Abstract
Background
Mesenchymal-to-epithelial transition (MET) exon 14 skipping mutations and MET gene amplification occur in 3–5% of non-small cell lung cancer (NSCLC) patients. Tyrosine kinase inhibitors (TKIs) targeting MET alterations have shown promising results in these patients.
Objective
The aim of this study was to describe the genomic profile, PD-L1 expression and clinicopathological features of MET dysregulated NSCLC.
Patients and Methods
We identified 188 patients with advanced-stage NSCLC with data on MET expression by immunohistochemistry (IHC). IHC for PD-L1 expression was performed in 131 patient samples, and next-generation sequencing (NGS) analysis was performed in 109 patient samples.
Results
MET exon 14 skipping alterations were identified in 16 (14.7%) samples, MET amplifications with cut-off ≥4 copy number variations were identified in 11 (10.1%) samples, and an oncogenic MET mutation (MET p.D1228N) was identified in 1 (0.9%) sample. 12/15 tumors (80.0%) harboring MET exon 14 alterations and 7/11 (63.6%) MET-amplified tumors expressed PD-L1 in ≥1% of tumor cells. Tumors harboring MET exon 14 skipping alterations expressed PD-L1 more frequently than MET wild-type IHC-positive tumors (p = 0.045). Twenty-five percent of MET exon 14-altered cases and 33% of MET-amplified cases harbored potentially targetable oncogenic co-mutations in KRAS, BRAF, and EGFR. The most frequent co-occurring mutations in all MET-altered tumors were TP53, KRAS, BRAF, and CDK4.
Conclusions
We demonstrated that MET exon 14 skipping alterations and MET amplification are not mutually exclusive to other oncogenic co-mutations, and report the association of genomic MET alterations with PD-L1 expression. Since genomic MET alterations are emerging targets requiring upfront treatment, optimal understanding of the co-mutational landscape for this patient population is needed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Understanding the co-mutational profile of mesenchymal-to-epithelial transition (MET)-altered tumors is crucial as MET-directed tyrosine kinase inhibitors have emerged as treatment options. |
In 188 patients with advanced-stage non-small cell lung cancer, MET exon 14 skipping alterations were associated with PD-L1 expression. |
Several MET exon 14-altered and MET-amplified tumors harbored oncogenic co-mutations in KRAS, BRAF, and EGFR. |
1 Introduction
The mesenchymal-to-epithelial transition (MET) proto-oncogene on chromosome 7 encodes for the membrane-spanning tyrosine kinase receptor MET and is activated by the hepatocyte growth factor (HGF) [1, 2]. HGF and MET signaling induce proliferation, epithelial-mesenchymal transition, scattering, and invasion of epithelial cells, and promote anti-apoptotic responses in the tumor microenvironment [3,4,5]. Aberrant MET and HGF signaling have been demonstrated in a wide range of carcinomas, including carcinomas of the gastrointestinal tract, head and neck, and lung [6,7,8,9].
MET alterations in non-small cell lung cancer (NSCLC) can occur at the genomic level through mutations, amplifications, or gene fusions. Mutations within the juxtamembrane domain of MET encoded by exon 14 occur in 3–4% of NSCLCs [10,11,12,13]. Mutations within splice sites flanking MET exon 14 result in MET exon 14 skipping. In this case, degradation of the MET receptor is hampered, whereas MET activation and signaling are sustained, resulting in oncogenic transformation. To investigate such alterations RNA sequencing can be applied to detect fusions between exon 13 and exon 15 (resulting in MET exon 14 skipping), whereas DNA sequencing detects mutations predicted to result in MET exon 14 skipping. Differences in the detection rate of MET exon skipping events between RNA- and DNA-based sequencing were reported [14, 15]. High-level amplification of the MET gene, present in about 3–5% of lung adenocarcinomas, may also lead to MET-dependent oncogenesis [16,17,18,19,20]. MET gene copy numbers are assessed by either fluorescence in situ hybridization (FISH) or next-generation sequencing (NGS) gene panels. At the protein level, MET overexpression is detected by immunohistochemistry (IHC) in approximately 22–24% of NSCLCs and may occur in the absence of genomic MET alterations [10, 16, 20,21,22,23]. Thus, it is not used as a screening tool for detecting MET amplification or MET exon 14 skipping. Importantly, available MET tyrosine kinase inhibitors (TKIs) have shown little success in tumors overexpressing MET in the absence of mutations within the MET gene [24,25,26,27]. Several TKIs in trials for MET-driven cancers inhibited cancer growth in MET-amplified as well as MET exon 14-mutated tumors [28,29,30,31,32,33,34]. Still, resistance to TKIs develops in the course of treatment and even pre-exists in certain tumors, and the mechanisms of such primary resistance remain unclear [35,36,37,38,39]. Earlier reports based on large NGS panels [11] and smaller polymerase chain reaction (PCR)-based gene panels [12, 40] stated that MET exon 14 alterations are mutually exclusive to other oncogenic driver mutations. Some studies indicated an association of MET dysregulation with PD-L1 expression [41, 42]. In order to characterize the genomic and clinical features of MET dysregulated tumors in patients with advanced NSCLC, we analyzed the genomic profile, PD-L1 expression, and clinicopathological features in relation to MET dysregulation.
2 Methods
2.1 Study Population
In this retrospective study, patients with advanced stage IV NSCLC treated at the Comprehensive Cancer Centre Zurich (C3Z), University Hospital Zurich between January 2011 and August 2020 with either c-MET staining performed at the time of diagnosis or available tissue and clinical follow-up data were included. Exclusion criteria were non-availability of tumor tissue for c-MET staining, rejection of general consent, or the lack of or incomplete clinical data. A total of 188 patients were included in this analysis and all medical records reviewed. Response Evaluation Criteria in Solid Tumors (RECIST) were applied to evaluate treatment response. Overall survival was collected from the medical records; when data were not available, patients or relatives were contacted by phone.
This study was conducted according to the law and regulations of the local Ethics Commission under reference number KEK ZH-2021-00381.
2.2 Immunohistochemistry and Fluorescence In Situ Hybridization
Formalin-fixed, paraffin-embedded 4 μm sections from tumor blocks were stained by IHC using rabbit anti-c-MET monoclonal antibody (clone SP44, dilution 1/50; Abcam, Cambridge, UK) and rabbit anti-PD-L1 monoclonal antibody (clone E1L3N, dilution 1/100; Cell Signaling Technology, Danvers, MA, USA). Staining was performed with an automated immunostainer (DiscoveryUltra; Roche Ventana). MET overexpression was scored semi-quantitatively as described by Spigel et al. [26], corresponding to an immunoscore of 3+ (≥ 50% of tumor cells stained exhibiting strong staining intensity) or 2+ (≥ 50% of tumor cells with moderate or higher staining intensity but < 50% strong intensity). PD-L1 expression was scored as described previously by Lacour et al. PD-L1 positivity was defined as ≥ 1% of tumor cells with stained membrane [43]. Scoring was performed by experienced pathologists.
For FISH analysis, MET was labeled with Abbott Molecular/Vysis MET SpectrumRed Probe (7q31.2) and the CEP7 region with Abbott Molecular/Vysis CEP (D7Z1) SpectrumGreen Probe (7p11.1-q11.1) [Abbott Molecular, Baar, Switzerland]. Staining with the fluorescent probes was performed according to the manufacturer’s protocol. The CEP7-labeled region was used as a reference for MET amplification. For scoring purposes, 100 tumor cells were evaluated and the mean MET/CEP7 ratio was calculated. Amplification of the MET gene was defined as a ratio of MET/CEP7 ≥ 2 according to the University of Colorado Cancer Center criteria [19].
2.3 Next-Generation Sequencing (NGS)
NGS was performed using the commercially available Oncomine Focus Assay (OFA) panel (Thermo Fisher Scientific, Carlsbad, CA, USA), enabling detection of variants in 52 genes, or with the DNA part of the Oncomine Comprehensive Assay (OCA) panel (Thermo Fisher Scientific), enabling detection of variants in 161 genes. Samples were analyzed according to the manufacturer’s protocol. Briefly, DNA and RNA were isolated from formalin-fixed, decalcified, paraffin-embedded tumor blocks with a Maxwell 16 FFPE Tissue LEV DNA/RNA Purification Kit (Promega, Fitchburg, WI, USA). Sequencing was performed on the Ion S5TM System using the Ion 540 Sequencing Kit (Thermo Fisher Scientific). Ion Reporter software 5.10 (Thermo Fisher Scientific) was used for alignment (hg19/GRChr37), variant calling and annotations (Oncomine Focus w2.4, DNA/fusions, single sample; filter chain: Oncomine 5% CI, copy number variation (CNV), ploidy greater than or equal to a gain of 2 over normal; and Oncomine Comprehensive v3—w3.2, DNA, single sample; filter chain: Oncomine 5% CI, CNV, ploidy greater than or equal to a gain of 2 over normal OR ≥ 0.5 over normal). MET amplification was reported at a cut-off of ≥ 4 CNVs and MET exon 14 skipping at a cut-off of 0.02% skipping reads in relation to total mapped fusion panel reads. For a number of patients, FoundationOne CDxTM (Foundation Medicine, Cambridge, MA, USA) was performed according to the manufacturer’s protocol and results incorporated in this study.
2.4 Statistics
Descriptive statistics were used to describe tumor and patient characteristics. The Fisher’s exact test was applied to test for significance and Kaplan–Meier survival curves were applied for survival analysis. P values < 0.05 were considered statistically significant. Statistical analyses were performed using IBM SPSS Statistics 26.0 (IBM Corporation, Armonk, NY, USA).
3 Results
3.1 Patient Characteristics
Overall, 188 patients with advanced-stage NSCLC (94.3% stage IV, 5.7% stage III) were included in this single-center, retrospective cohort study. Patient characteristics are summarized in Table 1. Median age at diagnosis was 62 years (range 33–89). The majority of patients were men (106/188 [56.4%]) and 156/187 patients were former or current smokers (83.4%; one patient smoking status was not available).
3.2 Immunohistochemistry
Tumor specimens of all patients with available tissue were stained for c-MET. Of 188 patient samples, 94 (50.0%) were positive for MET IHC. PD-L1 expression, either at diagnosis or for the purpose of this study, was available in 131 patients, of whom 69 (52.7%) were positive (cut-off 1% PD-L1 positivity on tumor cells). PD-L1 expression < 1% was detected in 62 patients (47.3%), expression between 1% and 49% was detected in 40 patients (30.5%), and expression ≥ 50% was detected in 29 patients (22.1%).
3.3 Characterization of MET Alterations
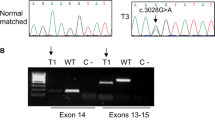
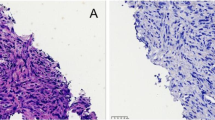
NGS data were available for 109 (58.0%) patients. Eighty-three (44.1%) patient samples were sequenced with the OFA, thereof 19 samples were additionally sequenced with the OCA, and 26 (13.8%) patient samples were analyzed using FoundationOne CDxTM. From these cases, 28/109 (25.7%) displayed genomic MET alterations. Sixteen cases (14.7%) harbored MET exon 14 skipping alterations and one case displayed a concurrent MET amplification (Fig. 1). Eleven cases were found with MET amplifications alone (10.1%) and a single case (0.9%) with an oncogenic MET mutation (MET p.D1228N) not leading to MET exon 14 skipping (Fig. 1). 26/28 (92.9%) cases with genomic MET alterations overexpressed c-MET. In two cases with MET exon 14 skipping alterations, MET expression was negative (one patient with squamous cell carcinoma and one patient with pleomorphic carcinoma). Fifty-one MET IHC-negative cases were tested for MET amplification by FISH but not NGS (at the time of diagnosis, not a diagnostic standard). No MET amplification was detected by FISH within this group. Further genomic MET dysregulation, in particular MET exon 14 mutations, cannot be excluded in those cases.
Characterization of MET alterations. MET mutations resulting in MET exon 14 skipping were reported for seven patients with positive MET IHC: P1 MET p.D1010N, P4 MET c.2942-20TTCTTTCTCTC>T, P7 MET splicesite_3, P10 MET c.2942-18CTTTCTCTCTGTTTT>C, P12 MET c.3029C>T, P16 MET c.2888-28_2888-13>A; in MET IHC negative: P9 MET c.2888-18_2888-17insAGn. A single case with an oncogenic MET mutation (MET p.D1228N) not leading to MET exon 14 skipping was observed (non-MET E14 mutation). MET mesenchymal-to-epithelial transition, IHC immunohistochemistry, NGS next-generation sequencing
3.4 PD-L1 Expression is Upregulated in Tumors Harboring MET Exon 14 Skipping Alterations Compared with MET Wild-Type IHC-Positive Tumors
PD-L1 status was evaluated in 27/28 MET-altered tumors, 58/61 MET wild-type with IHC-positive (MET wtIHCpositive) tumors, and 46/92 MET wild-type with IHC-negative (MET wtIHCnegative) tumors. Genomic MET alterations were associated with PD-L1 positivity, defined as PD-L1 expression in ≥1.0% of tumor cells. 12/15 (80.0%) tumors harboring MET exon 14 alterations and 7/11 (63.6%) MET-amplified tumors were PD-L1 positive, compared with 29/58 (50.0%) MET wtIHCpositive tumors and 20/46 (43.5%) MET wtIHCnegative tumors. The expression of PD-L1 was statistically different between the group of tumors harboring MET exon 14 skipping mutations and MET wtIHCpositive tumors (p = 0.045) when adjusted for multiple testing.
3.5 Co-Mutational Profile of MET Dysregulated Tumors Through NGS Analysis
In 4/16 (25.0%) MET exon 14-altered cases and 3/9 (33.3%) MET-amplified cases, mutations within other potentially targetable oncogenic driver genes such as KRAS (one MET exon 14-altered case; two MET-amplified cases), EGFR (one MET exon 14-altered case), BRAF (two MET exon 14-altered cases; one MET-amplified case), and ALK-Fusion (one MET exon 14-altered case) were found. In two patients (7.1%), MET amplifications occurred as acquired resistance to therapy with EGFR TKI (patients 23 and 24) (Fig. 2a). As a consequence, these two are excluded from calculation on co-mutations. In three patients, co-occurring EGFR alterations (one mutation and two amplifications) were diagnosed previous to treatment with EGFR TKIs (patients 7, 12, and 25) (Fig. 2a). Three patients showed common covariant in BRAF (p.D945G, p.V600E, p.D469R) diagnosed previous to BRAF-targeted therapy (patients 14, 17, and 24) (Fig. 2a).
a Co-mutational profile of MET exon 14-altered, MET-mutated (patient 2) and MET-amplified tumors. Specific co-mutations detected in EGFR (patient 7: amplification; patient 12: p.Pro772_His773insHisAla; patient 23: p.Leu858Arg; patient 24: Exon19del (A750_I759>PT); patient 25: amplification); KRAS (patient 8: amplification; patient 15: p.Gly12Ala; patients 16 and 21 p.Gly12Cys); BRAF (patient 14: p.Asp594Gly; patient 17: p.Val600Glu; patient 24: p.Gly469Arg; patient 25: amplification); ERBB3 (patient 13: p.Val104Leu); ATR (patient 10: p.Leu2208Te); ALK (patients 12 and 23: fusion). Variants with unknown significance detected with FoundationOne CDxTM are shown in electronic supplementary Table 1. * In patients 23 and 24, MET amplifications occurred as acquired resistance to therapy with EGFR TKIs. b Co-mutations in wild-type MET IHC-positive tumors. Patients with no detected mutations in NGS are not shown (patients 32, 60, 64, 69, 75, 80, 86, 87, 88). Variants with unknown significance detected with FoundationOne CDxTM are shown in electronic supplementary Table 1. MET mesenchymal-to-epithelial transition, IHC immunohistochemistry, EGFR epidermal growth factor receptor, TKIs tyrosine kinase inhibitors, NGS next-generation sequencing
TP53 was the most frequent co-occurring mutation in tumors harboring MET exon 14 skipping alterations or MET amplification (4/16 [25.0%] and 5/9 [55.5%] patients, respectively) (Fig. 2a). Further common co-occurring mutations or CNVs in MET-altered tumors were KRAS (12.5% of MET exon 14-altered cases; 22.0% of MET-amplified cases), BRAF (12.5% of MET exon 14-altered cases; 11.1% of MET-amplified cases), CDK4 (18.8% of MET exon 14-altered cases; 11.1% of MET-amplified cases), CDKN2A (33.3% of MET-amplified cases) and TSC2 (6.3% of MET exon 14-altered cases; 22.2% of MET-amplified cases) (Fig. 2a).
MET amplification is reported at a cut-off of ≥ 4 CNVs. Patient 20, with the highest CNV in this cohort (16.4 CNVs), had no co-occurring mutations. In the group of MET amplification with 4 \(\le\) CNVs < 11, no association between the level of MET amplification and the number of oncogenic co-mutations was observed. Patient 8, with MET exon 14 skipping and MET amplification (6.1 CNVs), did not harbor further oncogenic driver mutations. Two MET-amplified tumors displayed amplification of the HGF gene, the ligand for the MET tyrosine kinase receptor (Fig. 2a).
In 49/61 (80.3%) MET wtIHCpositive tumors, mutations or fusions in other potentially targetable oncogenic driver genes such as KRAS, EGFR, ALK, BRAF, ERBB2 were found. The most frequent oncogenic mutations within this group were KRAS (47.5%), followed by alterations in TP53 (18.0%) and CDKN2A (13.1%) (Fig. 2b). The most frequent single nucleotide variants were KRAS p.G12C (9/61, 14.8%) and KRAS p.G12V (7/61, 11.5%) on exon 2. In the group of MET wtIHCnegative tumors, 13/20 (65%) MET wtIHCnegative tumors with available NGS data harbored mutations or fusions in other potentially targetable oncogenic driver genes (electronic supplementary Fig. S1). EGFR (35.0%) was the most frequent oncogenic mutation, followed by alteration in KRAS (25.0%), CDKN2A (15.0%), SMARCA4 (15.0%) and ERBB2 (10%).
3.6 Outcome of Patients with MET Dysregulation
Median overall survival (mOS) for patients harboring MET exon 14 skipping alterations was 64 weeks (21–336 weeks) compared with a mOS of 351 weeks (12–414 weeks) for patients with MET amplification (electronic supplementary Fig. S2). Fifteen patients underwent treatment with a MET TKI (11 patients received crizotinib, 1 patient received capmatinib, 2 patients received crizotinib first followed by capmatinib, and 1 patient received cabozantinib) (Fig. 3). Median progression-free survival (PFS) on MET TKI therapy was 9 weeks (4–23 weeks) and 21 weeks (18–43 weeks, excluding patient 26, with recent therapy start) for patients with MET exon 14 skipping alterations and MET amplifications, respectively. Patients 29–31 received MET TKI therapy based on high MET overexpression and below cut-off level MET exon 14 skipping reads. Three patients with PD-L1 expression in ≥ 50% of tumor cells and one patient with PD-L1 expression < 1% harboring MET alterations showed long-lasting responses to immunotherapy with immune checkpoint inhibitors, with a minimum PFS of 2 years since the start of immunotherapy. Between these, in patient 9, with long-lasting response to immunotherapy, a PD-L1 amplification was detected by NGS.
Swimmer’s plot showing the course of systemic treatment, length of time on specific therapy, and response in individual patients. Three patients (patients 5, 13, and 20) with systemic treatment of < 1 month, or no systemic treatment, are not depicted. The treatment course for patient 29 is depicted from 2016, when stage IV disease progressed after the initial diagnosis in 2012. MET mesenchymal-to-epithelial transition, IHC immunohistochemistry
4 Discussion
Several MET-directed TKIs have recently emerged as treatment options for patients with oncogenic genomic alterations in the MET gene. Thus, it is crucial to analyze the genomic and clinical characteristics of patients harboring actionable MET alterations. In this study, we show that 25% of MET exon 14-altered cases and 33% of MET-amplified cases harbor potentially targetable oncogenic co-mutations. In addition, MET exon 14-altered tumors are significantly more likely to be PD-L1-positive. Of note, we observed long-lasting responses to immunotherapy in several patients with MET dysregulation, including a case of a durable response to an immune checkpoint inhibitor in a patient with MET overexpression and low-level MET exon 14 skipping reads.
To our knowledge, this is the first study that compares genomic profiles and PD-L1 expression in different subtypes of MET-dysregulated advanced-stage NSCLC. Studies on PD-L1 expression patterns in MET-dysregulated tumors are rare [41, 42, 44, 45]. In vivo studies indicated that downstream signaling of MET via AKT/GSK3β upregulates PD-L1 expression [44, 46]. In addition, MET activation also induces the upregulation of other immune suppressive genes such as PDCD1LG2 (PD-L2) and SOCS1 [44]. MET overexpression is also significantly associated with PD-L1 status in stage I–III lung adenocarcinoma with no reports on MET mutations and PD-L1 expression [41]. In this study, we observed a significant association between PD-L1 expression in MET exon 14-mutated tumors (80.0%) compared with MET wtIHCpositive tumors (50.0%). In contrast to other studies, we detected fewer cases co-expressing PD-L1 and MET in patients without MET alterations [41]. Therefore, our data support that MET dysregulation is associated with PD-L1 expression, which was also described by previous studies where PD-L1 expression in 63% of tumors with MET exon 14 mutations was detected [42].
Four NSCLC patients in our cohort with MET exon 14 alterations and MET amplification had long-lasting response to immunotherapy with immune checkpoint inhibitors administered as second- or further-line treatment. Such therapy has shown to prolong PFS in KRAS mutant tumors and tumors with high tumor mutational burden (TMB), but not in patients with tumors harboring unique oncogenic alterations such as EGFR [47,48,49,50,51]. Partial or complete responses of more than 18 months were seen in six patients (46.2%) with MET exon 14 skipping mutations but without concurrent oncogenic mutations or MET amplification [52]. Such responses might as well have occurred through the presence of several co-mutations [53]. In contrast, 36 patients from the IMMUNOTARGET registry with MET dysregulation (23 with exon 14 mutation, 13 with amplification) had an objective response rate to immunotherapy of 16%, probably reflecting a population of patients undergoing immunotherapy at later lines [54]. In another study, MET amplification was not associated with greater benefit from nivolumab treatment [55, 56]. Studies on the co-mutational profiles of MET-altered tumors are inconclusive. However, several studies, including one large NGS-based study [11], reported that MET exon 14 skipping alterations are mutually exclusive with other oncogenic mutations, except for MET amplification and MDM2 amplification [11, 12, 40]. In MET-amplified tumors, a negative correlation between the level of amplification and oncogenic co-mutations was shown with high-level MET-amplified tumors (MET/CEP7 ≥ 5) harboring concomitant drivers in 41% (11/27) of cases, and low-level MET-amplified tumors in 62% (32/52) of cases [57]. In this study, we demonstrate, in a subset of advanced-stage NSCLC, that MET amplification as well as MET exon 14 skipping alterations are not mutually exclusive events to KRAS, BRAF, and other oncogenic driver mutations. Twenty-five percent of MET exon 14-altered cases and 33% of MET-amplified cases harbored potentially targetable oncogenic co-mutations. In KRAS- and EGFR-mutated lung adenocarcinomas, MET amplification and mutations were detected [58]. In patients with MET exon 14-altered lung cancer concurrent MDM2 (35%), CDK4 (21%) and EGFR amplifications (6.4%) were the most frequent concurrent genetic alterations [13]. Of interest, we detected that two MET-amplified tumors displayed amplification of the HGF gene, the ligand for the MET tyrosine kinase receptor, and this finding warrants further investigation.
MET TKIs improved the survival of patients harboring MET exon 14 skipping mutations as well as MET amplifications [34]; however, resistance to MET TKIs is observed in a broad number of patients [59]. Patients with MET exon 14-mutated tumors showed a response rate of 32% to the TKI crizotinib, 46% to tepotinib, and 68% to capmatinib [30, 31, 39] with unknown resistance mechanisms. In the course of treatment, many patients further develop acquired resistance to treatment with TKIs and therefore response to TKIs is usually transient [59, 60]. Co-occurring genomic alterations to MET alterations might explain resistance mechanisms. To this end, co-mutations in KRAS might reduce treatment efficacy with MET TKIs, since activation of the RAS pathway was reported to be associated with poorer outcomes in patients with MET exon 14 skipping alterations [38, 61]. However, one of the four patients with KRAS co-mutation and amplification (Fig. 2a, patient 8) received crizotinib as second-line therapy, reaching 18 weeks of PFS compared with a median PFS of 9 weeks in the other patients in our cohort with MET exon 14-mutated tumors.
Previous studies have shown that MET IHC is an inefficient screening tool for genomic changes in MET [16, 20,21,22,23]. In our study, 7% of tumors with genomic alterations in MET were MET IHC-negative, all either squamous cell or pleomorphic carcinomas. However, in the subset of genomic MET-altered adenocarcinomas, all MET exon 14-altered or -amplified tumors were also positive by IHC. These results suggest that MET IHC screening could be considered for adenocarcinomas in resource-limited settings, where upfront NGS-based molecular testing is not readily available. However, all positive cases require an NGS-based confirmation, since only 29.9% of MET IHC-positive cases in our study harbored a MET genomic alteration.
Currently, molecular analysis does not include testing for MET alterations as part of routine. Our study also underlines the importance of such analysis in the presence of other alterations. We believe that over the next years, when NGS becomes part of routine testing for patients with NSCLC, better knowledge of such co-alterations and clinical outcome will allow to improve treatment for these patients and possibly understanding of resistance mechanisms to MET-targeted therapies.
This study has several limitations. It was a single-center, retrospective study and does not present universal MET NGS, MET FISH, and PD-L1 assessment. In the group of MET IHC-negative tumors, NGS data were available for a minority of patients, and consequently, characterization of those tumors and comparison with MET overexpressing could not be reported. For the purpose of this study, we analyzed all tumor specimens for c-MET in order to understand molecular features of tumors exhibiting MET overexpression. Therefore, we have enriched our cohort with MET IHC-positive cases (50%). In the literature, MET overexpression was detected by IHC in 22–24% of NSCLC tumors [10, 16, 20, 21]. Additionally, IHC was not confirmed by another experienced pathologist and PD-L1 results were obtained by EIL3N assay, which is not the most used assay worldwide and other assays were shown to be superior due to staining intensity, scoring range, and pathologist preference [62]. By enriching our cohort with MET IHC-positive cases, we also report higher prevalence of MET alterations (14.7% with MET exon 14 skipping, 10.1% with MET amplification). In the literature, mutations within the juxtamembrane domain of MET encoded by exon 14 were reported in 3–4% of NSCLCs [10,11,12] and high-level MET amplification in 3–5% of lung adenocarcinoma [16,17,18,19].
Moreover, this was a single-center study with a limited number of patients. Due to the small sample size of patients with MET exon 14 alterations and MET amplification, it is difficult to draw conclusions by comparing the molecular profiles of the four patients who received immunotherapy and responded to this treatment, with patients with a similar molecular profile not receiving immunotherapy. Finally, the latest MET TKIs capmatinib and tepotinib have been demonstrated to be superior to crizotinib in patients with MET exon 14 alterations, and only one patient in our cohort received capmatinib, therefore we cannot reach any conclusions to relate response to such targeted treatments and molecular features.
5 Conclusion
We were able to show the presence of oncogenic co-mutations in tumors with MET exon 14 skipping alterations and MET amplification, and described the association of MET exon 14 skipping alterations with PD-L1 expression.
References
Bottaro D, Rubin J, Faletto D, Chan A, Kmiecik T, Vande Woude G, et al. Identification of the hepatocyte growth factor receptor as the c-met proto-oncogene product. Science. 1991;251(4995):802–4.
Gherardi E, Youles ME, Miguel RN, Blundell TL, Iamele L, Gough J, et al. Functional map and domain structure of MET, the product of the c-met protooncogene and receptor for hepatocyte growth factor/scatter factor. Proc Natl Acad Sci USA. 2003;100(21):12039–44.
Weidner KM, Behrens J, Vandekerckhove J, Birchmeier W. Scatter factor: molecular characteristics and effect on the invasiveness of epithelial cells. J Cell Biol. 1990;111(5):2097–108.
Birchmeier C, Birchmeier W, Gherardi E, Vande Woude GF. Met, metastasis, motility and more. Nat Rev Mol Cell Biol. 2003;4(12):915–25.
Trusolino L, Bertotti A, Comoglio PM. MET signalling: principles and functions in development, organ regeneration and cancer. Nat Rev Mol Cell Biol. 2010;11(12):834–48.
Gherardi E, Birchmeier W, Birchmeier C, Woude GV. Targeting MET in cancer: rationale and progress. Nat Rev Cancer. 2012;12(2):89–103.
Deng N, Goh LK, Wang H, Das K, Tao J, Tan IB, et al. A comprehensive survey of genomic alterations in gastric cancer reveals systematic patterns of molecular exclusivity and co-occurrence among distinct therapeutic targets. Gut. 2012;61(5):673–84.
Pal SK, Ali SM, Yakirevich E, Geynisman DM, Karam JA, Elvin JA, et al. Characterization of clinical cases of advanced papillary renal cell carcinoma via comprehensive genomic profiling. Eur Urol. 2018;73(1):71–8.
Engelman JA, Zejnullahu K, Mitsudomi T, Song Y, Hyland C, Park JO, et al. MET amplification leads to gefitinib resistance in lung cancer by activating ERBB3 signaling. Science. 2007;316(5827):1039–43.
Awad MM, Oxnard GR, Jackman DM, Savukoski DO, Hall D, Shivdasani P, et al. MET exon 14 mutations in non–small-cell lung cancer are associated with advanced age and stage-dependent MET genomic amplification and c-Met overexpression. J Clin Oncol. 2016;34(7):721–30.
Frampton GM, Ali SM, Rosenzweig M, Chmielecki J, Lu X, Bauer TM, et al. Activation of MET via diverse exon 14 splicing alterations occurs in multiple tumor types and confers clinical sensitivity to MET inhibitors. Cancer Discov. 2015;5(8):850–9.
Tong JH, Yeung SF, Chan AWH, Chung LY, Chau SL, Lung RWM, et al. MET amplification and exon 14 splice site mutation define unique molecular subgroups of non-small cell lung carcinoma with poor prognosis. Clin Cancer Res. 2016;22(12):3048–56.
Schrock AB, Frampton GM, Suh J, Chalmers ZR, Rosenzweig M, Erlich RL, et al. Characterization of 298 patients with lung cancer harboring MET exon 14 skipping alterations. J Thorac Oncol. 2016;11(9):1493–502.
Davies KD, Lomboy A, Lawrence CA, Yourshaw M, Bocsi GT, Camidge DR, et al. DNA-based versus RNA-based detection of MET exon 14 skipping events in lung cancer. J Thorac Oncol. 2019;14(4):737–41.
Drilon A, Cappuzzo F, Ou SHI, Camidge DR. Targeting MET in lung cancer: will expectations finally be MET? J Thorac Oncol. 2017;12(1):15–26.
Bubendorf L, Dafni U, Schöbel M, Finn SP, Tischler V, Sejda A, et al. Prevalence and clinical association of MET gene overexpression and amplification in patients with NSCLC: Results from the European Thoracic Oncology Platform (ETOP) Lungscape project. Lung Cancer. 2017;111:143–9.
Cappuzzo F, Marchetti A, Skokan M, Rossi E, Gajapathy S, Felicioni L, et al. Increased MET gene copy number negatively affects survival of surgically resected non-small-cell lung cancer patients. J Clin Oncol. 2009;27(10):1667–74.
Schildhaus HU, Schultheis AM, Rüschoff J, Binot E, Merkelbach-Bruse S, Fassunke J, et al. MET amplification status in therapy-naïve adeno- and squamous cell carcinomas of the lung. Clin Cancer Res. 2015;21(4):907–15.
Go H, Jeon YK, Park HJ, Sung SW, Seo JW, Chung DH. High MET gene copy number leads to shorter survival in patients with non-small cell lung cancer. J Thorac Oncol. 2010;5(3):305–13.
Watermann I, Schmitt B, Stellmacher F, Müller J, Gaber R, Kugler C, et al. Improved diagnostics targeting c-MET in non-small cell lung cancer: expression, amplification and activation? Diagn Pathol. 2015;10:130.
Guo R, Berry LD, Aisner DL, Sheren J, Boyle T, Bunn PA, et al. MET IHC is a poor screen for MET amplification or MET exon 14 mutations in lung adenocarcinomas: data from a tri-institutional cohort of the lung cancer mutation consortium. J Thorac Oncol. 2019;14(9):1666–71.
Tsuta K, Kozu Y, Mimae T, Yoshida A, Kohno T, Sekine I, et al. c-MET/phospho-MET protein expression and MET gene copy number in non-small cell lung carcinomas. J Thorac Oncol. 2012;7(2):331–9.
Baldacci S, Figeac M, Antoine M, Descarpentries C, Kherrouche Z, Jamme P, et al. High MET overexpression does not predict the presence of MET exon 14 splice mutations in NSCLC: results from the IFCT PREDICT.amm study. J Thorac Oncol. 2020;15(1):120–4.
Schuler MH, Berardi R, Lim WT, Geel RV, De Jonge MJ, Bauer TM, et al. Phase (Ph) I study of the safety and efficacy of the cMET inhibitor capmatinib (INC280) in patients (pts) with advanced cMET+ non-small cell lung cancer (NSCLC). J Clin Oncol. 2016;34(15 Suppl):9067.
Guo R, Luo J, Chang J, Rekhtman N, Arcila M, Drilon A. MET-dependent solid tumours—molecular diagnosis and targeted therapy. Nat Rev Clin Oncol. 2020;17(9):569–87.
Spigel DR, Ervin TJ, Ramlau RA, Daniel DB, Goldschmidt JH, Blumenschein GR, et al. Randomized phase II trial of Onartuzumab in combination with erlotinib in patients with advanced non-small-cell lung cancer. J Clin Oncol. 2013;31(32):4105–14.
Rimassa L, Assenat E, Peck-Radosavljevic M, Pracht M, Zagonel V, Mathurin P, et al. Tivantinib for second-line treatment of MET-high, advanced hepatocellular carcinoma (METIV-HCC): a final analysis of a phase 3, randomised, placebo-controlled study. Lancet Oncol. 2018;19(5):682–93.
Drilon AE, Camidge DR, Ou SHI, Clark JW, Socinski MA, Weiss J, et al. Efficacy and safety of crizotinib in patients (pts) with advanced MET exon 14-altered non-small cell lung cancer (NSCLC). J Clin Oncol. 2016;34(15 Suppl):108.
Camidge DR, Ou SHI, Shapiro G, Otterson GA, Villaruz LC, Villalona-Calero MA, et al. Efficacy and safety of crizotinib in patients with advanced c-MET-amplified non-small cell lung cancer (NSCLC). J Clin Oncol. 2014;32(15 Suppl):8001.
Paik PK, Felip E, Veillon R, Sakai H, Cortot AB, Garassino MC, et al. Tepotinib in non–small-cell lung cancer with MET exon 14 skipping mutations. N Engl J Med. 2020;383(10):931–43.
Wolf J, Seto T, Han JY, Reguart N, Garon EB, Groen HJM, et al. Capmatinib in MET exon 14-mutated or MET-amplified non-small-cell lung cancer. N Engl J Med. 2020;383(10):944–57.
Camidge DR, Otterson GA, Clark JW, Ignatius Ou SH, Weiss J, Ades S, et al. Crizotinib in patients with MET-amplified NSCLC. J Thorac Oncol. 2021;16(6):1017–29.
Lamberti G, Andrini E, Sisi M, Rizzo A, Parisi C, Di Federico A, et al. Beyond EGFR, ALK and ROS1: current evidence and future perspectives on newly targetable oncogenic drivers in lung adenocarcinoma. Crit Rev Oncol Hematol. 2020;156: 103119.
Friedlaender A, Drilon A, Banna GL, Peters S, Addeo A. The METeoric rise of MET in lung cancer. Cancer. 2020;126(22):4826–37.
Bahcall M, Sim T, Paweletz CP, Patel JD, Alden RS, Kuang Y, et al. Acquired METD1228V mutation and resistance to MET inhibition in lung cancer. Cancer Discov. 2016;6(12):1334–41.
Dong HJ, Li P, Wu CL, Zhou XY, Lu HJ, Zhou T. Response and acquired resistance to crizotinib in Chinese patients with lung adenocarcinomas harboring MET Exon 14 splicing alternations. Lung Cancer. 2016;102:118–21.
Reis H, Metzenmacher M, Goetz M, Savvidou N, Darwiche K, Aigner C, et al. MET expression in advanced non-small-cell lung cancer: effect on clinical outcomes of chemotherapy, targeted therapy, and immunotherapy. Clin Lung Cancer. 2018;19(4):e441–63.
Rotow JK, Gui P, Wu W, Raymond VM, Lanman RB, Kaye FJ, et al. Co-occurring alterations in the RAS–MAPK pathway limit response to MET inhibitor treatment in MET exon 14 skipping mutation-positive lung cancer. Clin Cancer Res. 2020;26(2):439–49.
Drilon A, Clark JW, Weiss J, Ou SHI, Camidge DR, Solomon BJ, et al. Antitumor activity of crizotinib in lung cancers harboring a MET exon 14 alteration. Nat Med. 2020;26(1):47–51.
Onozato R, Kosaka T, Kuwano H, Sekido Y, Yatabe Y, Mitsudomi T. Activation of MET by gene amplification or by splice mutations deleting the juxtamembrane domain in primary resected lung cancers. J Thorac Oncol. 2009;4(1):5–11.
Kerr KM, Thunnissen E, Dafni U, Finn SP, Bubendorf L, Soltermann A, et al. A retrospective cohort study of PD-L1 prevalence, molecular associations and clinical outcomes in patients with NSCLC: Results from the European Thoracic Oncology Platform (ETOP) Lungscape Project. Lung Cancer. 2019;131:95–103.
Sabari JK, Leonardi GC, Shu CA, Umeton R, Montecalvo J, Ni A, et al. PD-L1 expression, tumor mutational burden, and response to immunotherapy in patients with MET exon 14 altered lung cancers. Ann Oncol. 2018;29(10):2085–91.
Lacour M, Hiltbrunner S, Lee SY, Soltermann A, Rushing EJ, Soldini D, et al. Adjuvant chemotherapy increases programmed death-ligand 1 (PD-L1) Expression in non-small cell lung cancer recurrence. Clin Lung Cancer. 2019;20(5):391–6.
Saigi M, Alburquerque-Bejar JJ, Mc Leer-Florin A, Pereira C, Pros E, Romero OA, et al. MET-oncogenic and JAK2-inactivating alterations are independent factors that affect regulation of PD-L1 expression in lung cancer. Clin Cancer Res. 2018;24(18):4579–87.
Yoshimura K, Inoue Y, Tsuchiya K, Karayama M, Yamada H, Iwashita Y, et al. Elucidation of the relationships of MET protein expression and gene copy number status with PD-L1 expression and the immune microenvironment in non-small cell lung cancer. Lung Cancer. 2020;141:21–31.
Ahn HK, Kim S, Kwon D, Koh J, Kim YA, Kim K, et al. MET receptor tyrosine kinase regulates the expression of co-stimulatory and co-inhibitory molecules in tumor cells and contributes to PD-L1-mediated suppression of immune cell function. Int J Mol Sci. 2019;20(17):4287.
Rizvi NA, Hellmann MD, Snyder A, Kvistborg P, Makarov V, Havel JJ, et al. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science. 2015;348(6230):124–8.
Lee CK, Man J, Lord S, Links M, Gebski V, Mok T, et al. Checkpoint inhibitors in metastatic EGFR-mutated non-small cell lung cancer: a meta-analysis. J Thorac Oncol. 2017;12(2):403–7.
Gainor JF, Shaw AT, Sequist LV, Fu X, Azzoli CG, Piotrowska Z, et al. EGFR mutations and ALK rearrangements are associated with low response rates to PD-1 pathway blockade in non-small cell lung cancer: a retrospective analysis. Clin Cancer Res. 2016;22(18):4585–93.
Lee CK, Man J, Lord S, Cooper W, Links M, Gebski V, et al. Clinical and molecular characteristics associated with survival among patients treated with checkpoint inhibitors for advanced non-small cell lung carcinoma: a systematic review and meta-analysis. JAMA Oncol. 2018;4(2):210.
Inoue Y, Yoshimura K, Mori K, Kurabe N, Kahyo T, Mori H, et al. Clinical significance of PD-L1 and PD-L2 copy number gains in non-small-cell lung cancer. Oncotarget. 2016;7(22):32113–28.
Mayenga M, Assié JB, Monnet I, Massiani MA, Tabeze L, Friard S, et al. Durable responses to immunotherapy of non-small cell lung cancers harboring MET exon-14-skipping mutation: a series of 6 cases. Lung Cancer. 2020;150:21–5.
Yoshimura K, Karayama M, Inoue Y, Kahyo T, Inui N, Maekawa M, et al. Heterogeneous MET gene copy number and EGFR mutation elicit discordant responses to crizotinib between primary and metastatic lesions in erlotinib-resistant lung adenocarcinoma. Lung Cancer. 2018;124:317–9.
Mazieres J, Drilon A, Lusque A, Mhanna L, Cortot AB, Mezquita L, et al. Immune checkpoint inhibitors for patients with advanced lung cancer and oncogenic driver alterations: results from the IMMUNOTARGET registry. Ann Oncol. 2019;30(8):1321–8.
Yoshimura K, Inoue Y, Inui N, Karayama M, Yasui H, Hozumi H, et al. MET amplification and efficacy of nivolumab in patients with NSCLC. JTO Clin Res Rep. 2021;2(11): 100239.
Inoue Y, Yoshimura K, Nishimoto K, Inui N, Karayama M, Yasui H, et al. Evaluation of programmed death ligand 1 (PD-L1) gene amplification and response to nivolumab monotherapy in non-small cell lung cancer. JAMA Netw Open. 2020;3(9): e2011818.
Noonan SA, Berry L, Lu X, Gao D, Barón AE, Chesnut P, et al. Identifying the appropriate FISH criteria for defining MET copy number-driven lung adenocarcinoma through oncogene overlap analysis. J Thorac Oncol. 2016;11(8):1293–304.
Scheffler M, Ihle MA, Hein R, Merkelbach-Bruse S, Scheel AH, Siemanowski J, et al. K-ras mutation subtypes in NSCLC and associated co-occuring mutations in other oncogenic pathways. J Thorac Oncol. 2019;14(4):606–16.
Recondo G, Bahcall M, Spurr LF, Che J, Ricciuti B, Leonardi GC, et al. Molecular mechanisms of acquired resistance to MET tyrosine kinase inhibitors in patients with MET exon 14-mutant NSCLC. Clin Cancer Res. 2020;26(11):2615–25.
Awad MM, Leonardi GC, Kravets S, Dahlberg SE, Drilon A, Noonan SA, et al. Impact of MET inhibitors on survival among patients with non-small cell lung cancer harboring MET exon 14 mutations: a retrospective analysis. Lung Cancer. 2019;133:96–102.
Suzawa K, Offin M, Lu D, Kurzatkowski C, Vojnic M, Smith RS, et al. Activation of KRAS mediates resistance to targeted therapy in MET Exon 14-mutant non-small cell lung cancer. Clin Cancer Res. 2019;25(4):1248–60.
Smith J, Robida MD, Acosta K, Vennapusa B, Mistry A, Martin G, et al. Quantitative and qualitative characterization of two PD-L1 clones: SP263 and E1L3N. Diagn Pathol. 2016;11(1):44.
Funding
Open access funding provided by University of Zurich.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This study was supported by the ‘Stiftung für angewandte Krebsforschung’ Zürich.
Conflicts of interest
Alessa Fischer, Lorenz Bankel, Stefanie Hiltbrunner, Markus Rechsteiner, Jan Rüschoff, Elisabeth Rushing, and Christian Britschgi declare they have no conflicts of interest that might be relevant to the contents of this manuscript. Alessandra Curioni-Fontecedro reports honoraria for lectures and advisory fees from Astra Zeneca, BMS, Boehringer Ingelheim, MSD, Pfizer, Roche, and Takeda.
Ethics approval
This study was conducted according to the law and regulations of the local Ethics Commission under reference number KEK ZH-2021-00381. All patients included in the study consented to their participation.
Availability of data and material
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
Code availability
Not applicable.
Authors' contributions
Conceptualization: AF, LB, SH, ACF. Methodology: AF, ACF, MR. Investigation: AF, JR. Data curation: AF, MR. Validation: LB, ACF. Formal analysis: AF. Visualization: AF, SH. Resources: SH, MR, ER, ACF. Writing: AF, ACF. Review and editing: LB, SH, MR, JR, ER, CB, ACF. Supervision: ACF. Funding acquisition: SH, ACF.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Fischer, A., Bankel, L., Hiltbrunner, S. et al. Mutational Landscape and Expression of PD-L1 in Patients with Non-Small Cell Lung Cancer Harboring Genomic Alterations of the MET gene. Targ Oncol 17, 683–694 (2022). https://doi.org/10.1007/s11523-022-00918-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11523-022-00918-6