A. 2007;104(45):17668–73.
Regad L, Martin J, Nuel G, Camproux A-C. Mining protein loops using a structural alphabet and statistical exceptionality. BMC Bioinformatics. 2010;11(1):75–75.
Siira SJ, Spåhr H, Shearwood A-MJMJ, Ruzzenente B, Larsson N-GG, Rackham O, Filipovska A. LRPPRC-mediated folding of the mitochondrial transcriptome. Nat Commun. 2017;8(1):1532–1532.
Oláhová M, Hardy SA, Hall J, Yarham JW, Haack TB, Wilson WC, Alston CL, He L, Aznauryan E, Brown RM, Brown GK, Morris AAM, Mundy H, Broomfield A, Barbosa IA, Simpson MA, Deshpande C, Moeslinger D, Koch J, Stettner GM, Bonnen PE, Prokisch H, Lightowlers RN, McFarland R, Chrzanowska-Lightowlers ZMA, Taylor RW. LRPPRC mutations cause early-onset multisystem mitochondrial disease outside of the French-Canadian population. Brain. 2015;138(12):3503–19.
Sasarman F, Nishimura T, Antonicka H, Weraarpachai W, Shoubridge EA. Tissue-specific responses to the LRPPRC founder mutation in French Canadian Leigh Syndrome. Hum Mol Genet. 2014;24(2):480–91.
Thompson Legault J, Strittmatter L, Tardif J, Sharma R, Tremblay-Vaillancourt V, Aubut C, Boucher G, Clish Clary BB, Cyr D, Daneault C, Waters Paula JJ, Morin C, Laprise C, Rioux John DD, Mootha VK, Des Rosiers C, Aliskashani A, Allen Bruce GG, Beauchamp C, Bemeur C, Burelle Y, Charron G, Coderre L, Deschênes S, Labarthe F, Landry J, Lavallée G, Lavoie P, Maranda B, Mukaneza Y, Nishimura T, Rivard M-ÈÈ, Sasarman F, Shoubridge Eric AA, Legault JT, Tremblay N, Vachon L, Villeneuve J, Thompson Legault J, Strittmatter L, Tardif J, Sharma R, Tremblay-Vaillancourt V, Aubut C, Boucher G, Clish Clary BB, Cyr D, Daneault C, Waters Paula JJ, Aliskashani A, Allen Bruce GG, Aubut C, Beauchamp C, Bemeur C, Burelle Y, Charron G, Coderre L, Des Rosiers C, Deschênes S, Labarthe F, Landry J, Laprise C, Lavallée G, Lavoie P, Maranda B, Morin C, Mukaneza Y, Nishimura T, Rioux John DD, Rivard M-ÈÈ, Sasarman F, Shoubridge Eric AA, Tardif J, Thompson Legault J, Tremblay N, Tremblay-Vaillancourt V, Vachon L, Villeneuve J, Vachon L, Morin C, Laprise C, Rioux John DD, Mootha VK, Des Rosiers C. A Metabolic signature of mitochondrial dysfunction revealed through a monogenic form of Leigh syndrome. Cell Reports. 2015;13(5):981–9.
Köhler F, Müller-Rischart AK, Condradt B, Rolland SG. The loss of LRPPRC function induces the mitochondrial unfolded protein response. Aging. 2015;7(9):701–12.
Rolland SG, Motori E, Memar N, Hench J, Frank S, Winklhofer KF, Conradt B. Impaired complex IV activity in response to loss of LRPPRC function can be compensated by mitochondrial hyperfusion. Proc Natl Acad Sci U S A. 2013;110(32):E2967–76.
Bennett CF, Kaeberlein M. The mitochondrial unfolded protein response and increased longevity: cause, consequence, or correlation? Exp Gerontol. 2014;56:142–6.
Mouchiroud L, Houtkooper Riekelt H, Moullan N, Katsyuba E, Ryu D, Cantó C, Mottis A, Jo Y-S, Viswanathan M, Schoonjans K, Guarente L, Auwerx J. The NAD+/sirtuin pathway modulates longevity through activation of mitochondrial UPR and FOXO signaling. Cell. 2013;154(2):430–41.
Villani G, Attardi G. In vivo control of respiration by cytochrome c oxidase in human cells. Free Radical Biol Med. 2000;29(3–4):202–10.
de Brito OM, Scorrano L. Mitofusin 2 tethers endoplasmic reticulum to mitochondria. Nature. 2008;456(7222):605–10.
Merkwirth C, Langer T. Mitofusin 2 builds a bridge between ER and mitochondria. Cell. 2008;135(7):1165–7.
Wang Y, Hekimi S. Understanding ubiquinone. Trends Cell Biol. 2016;26(5):367–78.
Züchner S, Vance JM. Mechanisms of disease: a molecular genetic update on hereditary axonal neuropathies. Nat Clin Pract Neurol. 2006;2(1):45–53.
Cantó C. Mitochondrial dynamics: shaping metabolic adaptation. Int Rev Cell Mol Biol. 2018;340:129–67.
Rojo M, Legros F, Chateau D, Lombès A. Membrane topology and mitochondrial targeting of mitofusins, ubiquitous mammalian homologs of the transmembrane GTPase Fzo. J Cell Sci. 2002;115(8):1663–74.
Chen Y, Dorn GW. PINK1-phosphorylated mitofusin 2 is a Parkin receptor for culling damaged mitochondria. Science (New York, NY). 2013;340(6131):471–5.
De Brito OM, Scorrano L. An intimate liaison: spatial organization of the endoplasmic reticulum-mitochondria relationship. EMBO J. 2010;29(16):2715–23.
Johnson SC, Rabinovitch PS, Kaeberlein M. mTOR is a key modulator of ageing and age-related disease. Nature. 2013;493(7432):338–45.
Johnson SC, Dong X, Vijg J, Suh Y. Genetic evidence for common pathways in human age-related diseases. Aging Cell. 2015;14(5):809–17.
Kennedy BK, Lamming DW. The mechanistic target of rapamycin: the grand conducTOR of metabolism and aging. Cell Metab. 2016;23(6):990–1003.
Morita M, Gravel SP, Chénard V, Sikström K, Zheng L, Alain T, Gandin V, Avizonis D, Arguello M, Zakaria C, McLaughlan S, Nouet Y, Pause A, Pollak M, Gottlieb E, Larsson O, St-Pierre J, Topisirovic I, Sonenberg N, Vr Chnard, Sikstrm K, Zheng L, Alain T, Gandin V, Avizonis D, Arguello M, Zakaria C, McLaughlan S, Nouet Y, Pause A, Pollak M, Gottlieb E, Larsson O, St-Pierre J, Topisirovic I, Sonenberg N. MTORC1 controls mitochondrial activity and biogenesis through 4E-BP-dependent translational regulation. Cell Metabolism. 2013;18(5):698–711.
Lerner C, Bitto A, Pulliam D, Nacarelli T, Konigsberg M, Van Remmen H, Torres C, Sell C. Reduced mammalian target of rapamycin activity facilitates mitochondrial retrograde signaling and increases life span in normal human fibroblasts. Aging Cell. 2013;12(6):966–77.
Tokunaga C, Yoshino K-i, Yonezawa K. mTOR integrates amino acid- and energy-sensing pathways. Biochem Biophys Res Com. 2004;313(2):443–6.
Ben-Sahra I, Hoxhaj G, Ricoult SJH, Asara JM, Manning BD. mTORC1 induces purine synthesis through control of the mitochondrial tetrahydrofolate cycle. Science (New York, NY). 2016;351(6274):728–33.
Mittelstrass K, Ried JS, Yu Z, Krumsiek J, Gieger C, Prehn C, Roemisch-Margl W, Polonikov A, Peters A, Theis FJ, Meitinger T, Kronenberg F, Weidinger S, Wichmann HE, Suhre K, Wang-Sattler R, Adamski J, Illig T. Discovery of sexual dimorphisms in metabolic and genetic biomarkers. PLoS Genet. 2011;7(8):e1002215–e1002215.
Matone A, Scott-Boyer M-P, Carayol J, Fazelzadeh P, Lefebvre G, Valsesia A, Charon C, Vervoort J, Astrup A, Saris WHM, Morine M, Hager J. Network analysis of metabolite GWAS hits: implication of CPS1 and the urea cycle in weight maintenance. PLoS ONE. 2016;11(3):e0150495–e0150495.
Hartiala JA, Wilson Tang WH, Wang Z, Crow AL, Stewart AFR, Roberts R, McPherson R, Erdmann J, Willenborg C, Hazen SL, Allayee H. Genome-wide association study and targeted metabolomics identifies sex-specific association of CPS1 with coronary artery disease. Nat Commun. 2016;7(1):10558–10558.
Ahuja V, Powers-Lee SG. Human carbamoyl-phosphate synthetase: insight into N-acetylglutamate interaction and the functional effects of a common single nucleotide polymorphism. J Inherit Metab Dis. 2008;31(4):481–91.
Díez-Fernández C, Gallego J, Häberle J, Cervera J, Rubio V. The study of carbamoyl phosphate synthetase 1 deficiency sheds light on the mechanism for switching on/off the urea cycle. J Genet Genomics. 2015;42(5):249–60.
Pekkala S, Martínez AI, Barcelona B, Gallego J, Bendala E, Yefimenko I, Rubio V, Cervera J. Structural insight on the control of urea synthesis: identification of the binding site for N-acetyl-L-glutamate, the essential allosteric activator of mitochondrial carbamoyl phosphate synthetase. Biochem J. 2009;424(2):211–20.
Jazwinski SM, Jiang JC, Kim S. Adaptation to metabolic dysfunction during aging: making the best of a bad situation. Exp Gerontol. 2018;107:87–90.
Keshet R, Szlosarek P, Carracedo A, Erez A. Rewiring urea cycle metabolism in cancer to support anabolism. Nat Rev Cancer. 2018;18(10):634–45.
Takakusa H, Mohar I, Kavanagh TJ, Kelly EJ, Kaspera R, Nelson SD. Protein tyrosine nitration of mitochondrial carbamoyl phosphate synthetase 1 and its functional consequences. Biochem Biophys Res Commun. 2012;420(1):54–60.
Latorre-Pellicer A, Moreno-Loshuertos R, Lechuga-Vieco AV, Sánchez-Cabo F, Torroja C, Acín-Pérez R, Calvo E, Aix E, González-Guerra A, Logan A, Bernad-Miana ML, Romanos E, Cruz R, Cogliati S, Sobrino B, Carracedo Á, Pérez-Martos A, Fernández-Silva P, Ruíz-Cabello J, Murphy MP, Flores I, Vázquez J, Enríquez JA. Mitochondrial and nuclear DNA matching shapes metabolism and healthy ageing. Nature. 2016;535(7613):561–5.
Rose G, Passarino G, Carrieri G, Altomare K, Franceschi C, Greco V, Bertolini S, Bonafe M, Benedictis GD, Bonafè M, Franceschi C, De Benedictis G. Paradoxes in longevity: sequence analysis of mtDNA haplogroup J in centenarians. Eur J Hum Genet. 2001;9(9):701–7.
Santoro A, Salvioli S, Raule N, Capri M, Sevini F, Valensin S, Monti D, Bellizzi D, Passarino G, Rose G, De Benedictis G, Franceschi C. Mitochondrial DNA involvement in human longevity. Biochim Biophys Acta. 2006;1757(9–10):1388–99.
Slagboom PE, van den Berg N. Deelen J 2018 Phenome and genome based studies into human ageing and longevity: an overview. Biochim Biophys Acta (BBA) Mol Basis Dis. 1864;9:2742–51.
Acknowledgements
We would like to thank Genomics Shared Facility at Albert Einstein College of Medicine for their assistance with the iPLEX MassArray assay. We also would like to thank Axeq Technologies for performing all Illumina sequencing.
Funding
This work was supported by NIH grants AG069750, DK127778, AG057433, AG061521, HL150521, AG055501, AG057341, AG057706, AG057909, and AG17242 (Y.S), a grant from The Paul F. Glenn Center for the Biology of Human Aging (Y.S.), a grant GCRLE-1320 (Y.S.) from the Global Consortium for Reproductive Longevity and Equality at the Buck Institute, made possible by the Bia-Echo Foundation, and a grant from The Simons Foundation (Y.S.). B. G. was supported by NIH pre-doctoral aging training grant 6T32AG023475-13. S.R. is the recipient of a Glenn/AFAR Scholarships for Research in the Biology of Aging. M.K. was supported by NIH grant P30AG013280.
Author information
Authors and Affiliations
Contributions
MK and YS conceived and designed the experiments. BG analyzed the data. AT and SR performed experiments. GA and NB collected and provided AJ samples. BG, SR, SJ, MK, and YS contributed to writing of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
The study was approved by the Committee on Clinical Investigations of the Albert Einstein College of Medicine.
Consent to participate
All participants provided written informed consent beforehand regarding participation.
Consent for publication
All participants provided written informed consent beforehand regarding publication of the data.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Gonzalez, B., Tare, A., Ryu, S. et al. High-throughput sequencing analysis of nuclear-encoded mitochondrial genes reveals a genetic signature of human longevity. GeroScience 45, 311–330 (2023). https://doi.org/10.1007/s11357-022-00634-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11357-022-00634-z