Abstract
Fulvifomes (Hymenochaetaceae) is a genus that was separated from Phellinus in the early 2000s based on internal transcribed spacer (ITS) and nuclear large ribosomal subunit (nLSU) analyses. Species recognition in the genus is challenging without molecular analysis due to general absence of discrete morphological characteristics. In this study, we examined Fulvifomes diversity in the Federated States of Micronesia (FS Micronesia), primarily found in the tropical and subtropical regions. Seven Fulvifomes species were confirmed: two new species, one unrecorded species and four species previously reallocated to Fulvifomes from other genera. We provide sequences including data on two protein-coding genes (RNA polymerase II; RPB2 and translation elongation factor-1 alpha; TEF1), nLSU and different conspecific ITS types.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Fulvifomes Murrill, typified by F. robiniae (Murrill) Murrill (Murrill 1914; Pyropolyporus robiniae Murrill, basionym), consists of bracket polypores with perennial basidiomes, rough to rimose pileal surfaces, and coloured basidiospores (Murrill 1903, 1904). The taxonomic status of this genus has long been debated. One argument supported Fulvifomes as a genus (Kotlaba and Pouzar 1978; Fiasson and Niemelä 1984), whilst the other considered it a synonym (Gilbertson 1980; Ryvarden and Johansen 1980; Ryvarden 1991; Ryvarden 1994) or a subgenus of Phellinus Quél. (Dai 1999). This debate was resolved by phylogenetic analyses of the nuclear large ribosomal subunit (nLSU) rDNA, which strongly supported Fulvifomes as an independent genus (Wagner and Fischer 2002).
The generic morphological diagnosis of Fulvifomes has been redefined based on the phylogenetic analysis of internal transcribed spacer (ITS) and nLSU sequences, which separated Fulvifomes species from the others. This revision encompassed characteristics such as annual to perennial growth, sessile, effused-reflexed to substipitate basidiomes with solitary to imbricate pilei, homogeneous or duplex context, a hyphal system ranging from monomitic to dimitic, the absence of hyphoid and hymenial setae, and subglobose to ellipsoid, flattened ventral side, yellowish to brown and slightly thin to thick-walled basidiospores (Zhou 2014; Salvador-Montoya et al. 2018).
Most Fulvifomes species are distributed in the tropical and subtropical regions of Asia (Zhou and Zhang 2012; Hattori et al. 2014; Zhou 2014, 2015; Ji et al. 2017), America (Ji et al. 2017; Salvador-Montoya et al. 2018; Zhou et al. 2023), and Africa (Olou et al. 2019). The Federated States of Micronesia (FS Micronesia) is a biodiversity hotspot in the tropical region of the Pacific Ocean. To date, four Fulvifomes species have been reported from the FS Micronesia, namely Aurificaria luteoumbrina (Romell) D.A. Reid (≡ Fulvifomes luteoumbrinus (Romell) Y.C. Dai & Vlasák), Phellinus fastuosus (Lév.) S. Ahmad (≡ Fulvifomes fastuosus (Lév.) Bondartseva & S. Herrera), P. mangrovicus (Imazeki) Imazeki (≡ Fulvifomes mangrovicus (Imazeki) T. Hattori), and P. merrillii (Murrill) Ryvarden (≡ Fulvifomes merrilli (Murrill) Baltazar & Gibertoni) (Zhou 2015; Gilbert et al. 2008). However, as various Fulvifomes species have been reported in tropical regions, the diversity of Fulvifomes species in FS Micronesia is expected to be higher.
In this study, we aimed to examine Phellinus-like specimens collected from the FS Micronesia based on ITS and nLSU phylogenetic analyses, and on morphological characteristics. We also generated sequence data for RNA polymerase II (RPB2) and translation elongation factor-1 alpha (TEF1) protein-coding genes. Three species of Fulvifomes were determined, two of which are proposed as new to science. Detailed morphological descriptions are provided for the new taxa. Our findings contribute to the diversity of Fulvifomes species in the FS Micronesia.
Material and methods
Morphological studies
Six fruiting bodies were collected from three islands in the FS Micronesia; Chuuk Weno, Kosrae Lelu and Kosrae Tafunsak, in 2016 and 2017 (Fig. 1). Sampling and transport processes were performed with the permission of the FS Micronesia Government. Dried fungal specimens were deposited at the Seoul National University Fungus Collection (SFC). Photographs of the fruiting bodies were captured in the field.
A Geographical map of Fulvifomes distribution around the world. Distribution of phylogenetically confirmed Fulvifomes species from Table 1 is shown. Red dots indicate the tropical region, green indicates the sub-tropical region, and blue indicates the temperate region. B Sampling locations of Fulvifomes species in FS Micronesia. Orange dots indicate the locations where specimens were found
Microscopic examination of the dried specimens was performed by sectioning the tissue into 1 cm × 1 cm pieces. Tissue sections were mounted in 5% KOH on a glass slide and then observed using a Nikon Eclipse 80i microscope (Nikon, Japan). Approximately 20 basidia and basidiospores were measured per specimen using ImageJ software (Collins 2007). The Q values of the basidiospores indicate their length-to-width ratios. Colours were described based on ‘Methuen Handbook of Colour’ (Kornerup and Wanscher 1967). The pore surface and pore size were measured using a Nikon SMZ1500 stereomicroscope (Nikon, Japan). The following abbreviations were used: IKI = Melzer’s reagent, IKI − = neither amyloid nor dextrinoid, CB = Cotton Blue, CB − = acyanophilous, L = mean spore length, W = mean spore width, Q = ratios of L/W of specimens studied, and n = number of spores measured from the given number of specimens.
DNA extraction, amplification and sequencing
Total genomic DNA was extracted from dried specimens (1 cm × 1 cm of tissue) using the CTAB (Rogers and Bendich 1994) protocol and the AccuPrep® Genomic DNA Extraction Kit (Bioneer, Daejeon, Republic of Korea). Polymerase Chain Reaction (PCR) was performed for four genetic regions. The ITS region was amplified using ITS1F/ITS4 (White et al. 1990) or ITS1/ITS4B primers (Gardes and Bruns 1993). LR0R/LR7 (Vilgalys and Hester 1990) or LR0R/LB-y (Tedersoo et al. 2008) primers were used for nLSU. PCR for the ITS region and nLSU was performed under the following conditions: 5 min initial denaturation at 95 °C, followed by 35 cycles of 40 s at 95 °C, 40 s at 54 °C and 90 s at 72 °C, with a final extension step for 5 min at 72 °C. For the protein-coding genes, RPB2 region was amplified with primers bRPB2-6F/bRPB2-7.1R (Matheny 2005), under the following conditions: initial denaturation at 94 °C for 2 min, 36 cycles of 94 °C for 45 s, 53 °C for 90 s, 72 °C for 90 s, and a final extension at 72 °C for 10 min. The TEF1 region was amplified with primers EF595F/EF1160R (Kauserud and Schumacher 2001) under the following conditions: initial denaturation at 95 °C for 4 min, 35 cycles of 95 °C for 30 s, 55 °C for 30 s, 72 °C for 1 min, and a final extension at 72 °C for 7 min.
The amplification of the PCR products was verified by gel electrophoresis using 1% agarose gel with EcoDye DNA staining solution (SolGent Co., Daejeon, Republic of Korea). The products were purified using an ExpinTM PCR Purification Kit (GeneAll Biotechnology, Seoul, Republic of Korea). Sequencing was performed by Macrogen (Seoul, Republic of Korea) using an ABI 3730xL DNA analyser (Life Technologies, Gaithersburg, MD, USA). All sequences were proofread and edited using MEGA version 11 (Kumar et al. 2016) and FinchTV v.1.4. All the newly generated sequences were deposited at GenBank (Table 1).
Multiple peaks were found in the ITS region for three specimens (SFC20170118-26, SFC20170120-06 and SFC20170120-09), which were separated into their respective ITS type through cloning. ITS amplicons were cloned into the pTOP TA V2 blunt end vector (Enzynomics, Korea) and transformed into competent cells (Escherichia coli, DH10β). Transformed clones were re-amplified as described by Hattori et al. (2022). Sequencing of each ITS type was performed as above described.
Phylogenetic analysis
Phylogenetic trees were constructed using ITS and nLSU sequences. Newly generated sequences and published reference sequences of Fulvifomes from GenBank were collected with GenMine (Seo et al. 2022), and used for phylogenetic analyses. Inocutis dryophilus and Fomitiporella americana were used as outgroups due to their close phylogenetic relationships with Fulvifomes (Guglielmo et al. 2008a, b; Pildain et al. 2018). Sequence data collection, outgroup selection, multiple sequence alignment, trimming, concatenation, model selection and maximum likelihood (ML) tree construction were performed with FunID ver 0.3.4 with ‘accurate’ preset (https://github.com/Changwanseo/FunID). The final sequence alignments from FunID are in Online Resource 1. Phylogenetic trees were visualised by a modified ‘tree_interpretation.py’ function of FunID. All processed data are shown in Table 1.
Results
Molecular phylogeny
The final combined dataset (ITS1 + 5.8S + ITS2 + nLSU) included 64 specimens from 40 species. The concatenated sequences comprised 1604 nucleotides, including gaps. The ITS1, 5.8S, ITS2 and nLSU regions comprised 365, 142, 216 and 881 bases, respectively. Protein-coding gene sequences (RPB2 and TEF1) were not included in the phylogenetic analyses because of the lack of reference sequences.
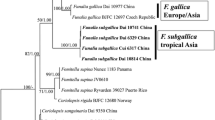
The six newly analysed specimens formed three well-supported clades in the ML analyses (Fig. 2). One specimen (SFC20170120-02) grouped with Fulvifomes halophilus with 100% bootstrap value. The other two clades were distinct from any described Fulvifomes species (bootstrap = 100). After comparing the morphological descriptions of these specimens with those of Fulvifomes species without molecular data (Lowe 1957; Ryvarden and Johansen 1980; Hattori 1999; Dai 2010; Hattori et al. 2014), we proposed them as two new species: Fulvifomes labyrinthus sp. nov. and Fulvifomes rhizophorus sp. nov. The two new species of Fulvifomes were further validated as novel based on their distinguishable morphological characteristics from other known Fulvifomes species.
Three ITS types were identified in Fulvifomes rhizophorus sp. nov. (SFC20170120-06, SFC20170120-09 and SFC20170118-26). Each specimen had two different types of ITS sequences: Type1 ITS was shared across all three specimens, Type2 ITS was found in SFC20170120-06, and Type3 ITS was found in SFC20170120-09 and SFC20170118-26. Two indels and 13 substitutions were detected (Fig. 3).
Taxonomy
Fulvifomes labyrinthus H. Suh, Y. Cho, D. Kim & Y.W. Lim, sp. nov. (Fig. 4)
MycoBank Number: MB849263
Diagnosis: Fulvifomes labyrinthus grows on the trunk of the mangrove tree, Xylocarpus granatum, and is characterised by irregular pores.
Holotype: Federated States of Micronesia, Chuuk State, Weno Island; on a rotting trunk of X. granatum, 26 January 2016. SFC20160126-30 (dried specimen). GenBank accession numbers: OR168711 (ITS) and OR168721 (nLSU).
Etymology: The epithet ‘labyrinthus’ referring to its partly irregular pores.
Description: Basidiome perennial, pileate, sessile, broadly attached, solitary, woody hard, without odour or taste. Pileus dimidiate, triquetrous to subungulate, projecting up to 10 cm, 14 cm wide, 7 cm thick at the base. Pileal surface reddish-brown (8E5) to dark-brown (8F5), velutinate, slightly rugose, concentrically sulcate with shallow grooves, irregularly cracked, sometimes covered with moss at the centre. Pore surface light brown (7D5) to reddish-gold (6C7); sterile margin distinct, up to 4 mm wide. Pores somewhat circular and regular near the sterile margin, 7‒8 pores per mm, but irregularly elongated to sinuous at the centre; dissepiments entire, thick. Tubes brown (6E7), stratified, woody hard, up to 3.5 cm thick in total, each annual layer up to 7 mm thick without layer of contextual tissue in between. Context darker than tubes, woody hard, up to 1.5 cm thick.
Hyphal system dimitic; generative hyphae hyaline to pale yellowish (3A3), thin- to slightly thick-walled, frequently septate, occasionally branched, present mostly in subhymenium and trama, rare in context, 1.8–3.3 μm wide; skeletal hyphae yellow (3B8) to dark brown (7F7), thick-walled, aseptate, unbranched, dominant in both trama and context, 2–3 μm wide in trama, 1.8–3.2 μm wide in context, IKI−, CB−.
Basidia rarely observed in the type material, barrel-shaped to subutriform, 4-sterigmate, simple septate, 11.8 × 4.8 μm. Basidiospores subglobose to broadly ellipsoid, yellowish orange (4A8) to brownish-yellow (5C8), thick-walled, smooth, IKI−, CB−, (3.7–)4.1–4.8(–5) × (2.9–)3.2–3.9(–4.1) μm, L = 4.44 μm, W = 3.57 μm, Q = 1.25 (n = 60/2). Cystidioles absent. Setae absent.
Additional specimen examined: SFC20160126-34; Federated States of Micronesia, Chuuk States, Weno Island; on a rotten trunk of Xylocarpus granatum, 26 January 2016. GenBank accession numbers: OR168712 (ITS) and OR168722 (nLSU).
Notes: Fulvifomes labyrinthus is phylogenetically closely related and morphologically similar to Fulvifomes xylocarpicola T. Hatt., Sakayaroj & E.B.G. Jones, they share the same host, Xylocarpus granatum (Hattori et al. 2014). However, F. labyrinthus can be distinguished from F. xylocarpicola by a reddish-brown pileus and partly irregular pores.
Fulvifomes rhizophorus H. Suh, Y. Cho, D. Kim & Y.W. Lim, sp. nov. (Fig. 5)
MycoBank Number: MB849264
Diagnosis: Fulvifomes rhizophorus grows on the trunk of the mangrove tree, Rhizophora apiculata, and is characterised by an ungulate basidiome.
Holotype: Federated States of Micronesia, Kosrae Lelu, 18 January 2017. SFC20170118-26 (dried specimen). GenBank accession numbers: OR168715 (ITS) and OR168725 (nLSU).
Etymology: The epithet ‘rhizophorus’ referring to the genus of its host tree.
Description: Basidiome perennial, pileate, sessile, broadly attached, solitary, woody hard and without odour or taste. Pileus dimidiate, applanate, projecting 6.5 cm up, 11.5 cm wide and 3 cm thick at the base. Pileal surface yellowish (5E8) to reddish-brown (8F8), furrowed, concentrically sulcate with deep grooves, radially cracked. Pore surface golden-brown (5D7) to linoleum-brown (5E7); sterile margin distinct, up to 4 mm wide. Pores primarily round in shape, featuring closed slits, typically not connected to adjacent pores, 6‒7 pores per mm; dissepiments entire, thick. Tubes light brown (6D8) to dark brown (6F8), stratified, woody hard, up to 1.4 cm thick in total, each annual layer up to 1 cm thick without layer of contextual tissue in between. Context darker than tubes, woody hard, duplex, the lower layer dark blond (5D4), up to 8 mm thick, the upper layer dark yellow (4C8), up to 9 mm thick.
Hyphal system dimitic; generative hyphae hyaline to pale yellowish (3A3), thin- to slightly thick-walled, frequently septate, occasionally branched, present mostly in subhymenium and trama, rare in context, 2.5–3 μm wide in trama, 3.3–5.2 wide in context; skeletal hyphae aseptae, unbranched, dominant in both trama and context, brown (6E8), in trama thick-walled with narrow lumen, 3.7–7.5 μm wide, dark brown (7F7), thick-walled with wide lumen, 6.2–8 μm in context, IKI−, CB−.
Basidia rarely observed in the type material, clavate to subutriform, 4-sterigmate, simple septate, 10.8 × 4.7 μm. Basidiospores subglobose to broadly ellipsoid, yellowish-orange (4A8) to brownish-yellow (5C8), thick-walled, smooth, IKI−, CB−, (5–)5.3–6(–6.8) × (4.3–)4.4–5.2(–6) μm, L = 5.6 μm, W = 5 μm, Q = 1.14 (n = 81/3). Cystidioles absent. Setae absent.
Additional specimens examined: SFC20170120-06; Federated States of Micronesia, Kosrae Tafunsak, on a trunk of R. apiculata, 20 January 2017. GenBank accession numbers: ITS OR168713 and nLSU OR168723. SFC20170120-09; Federated States of Micronesia, Kosrae Tafunsak, on a trunk of R. apiculata tree on 20 January 2017. GenBank accession numbers: OR168714 (ITS) and OR168724 (nLSU).
Notes: Fulvifomes rhizophorus is found on trunks of Rhizophora apiculata, which is submerged in the shallow waters of the mangrove forest. Amongst the Fulvifomes species in the FS Micronesia, Fulvifomes rhizophorus can be distinguished by larger basidiospores compared to those of Fulvifomes labyrinthus (4.1–4.8 × 3.2–3.9 μm) and Fulvifomes halophilus (4–5.5 × 3.5–5 μm) (Hattori et al. 2022).
Discussion
During the exploration of FS Micronesia in 2016 and 2017, six fruiting bodies of Fulvifomes were found. These were identified as three species, including two newly proposed species, based on morphological examination and phylogenetic analyses. Two specimens are described here as Fulvifomes labyrinthus sp. nov. and three are described as Fulvifomes rhizophorus sp. nov. The remaining specimen was identified as Fulvifomes halophilus, which is being reported for the first time in the FS Micronesia. Fulvifomes rhizophorus has three different types of ITS sequences, whereas other Fulvifomes species in FS Micronesia have only one ITS type. Various ITS types within a single species have been observed across Basidiomycota (Anderson and Kohn 2007; Lim et al. 2008). With various environmental changes occurring in mangrove forests, this phenomenon may indicate the initiation of sympatric speciation, where divergence occurs under non-disruptive selection (Giraud et al. 2008; Druzhinina et al. 2010). Therefore, our results suggest that Fulvifomes rhizophorus was introduced early in the FS Micronesia with some geographical and host restrictions, resulting in a diverse population (Burnett 1983; Griffith and Hedger 1994).
Fulvifomes species have relatively simple basidiome features, which make them difficult to distinguish amongst species. Species differentiation based on microscopic features is challenging because setae and cystidioles are absent. Furthermore, basidia are difficult to find, where some studies have noted that basidia are not observed (Zhou 2014; Olou et al. 2019; Salvador-Montoya et al. 2022). Similarly, basidia of these three species have been extensively searched. We observed basidia from the pore layer closer to the context of our perennial specimens by thin-sectioning in the vertical direction. Basidia may not be the key morphological character in distinguishing Fulvifomes species, however, providing the overall morphological profile of a species when designating a new species may ensure greater confidence in species delimitation.
According to Gilbert et al. (2008), four species were found in the FS Micronesia, namely Aurificaria luteoumbrina, Phellinus fastuosus, P. mangrovicus and P. merrillii, which have been transferred to Fulvifomes (Bondarceva et al. 1992; Hattori et al. 2014; Zhou 2015; Wu et al. 2022). As phylogenetic analyses had not been thoroughly conducted at the time, these species were classified and identified based only on morphological characteristics. Besides Fulvifomes species, other polypores were also transferred to other genera later on (Zmitrovich and Malysheva 2013). It is essential to re-identify the FS Micronesian specimens collected in the past using DNA sequencing and morphological characterisation to fully understand the Fulvifomes and other fungal distribution in this area.
Geographical and ecological features help to distinguish Fulvifomes from other genera in Hymenochaetaceae. Amongst the species that have validated sequences, three species were found in the temperate region: Fulvifomes robiniae from USA in the early 1900s (Murrill 1914), F. rimosus of which the lectotype was found in Tasmania, and F. imazekii from Kochi, Japan (Hattori et al. 2022). Other Fulvifomes species have been detected mainly in tropical and subtropical regions worldwide (Fig. 1). Fulvifomes halophilus, an unrecorded species in the FS Micronesia, also followed the distribution pattern of the majority of Fulvifomes; this species was also found in Thailand and Borneo, in the tropical and subtropical regions of Southeast Asia (Sakayaroj et al. 2012; Wu et al. 2022).
Limited protein-coding gene sequences have been found for Fulvifomes in open databases, such as for RPB2 and TEF1, which are commonly analysed in phylogenetic studies of Hymenochaetaceae. In this study, we provide sequences of protein-coding genes (RPB2 and TEF1) for the three Fulvifomes species, hoping to increase the resolution of phylogenetic analyses in future studies. Until now, a total of 40 Fulvifomes species have been discovered around the world. Two new species (Fulvifomes labyrinthus and Fulvifomes rhizophorus) and one previously unrecorded species (Fulvifomes halophilus) were identified in this study. Including species previously reported in the FS Micronesia, a total of seven Fulvifomes species are detected in the country. Considering the unique environments of the Pacific Islands in tropical and subtropical areas (Mueller-Dombois et al. 1998; Rehman et al. 2013), a more intensive study in these regions may reveal more Fulvifomes species.
Data availability
GenBank accession ITS: OR168710–5, nLSU: OR168720–5, TEF1: OR215495–8, RPB2: OR196705–7.
References
Anderson JB, Kohn LM (2007) Dikaryons, diploids, and evolution. Sex in fungi: molecular determination and evolutionary implications. ASM Press, Washington DC, pp 333–348
Bondarceva M, Herrera S, Sandoval D, Cejas F (1992) Taxonomical problems of the Cuban hymenochaetaceous fungi. Mikol Fitopatol 26(1):3–14
Burnett JH (1983) Presidential address: speciation in fungi. Trans Br Mycol Soc 81.1:IN1–14
Collins TJ (2007) ImageJ for microscopy. Biotechniques 43(S1):S25–S30
Dai YC (1999) Phellinus sensu lato (Aphyllophorales, Hymenochaetaceae) in east Asia. Acta Bot Fenn 166:1–115
Dai YC (2010) Hymenochaetaceae (Basidiomycota) in China. Fungal Divers 45(1):131–343
Druzhinina IS, Komoń-Zelazowska M, Atanasova L, Seidl V, Kubicek CP (2010) Evolution and ecophysiology of the industrial producer Hypocrea jecorina (anamorph Trichoderma reesei) and a new sympatric agamospecies related to it. PLoS ONE 5(2):e919
Ediriweera SS, Wijesundera RLC, Nanayakkara CM, Weerasena OVDSJ (2014) A new record of Fulvifomes fastuosus from Sri Lanka. J Natl Sci Found Sri Lanka 42(4):369–371
Fiasson JL, Niemelä T (1984) The Hymenochaetales: a revision of the European poroid taxa. Karstenia 24(1):14–28
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes-application to the identification of mycorrhizae and rusts. Mol Ecol 2(2):113–118
Guglielmo F, Gonthier P, Garbelotto M, Nicolotti G (2008) A PCR-based method for the identification of important wood rotting fungal taxa within Ganoderma, Inonotus s.l. and Phellinus s.l. FEMS Microbiol Lett 282(2):228–237
Gilbert GS, Gorospe J, Ryvarden L (2008) Host and habitat preferences of polypore fungi in Micronesian tropical flooded forests. Mycol Res 112(6):674–680
Gilbertson RL (1980) Wood-rotting fungi of North America. Mycologia 72(1):1–49
Gilbertson RL, Ryvarden L (1987) North American Polypores Vol. 2. Megasporoporia-Wrightoporia, pp 437–885
Giraud T, Refrégier G, Le Gac M, de Vienne DM, Hood ME (2008) Speciation in fungi. Fungal Genet Biol 45(6):791–802
Griffith G, Hedger J (1994) Spatial distribution of mycelia of the liana (L-) biotype of the agaric Crinipellis perniciosa (Stahel) Singer in tropical forest. New Phytol 127(2):243–259
Hattori T, Ota Y, Sotome K (2022) Two new species of Fulvifomes (Basidiomycota, Hymenochaetaceae) on threatened or near threatened tree species in Japan. Mycoscience 63(3):131–141
Hattori T, Sakayaroj J, Jones EBG, Suetrong S, Preedanon S, Klaysuban A (2014) Three species of Fulvifomes (Basidiomycota, Hymenochaetales) associated with rots on mangrove tree Xylocarpus granatum in Thailand. Mycoscience 55(5):344–354
Hattori T (1999) Phellinus setifer sp. nov. and P. acontextus, two noteworthy polypores from temperate areas of Japan, with notes on their allies. Mycoscience 40(6):483–490
Jayawardena RS, Hyde KD, Jeewon R, Ghobad-Nejhad M, Wanasinghe DN, Liu N, Kang JC (2019) One stop shop II: taxonomic update with molecular phylogeny for important phytopathogenic genera: 26–50 (2019). Fungal Divers 94:41–129
Ji XH, Wu F, Dai YC, Vlasák J (2017) Two new species of Fulvifomes (Hymenochaetales, Basidiomycota) from America. MycoKeys 22:1–13
Kauserud H, Schumacher T (2001) Outcrossing or inbreeding: DNA markers provide evidence for type of reproductive mode in Phellinus nigrolimitatus (Basidiomycota). Mycol Res 105(6):676–683
Kornerup A, Wanscher JH (1967) Methuen handbook of colour, 3rd edn. Eyre Methuen Ltd., London
Kotlaba F, Pouzar Z (1978) Notes on Phellinus rimosus complex (Hymenochaetaceae). Acta Bot Croat 37(1):171–182
Krauss KW, Keeland BD, Allen JA, Ewel KC, Johnson DJ (2007) Effects of season, rainfall, and hydrogeomorphic setting on mangrove tree growth in Micronesia. Biotropica 39(2):161–170
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874
Lim YW, Sturrock R, Leal I, Pellow K, Yamaguchi T, Breuil C (2008) Distinguishing homokaryons and heterokaryons in Phellinus sulphurascens using pairing tests and ITS polymorphisms. Antonie van Leeuwenhoek 93:99–110
Lowe JL (1957) Polyporaceae of North America. The genus Fomes, Technical Publication No. 80. New York State College of Forestry 97
Matheny PB (2005) Improving phylogenetic inference of mushrooms with RPB1 and RPB2 nucleotide sequences (Inocybe; Agaricales). Mol Phylogenet Evol 35(1):1–20
Mueller-Dombois D, Fosberg FR, Mueller-Dombois D, Fosberg FR (1998) Micronesia. Vegetation of the tropical Pacific Islands 199–313
Murrill WA (1914) Northen polypores. The New Era Printing Company, New York
Murrill WA (1915) Tropical polypores. USA, New York
Murrill WA (1903) A historical review of the genera of the Polyporaceae. J Mycol 9(2):87–102
Murrill WA (1904) The Polyporaceae of North America-IX. Inonotus, sesia and monotypic genera. Bull Torrey Bot Club 31(11):593–610
Núñez M, Stokland J (2000) Biogeographical affinities of East Asian polypores. Karstenia 40:123–128
Olou BA, Ordynets A, Langer E (2019) First new species of Fulvifomes (Hymenochaetales, Basidiomycota) from tropical Africa. Mycol Prog 18:1383–1393
Pildain MB, Cendoya RR, Ortiz-Santana B, Becerra J, Rajchenberg M (2018) A discussion on the genus Fomitiporella (Hymenochaetaceae, Hymenochaetales) and first record of F. americana from southern South America. MycoKeys 38:77
Rehman HU, Nakaya H, Kawai K (2013) Geological origin of the volcanic islands of the Caroline Group in the Federated States of Micronesia Western Pacific. South Pacific Stud 33(2):101–118
Rogers SO, Bendich AJ (1994) Extraction of total cellular DNA from plants, algae and fungi. Plant Molecular Biology Manual, pp 183–190
Ryvarden L (1991) Genera of polypores. Nomenclature and taxonomy. Synopsis Fungorum 5:1–363
Ryvarden L (1994) Phellinus umbrinellus—new to Europe. Mycologist 8(1):6
Ryvarden L, Johansen I (1980) A preliminary polypore flora of East Africa. Fungiflora, pp 1–636
Seo CW, Kim SH, Lim YW, Park MS (2022) Re-identification on Korean Penicillium sequences in GenBank collected by software GenMine. Mycobiology 50(4):231–237
Sakayaroj J, Preedanon S, Suetrong S, Klaysuban A, Jones EG, Hattori T (2012) Molecular characterization of basidiomycetes associated with the decayed mangrove tree Xylocarpus granatum in Thailand. Fungal Divers 56:145–156
Salvador-Montoya CA, Martínez M, Drechsler-Santos ER (2022) Taxonomic update of species closely related to Fulvifomes robiniae in America. Mycol Prog 21(12):95
Salvador-Montoya CA, Popoff OF, Reck M, Drechsler-Santos ER (2018) Taxonomic delimitation of Fulvifomes robiniae (Hymenochaetales, Basidiomycota) and related species in America: F. squamosus sp. nov. Plant Syst Evol 304:445–459
Tan YP, Bishop-Hurley SL, Shivas RG, Cowan DA, Maggs-Kölling G, Maharachchikumbura SS, Crous PW (2022) Fungal Planet description sheets: 1436–1477. Persoonia 49:261–350
Tchoumi JMT, Coetzee MPA, Rajchenberg M, Roux J (2020) Poroid Hymenochaetaceae associated with trees showing wood-rot symptoms in the Garden Route National Park of South Africa. Mycologia 112(4):722–741
Tedersoo L, Jairus T, Horton BM, Abarenkov K, Suvi T, Saar I, Kõljalg U (2008) Strong host preference of ectomycorrhizal fungi in a Tasmanian wet sclerophyll forest as revealed by DNA barcoding and taxon-specific primers. New Phytol 180(2):479–490
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 172(8):4238–4246
Wagner T, Fischer M (2002) Proceedings towards a natural classification of the worldwide taxa Phellinus s.l. and Inonotus s.l., and phylogenetic relationships of allied genera. Mycologia 94(6):998–1016
Wagner T, Ryvarden L (2002) Phylogeny and taxonomy of the genus Phylloporia (Hymenochaetales). Mycol Prog 1(1):105
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR protocols: a guide to methods and applications 18.1:315–322
Wu F, Zhou LW, Vlasák J, Dai YC (2022) Global diversity and systematics of Hymenochaetaceae with poroid hymenophore. Fungal Diversy 113(1):1–192
Zheng HF, Huang FC, Liu B, Shao YY, Qin PS (2021) Fulvifomes nonggangensis and F. tubogeneratus (Hymenochaetales, Basidiomycota): two new species from southern China based on morphological and molecular evidences. Mycobiology 49(3):213–222
Zhou LW (2014) Fulvifomes hainanensis sp. nov. and F. indicus comb. nov. (Hymenochaetales, Basidiomycota) evidenced by a combination of morphology and phylogeny. Mycoscience 55(1):70–77
Zhou LW (2015) Fulvifomes imbricatus and F. thailandicus (Hymenochaetales, Basidiomycota): two new species from Thailand based on morphological and molecular evidence. Mycol Prog 14:1–8
Zhou LW, Zhang WM (2012) A new species of Fulvifomes (Hymenochaetaceae) from Cambodia. Mycotaxon 119(1):175–179
Zhou M, Ji XH, Liu HG, Miller K, Yuan Y, Vlasák J (2023) Two new species of Hymenochaetaceae from tropical Asia and America. Front Cell Infect Microbiol 12:1985
Zmitrovich IV, Malysheva VF (2013) Towards a phylogeny of Trametes alliance (Basidiomycota, Polyporales). Mикoлoгия и фитoпaтoлoгия 47(6):358–380
Acknowledgements
We are grateful to Chuuk and Kosrae State Government, the Federated States of Micronesia, for allowing this research. This study results from a collaborative project between Micronesia and the Korean government. Therefore, there is no violation of the ABS protocol, and the results are being shared with Micronesia.
Funding
Open Access funding enabled and organized by Seoul National University. This research was supported by the grant of the Ministry of Oceans and Fisheries (PM59731).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation and data collection were performed by HS and YC. Phylogenetic analysis was performed by CS and drawings were done by DK. The first draft of the manuscript was written by HS and YC, and YWL commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Section Editor: Yu-Cheng Dai
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Suh, H., Cho, Y., Seo, C.W. et al. A taxonomic study of Fulvifomes (Hymenochaetales, Basidiomycota) in the Federated States of Micronesia and identification of two new species. Mycol Progress 23, 15 (2024). https://doi.org/10.1007/s11557-024-01946-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11557-024-01946-4