Abstract
Gene targeting is useful to isolate strains with mutations in a gene of interest for efficient breeding. In this study, we generated msh4 or mer3 single-gene disruptant monokaryons using a Pleurotus ostreatus Δku80 strain for efficient gene targeting. Dikaryons of P. ostreatus Δmsh4×Δmsh4 or Δmer3×Δmer3 were isolated via backcrosses, and the number of basidiospores produced was measured. The number of basidiospores fell by an average 1/13.7 in the P. ostreatus Δmsh4×Δmsh4 dikaryons versus the P. ostreatus msh4+×Δmsh4 dikaryons, and 1/82.6 in the P. ostreatus Δmer3×Δmer3 dikaryons versus the P. ostreatus mer3+×Δmer3 dikaryons. To demonstrate the effects of ku80 disruption, P. ostreatus Δku80×Δku80 dikaryon strains were isolated and no significant effects on basidiospore production were observed. Fluorescence microscopy showed meiotic progression was arrested during prophase I in the msh4 or mer3 disruptants. To our knowledge, this is the first report on molecular breeding of sporeless strains in cultivated mushrooms using an efficient method for targeted gene disruption.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Molecular breeding is a promising breeding strategy because one can isolate strains with desired phenotypes directly. Gene targeting is especially powerful for isolating strains with modifications in a gene of interest for efficient breeding purposes. Moreover, it is also effective to characterize the gene function for research purposes. However, the targeted molecular breeding of a cultivated mushroom strain is not common because of the low gene-targeting efficiency.
An efficient technique for gene targeting by homologous recombination was established in filamentous fungi by disrupting one or two of the genes encoding Ku70/Ku80 and Ligase 4 (Ruiz-Díez 2002; Ninomiya et al. 2004; Levy et al. 2008; de Jong et al. 2010; Kück and Hoff 2010; Nakazawa et al. 2011). These proteins play a major role in the non-homologous DNA end-joining (NHEJ) system, and their inactivation makes homologous recombination the relatively dominant repair system in the mycelia, which leads to the establishment of high-frequency gene targeting. The oyster mushroom Pleurotus ostreatus is one of the most economically important edible mushrooms in the world, and an efficient gene targeting was developed for the first time in the cultivated mushrooms by isolating a P. ostreatus Δku80 monokaryon (Salame et al. 2012).
A strain with low production of basidiospores is an important breeding target in the mushroom industry as the dispersal of basidiospores into cultivation facilities causes allergic reactions in workers/handlers as well as facility disruption (Baars et al. 2000). The msh4 and mer3 genes encode proteins required for meiosis (Snowden et al. 2004; Sugawara et al. 2009), and their inactivation is reported to cause a deficiency in spore formation in Coprinopsis cinerea and Pleurotus pulmonarius (Sugawara et al. 2009; Okuda et al. 2013). In this study, we generated msh4 or mer3 disruptants by gene knock out using the Δku80 strain of P. ostreatus and analyzed the effects on sporulation to demonstrate targeted molecular breeding of cultivated mushrooms.
Furthermore, it was suggested that NHEJ might be involved in the meiotic process of eukaryotes such as mouse and Coprinopsis cinerea (Namekawa et al. 2003; Liebe et al. 2006; Okuda et al. 2013). Disruption of ku80 may cause defects in sporulation in P. ostreatus; therefore, the effects of ku80 disruption on basidiospore production were also analyzed.
Materials and methods
Strains, culture conditions, and genetic techniques
P. ostreatus strains used in this study are listed in Table 1. Yeast and malt extract with glucose (YMG) medium (Rao and Niederpruem 1969) solidified with 2% (w/v) agar in 9-cm Petri dishes was used for routine cultures. The strains were grown at 28 °C under continuous darkness. The cultures were maintained at 4 °C for long-term storage. Dikaryotic strains of P. ostreatus were also grown on sawdust (Fagus crenata) media with 7.5% (w/w) wheat bran supplementation to induce fruiting body formation as described by Nakazawa et al. (2016). Sawdust and wheat bran were purchased from Shinkoen (Gifu, Japan) and Nisshin Seifun (Tokyo, Japan). Crosses and fruiting of P. ostreatus were performed as described by Inada et al. (2001) and Nakazawa et al. (2016), respectively. Dikaryon formation was confirmed by observation of clamp cells 6 days after mating.
Construction of msh4- and mer3-disrupting plasmids
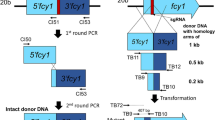
To generate a msh4-disrupting plasmid, upstream (1477 bp) and downstream (1452 bp) fragments of the msh4 gene, corresponding to Protein ID 65320 [msh4; Scaffold_19: 5841-9704 of P. ostreatus PC9 (https://mycocosm.jgi.doe.gov/PleosPC9_1/PleosPC9_1.home.html)], were amplified using a primer pair FY18/FY19 and FY20/FY21 (Table 2) from P. ostreatus 20b. A genomic fragment containing the hygromycin-B-resistance gene (hph) was also amplified using primers TN400/M13R from pTN24-1 (Nakazawa et al. 2017b). These three fragments were fused by overlap-extension polymerase chain reaction (PCR). The resulting msh4-disrupting cassette was cloned into pBluescript II KS+ digested with EcoRV, yielding pBS-Δmsh4 (Fig. 1a).
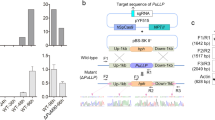
Targeted disruption of msh4 or mer3. a A diagram illustrating the procedure for the construction of pBS-Δmsh4. The red box indicates the 5′ region (1477 bp), and green the 3′ region (1452 bp) of the msh4 gene. The sequence of msh4-disrupting cassette is shown in Online Resource 1. b A diagram illustrating the procedure for the construction of pBS-Δmer3. The red box indicates the 5′-side ORF of the mer3 gene (1539 bp), and the green one the 3′-side ORF of the mer3 gene (1525 bp). The sequence of mer3-disrupting cassette is shown in Online Resource 2. c A diagram of the genomic locus of msh4. Arrows indicate the primers used for the PCR experiments in e. d A diagram of the genomic locus of mer3. Arrows indicate the primers used for the PCR experiments in f. e PCR experiments confirming msh4 disruption. Lane 1, negative control (genomic DNA was not added); Lane 2, P. ostreatus 20b; Lane 3, P. ostreatus Δmsh4#1; and Lanes M represent size markers: λHindIII marker on the upper, Ladder marker on the lower left, and another Ladder marker on the lower right. f Genomic PCR experiments confirming the mer3 disruption. Lane 1, negative control (genomic DNA was not added); Lane 2, P. ostreatus 20b; Lane 3, P. ostreatus Δmer3#1; and Lanes M on the left and right sides represent size markers: λHindIII marker on the upper, Ladder marker on the lower left, and another Ladder marker on the lower right
To generate a mer3-disrupting plasmid, a genomic fragment of the mer3 gene, corresponding to Protein ID 82484 [mer3; Scaffold_1: 2926299-2932521 of P. ostreatus PC9], was amplified using a primer pair FY31/FY34 and cloned into pBluescript II KS+ digested with EcoRV. Inverse PCR was performed using a primer pair FY32/FY33. A genomic fragment containing the hygromycin-B-resistance gene was also amplified using a primer pair TN400/M13R from pTN24-1. These two DNA fragments were fused using the Geneart Seamless Cloning and Assembly kit (Life Technologies, CA, USA). The resulting plasmid containing a mer3-disrupting cassette was designated as pBS-Δmer3 (Fig. 1b). The cassettes were amplified from pBS-Δmsh4 and pBS-Δmer3 using a primer pair FY18/FY21 and FY31/FY34, respectively, when used for transformation.
PCR experiments were performed using a KOD FX Neo (Toyobo, Japan) with the following program: an initial denaturating step at 94 °C for 2 min, followed by 35 cycles of 10 s at 98 °C, 30 s at 58 °C, and 30 s/kb at 68 °C, and a final extension step at 68 °C for 5 min.
Isolation of msh4 or mer3 single-gene disruptants
A mixture of pBS-Δmsh4 and msh4-disrupting cassette (about 2 μg and 1 μg, respectively) was introduced into protoplasts of P. ostreatus 20b (about 5 × 107) to obtain the disruptant of msh4. A mixture of pBS-Δmer3 and mer3-disrupting cassette (about 2 μg and 1 μg, respectively) was also introduced into protoplasts of P. ostreatus 20b (about 5 × 107) to obtain the disruptant of mer3. The hygromycin resistance transformation of P. ostreatus 20b, a Δku80 derivative of P. ostreatus PC9, was performed using protoplasts prepared from mycelial cells as described by Salame et al. (2012), except that heparin and single-strand lambda phage DNA were not used as described by Nakazawa et al. (2016).
Extraction of genomic DNA and PCR
Genomic DNA was extracted from each strain as described by Tsukihara et al. (2006) and Nakazawa et al. (2016), followed by genomic PCR using an EmeraldAmp MAX PCR Master Mix (TAKARA BIO, Japan). The program was 35 cycles of 10 s at 94 °C, 30 s at 58 °C, and 1 min/kb at 72 °C.
Basidiospore measurement
Fruiting bodies of each dikaryotic strain were formed on plastic-bottled sawdust media (Nakazawa et al. 2019). Bottles with fruiting bodies were placed on the center of empty Petri dishes. After 1 day (Table 3) or 3 days (Table 4), basidiospores falling from the fruiting body onto the Petri dish were collected in a 1.5-mL tube using 1 mL water. After that, the number of basidiospores was counted using a hemocytometer.
Fluorescence microscopy
For observation of basidia, gills were fixed with 4% formaldehyde in phosphate-buffered saline (PBS) (10 mM sodium phosphate, 150 mM NaCl, pH 7.5) for 2 h. Next, samples were stained with 4,6-diamidino-2-phenylindole (DAPI) after washing several times with PBS and observed under an AxioScope. A1 fluorescence microscope (ZEISS, Germany) equipped with a filter set 91 HE.
Results
Generation of msh4 or mer3 disruptants from P. ostreatus strain 20b
Approximately 10 hygromycin-resistant Δmsh4 or Δmer3 transformants were obtained after the transformation of P. ostreatus 20b with the respective disruption cassette/plasmid mixture. PCR-amplifications using primer sets A-D were used to determine whether msh4 or mer3 gene had been replaced with the respective hph cassette. This way, for the Δmsh4 transformants, disruption of Δmsh4 could be proven in one out of one transformant, and for the Δmer3 transformants, disruption of Δmer3 could be proven in three out of five transformants (Fig. 1c, d, e, and f). Primer sets A and B yielded amplicons from the gene disruptants, but not from P. ostreatus 20b. Primer set C produced an amplicon from P. ostreatus 20b, but not from the gene disruptants. As shown in Fig. 1c and e, the fragment was amplified from the transformant when primer set A (FY17/TN48) or B (TN378/FY22) was used, but not when primer set C (FY23/FY24) was used. In addition, it was expected that a 6864-bp genomic fragment was PCR-amplified from P. ostreatus 20b and 5332-bp from the transformant when primer set D (FY17/FY22) was used (Fig. 1c). As shown in Fig. 1e, the expected results were obtained from this genomic PCR experiment, although the possibility of the ectopic integration of the hph cassette could not be ruled out. These results indicate that the msh4 gene is replaced with the hph cassette in the transformant, which was designated as P. ostreatus Δmsh4#1. Similarly, the mer3 gene was shown to be disrupted in three out of the five transformants. PCR results for the mer3 disruptant, P. ostreatus Δmer3#1, are shown in Fig. 1d and f.
Effects of msh4 or mer3 disruption on basidiospore production in P. ostreatus
Dikaryotic strains P. ostreatus Δmsh4×Δmsh4, msh4+×Δmsh4, Δmer3×Δmer3, and mer3+×Δmer3 were obtained to measure the number of basidiospores. To create the dikaryons, P. ostreatus Δmsh4#1 was crossed with P. ostreatus #64, and P. ostreatus Δmer3#1 was crossed with P. ostreatus PC15 or P. ostreatus #64 (Table 1). F1 strains were obtained from each cross (P. ostreatus Δmsh4#1×#64, Δmer3#1×PC15, and Δmer3#1×#64). F1 progeny exhibiting resistance to hygromycin B (hygR) are considered as Δmsh4 or Δmer3, and those that were sensitive to hygromycin B (hygS) as msh4+ or mer3+. The F1 strains were mated with P. ostreatus Δmsh4#1 or Δmer3#1 to generate dikaryons (Table 1; three P. ostreatus msh4+×Δmsh4, four Δmsh4×Δmsh4, two mer3+×Δmer3, and five Δmer3×Δmer3). The number of basidiospores produced by fruiting bodies from the resulting dikaryotic strains was measured to analyze the effects of msh4 and mer3 disruption. The number of basidiospores fell by an average 1/13.7 in the P. ostreatus Δmsh4×Δmsh4 dikaryons versus the P. ostreatus msh4+×Δmsh4 dikaryons, and 1/82.6 in the P. ostreatus Δmer3×Δmer3 dikaryons versus the P. ostreatus mer3+×Δmer3 dikaryons (Table 3).
To determine the significance of these differences, a statistical analysis using the two-tailed t test was performed. The difference between P. ostreatus mer3+×Δmer3 and Δmer3×Δmer3 was shown to be significant (P = 0.0090). In the case of msh4, the difference was not significant (P = 0.081). The number of basidiospores produced by one of the four strains tested in this study (P. ostreatus Δmsh4×Δmsh4#2) was 1.32 × 108 and significantly differed from the other P. ostreatus Δmsh4×Δmsh4 strains.
Disruption of ku80 does not cause a decrease in the number of basidiospores in P. ostreatus
Effects of ku80 disruption on basidiospore production were also analyzed using the F1 progeny obtained from P. ostreatus Δmsh4#1×PC15 and P. ostreatus Δmsh4#1×#64. F1 progeny exhibiting resistance to carboxin (cbxR) are considered Δku80, and those sensitive to carboxin (cbxS) as ku80+. We attempted to obtain F1 progeny with A64B64, cbxR and hygS backgrounds from P. ostreatus Δmsh4#1×#64; however, we could not. Therefore, a monokaryon with A64B64, cbxR and hygR backgrounds, which is designated as P. ostreatus m4#1 (Table 1), was used in the following experiments. P. ostreatus m4#1 was mated with 20 cbxS/hygS and five cbxR/hygS F1 strains from P. ostreatus Δmsh4#1×PC15 (the former ones were designated as P. ostreatus Δku80Δmsh4×ku80+msh4+#1–#20 and the latter were designated P. ostreatus Δku80Δmsh4×Δku80msh4+#1–#5). As shown in Table 4, there was no significant difference in the number of basidiospores between P. ostreatus ku80+×Δku80 and Δku80×Δku80 (P = 0.092), suggesting that disruption of ku80 does not cause significant defects in P. ostreatus sporulation.
Meiotic progression is arrested during prophase I in the msh4 and mer3 single-gene disruptants
To observe meiotic progress in basidia, the gills were stained with DAPI and observed under a fluorescence microscope. The meiotic stages in a basidium are roughly divided into four stages in C. cinerea, a model agaric related to P. ostreatus: karyogamy, meiosis I, meiosis II, and sporulation (Fig. 2a; Sugawara et al. 2009). Basidia with tetrad nuclei at telophase II (Fig. 2b: 1, 2, and 5) and those with two nuclei at telophase I were observed in the P. ostreatus 20b×PC15, msh4+×Δmsh4, and mer3+×Δmer3 strains, but not in those of P. ostreatus Δmsh4×Δmsh4 and Δmer3×Δmer3 ones. In the P. ostreatus Δmsh4×Δmsh4 and Δmer3×Δmer3 strains, basidia with one large nucleus at prophase I (Fig. 2b: 3 and 6), which were not observed in the P. ostreatus 20b×PC15, msh4+×Δmsh4, and mer3+×Δmer3 strains, and those with no fluorescence after DAPI staining (Fig. 2b: 4 and 7) were observed.
Meiosis of P. ostreatus Δmsh4×Δmsh4 and Δmer3×Δmer3 basidia arrested at prophase I. a Four stages in meiosis of basidium: karyogamy, meiosis I, meiosis II, and sporulation (Sugawara et al. 2009). b Observation of basidia by fluorescence microscope. The scale bar represents 10 μm. 1: the wild type is P. ostreatus 20b×PC15. Normal tetrad nuclei in the basidium. 2: normal tetrad nuclei in a basidium of the P. ostreatus msh4+×Δmsh4. 3: one large nucleus in a basidium of the P. ostreatus Δmsh4×Δmsh4. 4: no nuclei in a basidium of the P. ostreatus Δmsh4×Δmsh4. 5: normal tetrad nuclei in a basidium of the P. ostreatus mer3+×Δmer3. 6: one large nucleus in a basidium of the P. ostreatus Δmer3×Δmer3. 7: no nuclei in a basidium of the P. ostreatus Δmer3×Δmer3
Discussion
Strains defective in the sporulation processes have been reported in other agaricomycetes. In Pleurotus pulmonarius, mutations in stpp1, an msh4 homolog, meiotic progression was arrested during prophase I, and basidiospore production reduced to less than 1/1000 of the wild-type control (Okuda et al. 2013). In C. cinerea, the expression of mer3, which encodes a putative protein involved in the formation of meiotic crossover (Nakagawa and Ogawa 1999), was suppressed by RNAi. As a result, meiosis progressed only to nuclear fusion, and then apoptosis was observed, and the mer3-suppressed homokaryotic strain had a basidiospore production of 1/125 compared with the wild-type strain (Sugawara et al. 2009).
In this study, the number of basidiospores fell by an average 1/13.7 in the P. ostreatus Δmsh4×Δmsh4 dikaryons versus the P. ostreatus msh4+×Δmsh4 dikaryons, and 1/82.6 in the P. ostreatus Δmer3×Δmer3 dikaryons versus the P. ostreatus mer3+×Δmer3 dikaryons (Table 3). However, the t test indicated that the effect of msh4 was not significant. These results suggest that both single-gene disruptions of mer3 and msh4 cause negative effects on sporulation in P. ostreatus, and Mer3 plays more crucial roles than Msh4 does as constant reduction in the numbers of basidiospores from different P. ostreatus Δmer3×Δmer3 strains whereas the numbers were reduced but varied widely in the P. ostreatus Δmsh4×Δmsh4 strains possibly depending on their genetic backgrounds.
Basidia harboring two nuclei were not observed in the P. ostreatus Δmsh4×Δmsh4 and Δmer3×Δmer3 strains. Moreover, one large nucleus was observed in some of these basidia (Fig. 2b: 3 and 6). These results suggest that an inactivation of msh4 or mer3 causes a defect of basidiospore formation in P. ostreatus due to arrest of meiotic progression during prophase I. A similar effect was reported in Schizophyllum commune mutants with constitutive activation of Ras1 (Knabe et al. 2013). Therefore, examining and comparing the effects of the constitutive activation of Ras1 in P. ostreatus to those in S. commune would be interesting to study in the future. Basidia without nuclei were also observed in the P. ostreatus Δmsh4×Δmsh4 and Δmer3×Δmer3 strains (Fig. 2b: 4 and 7). In the case of C. cinereal, where meiotic progression is synchronized (Kües 2000), apoptosis of fused nuclei was observed in basidia over time (Sugawara et al. 2009). However, it is not easy to distinguish whether the basidium without a nucleus is the result of nuclear apoptosis or the dispersal of sound basidiospores of P. ostreatus; since meiosis is asynchronous in P. ostreatus, basidia with various meiotic stages, but not a specific one, were observed at a certain time. However, taking together with very low basidiospore production in the P. ostreatus Δmsh4×Δmsh4 and Δmer3×Δmer3 strains, apoptosis of nuclei may occur in the meiosis arrested basidia in this fungus.
A few basidiospores were produced in some P. ostreatus Δmsh4×Δmsh4 or Δmer3×Δmer3 strains, indicating that basidiospore formation is not impaired completely, but maybe significantly delayed in these strains due to the defective meiosis. It is of interest whether double-gene disruption of msh4 and mer3 would completely inhibit basidiospore formation in this fungus.
In mice, it was reported that defects in NHEJ affect early prophase I in the meiosis (Liebe et al. 2006). In C. cinereal, it was indicated that Lig4 was expressed not only at the pre-meiotic S phase but also at meiotic prophase I (Namekawa et al. 2003). Considering these studies, NHEJ might have a function in the meiotic process in these organisms. However, in the present study, there was no significant difference in the number of basidiospores between P. ostreatus ku80+×Δku80 and Δku80×Δku80 dikaryons (Table 4), suggesting that disruption of ku80 does not cause significant defects in sporulation in P. ostreatus.
In conclusion, strains with low production of spores were successfully generated using gene targeting in P. ostreatus. The msh4 and mer3 disruptant strains isolated in this study showed impaired meiosis that may cause low production of basidiospores. It is also demonstrated that disruption of ku80 has a negligible effect on basidiospore production in this fungus.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Baars JJP, Sonnenberg ASM, Mikosch TSP, Van Griensven LJLD (2000) Development of a sporeless strain of oyster mushroom Pleurotus ostreatus. In: Gnensven V (ed) Science and cultivation of edible fungi. Balkema, Rotterdam, pp 317–323
de Jong JF, Ohm RA, de Bekker C, Wösten HAB, Lugones LG (2010) Inactivation of ku80 in the mushroom-forming fungus Schizophyllum commune increases the relative incidence of homologous recombination. FEMS Microbiol Lett 310:91–95. https://doi.org/10.1111/j.1574-6968.2010.02052.x
Inada K, Morimoto Y, Arima T, Murata Y, Kamada T (2001) Homologous expression of recombinant manganese peroxidase genes in ligninolytic fungus Pleurotus ostreatus. Appl Microbiol Biotechnol 65:287–294. https://doi.org/10.1007/s002530000540
Knabe N, Jung EM, Freihorst D, Hennicke F, Horton JS, Kothe E (2013) A central role for Ras1 in morphogenesis of the Basidiomycete Schizophyllum commune. Eukaryot Cell 12:941–952. https://doi.org/10.1128/EC.00355-12
Kück U, Hoff B (2010) New tools for the genetic manipulation of filamentous fungi. Appl Microbiol Biotechnol 86:51–62. https://doi.org/10.1007/s00253-009-2416-7
Kües U (2000) Life history and developmental processes in the Basidiomycete Coprinus cinereus. Microbiol Mol Biol Rev 64:316–353. https://doi.org/10.1128/MMBR.64.2.316-353.2000
Larraya LM, Perez G, Pe nas MM, Baars JJP, Mikosch TSP, Pisabarro AG, Ramirez L (1999) Molecular karyotype of the white rot fungus Pleurotus ostreatus. Appl Environ Microbiol 65:3413–3417
Levy M, Erental A, Yarden O (2008) Efficient gene replacement and direct hyphal transformation in Sclerotinia sclerotiorum. Mol Plant Pathol 9:719–725. https://doi.org/10.1111/J.1364-3703.2008.00483.X
Liebe B, Petukhova G, Barchi M, Bellani M, Braselmann H, Nakano T, Pandita TK, Jasin M, Fornace A, Meistrich ML, Baarends WM, Schimenti J, de Lange T, Keeney S, Camerini-Otero RD, Scherthan H (2006) Mutations that affect meiosis in male mice influence the dynamics of the mid-preleptotene and bouquet stages. Exp Cell Res 312:3768–3781. https://doi.org/10.1016/j.yexcr.2006.07.019
Nakagawa T, Ogawa H (1999) The Saccharomyces cerevisiae MER3 gene, encoding a novel helicase-like protein, is required for crossover control in meiosis. EMBO J 18:5714–5723. https://doi.org/10.1093/emboj/18.20.5714
Nakazawa T, Ando Y, Kitaaki K, Nakahori K, Kamada T (2011) Efficient gene targeting in ΔCc.ku70 or ΔCc.lig4 mutants of the agaricomycete Coprinopsis cinerea. Fungal Genet Biol 48:939–946. https://doi.org/10.1016/j.fgb.2011.06.003
Nakazawa T, Tsuzuki M, Irie T, Sakamoto M, Honda Y (2016) Marker recycling via 5-fluoroorotic acid and 5-fluorocytosine counter-selection in the white-rot agaricomycete Pleurotus ostreatus. Fungal Biol 120:1146–1155. https://doi.org/10.1016/j.funbio.2016.06.011
Nakazawa T, Izuno A, Kodera R, Miyazaki Y, Sakamoto M, Isagi Y, Honda Y (2017a) Identification of two mutations that cause defects in the ligninolytic system through an efficient forward genetics in the white-rot agaricomycete Pleurotus ostreatus. Environ Microbiol 19:261–272. https://doi.org/10.1111/1462-2920.13595
Nakazawa T, Izuno A, Horii M, Kodera R, Nishimura H, Hirayama Y, Tsunematsu Y, Miyazaki Y, Awano T, Muraguchi H, Watanabe K, Sakamoto M, Takabe K, Watanabe T, Isagi Y, Honda Y (2017b) Effects of pex1 disruption on wood lignin biodegradation, fruiting development and the utilization of carbon sources in the white-rot Agaricomycete Pleurotus ostreatus and non-wood decaying Coprinopsis cinerea. Fungal Genet Biol 109:7–15. https://doi.org/10.1016/j.fgb.2017.10.002
Nakazawa T, Morimoto R, Wu HL, Kodera R, Sakamoto M, Honda Y (2019) Dominant effects of gat1 mutations on the ligninolytic activity of the white-rot fungus Pleurotus ostreatus. Fungal Biol 123:209–217. https://doi.org/10.1016/j.funbio.2018.12.007
Namekawa S, Ichijima Y, Hamada F, Kasai N, Iwabata K, Nara T, Teraoka H, Sugawara F, Sakaguchi K (2003) DNA ligase IV from a basidiomycete, Coprinus cinereus, and its expression during meiosis. Microbiol 149:2119–2128. https://doi.org/10.1099/mic.0.26311-0
Ninomiya Y, Suzuki K, Ishii C, Inoue H (2004) Highly efficient gene replacements in Neurospora strains deficient for nonhomologous end-joining. PNAS 101:12248–12253. https://doi.org/10.1073/pnas.0407377101
Okuda Y, Murakami S, Honda Y, Matsumoto T (2013) An MSH4 homolog, stpp1, from Pleurotus pulmonarius is a “silver bullet” for resolving problems caused by spores in cultivated mushrooms. Appl Environ Microbiol 79:4520–4527. https://doi.org/10.1128/AEM.00561-13
Rao PS, Niederpruem DJ (1969) Carbohydrate metabolism during morphogenesis of Coprinus lagopus (sensu Buller). J Bacteriol 100:1222–1228. https://doi.org/10.1128/JB.100.3.1222-1228.1969
Ruiz-Díez B (2002) Strategies for the transformation of filamentous fungi. J Appl Microbiol 92:189–195. https://doi.org/10.1046/j.1365-2672.2002.01516.x
Salame TM, Knop D, Tal D, Levinson D, Yarden O, Hadar Y (2012) Predominance of a versatile-peroxidase-encoding gene, mnp4, as demonstrated by gene replacement via a gene targeting system for Pleurotus ostreatus. Appl Environ Microbiol 78:5341–5352. https://doi.org/10.1128/AEM.01234-12
Snowden T, Acharya S, Butz C, Berardini M, Fishel R (2004) hMSH4-hMSH5 recognizes Holliday junctions and forms a meiosis-specific sliding clamp that embraces homologous chromosomes. Mol Cell 15:437–451. https://doi.org/10.1016/j.molcel.2004.06.040
Sugawara H, Iwabata K, Koshiyama A, Yanai T, Daikuhara Y, Namekawa S, Hamada F, Sakaguchi K (2009) Coprinus cinereus Mer3 is required for synaptonemal complex formation during meiosis. Chromosoma 118:127–139. https://doi.org/10.1007/s00412-008-0185-1
Tsukihara T, Honda Y, Sakai R, Watanabe T (2006) Exclusive overproduction of recombinant versatile peroxidase MnP2 by genetically modified white rot fungus, Pleurotus ostreatus. J Biotechnol 126:431–439. https://doi.org/10.1016/j.jbiotec.2006.05.013
Acknowledgments
We thank Prof. Yitzhak Hadar and Dr. Tomer M, Salame (Hebrew University of Jerusalem) for providing P. ostreatus strain 20b.
Funding
This study was supported in part by JSPS KAKENHI (Grant No. 18KK0178 to Y.H.).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Fuga Yamasaki, Takehito Nakazawa and Yoichi Honda. The first draft of the manuscript was written by Fuga Yamasaki and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Section Editor: Martin Rühl
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yamasaki, F., Nakazawa, T., Sakamoto, M. et al. Molecular breeding of sporeless strains of Pleurotus ostreatus using a non-homologous DNA end-joining defective strain. Mycol Progress 20, 73–81 (2021). https://doi.org/10.1007/s11557-020-01661-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-020-01661-w