Abstract
We present a new approach for relating nucleic-acid content to fluorescence in a real-time Polymerase Chain Reaction (PCR) assay. By coupling a two-type branching process for PCR with a fluorescence analog of Beer’s Law, the approach reduces bias and quantifies uncertainty in fluorescence. As the two-type branching process distinguishes between complementary strands of DNA, it allows for a stoichiometric description of reactions between fluorescent probes and DNA and can capture the initial conditions encountered in assays targeting RNA. Analysis of the expected copy-number identifies additional dynamics that occur at short times (or, equivalently, low cycle numbers), while investigation of the variance reveals the contributions from liquid volume transfer, imperfect amplification, and strand-specific amplification (i.e., if one strand is synthesized more efficiently than its complement). Linking the branching process to fluorescence by the Beer’s Law analog allows for an a priori description of background fluorescence. It also enables uncertainty quantification (UQ) in fluorescence which, in turn, leads to analytical relationships between amplification efficiency (probability) and limit of detection. This work sets the stage for UQ-PCR, where both the input copy-number and its uncertainty are quantified from fluorescence kinetics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Polymerase Chain Reaction (PCR) is a hallmark of molecular biology and applied genetics. When the dynamics of PCR are monitored by a fluorescent probe, the initial amount of target sequence can be quantified (qPCR) by a computational algorithm equipped with a mathematical model and a set of control experiments (Ruijter et al. 2009; Lievens et al. 2012; Zhao and Fernald 2005; Peirson et al. 2003; Tichopad et al. 2003; Boggy and Woolf 2010; Guescini et al. 2008; Ruijter et al. 2013). Quantification by PCR is routinely exploited in many applications, including analysis of forensic evidence (Nicklas and Buel 2003; Bauer 2007), monitoring of food safety (Elizaquível et al. 2014), and clinical diagnostics (Kaltenboeck and Wang 2005; Bustin et al. 2021).
The accuracy and precision of the quantification process is limited by the mathematical model relating DNA content to fluorescence. Current models possess subjective and systematic bias and do not account for the uncertainty in fluorescence that arises from imperfect amplification and pipetting errors.
Systematic bias originates from assuming that the initial DNA type is double-stranded and that the fluorescence increases each time either complementary strand is replicated. The former is obviously not true when the initial DNA is produced by reverse-transcription of single-stranded RNA (RT, as in RT-qPCR), as only one of the complementary DNA strands is present at the beginning of PCR. The second statement is not true for common probes that possess a fluorophore covalently attached to an oligonucleotide. Since the oligonucleotide only hybridizes to one of the complementary strands, the fluorescence only increases when one of the complementary strands is replicated. The second assumption also does not appear to be true for fluorescent dyes that bind non-specifically to DNA, as the amount of dye bound to each DNA strand depends on the amount of DNA in solution.
The impact of several of these assumptions was assessed by Ruijter et al. (2014). The authors found that, depending on the initial DNA type and fluorescent probe chemistry, the background-subtracted fluorescence could differ by up to a factor of 2 in the exponential phase. However, the authors’ analysis was rooted in the assumption of perfect amplification. They also noted that what actually occurs during the first few cycles of PCR is unknown.
Another source of systematic bias in the mathematical description of fluorescence arises when the initial DNA content is very small. Current approaches are deterministic and do not take into account the fact that the number of DNA strands is an integer. While the kinetics of PCR have been investigated in the framework of stochastic branching processes (Nedelman et al. 1992; Sun 1995; Weiss and von Haeseler 1995; Stolovitzky and Cecchi 1996; Jacob and Peccoud 1996b, a), the first of which was published in this journal, such models have not been linked to the fluorescence reported by probes. Like the deterministic approaches described above, these stochastic models neither discriminate between complementary strands nor describe initial conditions encountered in RT-qPCR.
A mathematical model that discriminates between complementary DNA strands can investigate another source of bias: the assumption that the efficiency of synthesis is independent of directionality (i.e., reverse or forward). Since primers are complementary to different ends of the target sequence, and are specifically chosen not to be complementary to each other (i.e., avoiding dimerization), the formation of one primer–target complex may be more efficient than the other. In addition, the yield of the strand whose replication is being monitored by the fluorescent probe may be affected by the monitoring process. These arguments are also supported by the fact that optimal concentrations of each primer can be different (see, for example, Bustin (2004), where the two concentrations differ by a factor of 3).
To address these challenges, we present a two-type stochastic branching process model in Sect. 2.1 that differentiates between complementary strands and amplification probabilities. Analysis of the expected value in Sect. 2.2 identifies a new timescale that is prevalent during the first few cycles. This timescale explains some of the unknown behavior that occurs during the first few cycles of PCR, explaining some of the aforementioned unknown behavior. At short times, there is a lag in exponential growth where the ratio of expected strand counts changes from its initial to critical value. The critical ratio is related to the amplification probability of each complementary strand, being unity when the probabilities are identical. The analysis also demonstrates that the popular parameter describing PCR efficiency (or amplification probability) is really the geometric mean of the efficiencies of both complementary strands.
Quantification by real-time PCR is also limited by a subjective and empirical description of the fluorescence that is not associated with amplification, or the background fluorescence. The description of background fluorescence is usually taken a posteriori (i.e., after measurements of fluorescence monitoring amplification). Without a clear connection to the chemical and physical processes occurring in solution, the background fluorescence is often assumed to be a linear function of cycle, or a ‘baseline.’
In Sect. 3.1, we address these concerns by using the fluorescence analog of Beer’s Law to relate fluorescence to the concentration of each fluorescent species. While such expressions have often been used to describe the fluorescence of dyes interacting with known amounts of DNA (Biver et al. 2003, 2005), we are not aware of any adaptation to real-time PCR. We discriminate between the fluorescent species by referring to the form present before PCR as the inactive species and the form activated by PCR as the active species. For hydrolysis probes, we show how the relevant parameters can be extracted from a few control experiments (see Sect. 3.2). In contrast to other approaches, we can quantify the validity of the model of background fluorescence. The relevant contributions can be determined without interrogating or adjusting fluorescence data associated with amplification. We find that the model agrees well with experiment and observe that the incremental increase in fluorescence is not independent of cycle.
A final limitation of real-time PCR is the lack of a mathematical expression relating errors arising from pipetting and imperfect amplification to uncertainty in fluorescence. To quantify the variance in copy number (Peccoud and Jacob 1996), we investigate the stochastic branching process (Sect. 2.3) in a manner similar to previous reports analyzing error in high-throughput sequencing (Kebschull and Zador 2015; Schwabe and Falcke 2022). After validating that the fluorescence parameter (the fluorescence per mole) for each species is approximately constant for each cycle and well (Sect. 3.2), the models for PCR and fluorescence are combined (Sect. 3.3). This yields analytical expressions of the first two central moments of fluorescence in terms of reaction efficiencies, input content, and input type (i.e., double-stranded DNA, forward-stranded RNA, and reverse-stranded RNA).
Together with the parameters determined from experiment, fluorescence curves computed with uncertainty identify regimes under which certain sources of error are more prevalent than others (see Sect. 3.3). When the expected initial-strand-number is sufficiently large, or the cycle number sufficiently small, the error in fluorescence in a specific well is less than the well-to-well variation in expected value. As the initial strand count decreases and the fluorescence rises above initial levels, however, the variance in input copy-number and imperfect amplification become the dominant contributions to error. Finally, in Sect. 3.4, we use the fluorescence model to develop analytical expressions for the limit of detection as a function of amplification efficiency and nucleic acid type. These expressions may be particularly useful for application in epidemic diseases, as false positives or false negatives may be instead termed inconclusive.
2 Strand-Specific Branching Process
In this section, we model PCR as a two-type branching-process. The model distinguishes between complementary DNA strands and amplification efficiencies. We then derive analytical expressions relating the first two central-moments of strand counts before PCR to those after each cycle has completed. Compartmentalizing DNA amplification and fluorescence, the linking of the two phenomena is postponed until Sect. 3.
Illustration of strand-specific model of DNA amplification. (top) After each cycle i, the outcome of synthesis of a forward strand (orange, solid) from each reverse strand (blue, dashed) present in the previous cycle is modeled as a Bernoulli random-variable with probability of success \(p_\text {rf}\). (bottom) Similarly, the outcome of synthesis of a reverse strand (blue, dashed) from each forward strand (orange, solid) present in the previous cycle is modeled as a Bernoulli random-variable with probability of success \(p_\text {fr}\) (Color figure online)
2.1 Mathematical Model
PCR consists of a series of n cycles, with n usually ranging between 35 and 50. Each cycle consists of a melting, annealing, and elongation step to synthesize new DNA from existing DNA (i.e., a chain reaction). A variety of resources on PCR are available online for further information (e.g., National Institutes of Health, National Human Genome Research Institute 2023).
To distinguish between the two complementary strands of DNA, we refer to one as the forward strand and the other as the reverse strand. We let the discrete random variables \(X_i\) and \(Y_i\) represent the number of forward and reverse strands, respectively, present after \(i=0\) to n cycles have been completed. We represent the strand counts after completion of i cycles as the random vector
and refer to \(\varvec{U}_0\) as the initial strand count. However, we distinguish \(\varvec{U}_0\) from the strand count input to the reaction mixture, denoting the latter by the random vector \(\displaystyle \begin{pmatrix} I_X \\ I_Y \end{pmatrix} \), where \(I_X\) and \(I_Y\) are discrete random variables representing the number of forward and reverse strands input, respectively. This is important to distinguish between assays targeting DNA and RNA sequences, as discussed in Sect. 2.1.2.
2.1.1 Relationship Between Consecutive Cycles
After completing \(i-1\) cycles, the biochemical events occurring during the next cycle involve the attempt to produce one forward strand from each of the \(Y_{i-1}\) reverse strands and the attempt to produce one reverse strand from each of the \(X_{i-1}\) forward strands (see Fig. 1). The outcome of synthesis of a forward strand from each reverse strand is modeled as a Bernoulli random-variable with probability of success \(p_\text {rf}\in (0, 1)\). Similarly, the outcome of synthesis of a reverse strand from each forward strand is modeled as a Bernoulli random-variable with probability of success \(p_\text {fr}\in (0, 1)\). (The subscripts rf and fr denote the direction reverse-to-forward and forward-to-reverse, respectively.) This corresponds to the mathematical model
after completing i cycles, where
and \(\textsf{B}\left( a; b\right) \) denotes a Binomial random-variable of a trials with probability of success b. All Bernoulli trials, whether associated with the outcome of synthesis of a forward or reverse strand, are taken to be independent.
To compare to previous approaches that do not discriminate between complementary strands and efficiencies, we will use
to denote the total number of strands after i cycles have been completed. We will see that an appropriate characterization of the average amplification efficiency of both complementary strands is
and that an appropriate parameter for the deviation in efficiencies is
To avoid changing notation, we will subsequently only investigate \(\displaystyle p_\text {fr}= \frac{{\bar{p}}}{R}\) and \(p_\text {rf}= {\bar{p}}R\) in terms of \({\bar{p}}\) and R, so that
2.1.2 Relationship Between Initial and Input Condition
The relationship between the input number \(\displaystyle \begin{pmatrix}I_X \\ I_Y \end{pmatrix}\) and initial number \(\varvec{U}_0\) of strands depends on whether the nucleic acids input to the reaction mixture are forward-stranded RNA (fs-RNA, referred to as Case RF), reverse-stranded RNA (rs-RNA, referred to as Case RR), or double-stranded DNA (ds-DNA, which consists of both fs-DNA and rs-DNA, referred to as Case D). If ds-DNA is generated by transferring rs-DNA and fs-DNA separately into the reaction mixture, \(X_0\) and \(Y_0\) can be modeled as independent and identically distributed (i.i.d.). Here,
When RNA is input to the reaction mixture, on the other hand, the nucleic acids all possess the same strandedness (i.e., they are all fs-RNA or rs-RNA). The RT step yields DNA strands that are complementary to the RNA. As all RNA strands are fs-RNA (or rs-RNA), the RT step yields rs-DNA (or fs-DNA). Modeling the outcome of synthesis of each cDNA from each RNA strand as a Bernoulli random-variable with probability of success \(r\in (0, 1)\), \(\varvec{U}_0\) is related to \(\displaystyle \begin{pmatrix}I_X \\ I_Y\end{pmatrix}\) via either
or
In comparison to the conventional PCR amplification efficiency \(p:=p_\text {fr}=p_\text {rf}\) when \(R=1\), which is usually between 0.8 and 0.99, the RT efficiency r can adopt a relatively large range of values (Bustin et al. 2015; Schwaber et al. 2019).
The nucleic acids are input to the reaction mixture by transferring liquids from one container to another using a pipette. Since the process of transferring such liquids is independent of the type of nucleic acids (i.e., independent of whether they are fs-RNA, rs-RNA, fs-DNA, or rs-DNA), the number of strands of each type input to the reaction mixture obey the same distribution. We will let this distribution be obeyed by the discrete random variable I. As a result, it follows that
where i.d. denotes identically distributed.
2.2 Expected Value
In this section, we derive relationships between \(\mathbb {E}\left[ I\right] \) and expected copy-numbers after i cycles have been completed. After using the total law of expectation to obtain a relationship in expected copies between two sequential cycles, we use induction to generate the desired results. We first investigate the conventional branching process for PCR which results from assuming \(R=1\) and does not distinguish between fs-DNA and rs-DNA. Subsequently, we consider the two-type branching process (1) where R may not be 1 and fs-DNA is distinguished from rs-DNA.
2.2.1 Conventional Branching Process
The conventional branching process model for PCR occurs when \(R=1\), implying from (3) that \(p_\text {rf}=p_\text {fr}=:p\). In this case, the system (1) can be summed to yield
as has been investigated elsewhere (Nedelman et al. 1992; Sun 1995; Weiss and von Haeseler 1995; Peccoud and Jacob 1996; Jacob and Peccoud 1996b, a; Stolovitzky and Cecchi 1996). Using the law of total expectation and (4), one finds that
for any two consecutive cycles. From induction, it follows that (5) is identical to
as reported elsewhere (Nedelman et al. 1992; Sun 1995; Weiss and von Haeseler 1995; Peccoud and Jacob 1996; Jacob and Peccoud 1996b, a; Stolovitzky and Cecchi 1996).
2.2.2 Strand-Specific Branching Process
Below, we develop relationships between \(\mathbb {E}\left[ I\right] \) and each \(\mathbb {E}\left[ \varvec{U}_i\right] \) for the more general case of (1) where forward strands are distinguished from reverse strands and R may not be 1. The law of total expectation and (1) lead to the relation
where \(\varvec{A}\) is defined as
and the last step follows from induction. The relationship between \(\mathbb {E}\left[ \varvec{U}_0\right] \) in (7) depends on \(\mathbb {E}\left[ I\right] \) through
which follows from \(\mathbb {E}\left[ \varvec{U}_0\right] = \mathbb {E}\left[ \mathbb {E}\left[ \varvec{U}_0 \mid I_X, I_Y\right] \right] \) and the case-by-case relationships presented in Sect. 2.1.2.
We will see that the matrix \(\varvec{A}\) plays a central role in dynamics of the first two central-moments of strand counts. \(\varvec{A}\) has two distinct eigenvalues,
and can be decomposed as
where
and
If \(R=1\), the scale factor \(\displaystyle \frac{1}{\sqrt{2}}\) in the definition of \(\varvec{X}\) implies that
where \(\Vert \cdot \Vert \) denotes the Euclidean norm. If \(R\ne 1\), on the other hand, no scale factor can be chosen to satisfy (10).
Since \(\varvec{Z}:=\varvec{X}^{-1}\),
for \(i,j\in \{1,2\}\), and (after substitution of (9))
Substituting (12) into (7) leads to
where second expression aids in physical interpretation (below) of Case D, RF, and RR.
Where Equation (6) has \(1 + p\), Equation (13) has two values \(\lambda _1\) and \(\lambda _2\). In the case that \(R=1\), \(\lambda _1:=1 + {\bar{p}}=1+p\), as is present in (6). The eigenvalue \(\lambda _2\) does not have an analog in (6).
The quantities \(X_i + RY_i\) and \(X_i - RY_i\) in the second expression of (13) arise frequently in the investigation of the first two central-moments of \(\varvec{U}_i\). The quantity \(X_i + RY_i\) is referred to as the weighted sum of strand counts after i cycles have been completed, while \(X_i - RY_i\) is referred to as the weighted difference. Multiplying each side of the equation (13) by the row vector \(\displaystyle \begin{pmatrix}1 \\ \pm R\end{pmatrix}^\top \) demonstratesFootnote 1 that
As \(\lambda _1 > 1\), the expected weighted-sum exhibits exponential growth. As \(\lambda _2 < 1\), on the other hand, the expected weighted-difference exhibits exponential decay.
The term in (13) associated with \(\lambda _2\) is always present when the input is RNA (i.e., for Case RF or RR and any \({\bar{p}}\in (0, 1)\) and \(R\in (0, \infty )\)), as \(\mathbb {E}\left[ X_0\right] \ne R\mathbb {E}\left[ Y_0\right] \). However, it may not be present if the input is DNA, as the term vanishes (\(\mathbb {E}\left[ X_0-RY_0\right] = (1-R)\mathbb {E}\left[ I\right] \)) for Case D when \(R=1\).
When \(\mathbb {E}\left[ X_0\right] \ne R \mathbb {E}\left[ Y_0\right] \), the term involving \(\lambda _2^i\) is usually negligible after a few cycles, as \({\bar{p}}\) is usually above 0.8. After the lag time is over (i is large enough for \(\lambda _2^i\) to be negligible), the ratio of expected forward to reverse strand counts reaches its critical value, as
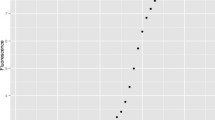
where \(\displaystyle \varvec{e}_1 = \begin{pmatrix}1 \\ 0 \end{pmatrix}\) and \(\displaystyle \varvec{e}_2=\begin{pmatrix}0 \\ 1\end{pmatrix}\) are the standard unit vectors. An illustration of the transition of \(\displaystyle \frac{\mathbb {E}\left[ X_i\right] }{\mathbb {E}\left[ Y_i\right] }\) (when \(\mathbb {E}\left[ Y_i\right] > 0\)) to R is depicted in Fig. 2.
Visualization of convergence of the ratio of expected-forward to expected-reverse strands to R. The initial condition \(\mathbb {E}\left[ \varvec{U}_0\right] \) is taken from (8) with Case D, RF, or RR. The values of \({\bar{p}}\) and R are annotated, while r and \(\mathbb {E}\left[ I\right] \) are chosen so that \(\mathbb {E}\left[ N_0\right] = \mathbb {E}\left[ X_0 + Y_0\right] = 10\)
To compare the two-type branching process to (6), we compute from (13)
and, for Case D,
as (8) implies that \(\mathbb {E}\left[ I\right] =\mathbb {E}\left[ X_0\right] =\mathbb {E}\left[ X_0 + Y_0\right] /2 = \mathbb {E}\left[ N_0\right] /2\). Equation (15) demonstrates that the two approaches are equal if \(R=1\). Otherwise, the absolute difference between \(\mathbb {E}\left[ N_i\right] \) calculated from (6) and (15) increases exponentially with increasing i. In the future, determining R and using (15) instead of (6) may therefore be important to reduce bias in quantification.
On the other hand, when \(R=1\), the expected amount of each strand is eventually independent of the composition at the start of PCR. That is,
only depends on the expected initial-sum, or \(\mathbb {E}\left[ N_0\right] \). This result is in contrast to the report of Ruijter et al. (2014), who used a deterministic model with perfect amplification efficiency. When \(R=1\) and the input is RNA, the directionality of RNA (i.e., fs-RNA or rs-RNA) only dictates the characteristic length of the lag time.
2.3 Variance
In this section, we derive the relationship between \(\textsf{Var}\left[ I\right] \) and the variance after i cycles have been completed. The procedure is similar to the previous section, except the law of total variance is used instead of the law of total expectation.
2.3.1 Conventional Branching Process
Using the law of total variance and (4), one obtains
Induction can be used to show that (17) is equivalent to
Substitution of (6), simplification of the resultant geometric series, and rearrangement leads to the expression
When \(\textsf{Var}\left[ N_0\right] = 0\), Equation (19) becomes identical to what has been reported elsewhere (Sun 1995; Weiss and von Haeseler 1995; Jacob and Peccoud 1996b, a; Stolovitzky and Cecchi 1996) (Nedelman et al. (1992); Peccoud and Jacob (1996) report the leading-order approximation for large i). The more general result, Equation (19), demonstrates that \(\textsf{Var}\left[ N_0\right] \) can significantly impact \(\textsf{Var}\left[ N_i\right] \), as it is part of the dominant term.
The growth in variance with increasing i by an exponent twice that of the expected value explains why very large cycles, where the expected copy-number is also very large, are not of interest. It also explains why quantification by real-time PCR is more reproducible than end-point PCR.
2.3.2 Strand-Specific Branching Process
The variance–covariance matrix of \(\varvec{U}_i\) is defined as
for each i. From the law of total variance,
In a manner similar to (7), it follows that
In addition, after substitution of (1), it follows that
Combining the two expressions, we obtain the two-type analog of (17),
Before using induction to relate \(\textsf{Var}\left[ \varvec{U}_i\right] \) to \(\textsf{Var}\left[ \varvec{U}_0\right] \), it is useful to simplify (20) by substituting \(\mathbb {E}\left[ Y_{i-1}\right] =\varvec{e}_2^\top \mathbb {E}\left[ \varvec{U}_{i-1}\right] \) and \(\mathbb {E}\left[ X_{i-1}\right] =\varvec{e}_1^\top \mathbb {E}\left[ \varvec{U}_{i-1}\right] \) with \(\mathbb {E}\left[ \varvec{U}_{i-1}\right] \) provided by the first expression of (13). Collecting terms multiplying each eigenvalue, we find that
where
From induction, it follows that (21) is identical to
Substitution of (12) and simplification of the resultant geometric series leads to (see Sect. B.1)
where
In (24), the dependence of \(\textsf{Var}\left[ \varvec{U}_i\right] \) on I arises through \(\textsf{Var}\left[ \varvec{U}_0\right] \) in each \(\nu _{j,k}\) and \(\mathbb {E}\left[ \varvec{U}_0\right] \) in each \(\eta _{j,k}^\ell \) (through (22)). While the dependence of \(\mathbb {E}\left[ \varvec{U}_0\right] \) on \(\mathbb {E}\left[ I\right] \) is given in (8), the relationship between \(\textsf{Var}\left[ \varvec{U}_0\right] \) and \(\textsf{Var}\left[ I\right] \) for Case D, RF, or RR is given by
where (26b) utilizes the law of total variance. Since \(X_0\) and \(Y_0\) are independent for any case, \(\textsf{Cov}\left[ X_0,Y_0\right] = 0\). As such, we will often use the substitution
without specifying a particular case.
In contrast to (19), which only has terms proportional to \((1 + p)^{2i}\) and \((1 + p)^{i-1}\), Equation (24) has terms proportional to \(\lambda _1^{2i}> \lambda _1^i> (\lambda _1\lambda _2)^i> \lambda _2^i > \lambda _2^{2i}\) (if \(i > 0\)). While physical interpretation of all terms can be complicated, it is useful to examine the leading-order expression as \(i\rightarrow \infty \),
and compute the terms explicitly as
where we have used (27). Note that the moments of \((X_0 \pm RY_0)/2\) arise in (28) as they did in (13). The term \(\nu _{1,1}\) represents the contribution from the variance at the start of PCR. The term \(\eta _{1,1}^{(1)}\) accounts for the variance due to imperfect amplification, as \({\bar{p}}\approx 1\) implies that \(\lambda _2 \approx 0 \approx \eta _{1,1}^{(1)}\). If \(R=1\),
one-half the second term in brackets of (19). The third term \(\eta _{1,1}^{(2)}\) does not have a counterpart in (19). It accounts for differences in strand-specific amplification, as it vanishes when \(R\rightarrow 1\).
The expressions developed for expected value and variance can be used to produce expressions in the squared coefficients of variation, defined as
where \(\oslash \) denotes element-wise division. Substitution of (13) and (28) into (30) yields
As the term \(O\left( \lambda _1^{-i}\right) \) rapidly approaches zero with increasing i, and \(\lambda _1\) is usually more than 1.8, the dominant term in (31) is a useful approximation. The dominant term is independent of i and is therefore a very practical tool for estimating the error present in PCR. As such, it is useful to express (31) as
for any \(j,k\in \{1, 2\}\), where (27) is used in the first step and the second step follows from simplification with (8) and (26). The quantities \(\alpha \) and \(\beta \), defined as
relate the reaction efficiencies to the coefficient of variation, as \(\alpha =\alpha \left( R\right) \) and \(\beta = \beta \left( R, {\bar{p}}, r\right) \). In Sect. 3.4, we will see that (32) also plays a central role in the limit of detection.
3 Fluorescence Dynamics
3.1 Mathematical Model
To adapt the approach to the fluorescence measured in real-time PCR, it is necessary to relate the DNA content in solution to the monitoring chemistry. When a fluorescent probe is used to monitor the kinetics of PCR, the inactive and active probe species usually make significant contributions to fluorescence. With these two fluorescent species, the fluorescence analog of Beer’s Law is
afterFootnote 2 each cycle \(i=1\) to n and for each well \(w=1\) to m. Here, \(F_{i,w}\) is the fluorescence measured, \(f_{i,w}^{-}\) (or \(f_{i,w}^{+}\)) is a constant representing the fluorescence per mole of inactive (or active) probe, and \(C_{i,w}^{-}\) (or \(C_{i,w}^{+}\)) is the molar concentration of inactive (or active) probes. Each molar fluorescence, \(f_{i,w}^{-}\) or \(f_{i,w}^{+}\) (also denoted as \(f_{i,w}^\pm \)), may depend on i due to photobleaching. Each may also depend on w due to spatial variation in electronics and temperature. The terms \(F_{i,w}\), \(C_{i,w}^{-}\), and \(C_{i,w}^{+}\) are random variables through their dependence on DNA content (see below).
Illustration of relationship between successful DNA replication and changes in fluorescence associated with a hydrolysis probe binding to the reverse strand. (a) As polymerization begins, the hydrolysis probe binds to the reverse strand (blue, dashed line). The probe is in its inactive state, where fluorescence emitted by the fluorophore (‘F’) is quenched by the quencher (‘Q’) in close proximity. (b) As polymerization (green arrow) reaches the location of the probe, the probe is hydrolyzed. (c) After successful production of a forward strand (orange, solid line) from a reverse strand, the fluorophore is activated, as it is no longer in close proximity to the quencher. When a reverse strand is produced from a forward strand, however, the fluorescence does not change (Color figure online)
Assuming that probe is not degraded during cycling,
for all i and w, where C is a known constant representing the total concentration of probe in solution. As DNA is replicated, inactive probe is converted to active probe in a manner that depends on the reaction stochiometry. For hydrolysis probes, an inactive probe is activated, or hydrolyzed, each time one of the complementary strands is replicated. Without loss of generality, we consider the case where the hydrolysis probe binds to the reverse strand (see Fig. 3). After completing i PCR cycles, the concentration of active probe in each well w is then
where \(\mathcal {V}\) and \(\textsf {N}_\textsf {a}\) are constants representing the volume of solution and Avogadro’s number, respectively, and \(\Delta X_{i,w}:=X_{i,w} - X_{0,w}\). We assume for each \(i=0\) to n that \(X_{i,1},\ldots , X_{i,m}\) are independent and distributed identically to \(X_i\), and similarly that \(Y_{i,1},\ldots , Y_{i,m}\) are independent and distributed identically to \(Y_i\). By combining (34), (35), and (36), the fluorescence model becomes
where
represents the contribution of background fluorescence, and
represents the increase in fluorescence per synthesis of forward strand.
From (37), the first two central-moments of \(F_{i,w}\) are
where \(\Delta X_i :=X_i - X_0\). Here, \(\mathbb {E}\left[ \Delta X_i\right] \) can be viewed as a function of \({\bar{p}}\), R, r, and \(\mathbb {E}\left[ I\right] \) by substituting (8) into (13). Similarly, \(\textsf{Var}\left[ X_i\right] \) can be viewed as a function of \({\bar{p}}\), R, r, \(\mathbb {E}\left[ I\right] \), and \(\textsf{Var}\left[ I\right] \) by substituting (8) and (26) into (24). While \(\textsf{Var}\left[ X_0\right] \) is a function of r, \(\mathbb {E}\left[ I\right] \), and \(\textsf{Var}\left[ I\right] \) through (26), an expression for \(\textsf{Cov}\left[ X_i, X_0\right] \) is needed for (40b). The methods described in Sect. 2.2 and 2.3 can be readily adapted to the cross-covariance matrix (see Sect. B.2) to obtain
From (13), (28), and (41), it follows that (40) can be expressed as
and
as \(i\rightarrow \infty \). As the dominant term of \(\textsf{Var}\left[ F_{i,w}\right] \) is proportional to \(\lambda _1^{2i} \gg \lambda _1^i\), it is independent of \(\textsf{Cov}\left[ X_i, X_0\right] = O\left( \lambda _1^i\right) \). Instead, the dominant term of \(\textsf{Var}\left[ F_{i,w}\right] \) arises from \(\textsf{Var}\left[ X_i\right] \) as in (28). As in (32), Equations (42) and (43) imply
where \(\alpha =\alpha \left( R\right) \) and \(\beta =\beta \left( R, {\bar{p}}, r\right) \) were defined in (33) for Case D, RF, and RR.
In contrast to other models (e.g., Ruijter et al. 2013, 2014), the fluorescence model (37) is consistent with stoichiometric reactions involving hydrolysis probes and DNA polymerase. By using the fluorescence analog of Beer’s Law, it provides a physical basis for description of the background fluorescence \(b_{i,w}\). Finally, unlike conventional approaches, the more mechanistic model demonstrates that \(d_{i,w}\) may depend on cycle i.
When \(R\ne 1\), R complicates the relationship between DNA content and fluorescence. However, for Case RR, Equations (8), (13), and (40a) lead to
which is independent of R. Since the choice of forward and reverse strands was arbitrary, this demonstrates that the monitoring probe can be chosen so that \(\mathbb {E}\left[ F_{i,w}\right] \) is independent of R, and may be a useful design-rule for RT-qPCR assays.
3.2 Extraction of Molar Fluorescence
In real-time PCR, control experiments containing all reagents except nucleic acid template are often performed to check for contamination. In this section, we will show how they can also be used to calculate \(f_{i,w}^\pm \).
Since template is not present (i.e., \(\varvec{U}_0= 0\)), amplification does not occur and the probe cannot be activated. Equation (37) with (38) becomes
where \(F_{i,w}\) is instead deterministic. After filling m wells of a PCR plate with inactive probe at known C (and appropriate solvation environment) and measuring \(F_{i,w}\) after each \(i \ge 1\), \(f_{i,w}^{-}\) can be calculated pointwise from (45) by division. However, to get a more realistic estimate of each \(f_{i,w}^{-}\), the experiment can be repeated with q different plates having a total probe concentration \(C^1< \cdots < C^q\) in all m wells. With
these experiments yield the measurements
for each cycle i and well w. Under the assumption that \(\displaystyle \frac{F^j_{i,w}}{C^j}\) for \(j=1\) to q are i.i.d. to a normal distribution, we can compute \(f_{i,w}^{-}\) via
The same procedure can be used to calculate \(f_{i,w}^{+}\) after performing the identical experiments with active probe instead of inactive probe. The standard deviation in \(f_{i,w}^\pm \) can be estimated pointwise by
Using hydrolysis probes, we selected the fluorophore (‘F’ in Fig. 3) to represent the active probe. We used \(q=4\) concentrations for the active probe, and \(q=3\) concentrations for the inactive probe. Additional details of the experimental procedure can be found in Appendix A. Visual comparisons between the model and experimental data can be found in Figs. S1 to S96 in the Supplementary Information (SI). The pointwise values of \(f_{i,w}^{\pm }\) and \(\sigma _{i,w}^{\pm }\) are tabulated in Tables S1 to S96 of the SI.
Total number of cycles and wells for all plates (count, vertical axis) possessing values of \(\sigma _{i,w}^{\pm }/f_{i,w}^\pm \) in a certain interval (bin, horizontal axis). The bin widths are obtained from the Freedman–Diaconis rule. All counts for \(\sigma _{i,w}^{-}/f_{i,w}^{-} > 0.022\) correspond to \(w=2\) (well A2, see Fig. S2 and Table S2 in the SI)
To assess the validity of approximating the measured fluorescence by \(f_{i,w}^\pm C\) for each cycle i and well w, the coefficient of variation, or \(\displaystyle \frac{\sigma _{i,w}^{\pm }}{f_{i,w}^{\pm }},\) was calculated. A histogram of all values (i.e., all n cycles, all m wells, and all q plates) for each probe species is depicted in Fig. 4. For the active probe, the coefficient of variation is often very small, typically much less than 0.01. For the inactive probe, the coefficient of variation can be larger but is still often less than 0.02. This is an indication that the model is realistic.
Background (top) and incremental increase (bottom) as a function of cycle i for a selection of eight different wells (colors, see legend) with \(C=0.125\) pmol/L and \(\mathcal {V}=20\) \(\mu \)L. The partially transparent shadings at each i depict an estimate of a 95 % confidence interval (Color figure online)
Having validated the use of the Beer’s Law analog in describing the fluorescence, we used the molar-fluorescence parameters in Fig. 5 to assess how the background \(b_{i,w}\) and incremental increase \(d_{i,w}\) change with cycle i (for a fixed well w). Here, we see that \(b_{i,w}\) is not a linear function of i. In fact, for several w, \(b_{i,w}\) possesses a maximum in cycle around \(i=2\) or \(i=3\). The decreases in \(b_{i,w}\) with increasing i are attributed to result from photobleaching.
Since the fluorescence of an active probe is larger than the fluorescence of an inactive probe, \(d_{i,w}\) is positive. Figure 5 also illustrates that \(d_{i,w}\) is not independent of cycle; instead it often increases with i. This reveals another source of systematic error, as most models assume that \(d_{i,w}\) is independent of cycle (see, e.g., Ruijter et al. (2009), Equation (4); Lievens et al. (2012), Equation (7); Liu and Saint (2002), Equation (2)).
3.3 Calculation of Fluorescence Profiles
Having calculated \(f_{i,w}^-\) and \(f_{i,w}^+\) pointwise through (46), we leverage (40) with (8), (13), (24), (26), and (41) to compute fluorescence curves with uncertainty. We prescribe common values for assay parameters C, \(\mathcal {V}\), \({\bar{p}}\), and R and assume \(\mathbb {E}\left[ I\right] \) is known. However, we also need to specify the relationship between \(\textsf{Var}\left[ I\right] \) and \(\mathbb {E}\left[ I\right] \), as well as the distribution of each \(F_{i,w}\).
We will assume for simplicity that
for some constant \(\chi > 0\). If we make the conventional assumption (Nedelman et al. 1992; Sundberg et al. 2010; Tellinghuisen and Spiess 2015; Ruiz-Villalba et al. 2021) that I is a Poisson random-variable,Footnote 3 then \(\chi = 1\). However, (48) can also correspond to different probability distributions. If I obeys a negative binomial distribution with probability of success \(\varphi \in (0, 1)\), for example, then \(\chi = 1/\varphi > 1\).
Our assumption on the distribution of each \(F_{i,w}\) is rooted in the characteristic values of the fluorescence parameters \(b_{i,w}\) and \(d_{i,w}\). As Fig. 5 demonstrates that \(b_{i,w}\) and \(d_{i,w}\) are typically around 1 and \(10^{-6}\), respectively, this implies with (40a) that \(\mathbb {E}\left[ \Delta X_i\right] \) should be more than \(10^6\) for \(\mathbb {E}\left[ F_{i,w}\right] > b_{i,w}\). That is, the expected number of successful Bernoulli trials over all i cycles should be more than \(10^6\) for the fluorescence to reach levels above background. With such a large sample size, it is natural to invoke the central limit theorem and assume that \(F_{i,w}\) obeys a normal distribution with mean \(\mathbb {E}\left[ F_{i,w}\right] \) from (40a) and variance \(\textsf{Var}\left[ F_{i,w}\right] \) from (40b).
Fluorescence curves calculated from (40) with (8), (13), (24), (26), and (41) for \(w=13\). Each subplot is associated with a different input copy-number \(\mathbb {E}\left[ I\right] \), as depicted in each bottom-left corner, while all subplots have \(C=0.125\) pmol/L, \(\mathcal {V}=20\) \(\mu \)L, \({\bar{p}}=0.9\), and \(R=1\). The initial distribution is taken as Case D with (48) and \(\chi = 1\). The light-grey shadings at each i depict the region between \(\min \limits _{1\le w\le 96}\mathbb {E}\left[ F_{i,w}\right] \) and \(\max \limits _{1\le w\le 96}\mathbb {E}\left[ F_{i,w}\right] \)
In Fig. 6, fluorescence curves are computed with uncertainty for ds-DNA for \(w=13\). Different \(\mathbb {E}\left[ I\right] \) ranging from 64 (top-left subplot) down to 4 (bottom-right subplot) are investigated. The expected fluorescence depicts behavior that is characteristic of the background and exponential phase observed in typical measurements. During the initial cycles, the term \(b_{i,w}\) is much larger than \(d_{i,w}\mathbb {E}\left[ \Delta X_i\right] \) and only small changes in fluorescence are observed. Here, the fluorescence is in the background regime. After more cycles are performed, however, the expected value of fluorescence increases exponentially with cycle. The change in \(\mathbb {E}\left[ F_{i,13}\right] \) with \(\mathbb {E}\left[ I\right] \) is also in line with typical trends. As \(\mathbb {E}\left[ I\right] \) is decreased, the expected value of the fluorescence in the exponential phase shifts to the right. In other words, more cycles are required to reach the same expected fluorescence value.
This approach provides quantitative estimates of sources and magnitudes of uncertainty in different regimes, which are difficult to determine from replicate experiments alone. For large \(\mathbb {E}\left[ I\right] \) and small i, the well-to-well variation in expected value (light-grey, shaded regions in Fig. 6) is larger or comparable to the error in fluorescence in each well. As such, spatial variation has a significant impact on the error. After many cycles have been completed, on the other hand, the uncertainty in fluorescence is strongly dependent on the expected initial copy-number, increasing drastically with decreasing \(\mathbb {E}\left[ I\right] \). At \(\mathbb {E}\left[ I\right] =4\), the fluorescence does not reach values that are larger than the background fluorescence by an amount that is statistically significant (for \(\kappa = 3\)). This observation suggests that quantifying the uncertainty in fluorescence can provide limitations on the measurement.
3.4 Limit of Detection
After performing n PCR cycles, the fluorescence produced by PCR is only useful if it is larger than background levels by a statistically significant amount. Requiring the increase to be at least some \(0< \kappa < \infty \) standard deviations, this amounts to the constraint
for some well w. Equation (49) describes a feasible region of design space for a real-time PCR assay. It can be considered to depend on n, \(b_{n,w}\), \(d_{n,w}\), \({\bar{p}}\), R, r, \(\mathbb {E}\left[ I\right] \), and \(\textsf{Var}\left[ I\right] \) through (8), (13), (24), (26), and (41). However, (49) can be further simplified by substituting (40) and rearranging, leading to
an expression that is no longer dependent on w. Since \(\textsf{CV}\left[ \Delta X_n\right] \ge 0\) and \(\kappa > 0\), it follows that
In addition, since
by (13), (28), and (41), and n typically ranges from 35 to 50, the error in approximating \(\textsf{CV}\left[ \Delta X_n\right] ^2\) by \(\textsf{CV}\left[ X_n\right] ^2\) is extremely small, often less than machine precision. As a result, the left-hand-side of (50) is expressed as the term on the right-hand-side of (32), or
where \(\alpha \) and \(\beta \) are defined in (33) for Case D, where the input is ds-DNA; Case RF, where the input is fs-RNA; and Case RR, where the input is rs-RNA. If we let I satisfy (48), Equation (51) can be rearranged to
The limit of detection, L, or the smallest expected-initial-copy-number that can be detected reliably, is then
The largest coefficient of variation in I that can be detected, M, is estimated from (53) and (48), or
To compute typical values of L and M, we evaluated them as in (53) and (54) with \(\chi = 1\), \(\kappa = 3\), for 100 equally-spaced \({\bar{p}}\in [0.8, 0.99]\), \(R\in [0.9, 1.1]\), and \(r \in [0.2, 0.99]\) (including endpoints). For I representing ds-DNA (Case D), we find that L is either 5 or 6, corresponding to M of 0.447 and 0.408, respectively. For I representing RNA as in Case RF or Case RR, L ranges between 9 and 52, corresponding to M of 1/3 and 0.139, respectively. The range of L is the same for fs-RNA and rs-RNA.
4 Conclusions and Future Work
In this work, we presented a new model for fluorescence in real-time PCR that reduced bias and quantified uncertainty. Distinguishing between complementary strands provided a stoichiometric description of fluorescence reported by hydrolysis probes and permitted application to initial conditions encountered in RT-qPCR. Viewing the fluorescence as a Beer’s Law analog enabled the background fluorescence to be determined without extrapolation or assuming a certain relationship with cycle. It also allowed for measurement and calculation of background fluorescence without adjusting amplification data. Incorporating the variance in copy number into the fluorescence model enabled quantification of fluorescence uncertainty and analytical expressions for the limit of detection.
In addition to their practical utility, the two-type branching-process and repurposed fluorescence-model provided new intuition on the physics and chemistry in PCR. At short times, there is a lag in exponential growth (usually at most 5 cycles) as the ratio of expected strand counts changes from its initial to critical value, R. The quantity R represents the square root of the ratio of the two synthesis efficiencies (see (3)). In constrast to a previous report investigating deterministic models, we found that the initial composition only impacts the dynamics after the lag phase if \(R\ne 1\).
The variance in the fluorescence is dominated by a term that increases exponentially by twice the factor of the expected value. This explains, in part, why quantification by end-point PCR is not reproducible. The three terms dominating the variance were attributed to arise from the initial variance (\(\nu _{1,1}\), see (29a)), imperfect amplification (\(\eta _{1,1}^{(1)}\), see (29b)), and deviation in directional efficiencies (i.e., \(R\ne 1\); see \(\eta _{1,1}^{(2)}\) in (29c)). The fluorescence model for hydrolysis probes demonstrated that the background fluorescence originates from the molar fluorescence of the inactive probe times the total concentration of probe. The incremental increase in fluorescence is proportional to the difference in molar fluorescence between active and inactive probe and, like the background fluorescence, is neither independent of cycle nor a linear function of cycle.
The stochastic view of PCR explains, in part, why deterministic methods that use reaction-specific amplification probabilities are generally less accurate (Ruijter et al. 2013). This is because, for each well w and cycle i, \(N_{i,w}/N_{i-1,w}\ne 1 + p\) (see Equation (5)). Even if \(R=1\) and \(N_{i,1},\ldots , N_{i,m}\) are independent and distributed identically to \(N_i\) for each i, this is not necessarily true because not every realization of a random variable is equal to its expected value.
While this work applied the stochastic model of PCR to the fluorescence reported by hydrolysis probes, it can readily be extended to other chemistries. For example, for probes that anneal to forward-stranded DNA, (37) instead becomes
For these probes, the fluorescence is measured during the annealing portion of each cycle where only \(i-1\) cycles of PCR have been completed.
To capture the fluorescence reported by DNA-binding dyes, on the other hand, an extension to the model is needed. This is because the amount of dye bound to a DNA strand depends on the total amount of DNA present in solution (this includes DNA that is not template, like primers (Ruijter et al. 2009)). Application to fluorescent dyes represents an interesting direction for future generalizations of the fluorescence model.
Finally, our approach in this work focused on quantifying the dynamics and uncertainty of fluorescence when the initial amount of each complementary strand is known, as well as their amplification probabilities. (That is, we assumed that \(\mathbb {E}\left[ I\right] \), \(\textsf{Var}\left[ I\right] \), \({\bar{p}}\), R, and r were known.) However, the ultimate goal of monitoring the kinetics of PCR by fluorescent probes is to infer \(\mathbb {E}\left[ I\right] \), the expected input number. As such, it is of interest to extend the approach to UQ-PCR, or uncertainty quantification of the initial amount of DNA. To this end, another direction for future work is the investigation of the probabilistic nature of PCR in backwards time (see Fig. 7).
Data Availability
The experimental data and software associated with this study are available at https://github.com/usnistgov/bias-uq-pcr. Additional parameters computed from experimental data are tabulated in the Supplementary Information.
Notes
The expression which follows could have also been developed by substituting the first expression of (9) into the final expression of (7) and left-multiplying by \(\varvec{Z}\), yielding
$$\begin{aligned} \varvec{Z}\mathbb {E}\left[ \varvec{U}_i\right] = \varvec{\Lambda }^i\varvec{Z}\mathbb {E}\left[ \varvec{U}_0\right] . \end{aligned}$$In this work, we assume that the fluorescence is measured after each PCR cycle. This is typical for assays utilizing hydrolysis probes, but is not always the case.
There is some limited experimental evidence for assuming that I is a Poisson random-variable (Beer et al. 2007); however, its validity is challenging to assess experimentally. (If the initial number of DNA strands could be easily measured, there would be no need for qPCR.) In the future, it would be interesting to compare (48) to a result obtained from error propagation for a conventional experimental procedure (see, e.g., Hedges (2002)).
References
Bauer M (2007) RNA in forensic science. Forensic Sci Int Genet 1:69–74. https://doi.org/10.1016/j.fsigen.2006.11.002
Beer NR, Hindson BJ, Wheeler EK et al (2007) On-chip, real-time, single-copy polymerase chain reaction in picoliter droplets. Anal Chem 79:8471–8475. https://doi.org/10.1021/ac701809w
Biver T, Secco F, Tiné MR et al (2003) Equilibria and kinetics of the intercalation of Pt-proflavine and proflavine into calf thymus DNA. Arch Biochem Biophys 418:63–70. https://doi.org/10.1016/S0003-9861(03)00384-9
Biver T, De Biasi A, Secco F et al (2005) Cyanine dyes as intercalating agents: Kinetic and thermodynamic studies on DNA/Cyan40 and DNA/CCyan2 systems. Biophys J 89:374–383. https://doi.org/10.1529/biophysj.105.059790
Boggy GJ, Woolf PJ (2010) A mechanistic model of PCR for accurate quantification of quantitative PCR data. PLoS ONE 5:e12355. https://doi.org/10.1371/journal.pone.0012355
Bustin S, Dhillon HS, Kirvell S et al (2015) Variability of the reverse transcription step: Practical implications. Clin Chem 61:202–212. https://doi.org/10.1373/clinchem.2014.230615
Bustin S, Mueller R, Shipley G et al (2021) COVID-19 and diagnostic testing for SARS-CoV-2 by RT-qPCR—facts and fallacies. Int J Mol Sci 22:2459. https://doi.org/10.3390/ijms22052459
Bustin SA (2004) Getting started–the basics of setting up a QPCR assay. IUL Biotechnol, La Jolla, CA, chap 13:530–543
Elizaquível P, Aznar R, Sanchez G (2014) Recent developments in the use of viability dyes and quantitative PCR in the food microbiology field. J Appl Microbiol 116:1–13. https://doi.org/10.1111/jam.12365
Guescini M, Sisti D, Rocchi MB et al (2008) A new real-time PCR method to overcome significant quantitative inaccuracy due to slight amplification inhibition. BMC Bioinform 9:326. https://doi.org/10.1186/1471-2105-9-326
Hedges AJ (2002) Estimating the precision of serial dilutions and viable bacterial counts. Int J Food Microbiol 76:207–214. https://doi.org/10.1016/S0168-1605(02)00022-3
Jacob C, Peccoud J (1996) Estimation of the offspring mean for a supercritical branching process from partial and migrating observations. C R Acad Sci Paris Ser I 322:763–768
Jacob C, Peccoud J (1996) Inference on the initial size of a supercritical branching process from migrating binomial observations. C R Acad Sci Paris Ser I 322:875–880
Kaltenboeck B, Wang CM (2005) Advances in real-time PCR: application to clinical laboratory diagnostics. Adv Clin Chem 40:219–259. https://doi.org/10.1016/S0065-2423(05)40006-2
Kebschull JM, Zador AM (2015) Sources of PCR-induced distortions in high-throughput sequencing data sets. Nucleic Acids Res 43:e143. https://doi.org/10.1093/nar/gkv717
Lievens A, Van Aelst S, Van den Bulcke M et al (2012) Enhanced analysis of real-time PCR data by using a variable efficiency model: FPK-PCR. Nucleic Acids Res 40:e10. https://doi.org/10.1093/nar/gkr775
Liu W, Saint DA (2002) A new quantitative method of real time reverse transcription polymerase chain reaction assay based on simulation of polymerase chain reaction kinetics. Anal Biochem 302:52–59. https://doi.org/10.1006/abio.2001.5530
National Center for Immunization and Respiratory Diseases (U.S.). Division of Viral Diseases (2020) 2019-Novel Coronavirus (2019-nCoV) Real-time rRT-PCR Panel Primers and Probes. https://stacks.cdc.gov/view/cdc/84525
National Institutes of Health, National Human Genome Research Institute (2023) Polymerase chain reaction (PCR). https://www.genome.gov/genetics-glossary/Polymerase-Chain-Reaction
Nedelman J, Heagerty P, Lawrence C (1992) Quantitative PCR: Procedures and precisions. Bull Math Biol 54:477–502
Nicklas JA, Buel E (2003) Quantification of DNA in forensic samples. Anal Bioanal Chem 376:1160–1167. https://doi.org/10.1007/s00216-003-1924-z
Peccoud J, Jacob C (1996) Theoretical uncertainty of measurements using quantitative polymerase chain reaction. Biophys J 71:101–108. https://doi.org/10.1016/S0006-3495(96)79205-6
Peirson SN, Butler JN, Foster RG (2003) Experimental validation of novel and conventional approaches to quantitative real-time PCR data analysis. Nucleic Acids Res 31:e73. https://doi.org/10.1093/nar/gng073
Ruijter JM, Ramakers C, Hoogaars WMH et al (2009) Amplification efficiency: linking baseline and bias in the analysis of quantitative PCR data. Nucleic Acids Res 37:e45. https://doi.org/10.1093/nar/gkp045
Ruijter JM, Pfaffl MW, Zhao S et al (2013) Evaluation of qPCR curve analysis methods for reliable biomarker discovery: Bias, resolution, precision, and implications. Methods 59:32–46. https://doi.org/10.1016/j.ymeth.2012.08.011
Ruijter JM, Lorenz P, Tuomi JM et al (2014) Fluorescent-increase kinetics of different fluorescent reporters used for qPCR depend on monitoring chemistry, targeted sequence, type of DNA input and PCR efficiency. Microchim Acta 181:1689–1696. https://doi.org/10.1007/s00604-013-1155-8
Ruiz-Villalba A, Ruijter JM, van den Hoff MJB (2021) Use and misuse of \(C_{{\rm q }}\) in qPCR data analysis and reporting. Life 11:496. https://doi.org/10.3390/life11060496
Schwabe D, Falcke M (2022) On the relation between input and output distributions of scRNA-seq experiments. Bioinformatics 38:1336–1343. https://doi.org/10.1093/bioinformatics/btab841
Schwaber J, Andersen S, Nielsen L (2019) Shedding light: The importance of reverse transcription efficiency standards in data interpretation. Biomol Detect Quantif 17:100077. https://doi.org/10.1016/j.bdq.2018.12.002
Stolovitzky G, Cecchi G (1996) Efficiency of DNA replication in the polymerase chain reaction. Proc Natl Acad Sci USA 93:12947–12952. https://doi.org/10.1073/pnas.93.23.12947
Sun F (1995) The polymerase chain reaction and branching processes. J Comput Biol 2:63–86. https://doi.org/10.1089/cmb.1995.2.63
Sundberg SO, Wittwer CT, Gao C et al (2010) Spinning disk platform for microfluidic digital polymerase chain reaction. Anal Chem 82:1546–1550. https://doi.org/10.1021/ac902398c
Tellinghuisen J, Spiess AN (2015) Absolute copy number from the statistics of the quantification cycle in replicate quantitative polymerase chain reaction experiments. Anal Chem 87:1889–1895. https://doi.org/10.1021/acs.analchem.5b00077
Tichopad A, Dilger M, Schwarz G et al (2003) Standardized determination of real-time PCR efficiency from a single reaction set-up. Nucleic Acids Res 31:e122. https://doi.org/10.1093/nar/gng122
Weiss G, von Haeseler A (1995) Modeling the polymerase chain reaction. J Comput Biol 2:49–61. https://doi.org/10.1089/cmb.1995.2.49
Zhao S, Fernald RD (2005) Comprehensive algorithm for quantitative real-time polymerase chain reaction. J Comput Biol 12:1047–1064. https://doi.org/10.1089/cmb.2005.12.1047
Acknowledgements
The authors acknowledge D. Duewer for helpful discussions. R. F. D. and M. J. R. acknowledge support from a National Research Council fellowship. R. F. D. also acknowledges support from the NIST Professional Research Experience Program through University of Maryland.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclaimer
Certain equipment, instruments, software, or materials, commercial or non-commercial, are identified in this paper in order to specify the experimental procedure adequately. Such identification is not intended to imply recommendation or endorsement of any product or service by the National Institute of Standards and Technology, nor is it intended to imply that the materials or equipment identified are necessarily the best available for the purpose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Appendices
Appendix A Experimental
Reagents were obtained from suppliers for use in experiments. A DNA suspension buffer (TE\(^{-4}\)) of pH 8.0 with 10 mmol/L Tris and 0.1 mmol/L EDTA was obtained from Teknova (Hollister, CA). 6-Carboxyfluorescein (6-FAM), single isomer, was purchased in solid form from Thermo Fisher Scientific (Waltham, MA). The working solution of 6-FAM was prepared by dissolving 10.1 mg 6-FAM in 1.75 mL of absolute ethanol (Sigma-Aldrich, St. Louis, MO). Subsequently, 10.0 \(\mu \)L of the ethanolic solution was dissolved in 500 mL of TE\(^{-4}\). A TaqMan minor groove binder probe with non-fluorescent quencher (Applied Biosystems, Waltham, MA), possessing the sequence \(5^\prime -\)ACCCCGCATTACGTTTGGTGGACC\(-3^\prime \) and reporter (6-FAM) of the National Center for Immunization and Respiratory Diseases (U.S.). Division of Viral Diseases (2020) 2019-nCoV_N1 assay, was obtained from Thermo Fisher Scientific. Working solutions of probe were prepared by adding TE\(^{-4}\) to a portion of the stock solution to yield a solution of concentration 5 \(\mu \)mol/L probe.
In a typical experiment, a solution was prepared with 25 vol. % TE\(^{-4}\) at a specific concentration of either 6-FAM (active probe) or TaqMan probe (inactive probe). After mixing, 20 \(\mu \)L was transferred into each well of a 96-well plate. The plate was covered with adhesive film and centrifuged. After assessing that air bubbles were not visible, the plate was placed in an Applied Biosystems 7500 HID Real-Time PCR instrument. The thermal cycling protocol consisted of a 2 min holding stage at 55 \(^\circ \)C (328 K), followed by 45 cycles. Each cycle consisted of 30 s at 55 \(^\circ \)C (328 K), followed by 3 s at 95 \(^\circ \)C (368 K). Data collection occurred during the 55 \(^\circ \)C step in each cycle. The raw data through filter 1 was exported with HID Real-Time PCR Analysis Software, Version 1.2 (Applied Biosystems). The fluorescence values were divided by \(10^6\) before subsequent analysis.
Appendix B Extended Derivations
1.1 B.1 Derivation of (24) from (23)
Consider some \(\varvec{B}\in \mathbb {R}^{2\times 2}\). From (12), we can write
where \(\varvec{T}_{j,k}:\mathbb {R}^{2\times 2}\mapsto \mathbb {R}^{2\times 2}\) is the linear operator
Application of (B1) to (23) leads to
For each \(b\in \{\lambda _1^2, \lambda _1\lambda _2, \lambda _2^2\}\), the geometric series in (B3) can be simplified via
This admits the expression
it is evident that (B4) is identical to (24).
1.2 B.2 Derivation of (41)
In this section, the derivation of (41) is completed by investigating the cross-covariance matrix, defined as
for any two random vectors \(\varvec{X}\) and \(\varvec{Y}\). Notice that \(\mathcal {K}\left[ \varvec{X}, \varvec{X}\right] = \textsf{Var}\left[ \varvec{X}\right] \).
From the law of total expectation,
and
The latter expression follows because \(\varvec{U}_i\) and \(\varvec{U}_0\) are independent when conditioned on \(\varvec{U}_{i-1}\). Thus,
where the second, third, and fourth step follow from induction, Equation (12), and the independence of \(X_0\) and \(Y_0\), respectively. In particular, it follows that
which is the same as (41).
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
DeJaco, R.F., Roberts, M.J., Romsos, E.L. et al. Reducing Bias and Quantifying Uncertainty in Fluorescence Produced by PCR. Bull Math Biol 85, 83 (2023). https://doi.org/10.1007/s11538-023-01182-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11538-023-01182-z