Abstract
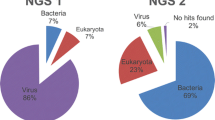
Many viruses can cause respiratory diseases in humans. Although great advances have been achieved in methods of diagnosis, it remains challenging to identify pathogens in unexplained pneumonia (UP) cases. In this study, we applied next-generation sequencing (NGS) technology and a metagenomic approach to detect and characterize respiratory viruses in UP cases from Guizhou Province, China. A total of 33 oropharyngeal swabs were obtained from hospitalized UP patients and subjected to NGS. An unbiased metagenomic analysis pipeline identified 13 virus species in 16 samples. Human rhinovirus C was the virus most frequently detected and was identified in seven samples. Human measles virus, adenovirus B 55 and coxsackievirus A10 were also identified. Metagenomic sequencing also provided virus genomic sequences, which enabled genotype characterization and phylogenetic analysis. For cases of multiple infection, metagenomic sequencing afforded information regarding the quantity of each virus in the sample, which could be used to evaluate each viruses’ role in the disease. Our study highlights the potential of metagenomic sequencing for pathogen identification in UP cases.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Assiri, A., McGeer, A., Perl, T.M., Price, C.S., Al Rabeeah, A.A., Cummings, D.A.T., Alabdullatif, Z.N., Assad, M., Almulhim, A., Makhdoom, H., Madani, H., Alhakeem, R., Al-Tawfiq, J.A., Cotten, M., Watson, S.J., Kellam, P., Zumla, A.I., Memish, Z.A., and Memish, Z.A. (2013). Hospital outbreak of middle east respiratory syndrome coronavirus. N Engl J Med 369, 407–416.
Briese, T., Renwick, N., Venter, M., Jarman, R.G., Ghosh, D., Köndgen, S., Shrestha, S.K., Hoegh, A.M., Casas, I., Adjogoua, E.V., Akoua-Koffi, C., Myint, K.S., Williams, D.T., Chidlow, G., van den Berg, R., Calvo, C., Koch, O., Palacios, G., Kapoor, V., Villari, J., Dominguez, S.R., Holmes, K.V., Harnett, G., Smith, D., Mackenzie, J.S., Ellerbrok, H., Schweiger, B., Schønning, K., Chadha, M.S., Leendertz, F.H., Mishra, A.C., Gibbons, R.V., Holmes, E.C., and Lipkin, W.I. (2008). Global distribution of novel rhinovirus genotype. Emerg Infect Dis 14, 944–947.
Dabisch-Ruthe, M., Vollmer, T., Adams, O., Knabbe, C., and Dreier, J. (2012). Comparison of three multiplex PCR assays for the detection of respiratory viral infections: evaluation of xTAG respiratory virus panel fast assay, RespiFinder 19 assay and RespiFinder SMART 22 assay. BMC Infect Dis 12, 163.
Ding, Y., Wang, H., Shen, H., Li, Z., Geng, J., Han, H., Cai, J., Li, X., Kang, W., Weng, D., Lu, Y., Wu, D., He, L., and Yao, K. (2003). The clinical pathology of severe acute respiratory syndrome (SARS): a report from China. J Pathol 200, 282–289.
Du, J., Yang, L., Ren, X., Zhang, J., Dong, J., Sun, L., Zhu, Y., Yang, F., Zhang, S., Wu, Z., and Jin, Q. (2016). Genetic diversity of coronaviruses in Miniopterus fuliginosus bats. Sci China Life Sci 59, 604–614.
Graf, E.H., Simmon, K.E., Tardif, K.D., Hymas, W., Flygare, S., Eilbeck, K., Yandell, M., and Schlaberg, R. (2016). Unbiased detection of respiratory viruses by use of RNA sequencing-based metagenomics: a systematic comparison to a commercial PCR panel. J Clin Microbiol 54, 1000–1007.
Heikkinen, T., and Järvinen, A. (2003). The common cold. Lancet 361, 51–59.
Hu, A., Cattaneo, R., Schwartz, S., and Norrby, E. (1994). Role of N-linked oligosaccharide chains in the processing and antigenicity of measles virus haemagglutinin protein. J Gen Virol 75, 1043–1052.
Hu, A., and Norrby, E. (1994). Role of individual cysteine residues in the processing and antigenicity of the measles virus haemagglutinin protein. J Gen Virol 75, 2173–2181.
Hu, A., Sheshberadaran, H., Norrby, E., and Kövamees, J. (1993). Molecular characterization of epitopes on the measles virus hemagglutinin protein. Virology 192, 351–354.
Huson, D.H., Auch, A.F., Qi, J., and Schuster, S.C. (2007). MEGAN analysis of metagenomic data. Genome Res 17, 377–386.
Ishikane, M., Kato, H., Kawabata, K., Ito, H., Kanayama, A., Matsui, T., and Oishi, K. (2015). The Cutting-edge of Medicine; A recent trend of emerging infections: MERS and Avian influenza A (H7N9). Nihon Naika Gakkai Zasshi 104, 114–119.
Jianzhi, X., and Zhihui, C. (1983). Measles vaccine in the People’s Republic of China. Clin Infect Dis 5, 506–510.
Khetsuriani, N., Lu, X., Teague, W.G., Kazerouni, N., Anderson, L.J., and Erdman, D.D. (2008). Novel human rhinoviruses and exacerbation of asthma in children1. Emerg Infect Dis 14, 1793–1796.
Kiliç, A., Altinkaynak, S., Ertekin, V., and Inandi, T. (2003). The duration of maternal measles antibodies in Children. J Trop Pediatr 49, 302–305.
Kohl, C., Brinkmann, A., Dabrowski, P.W., Radonic, A., Nitsche, A., and Kurth, A. (2015). Protocol for metagenomic virus detection in clinical specimens. Emerg Infect Dis 21, 48.
Lu, Q.B., Zhang, X.A., Wo, Y., Xu, H.M., Li, X.J., Wang, X.J., Ding, S.J., Chen, X.D., He, C., Liu, L.J., Li, H., Yang, H., Li, T.Y., Liu, W., and Cao, W.C. (2012). Circulation of coxsackievirus A10 and A6 in handfoot- mouth disease in China, 2009–2011. PLoS ONE 7, e52073.
Palacios, G., Druce, J., Du, L., Tran, T., Birch, C., Briese, T., Conlan, S., Quan, P.L., Hui, J., Marshall, J., Simons, J.F., Egholm, M., Paddock, C.D., Shieh, W.J., Goldsmith, C.S., Zaki, S.R., Catton, M., and Lipkin, W.I. (2008). A new arenavirus in a cluster of fatal transplant-associated diseases. N Engl J Med 358, 991–998.
Pierce, V.M., and Hodinka, R.L. (2012). Comparison of the GenMark Diagnostics eSensor respiratory viral panel to real-time PCR for detection of respiratory viruses in children. J Clin Microbiol 50, 3458–3465.
Plemper, R.K., Hammond, A.L., and Cattaneo, R. (2000). Characterization of a region of the measles virus hemagglutinin sufficient for its dimerization. J Virol 74, 6485–6493.
Qian, Y.H., Su, J., Shi, P., He, E.Q., Shao, J., Sun, N., Zu, R.Q., and Yu, R.B. (2011). Attempted early detection of influenza A (H1N1) pandemic with surveillance data of influenza-like illness and unexplained pneumonia. Influenza Other Respir Viruses 5, e479–e486.
Rudan, I., O’Brien, K.L., Nair, H., Liu, L., Theodoratou, E., Qazi, S., Luksic, I., Fischer Walker, C.L., Black, R.E., Campbell, H., and Child Health Epidemiology Reference, G. (2013). Epidemiology and etiology of childhood pneumonia in 2010: estimates of incidence, severe morbidity, mortality, underlying risk factors and causative pathogens for 192 countries. J Glob Health 3, 010401.
Xie, Q., Cao, Y.J., Su, J., Wu, X.B., Wan, C.S., Ke, C.W., Zhao, W., and Zhang, B. (2015). Genomic sequencing and analysis of the first imported Middle East Respiratory Syndrome Coronavirus (MERS CoV) in China. Sci China Life Sci 58, 818–820.
Xu, S., Zhang, Y., Rivailler, P., Wang, H., Ji, Y., Zhen, Z., Mao, N., Li, C., Bellini, W.J., Xu, W., and Rota, P.A. (2014). Evolutionary genetics of genotype H1 measles viruses in China from 1993 to 2012. J Gen Virol 95, 1892–1899.
Yang, Q., Ding, J., Cao, J., Huang, Q., Hong, C., and Yang, B. (2015). Epidemiological and etiological characteristics of hand, foot, and mouth disease in Wuhan, China from 2012 to 2013: Outbreaks of Coxsackieviruses A10. J Med Virol 87, 954–960.
Zhang, Y., Xu, S., Wang, H., Zhu, Z., Ji, Y., Liu, C., Zhang, X., Sun, L., Zhou, J., Lu, P., Hu, Y., Feng, D., Zhang, Z., Wang, C., Fang, X., Zheng, H., Liu, L., Sun, X., Tang, W., Wang, Y., Liu, Y., Gao, H., Tian, H., Ma, J., Gu, S., Wang, S., Feng, Y., Bo, F., Liu, J., Si, Y., Zhou, S., Ma, Y., Wu, S., Zhou, S., Li, F., Ding, Z., Yang, Z., Rota, P.A., Featherstone, D., Jee, Y., Bellini, W.J., and Xu, W. (2012). Single endemic genotype of measles virus continuously circulating in China for at least 16 years. PLoS ONE 7, e34401.
Zoll, J., Rahamat-Langendoen, J., Ahout, I., De Jonge, M.I., Jans, J., Huijnen, M.A., Ferwerda, G., Warris, A., and Melchers, W.J.G. (2015). Direct multiplexed whole genome sequencing of respiratory tract samples reveals full viral genomic information. J Clin Virol 66, 6–11.
Acknowledgements
We thank Dr. Zhen Zhu in China Institute of Veterinary Drug Control (IVDC) for her help on sequence characterization of the measles virus. This study was supported by the National Mega-projects for Infectious Diseases (2014ZX10004002), and Beijing Municipal Science and Technology Commission grant (D151100002115003).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Contributed equally to this work
This article is published with open access at springerlink.fh-diploma.de
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0), which permits use, duplication, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Zou, X., Tang, G., Zhao, X. et al. Simultaneous virus identification and characterization of severe unexplained pneumonia cases using a metagenomics sequencing technique. Sci. China Life Sci. 60, 279–286 (2017). https://doi.org/10.1007/s11427-016-0244-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-016-0244-8