Abstract
Purpose
Glioblastoma Multiforme (GBM) represents the predominant aggressive primary tumor of the brain with short overall survival (OS) time. We aim to assess the potential of radiomic features in predicting the time-to-event OS of patients with GBM using machine learning (ML) algorithms.
Materials and methods
One hundred nineteen patients with GBM, who had T1-weighted contrast-enhanced and T2-FLAIR MRI sequences, along with clinical data and survival time, were enrolled. Image preprocessing methods included 64 bin discretization, Laplacian of Gaussian (LOG) filters with three Sigma values and eight variations of Wavelet Transform. Images were then segmented, followed by the extraction of 1212 radiomic features. Seven feature selection (FS) methods and six time-to-event ML algorithms were utilized. The combination of preprocessing, FS, and ML algorithms (12 × 7 × 6 = 504 models) was evaluated by multivariate analysis.
Results
Our multivariate analysis showed that the best prognostic FS/ML combinations are the Mutual Information (MI)/Cox Boost, MI/Generalized Linear Model Boosting (GLMB) and MI/Generalized Linear Model Network (GLMN), all of which were done via the LOG (Sigma = 1 mm) preprocessing method (C-index = 0.77). The LOG filter with Sigma = 1 mm preprocessing method, MI, GLMB and GLMN achieved significantly higher C-indices than other preprocessing, FS, and ML methods (all p values < 0.05, mean C-indices of 0.65, 0.70, and 0.64, respectively).
Conclusion
ML algorithms are capable of predicting the time-to-event OS of patients using MRI-based radiomic and clinical features. MRI-based radiomics analysis in combination with clinical variables might appear promising in assisting clinicians in the survival prediction of patients with GBM. Further research is needed to establish the applicability of radiomics in the management of GBM in the clinic.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Glioblastoma Multiforme (GBM) represents the predominant aggressive tumor of the brain and spinal cord [1]. Unfortunately, this aggressive tumor has a mean survival time of fewer than 15 months. Moreover, its inadequate response to current treatment options and complex progression patterns makes clinical decision-making harder for physicians [2, 3]. Therefore, overall survival (OS) prediction, as a component of prognostication, is a major area of interest in oncological studies. It is evident from the literature that several studies have assessed the predictive power of demographical, clinical, or laboratory data in GBM patient survival analysis. For example, a landmark study by Czipska et al. [4] assessed the prognostic factors of long-term survival for patients with GBM. In another systematic review [5], three types of outcomes for OS prediction, including continuous, binary, and time-to-event, were reported as defined in the reviewed articles.
Despite the decent predictive power of clinical-related features shown by previous studies, most studies intended to examine the prognostication ability of imaging biomarkers [6]. Magnetic Resonance Imaging (MRI) is the most frequent diagnostic procedure utilized in the detection and evaluation of GBM. Radiologists integrate several conventional qualitative and quantitative MRI assessment methods [7, 8]. However, conventional qualitative evaluation might have some limitations and overlook the hidden layers of information within images. In this light, the term “radiomics” can be put forward as it fundamentally describes the analysis of medical images via computational data extraction and, in other words, transforming images into minable biomarkers [9,10,11]. Radiomics can help us prepare data hidden in images that cannot be seen with conventional image assessment methods with the naked eye in different diseases [12,13,14,15,16]. Whether in the diagnostic or the prognostic area, cancer research has always been of interest to radiomics researchers, and GBM is not an exception [17, 18].
A growing body of research recognizes the efficiency of radiomics analyses for GBM tumors [19, 20]. A recent study by Artzi et al. [21] studied the efficiency of radiomics for differentiating brain metastasis versus GBM. The study of Buchlak et al. [22] is also worth mentioning as they systematically reviewed previous studies focusing on GBM diagnosis via radiomics assessment. In a study by Bae et al. [23], the authors found that adding radiomic features to patients’ genetic and clinical profiles can improve survival prediction. Other studies reached a similar conclusion as MRI-based radiomics and machine learning (ML) algorithms could predict the OS of patients with decent statistics [24,25,26,27]. While several studies reported on the use of radiomics in neurodegenerative disorders or GBM, most have utilized ML to binary (high vs. low risk or short vs. long survival time) or multiple (high, intermediate, and low risk or short, intermediate, and long survival time) classification of OS in GBM patients. Therefore, our study's contribution is in using preprocessing methods, feature selectors, and MLs to predict time-to-event survival using MRI-based radiomics features. Therefore, clinicians need to have a more comprehensive knowledge of GBM management by further elaborating on patients’ prognoses. In this study, we aimed to evaluate the applicability of ML algorithms to MRI-based radiomic features along with clinical variables to predict patients’ time-to-event OS.
Materials and methods
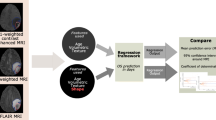
Figure 1 illustrates a flowchart of the various steps followed in the present study protocol.
Flowchart of the various steps followed in the present study protocol. Feature selection methods include Univariate C-Index (UCI), Boruta, Variable hunting Variable Importance (VH.VIMP), Variable hunting (VH), Minimal Depth (MD), Mutual Information (MI), and Iterated Bayesian Model Averaging (IBMA). Machine learning methods include Cox Proportional Hazard regression (CoxPH), Cox Boost (CB), Generalized Linear Model Network (GLMN), Random Survival Forest (RSF), GLM Boosting (GLMB), and Gradient Boosting (GB)
Data and image acquisition
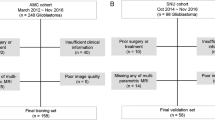
The Cancer Imaging Archive (TCIA) was used to download images and clinical information of the enrolled patients [28, 29]. The following were the inclusion criteria for enrolment in this study protocol: (1) All GBM patients who underwent T1-weighted contrast-enhanced (T1w-CE) and T2-FLAIR MRI sequences AND (2) Images acquired before therapy AND (3) All patients have clinical data including gender, age, Karnofsky Performance Score (KPS), radiation treatment adjuvant (RTA), pharmaceutical treatment adjuvant (PTA), and survival time with follow-up event status (expired and alive). The exclusion criteria for this study include: (1) Missing one of the MRI sequences; (2) Images presenting with poor quality or motion artifacts; (3) Images acquired after treatment, and (4) Missing clinical data. Figure 2 shows a flowchart of inclusion and exclusion criteria.
After applying inclusion and exclusion criteria, 119 GBM patients with T1w-CE and T2-FLAIR MRI sequences, clinical data, and survival time were enrolled in this study protocol. MR image sequence specifications are summarized in Table 1. Patients collected clinical and demographical information, including gender, age, MGMT status, Karnofsky Performance Score (KPS), tumor histology subtype, RTA, and PTA used as clinical variables and time-to-event OS as a target. Due to the unavailability of the MGMT status and tumor histology subtype variables, these variables were removed from this study. KPS is an index reflecting the functional performance of cancer patients with 11 levels ranging from 0 (expired) to 100 (normal) [30]. The clinical and demographic data of patients are shown in Table 2.
Image preprocessing
A 64 bin discretization, Laplacian of Gaussian (LOG), and Wavelet Transform were used to preprocess MR images prior to feature extraction to achieve multiple sets of image features. Sigma values of 1, 1.5, and 2.5 mm were utilized for applying the LOG filter as fine, medium, and coarse filters, respectively [31]. The Wavelet Transform was performed via eight distinct decompositions, including LLL, LLH, LHL, LHH, HLL, HLH, HHL, and HHH, in which L and H stand for low-pass and high-pass filters, respectively, applied to the three-dimensional region. Overall, we utilized 12 distinct preprocessing methods [31].
Segmentation
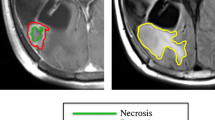
Following the aforementioned preprocessing methods and using the 3D Slicer version 4.9 [32], we manually delineated and segmented four volumes of interest (VOI) for each patient, including core tumor (VOIT), active enhanced tumor (VOIA) and necrotic tumor (VOIN) regions from T1w-CE and edema (VOIE) region from T2-FLAIR. VOIs. Manual segmentation was performed by two experienced medical physicists with 4–6 years of experience. In addition, the manual segmentation was visually inspected and manually corrected, when necessary, by an experienced radiologist with 8 years of experience. The corrected VOIs were then used in subsequent analyses.
Feature extraction
Three feature types were extracted from each of the four VOIs, including shape features (n = 13), first-order (FO) features (n = 18), and texture features (n = 74) (Gray Level Size Zone Matrix (GLSZM), Gray Level Co-occurrence Matrix (GLCM), Neighboring Gray Tone Difference Matrix (NGTDM), Gray Level Run Length Matrix (GLRM), and Gray Level Dependence Matrix (GLDM)). For each preprocessing method, 92 features, including FO and texture features, were extracted (shape features were the same for all preprocessing methods). The whole tumor was also used to calculate three shape features, including A/T, N/T, and T/E volume ratios. In total, 4471 features ([92 × 12 × 4] + [13 × 4] + 3) were extracted from 4 VOIs and 12 image preprocessing methods. Radiomic features were extracted by the Pyradiomics python library [33], which is compliant with IBSI guidelines [31, 34].
Feature selection
A careful selection process took place, during which a group of extracted features were chosen with the assistance of various feature selection (FS) algorithms. The seven FS methods used in this study included Univariate C-Index (UCI), Boruta, Variable hunting Variable Importance (VH.VIMP), Variable hunting (VH), Minimal Depth (MD), Mutual Information (MI), and Iterated Bayesian Model Averaging (IBMA). Further details about these FS methods are provided in the Supplementary section.
Machine learning models
The preprocessing procedures and FS methods along with six ML models (Cox Proportional Hazard regression (CoxPH), Cox Boost (CB), Generalized Linear Model Network (GLMN), Random Survival Forest (RSF), GLM Boosting (GLMB), and Gradient Boosting (GB)) were evaluated by multivariate analysis using the C-index [35]. More details about the ML methods are provided in the Supplementary section.
Modeling and statistical analysis
In the first step, data were split into training (70%) and test (30%) datasets. Subsequently, FS methods were implemented on the training dataset, and the selected features were fed to ML models. Next, hyperparameter optimization with grid search and threefold cross-validation (fivefold cross-validation in supplemental data) was performed for ML models in the training dataset. The details of hyperparameter optimization are shown in Table 3. 1000 bootstraps were performed on the testing dataset to assess the models, and mean ± standard deviation (SD) and 95% confidence interval (95% CI) were reported for each model.
We also performed the Wilcoxon rank sum test among different preprocessing, FS, and ML algorithms and reported the results as box plots and p value tile plots. Kaplan–Meier of the best models with a log-rank p value for the training and test datasets was drawn. A p value of 0.05 was considered as the cut-off for statistically significant differences. Modeling and statistical analysis were implemented in R version 4.0 (R Foundation for Statistical Computing, Vienna, Austria) [36].
Results
Characteristics of GBM patients
In this study, we used data from 119 GBM patients with a mean age (± SD) of 57.83 (± 13.65), with 73 (61.34%) of them being male. Nighty-two patients (77.3%) with a mean survival (± SD) of 13.42 ± 11.91 months passed, while 27 patients (22.7%) with a mean survival (± SD) (i.e., follow-up time) of 17.60 ± 15.45 were alive and considered as censor data.
Feature selection analysis
The feature selectors had decent power for selecting the most important features related to our study purpose. The popularity of the selected features for each FS method can be found in Fig. 3. Our results showed that seven FS methods, including Boruta, UCI, IBMA, MD, MI, VH, and VH.VIMP selected and ranked 38, 120, 120, 120, 120, 120, and 59 features, respectively. Among the features which were selected, more than 10 times out of 697 features (151 unique features), MinorAxis (n = 51), Maximum 2D diameter slice (M2DDS) (n = 39), Sphericity (n = 26), SurfaceArea (n = 20), Maximum 2D diameter column (M2DDC) (n = 15) and Elongation (n = 13) from shape features of VOIT, KPS (n = 38), age (n = 24), RTA (n = 20), PTA (n = 18) and gender (n = 13) from clinical features, M2DDS (n = 21), M2DDC (n = 18), MinorAxis (n = 17), SurfaceArea (n = 13) and A/T ratio (n = 16) from shape features of VOIA, and NGTDM_Coarseness (n = 12) and GLSZM_LAHGLE (n = 12) from texture features of VOIA were repeatedly selected by most of the seven FS algorithms. Figure 4 shows Spearman’s correlation coefficient of shape features.
Model analysis
A heat map of mean C-indices of 1000 bootstraps for each model is shown in Fig. 5. We also generated a heat map of 95% CI and mean ± SD in supplemental Figs. 1S and 2S, respectively. We also generated a heat map of the mean C-indices of these models with fivefold cross-validation for hyperparameter optimization in Fig. 3S. A difference heat map of threefold cross-validation with respect to fivefold cross-validation C-indices for each model is depicted in Fig. 4S. It can be seen that the range of C-index difference is 0 to 0.25 (less than 0.1 in most algorithms) and that the threefold cross-validation for hyperparameter optimization is slightly better on most of the models. As such, the rest of the analysis of this study is based on a threefold cross-validation scheme. Our multivariate model analysis showed that the best prognostic FS/ML combination is the MI/CB applied to the LOG (Sigma = 1 mm) preprocessing method (C-index = 0.77 ± 0.05) (mean ± SD), 0.77–0.77 (95% CI)). The second best-performing combinations were MI/GLMB and MI/GLMN applied to the LOG (Sigma = 1 mm) preprocessing method (C-index = 0.77 ± 0.05, 0.76–0.77). Figure 6 shows Kaplan–Meier curves of the eight best models. All these models had significant log-rank p values in the training and testing datasets.
Statistical analysis
A comparison of the different preprocessing, FS, and ML methods are depicted in Fig. 7 as box plots (left) and Wilcoxon p value tile plots (right). For preprocessing methods (Fig. 7a), the LOG filter with Sigma of 1.0 mm had a higher significant p value (mean C-index = 0.65) than other methods. Wavelet transforms (HLH form) and LOG filter with Sigma of 2.5 with a mean C-index of 0.64 had a higher significant p value than other preprocessing methods (except LOG filter with Sigma = 1.0 mm). Figure 7b indicates that MI performed better (mean C-index = 0.70) than other FS methods. In the second place, the IBMA method, with a mean C-index of 0.65, performed better than other FS methods except for the MI method. Figure 7c demonstrates that the GLMB, and GLMN model with a mean C-index of 0.64 had a higher significant p value than other models. CoxPH is the second best model (mean C-index = 0.64) with a significantly higher p value than other models, except the GLMB and GLMN models.
Discussion
Our study demonstrated that ML algorithms are capable of predicting the time-to-event OS of patients through the use of MRI-based radiomic features. In an extensive systematic review, the 2-year OS of patients was comprehensively reviewed and analyzed considering a large number of articles [37]. It has been reported that the 2-year and 3-year OS of GBM patients is 18% and 11%, respectively. Other clinical trials indicated a 2-year survival of 26–33% and an approximate median OS of 15 months [37, 38]. Another study reported that patients with confirmed GBM had a 5.6% 5-year survival rate [1]. In our study, 77% of the patients decreased after 13.42 ± 11.91 (mean ± SD) months, while 23% were in the survivor group after 17.60 ± 15.45 months. We used seven distinct feature selectors and six ML methods applied to radiomic features derived from MR images preprocessed with twelve different methods. This leads to 504 models, the performance of which was assessed using the C-index. Many research and development efforts focused on assessing OS in GBM patients. The best model was the MI/CB applied to the LOG (Sigma = 1 mm) preprocessing method (C-index = 0.77 ± 0.05) (mean ± SD), 0.77–0.77 (95% CI)). LOG with Sigma = 1 mm, also known as the fine filter, had better performance, significantly higher than other methods. Among FS methods, MI had the best performance. Conversely, the GLMB, and GLMN models had significantly better performance than other ML algorithms.
Regarding the use of radiomics for OS prediction, the models perform decently and satisfactorily, according to the latest studies. Bakas et al. [26] used MRI-based radiomic features to predict survival status with 74.26% accuracy. In another study, it was claimed that PET-based radiomic features could contribute to the prognosis and indicate whether patients benefit from re-irradiation (p value for OS prediction < 0.05) [44]. Jajroudi et al. [39] examined the ability of MRI-extracted radiomic features to determine GBM survival. The mean percentage of sensitivity, accuracy, AUC, and specificity of the four implemented ML classifier was 82.54, 80.47, 85, and 79.78, respectively. In a retrospective ML/radiomics study, Cepeda et al. [40] reported that RSF achieved an integrated AUC (iAUC) of 0.76 and a C-Index of 0.61 with six months follow-up. Moreover, Chaddad et al. [41] found a decent predictive value of radiomics signature for determining patients’ survival time with an AUC of 78.56%. In addition, the radiomic features proved to be reliable and repeatable among different centers and datasets [45]. To estimate the survival time in GBM patients, Hedyehzadeh et al. [42] studied a dataset of 118 GBM patients. They computed C-index using eight traditional and three convolutional neural networks (CNN)-based regression techniques to predict survival periods. GoogleNet achieved the highest performance with a C-index of 0.89.
As mentioned earlier, we used only radiomic features and clinical variables for OS prediction. Similar to our approach, Ammari et al. [24] utilized radiomics and clinical data to stratify patients based on their survival. Using ML regression algorithms, they achieved a C-index of 0.64 for the prediction of OS. Although these decent results arise from relatively comprehensive models (radiomics + clinical and/or genomic data), our study could reach fair prediction accuracies using radiomics + clinical data. Chen et al. [43] showed that radiomics-only models performed better than clinical-only models. In our study, the mean C-index of the combination of preprocessing/FS/ML algorithms was in the range of 0.50–0.77 in the test dataset.
Nevertheless, several previous studies combined radiomics with features other than clinical for model construction. For example, a study by Bae et al. [23] extracted radiomic features and constructed a model using clinical, genetic, and radiomic features. They compared the model’s performance with another one consisting of clinical and genetic data and reported promising results demonstrating that adding radiomic features can significantly improve the OS and progression-free survival (PFS) prediction power of ML models with p values of 0.04 and 0.03, respectively. In addition, the iAUC of their model plus RSF algorithm was 0.65 (radiomics only) and 0.76 (radiomics + clinical) for OS prediction.
We compared the latest studies to ours in Table 4. These studies used random split and cross-validation for the validation of their models. None of these studies used external validation dataset. These studies used two types of models, including classification and time-to-event OS prediction. In our study, we used random split for model validation. We also used cross-validation for hyperparameter optimization and bootstrap on the testing dataset. While we used time-to-event OS prediction, the results of our best model is comparable with these studies.
We had four feature types for overall survival prediction in GBM patients. The number (percent) of clinical, shape, FO, and texture features selected by feature selectors were 113 (16%), 366 (53%), 43 (6%), and 175 (25%), respectively, more than half of which belonged to the shape category. We also had four VOIs for feature extraction. The number (percent) of features selected by feature selectors in VOIT, VOIA, VOIN, and VOIE was 207 (37%), 156 (28%), 54 (10%), and 141 (25%), respectively. The results indicated that core tumor (VOIT) in GBM patients greatly impacted survival prediction. MinorAxis from VOIT was the most repeated feature among the top selected features. Based on IBSI guidelines [34, 46], MajorAxis, MinorAxis, LeastAxis, Flatness, and Elongation are computed by principal component analysis (PCA) of VOI. MajorAxis, MinorAxis, and LeastAxis are the largest, second largest, and smallest eigenvalues, respectively, calculated by PCA [34, 46]. Flatness is the ratio of LeastAxis to MajorAxis, whereas Elongation is the ratio of MinorAxis to MajorAxis. In several studies [47, 48], MajorAxis had a significant log-rank p value. In our study, this feature was selected by UCI and IBMA feature selection techniques. While MinorAxis had no significant log-rank p value in a previous study [48], our study selected all feature selectors to cross several preprocessing sets. M2DDS of VOIT is the maximal diameter on the axial plane [34, 46], equivalent to Response Evaluation Criteria in Solid Tumors (RECIST) [49]. Galanis et al. [50] showed that this feature could predict OS, while our study selected 6 FSs. The sphericity of VOIT is another top-selected feature. This feature measured how the VOI is sphere-like [34, 46]. In Sanghani et al. [51] study, sphericity was predictive of OS prognosis. In another study [52], sphericity or surface regularity and age were predictors of overall survival, but the surface area had no significant p value. Figure 4 indicates that the shape feature of VOIT and VOIA had a Spearman’s coefficient of over 0.90. MinorAxis, M2DDS, M2DDC, and SurfaceArea from VOIA had the same interpretation as VOIT features. The Coarseness feature of NGTDM from VOIA implies that on the spatial rate of change, that lower value shows a higher spatial change rate and non-uniform texture [34, 46]. This pattern is an enhanced area of the tumor correlates with lower OS in GBM patients. Large Area High Gray Level Emphasis (LAHGLE) of GLSZM from VOIA was selected by 5 FSs. This feature emphasizes the high gray level with a large size zone [34, 46]. Larger values indicate lower OS, and those patients are at high risk. A/T volume ratio is the proportion of contrast-enhanced tumors. This feature was selected by 4 FSs and associated with OS in several studies [47, 48, 53]. In the clinical features, KPS and age were selected by 5 and 4 FSs, respectively. Gender was selected by MI and IBMA FS methods. Various studies reported that age and KPS were predictors for OS, but gender was not an OS predictor [54,55,56]. The PTA and RTA are two important variables that demonstrate how patients were treated. The RTA was selected by four FS methods, while the PTA was selected by MI and IBMA FS techniques. These two variables were selected by the MI method across all 12 feature sets, and this FS method outperformed all other strategies. A number of studies reported that using PTA and RTA with total resection improves survival in GBM patients [57,58,59]. As shown in Table 2, 93 (78%) of GBM patients received both treatments, while 22 (18%) received at least one treatment. Only four patients did not receive treatment.
Our study is not exempt from limitations. First of all, the nature of our study is retrospective, and as such, it is inherently prone to some bias. Secondly, we had no external validation, and our sample size was insufficient to expand our results' generalizability. To overcome this limitation, we used threefold cross-validation for hyperparameter optimization and 1000 bootstrap on the testing dataset. Last, we did not include genetic variables and tumor histology subtype in our modeling to compare it with radiomic features predictions.
Conclusion
This study showed that the radiomics framework plus clinical data could be utilized in the management of GBM patients. MRI-extracted radiomic and clinical features could predict the patient's time-to-event OS by applying feature selection methods and ML algorithms. MRI-based radiomics combined with clinical variables might be promising in terms of assisting clinicians in the survival prediction of patients with GBM.
Abbreviations
- CoxPH:
-
Cox Proportional Hazard regression
- CB:
-
Cox Boost
- FS:
-
Feature Selection
- GBM:
-
Glioblastoma Multiforme
- GB:
-
Gradient Boosting
- GLMB:
-
Generalized Linear Model Boosting
- GLMN:
-
Generalized Linear Model Network
- IBMA:
-
Iterated Bayesian Model Averaging
- LOG:
-
Laplacian of Gaussian
- MD:
-
Minimal Depth
- MI:
-
Mutual Information
- ML:
-
Machine Learning
- MRI:
-
Magnetic Resonance Imaging
- OS:
-
Overall Survival
- RSF:
-
Random Survival Forest
- TCIA:
-
The Cancer Imaging Archive
- UCI:
-
Univariate C-index
- VH:
-
Variable Hunting
- VH.VIMP:
-
Variable Hunting Variable Importance
- VOI:
-
Volume of Interest
References
Ostrom QT, Gittleman H, Truitt G, Boscia A, Kruchko C, Barnholtz-Sloan JS (2018) CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the United States in 2011–2015. Neuro Oncol 20(suppl_4):iv1–iv86
Tamimi AF, Juweid M (2017) Epidemiology and outcome of glioblastoma. Exon Publications, Brisbane City, pp 143–153
Bette S, Barz M, Huber T et al (2018) Retrospective analysis of radiological recurrence patterns in glioblastoma, their prognostic value and association to postoperative infarct volume. Sci Rep 8(1):4561
Czapski B, Baluszek S, Herold-Mende C, Kaminska B (2018) Clinical and immunological correlates of long term survival in glioblastoma. Contemp Oncol 22(1A):81
Tewarie IA, Senders JT, Kremer S et al (2021) Survival prediction of glioblastoma patients—are we there yet? A systematic review of prognostic modeling for glioblastoma and its clinical potential. Neurosurg Rev 44(4):2047–2057
Shukla G, Alexander GS, Bakas S et al (2017) Advanced magnetic resonance imaging in glioblastoma: a review. Chin Clin Oncol 6(4):40
Bohman LE, Swanson KR, Moore JL et al (2010) Magnetic resonance imaging characteristics of glioblastoma multiforme: implications for understanding glioma ontogeny. Neurosurgery 67(5):1319–1327 (discussion 27-8)
Itakura H, Achrol AS, Mitchell LA et al (2015) Magnetic resonance image features identify glioblastoma phenotypic subtypes with distinct molecular pathway activities. Sci Transl Med 7(303):303ra138
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278(2):563–577
Manafi-Farid R, Askari E, Shiri I et al (2022) [(18)F]FDG-PET/CT radiomics and artificial intelligence in lung cancer: technical aspects and potential clinical applications. Semin Nucl Med 52(6):759–780
Shiri I, Amini M, Nazari M et al (2022) Impact of feature harmonization on radiogenomics analysis: prediction of EGFR and KRAS mutations from non-small cell lung cancer PET/CT images. Comput Biol Med 142:105230
Hosseini SA, Hosseini E, Hajianfar G et al (2023) MRI-based radiomics combined with deep learning for distinguishing IDH-mutant WHO grade 4 astrocytomas from IDH-wild-type glioblastomas. Cancers 15(3):951
Sabouri M, Hajianfar G, Hosseini Z et al (2022) Myocardial perfusion SPECT imaging radiomic features and machine learning algorithms for cardiac contractile pattern recognition. J Digit Imaging 36:497–509
Moradi Khaniabadi P, Bouchareb Y, Al-Dhuhli H et al (2022) Two-step machine learning to diagnose and predict involvement of lungs in COVID-19 and pneumonia using CT radiomics. Comput Biol Med 150:106165
Shiri I, Mostafaei S, Haddadi Avval A et al (2022) High-dimensional multinomial multiclass severity scoring of COVID-19 pneumonia using CT radiomics features and machine learning algorithms. Sci Rep 12(1):14817
Khodabakhshi Z, Amini M, Mostafaei S et al (2021) Overall survival prediction in renal cell carcinoma patients using computed tomography radiomic and clinical information. J Digit Imaging 34(5):1086–1098
Taha B, Boley D, Sun J, Chen CC (2021) State of radiomics in glioblastoma. Neurosurgery 89(2):177–184
Wu G, Chen Y, Wang Y et al (2018) Sparse representation-based radiomics for the diagnosis of brain tumors. IEEE Trans Med Imaging 37(4):893–905
Gore S, Chougule T, Jagtap J, Saini J, Ingalhalikar M (2021) A review of radiomics and deep predictive modeling in glioma characterization. Acad Radiol 28(11):1599–1621
Wan Y, Zhou S, Zhang Y, Deng X, Xu L (2022) Radiomic analysis of contrast-enhanced MRI predicts DNA copy-number subtype and outcome in lower-grade gliomas. Acad Radiol 29(9):e189–e196
Artzi M, Bressler I, Ben BD (2019) Differentiation between glioblastoma, brain metastasis and subtypes using radiomics analysis. J Magn Reson Imaging 50(2):519–528
Buchlak QD, Esmaili N, Leveque JC, Bennett C, Farrokhi F, Piccardi M (2021) Machine learning applications to neuroimaging for glioma detection and classification: an artificial intelligence augmented systematic review. J Clin Neurosci 89:177–198
Bae S, Choi YS, Ahn SS et al (2018) Radiomic MRI phenotyping of glioblastoma: improving survival prediction. Radiology 289(3):797–806
Ammari S, Sallé de Chou R, Assi T et al (2021) Machine-learning-based radiomics MRI model for survival prediction of recurrent glioblastomas treated with bevacizumab. Diagnostics (Basel, Switzerland) 11(7):1263
Baid U, Rane SU, Talbar S et al (2020) Overall survival prediction in glioblastoma with radiomic features using machine learning. Front Comput Neurosci 14:61
Bakas S, Shukla G, Akbari H et al (2020) Overall survival prediction in glioblastoma patients using structural magnetic resonance imaging (MRI): advanced radiomic features may compensate for lack of advanced MRI modalities. J Med Imaging (Bellingham, Wash) 7(3):031505
Hajianfar G, Haddadi Avval A, Hosseini SA, Oveisi M, Shiri I, Zaidi H (2022) Machine learning based overall survival prediction in GBM patients using MRI radiomics. IEEE NSS/MIC
Clark K, Vendt B, Smith K et al (2013) The Cancer Imaging Archive (TCIA): maintaining and operating a public information repository. J Digit Imaging 26(6):1045–1057
Scarpace L, Mikkelsen T, Cha S, Rao S, Tekchandani S, Gutman D, Saltz JH, Erickson BJ, Pedano N, Flanders AE, Barnholtz-Sloan J, Ostrom Q, Barboriak D, Pierce LJ (2016) The Cancer Genome Atlas Glioblastoma Multiforme Collection (TCGA-GBM) (Version 4). The Cancer Imaging Archive
Mor V, Laliberte L, Morris JN, Wiemann M (1984) The Karnofsky performance status scale: an examination of its reliability and validity in a research setting. Cancer 53(9):2002–2007
Depeursinge A, Andrearczyk V, Whybra P et al (2020) Standardised convolutional filtering for radiomics. arXiv preprint arXiv:2006.05470
Fedorov A, Beichel R, Kalpathy-Cramer J et al (2012) 3D slicer as an image computing platform for the quantitative imaging network. Magn Reson Imaging 30(9):1323–1341
Van Griethuysen JJ, Fedorov A, Parmar C et al (2017) Computational radiomics system to decode the radiographic phenotype. Cancer Res 77(21):e104–e107
Zwanenburg A, Vallières M, Abdalah MA et al (2020) The image biomarker standardization initiative: standardized quantitative radiomics for high-throughput image-based phenotyping. Radiology 295(2):328
Amini M, Hajianfar G, Hadadi Avval A et al (2022) Overall survival prognostic modelling of non-small cell lung cancer patients using positron emission tomography/computed tomography harmonised radiomics features: the quest for the optimal machine learning algorithm. Clin Oncol (R Coll Radiol) 34(2):114–127
Team RC (2021) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Poon MTC, Sudlow CLM, Figueroa JD, Brennan PM (2020) Longer-term (≥ 2 years) survival in patients with glioblastoma in population-based studies pre- and post-2005: a systematic review and meta-analysis. Sci Rep 10(1):11622
Gilbert MR, Wang M, Aldape KD et al (2013) Dose-dense temozolomide for newly diagnosed glioblastoma: a randomized phase III clinical trial. J Clin Oncol 31(32):4085–4091
Jajroudi M, Enferadi M, Homayoun AA, Reiazi R (2022) MRI-based machine learning for determining quantitative and qualitative characteristics affecting the survival of glioblastoma multiforme. Magn Reson Imaging 85:222–227
Cepeda S, Perez-Nuñez A, Garcia-Garcia S et al (2021) Predicting short-term survival after total resection in glioblastomas by machine learning-based radiomic analysis of preoperative MRI. Cancers (Basel) 13(20):5047
Chaddad A, Daniel P, Desrosiers C, Toews M, Abdulkarim B (2019) Novel radiomic features based on joint intensity matrices for predicting glioblastoma patient survival time. IEEE J Biomed Health Inform 23(2):795–804
Hedyehzadeh M, Pu J, Leilizadeh S, Gezer S, Dresser C, Beeche CA. A comparison of deep and conventional regression methods for mri-based estimation of survival time in GBM patients. Available at SSRN 4001112
Chen X, Fang M, Dong D et al (2019) Development and validation of a MRI-based radiomics prognostic classifier in patients with primary glioblastoma multiforme. Acad Radiol 26(10):1292–1300
Carles M, Popp I, Starke MM et al (2021) FET-PET radiomics in recurrent glioblastoma: prognostic value for outcome after re-irradiation? Radiat Oncol 16(1):46
Shiri I, Hajianfar G, Sohrabi A et al (2020) Repeatability of radiomic features in magnetic resonance imaging of glioblastoma: test–retest and image registration analyses. Med Phys 47(9):4265–4280
Zwanenburg A, Leger S, Vallières M, Löck S (2016) Image biomarker standardisation initiative. arXiv preprint arXiv:1612.07003
Gutman DA, Cooper LA, Hwang SN et al (2013) MR imaging predictors of molecular profile and survival: multi-institutional study of the TCGA glioblastoma data set. Radiology 267(2):560
Mazurowski MA, Zhang J, Peters KB, Hobbs H (2014) Computer-extracted MR imaging features are associated with survival in glioblastoma patients. J Neurooncol 120(3):483–488
Eisenhauer EA, Therasse P, Bogaerts J et al (2009) New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur J Cancer 45(2):228–247
Galanis E, Buckner JC, Maurer MJ et al (2006) Validation of neuroradiologic response assessment in gliomas: measurement by RECIST, two-dimensional, computer-assisted tumor area, and computer-assisted tumor volume methods. Neuro Oncol 8(2):156–165
Sanghani P, Ti AB, King NKK, Ren H (2019) Evaluation of tumor shape features for overall survival prognosis in glioblastoma multiforme patients. Surg Oncol 29:178–183
Pérez-Beteta J, Molina-García D, Ortiz-Alhambra JA et al (2018) Tumor surface regularity at MR imaging predicts survival and response to surgery in patients with glioblastoma. Radiology 288(1):218–225
Nicolasjilwan M, Hu Y, Yan C et al (2015) Addition of MR imaging features and genetic biomarkers strengthens glioblastoma survival prediction in TCGA patients. J Neuroradiol 42(4):212–221
Ingrisch M, Schneider MJ, Nörenberg D et al (2017) Radiomic analysis reveals prognostic information in T1-weighted baseline magnetic resonance imaging in patients with glioblastoma. Investig Radiol 52(6):360–366
Prasanna P, Patel J, Partovi S, Madabhushi A, Tiwari P (2017) Radiomic features from the peritumoral brain parenchyma on treatment-naive multi-parametric MR imaging predict long versus short-term survival in glioblastoma multiforme: preliminary findings. Eur Radiol 27(10):4188–4197
Zinn PO, Sathyan P, Mahajan B et al (2012) A novel volume-age-KPS (VAK) glioblastoma classification identifies a prognostic cognate microRNA-gene signature. PLoS ONE 7(8):e41522
Stupp R, Dietrich PY, Ostermann Kraljevic S et al (2002) Promising survival for patients with newly diagnosed glioblastoma multiforme treated with concomitant radiation plus temozolomide followed by adjuvant temozolomide. J Clin Oncol 20(5):1375–1382
Hamisch C, Ruge M, Kellermann S et al (2017) Impact of treatment on survival of patients with secondary glioblastoma. J Neurooncol 133(2):309–313
Salcman M (1980) Survival in glioblastoma: historical perspective. Neurosurgery 7(5):435–439
Acknowledgements
The results shown here are in whole or part based upon data generated by the TCGA Research Network: http://cancergenome.nih.gov/.
Funding
Open access funding provided by University of Geneva. This work was supported by the Swiss National Science Foundation under Grant SNRF 320030_176052.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no potential conflicts of interest.
Ethical statement
The study was conducted using the TCIA dataset (https://cancerimagingarchive.net), and as such, ethics approval and participants consent were not required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hajianfar, G., Haddadi Avval, A., Hosseini, S.A. et al. Time-to-event overall survival prediction in glioblastoma multiforme patients using magnetic resonance imaging radiomics. Radiol med 128, 1521–1534 (2023). https://doi.org/10.1007/s11547-023-01725-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11547-023-01725-3