Abstract
Polymyxin acts as an ultimate line of refuge against the severe infections by multidrug-resistant Gram-negative pathogens. This conventional idea is challenged dramatically by the recent discovery of mobile colistin resistance gene (mcr-1) is prevalent in food animals and human beings worldwide. More importantly, the mcr-1 gene was found to be co-localized with other antibiotic resistance genes, raising the possibility that super-bugs with pan-drug resistance are emerging. However, little is reported on the genomes of the mcr-1-positive bacterial host reservoirs. Here we report genome sequencing of three human isolates of the mcr-1-positive Escherichia coli (E15004, E15015 and E15017) and define general features through analyses of bacterial comparative genomics. Further genomic mining together with sequence typing allowed us to elucidate that the MCR-1-carrying E. coli E15017 belongs to the sequence type ST648 and coproduces extended-spectrum β-lactamase (ESBL). Given the fact that ST648 has been known to associate with either New Delhi metallo-β-lactamase 1 or ESBL, our results highlighted the possibility of ST648 as an epidemic clone with multidrug resistances.
摘要
通常认为多粘菌素是抵御多药耐受的革兰阴性病原菌严重感染的最后一道防线。 最新研究表明在动物类食品和人体肠道菌群中存在一种多粘菌素抗性基因 (mcr-1)。这一惊人发现对上述传统观点提出了严重挑战。更为惊讶的是, 该 mcr-1 基因可与其他抗生素抗性基因共存, 极大增加了产生具有广泛耐药的超级 细菌的可能性。但是, 关于mcr-1 阳性细菌的基因组学特征鲜见报道。本研究对 3 株 mcr-1 阳性人源大肠杆菌(E15004, E15015 和E15017)开展了全基因组 测序, 并通过比较基因组学描述了其基因组学特征。基因组深入挖掘和序列分型 实验揭示了序列型为ST648 的大肠杆菌E15017 不仅编码MCR-1 多粘菌素耐药 基因, 而且携带超广谱β-内酰胺耐药基因(ESBL)。鉴于序列型ST648 的大肠杆 菌通常与新德里β-内酰胺耐药-1 (NDM-1) 或ESBL 紧密关联这一事实, 我们的 研究结果提示了具有MCR-1 和ESBL 耐药的ST648 细菌发展成流行性克隆的潜 在风险。
Similar content being viewed by others
Avoid common mistakes on your manuscript.
The identification of the mobilized colistin resistance gene mcr-1 recently attracted extensive attention from the scientific community. MCR-1 confers resistance to polymyxins, a group of polypeptide antibiotics that are currently considered the last refuge of therapeutics against lethal challenges by Gram-negative pathogens with multi-drug resistance [1, 2]. Very recently, two separate groups reported the co-occurrence of MCR-1 and extended-spectrum β-lactamase (ESBL) on plasmids in Enterobacteriaceae [3–6]. However, genomic hallmarks of the bacterial host reservoir for the mcr-1-harbouring plasmids remain unclear. Here we report on their genomic compositions.
After three mcr-1-positive E. coli isolates (E15004, E15015 and E15017) were successfully screened from the microbiota of clinical diarrhea patients [7], we applied next-generation Illumina MiSeq sequencing to decode their genomic sequences. The pool of paired-end reads produced here were assembled with GS De Novo Assembler into a collection of contigs. Then the individual contigs were ordered into draft genomes with the prototypical strain of E. coli MG1655 as the reference (Fig. 1, S1). Relative to the paradigm version of E. coli, MG1655 (4,641,425 bp), the three mcr-1-positive clinical E. coli isolates exhibited variations in the size of sequenced genomes (i.e., 4,643,275 bp for strain E15004; 4,637,424 bp for strain E15015, and 4,780,540 bp for strain E15017) (Table S1). The values of their GC percentages are all approximately 50 % (Table S1), although the draft genomes identified several regions with a strong GC skew, indicative of novel insertions of genomic material.
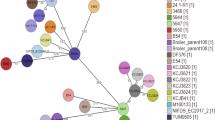
Genomics-based discovery of multidrug-resistant genes in the mcr-1-positive ST648 E. coli coproducing extended-spectrum β-lactamase. Circular comparison of the three sequenced genomes (E15004, E15015 and E15017) with the paradigm strain MG1655 as the reference. Individual rings range from 1 (inner ring) to 4 (outer ring). (Ring 1—red) Strain 15005 conservation plot. (Ring 2—green) Strain 15015 conservation plot. (Ring 3—blue) Strain 15015 conservation plot. (Ring 4—magenta/green) GC Skew of MG1655 reference genome [(G−C)/(G+C)] magenta > 0, green < 0
Further comparative genomics suggests that genetic heterogeneity is present in the three mcr-1-positive E. coli isolates (Fig. 1, S2). We retrieved the sequences of seven house-keeping genes (adk, fumC, gyrB, icd, mdh, purA and recA) from the above three sequenced genomes and subjected them to analyses of Multi-Locus Sequence Typing (MLST) (http://mlst.warwick.ac.uk/mlst/dbs/Ecoli). Unlike the epidemic spreading clone, E. coli ST131 that carried the mcr-1 gene in Denmark [8], the three mcr-1-harbouring clinical strains belong to different sequence types (i.e., E15004 is in ST40, E15015 is in ST642, and E15017 is in ST648) (Table 1, Fig. S3), which is generally consistent with our findings from comparative genomics (Fig. 1, S2). The fact that mcr-1-harbouring E. coli isolates are classified into different sequence types argues that the dissemination of mcr-1 colistin resistance gene is ongoing by clonal expansion [9]. Given the fact that E. coli ST648 was associated with ESBL [10, 11] and two variants of New Delhi metallo-β-lactamase 1 (NDM-1), NDM-5 [12] and NDM-7 [13]), we thereby were interested in determining whether or not the genes of ESBL and NDM would also be found with the mcr-1 gene in the ST648 strain, E15017.
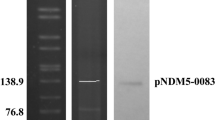
Using ResFinder2.1, a newly-improved database for identifying antibiotic resistance genes (https://cge.cbs.dtu.dk/services/ResFinder), we screened the above three genomic sequences, as well as the remaining unordered contigs, which likely encode additional plasmids, for the presence of antibiotic resistance genes esp. ESBL and NDM-1 (and/or its variants). As anticipated, a 100 % identical mcr-1 gene was observed in the unordered contigs in each of the three strains. NDM-1 variants were not found, which we then verified by PCR-based detection (not shown). Unexpectedly, no other antibiotic resistance gene besides mcr-1 is found in the strain E15004 (ST40) (not shown), whereas multiple drug-resistance genes apart from mcr-1 were identified in the unordered contigs from the other two strains, E15015 (ST642) and E15017 (ST648) (Table 2, S2). In particular, the blaCTX-M-15 gene that encodes ESBL was found to be present in the ST648 strain, E15017 (Table 2). Additionally, we noted that the mcr-1 and blaCTX-M-15 are located inside distinct unordered contigs, suggesting the possibility that they are encoded on different plasmids. This represents the first example of a clinical clone of E. coli with a sequence type of ST648 that has the potential to spread MCR-1 colistin resistance together with ESBL resistance.
In summary, our data provides genomic insights into three strains of mcr-1-positive E. coli with multiple drug resistance, which reveals the increasing possibility of ST648 becoming an epidemic vector for circulation/spread of the mcr-1 colistin resistance gene in China. As the inter/intra-species dissemination of the mcr-1 gene has been linked to the spread of other drug resistance including ESBL [11] and NDM-1 variants [12, 13], our findings underscore the urgent need to modulate and control the use of colistin in veterinary/clinical practices, which might facilitate prevention of the further emergence of superbugs with multi-drug resistance.
References
Paterson DL, Harris PN (2015) Colistin resistance: a major breach in our last line of defence. Lancet Infect Dis 16(2):132–133
Nation RL, Li J, Cars O et al (2015) Framework for optimisation of the clinical use of colistin and polymyxin B: the Prato polymyxin consensus. Lancet Infect Dis 15:225–234
Falgenhauer L, Waezsada SE, Yao Y et al (2016) Colistin resistance gene mcr-1 in extended-spectrum β-lactamase-producing and carbapenemase-producing Gram-negative bacteria in Germany. Lacet Infect Dis 16:282–283
Haenni M, Poirel L, Kieffer N et al (2016) Co-occurrence of extended spectrum β-lactamase and MCR-1 encoding genes on plasmids. Lancet Infect Dis 16:281–282
Kluytmans-van den Bergh MF, Huizinga P, Bonten MJ et al (2016) Presence of mcr-1-positive Enterobacteriaceae in retail chicken meat but not in humans in the Netherlands since 2009. Euro Surveill. doi:10.2807/1560-7917.ES.2016.21.9.30149
Zurfuh K, Poirel L, Nordmann P et al (2016) Occurrence of the plasmid-borne mcr-1 colistin resistance gene in extended-spectrum-beta-lactamase-producing Enterobacteriaceae in river water and imported vegetable samples in Switzerland. Antimicrob Agents Chemother 60:2594–2595
Ye H, Li Y, Li Z et al (2016) Diversified mcr-1-harbouring plasmid reservoirs confer resistance to colistin in human gut microbiota. MBio pii:e00177-16
Hasman H, Hammerum AM, Hansen F et al (2015) Detection of mcr-1 encoding plasmid-mediated colistin-resistant Escherichia coli isolates from human bloodstream infection and imported chicken meat, Denmark 2015. Euro Surveill. doi:10.2807/1560-7917.ES.2015.20.49.30085
Hu Y, Liu F, Lin IY et al (2015) Dissemination of the mcr-1 colistin resistance gene. Lancet Infect Dis 16:146–147
Zhao SY, Wang YC, Xiao SZ et al (2015) Drug susceptibility and molecular epidemiology of Escherichia coli in bloodstream infections in Shanghai, China, 2011–2013. Infect Dis 47:310–318
Sherchan JB, Hayakawa K, Miyoshi-Akiyama T et al (2015) Clinical epidemiology and molecular analysis of extended-spectrum-beta-lactamase producing Escherichia coli in Nepal: characteristics of sequence types 131 and 648. Antimicrob Agents Chemother 59:3424–3432
Hornsey M, Phee L, Wareham DW (2011) A novel variant, NDM-5, of the New Delhi metallo-beta-lactamase in a multidrug-resistant Escherichia coli ST648 isolate recovered from a patient in the United Kingdom. Antimicrob Agents Chemother 55:5952–5954
Mizuno Y, Yamaguchi T, Matsumoto T (2014) A first case of New Delhi metallo beta-lactamase-7 in an Escherichia coli ST648 isolate in Japan. J Infect Chemother 20:814–816
Acknowledgments
This work was supported by Zhejiang Provincial Natural Science Foundation for Distinguished Young Scholars (LR15H190001), the National Natural Science Foundation of China (31570027), and a start-up package from Zhejiang University (Y.F.). Dr. Feng is a recipient of the “Young 1000 Talents” Award.
Author contributions
Y.F. designed this project; Y.F., H.Z., C.S., Z.W., and H.Y. performed experiments and analyzed the data; Y.F. H.Z., C.S., and Z.W. contributed reagents and tools; Y.F. and C.S. prepared this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Huimin Zhang and Christopher H. Seward contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Zhang, H., Seward, C.H., Wu, Z. et al. Genomic insights into the ESBL and MCR-1-producing ST648 Escherichia coli with multi-drug resistance. Sci. Bull. 61, 875–878 (2016). https://doi.org/10.1007/s11434-016-1086-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-016-1086-y