Abstract
Homeodomain-leucine zipper (HD-Zip) transcription factor genes are unique in the plant kingdom. Studies have shown that HD-Zip transcription factor has four subfamilies (subfamilies I–IV), and each subfamily exerts similar or diverse functions in the growth and development as well as environmental stress responses of plants. Although a genome-wide analysis of this transcription factor has been performed in some species, systematic identification of sequences and expression patterns under biotic and abiotic stress have not been carried out in pear. In this study, we identified 52 putative HD-Zip genes within the pear genome. Based on the phylogenetic analysis, HD-Zip transcription factors were classified into four subfamilies: 18, 13, 5, and 16 members were identified separately in subfamilies I, II, III, and IV, respectively. Genes in the same subfamily shared the constituent structure and conserved motifs, thereby implying that the HD-Zip genes that shared similar structures may exert similar functions. Moreover, abundant stress-related and pathogen-related cis-elements were observed in the promoter region of the pear HD-Zip I and II genes. Transcript abundance level of 20 selected HD-Zip I and II genes were analyzed by quantitative real-time RT-PCR (qRT-PCR) during the infection of Alternaria alternata, and abiotic stress conditions such as drought and salinity treatment. The results confirmed that some genes display stress-inducible and pathogen-inducible expression patterns.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants launch complex cellular, physiological, and molecular responses during their developmental processes (Sultan 2010). These responses are usually regulated by transcription factors (Nakashima et al. 2009). The current studies on transcription factors are focused on NAC, AP2/EREBP, MYB, WRKY, and HD-Zip. Homeobox is a conserved 60-amino acid motif called the homeodomain (HD), which is able to bind specific DNA sequences (Moens and Selleri 2006; Ariel et al. 2010). The homeodomain has a three-helix structure and executes function in all the eukaryotic organisms associated with development (Moens and Selleri 2006; Ariel et al. 2010). According to the difference of HD sequence, HD size, and HD location, which are associated with other domains and gene structure, the HD-containing proteins are classified into six members including homeodomain-leucine zipper (HD-Zip), Wuschel-related homeobox (WOX), plant homeodomain associated to a finger domain (PHD-finger), bell domain (BELL), knotted related homeobox (KNOX), and zinc finger-homeodomain (ZF-HD) (Ariel et al. 2010). The HD-Zip transcription factors are unique to the plant kingdom (Ariel et al. 2010). They contain not only a highly conserved HD, but also a Zip element (Schena and Davis 1992). The homeodomain is at the C-terminal end and binds specifically to DNA, and the Zip element mediates the formation of dimerization (Ariel et al. 2010). Based on the conserved HD-Zip domain, structure features, additional conserved motifs, and physiological functions, the transcription factors of HD-Zip is classified into four subfamilies, HD-Zip I–IV. Studies show that HD-Zip family genes participate in different biological prRocesses of growth and development of plants as well as adaptation response to environmental stress (Ariel et al. 2010).
HD-Zip subfamily I has the shortest sequences and contains only HD domain and LZ motif (Chen et al. 2014). HD-Zip II proteins contain five additional conserved amino acids, Cys, Pro, Ser, Cys, and Glu, known as “CPSCE”, which is located downstream of the LZ motif (Tron et al. 2002; Chen et al. 2014). Members of these two subfamilies bind similar pseudo-palindromic cis-elements, CAATNATTG (Sessa et al. 1993; Frank et al. 1998; Johannesson et al. 2001). Arabidopsis HD-Zip I proteins consist of 17 members: AtHB1/HAT5, AtHB3/HAt7, AtHB5–7, AtHB12, AtHB13, AtHB16, AtHB20–23, AtHB40, AtHB51–54 (Ariel et al. 2010; Hu et al. 2012); Arabidopsis HD-ZIP II proteins consist of nine members: AtHB2/HAT4, AtHB4, HAT1–HAT3, HAT9, HAT14, HAT17, and HAT22 (Ariel et al. 2010; Hu et al. 2012). In summary, members of HD-Zip subfamilies I and II act as the integrators of development and environmental cues and endogenous hormone signal pathway to regulate targeted gene expression and plant adaptation response (Son et al. 2010; Wang et al. 2013c). For example, the expression of HD-Zip I and II genes was induced by the following three environmental cues: drought/salt/ABA/cold (Harris et al. 2011), light stimulation (Ariel et al. 2010; Harris et al. 2011), and biotic stress such as Pseudomonas syringae (Dezar et al. 2011). AtHB5, AtHB6, AtHB7, and AtHB12 participate in ABA-dependent abiotic stress (Soderman et al. 1999; Johannesson et al. 2003; Olsson et al. 2004). The treatment of 100 mM NaCl induces 12- and 25-fold increase of AtHB7 and AtHB12 transcript abundance relative to controls (Henriksson et al. 2005). The expression of AtHB6, AtHB21, AtHB40, AtHB35 is also activated by salt-induced stress (Skinner and Gasser 2009). In ABA signal pathway, AtHB7 and AtHB12 are positive regulators (Olsson et al. 2004); however, AtHB6 is as a negative regulator (Deng et al. 2006; Ariel et al. 2010). In addition, genes of HD-Zip I and II are induced by biotic stress such as P. syringae. HaHB10, a member of HD-Zip II subfamily in H. annuus, was involved in inducing the particular flowering transition genes and promoting the accumulation of salicylic acid (SA), when transgenic Arabidopsis was attacked by wounding and a virulent strain of P. syringae (Dezar et al. 2011). The transcript abundance of HaHB4, a member of HD-Zip I in H. annuus, is activated by Spodoptera littoralis or Spodoptera frugiperda larvae (Dezar et al. 2005). Transgenic Arabidopsis and Z. mays with HaHB4 overexpression displayed higher expression level of defense-related genes when these transgenic plants were attacked by herbivores (Dezar et al. 2005).
Pear (Pyrus spp) is one of the most cultivated and nutritionally vital fruit trees in the world. However, pear production is limited by drought, salinity, and pathogenic infection (Suzuki et al. 2003). To improve fruit quality and enhance production, it is necessary to know more about pear. Sequencing of the pear genome provides a possibility for gene prediction and annotation. Pyrus betulifolia Bunge is one the most widely used rootstock in China because of its excellent resistance and tolerance to unfavorable environmental factors. The knowledge about the expression patterns of genes in P. betulifolia Bunge under stress may help in narrowing down on candidate genes, which can be used during cultivation in breeding in the future.
The HD-Zip family members were widely investigated in A. thaliana (Schena and Davis 1992), Z. mays (Zhao et al. 2011), O. sativa (Agalou et al. 2008), P. trichocarpa (Hu et al. 2012), and other species (Liu et al. 2013; Chen et al. 2014; Zhang et al. 2014). Until now, no genome-wide research, including gene identification, genome organization, and phylogenetic analysis, has been reported in pear. In this study, 52 putative genes of the HD-Zip family were identified based on current pear genome database, apple genome database, and The Arabidopsis Information Resource (TAIR). First, we identified gene members, analyzed phylogenetic and gene structure, found stress-related and pathogen-related cis-elements in the promoter region, and searched conserved motifs. Second, we investigated the expression patterns of selected HD-Zip I and II members under both abiotic stress, including salt and drought conditions, and biotic stress, such as Alternaria alternata.
Materials and methods
Identification of HD-Zip genes in pear
Sequences of Arabidopsis HD-Zip proteins were downloaded from the Arabidopsis Information Resource (TAIR, http://arabidopsis.org/, release 10.0). Pear genome sequence was downloaded from Pear Genome Project (http://peargenome.njau.edu.cn:8004/) and Rosaceae Genome Database (http://www.rosaceae.org/species/pyrus/pyrus_communis). The Hidden Markov Model (HMM) profile of the Homeobox (PF00046) and HD-Zip domain (PF04618) was obtained from Pfam database (http://pfam.sanger.ac.uk) (Finn et al. 2006; Zhao et al. 2011) and SMART (http://smart.embl-heidelberg.de) (Letunic et al. 2009). All proteins containing the PF00046 domain were acquired from pear genome data. Apple (Malus × domestica) HD-Zip protein sequences (IPR001356) were obtained from Apple Genome Annotation Database (http://www.applegene.org/, release 1.0) and Plant Transcription Factor Database (http://planttfdb.cbi.pku.edu.cn, release 3.0). Afterward, all of the confirmed HD-ZIP proteins from pear were aligned using MUSCLE v3.8.31 (Edgar 2004). The gene family tree was generated by analyzing the multiple-sequence alignment results based on MUSCLE (Guo et al. 2014) using the Clustal W method with the MEGA 5.0 software (Tamura et al. 2011). Local blast was carried out using Arabidopsis HD-ZIP proteins as queries, and all identical sequences were checked manually to remove redundant sequences.
Phylogenetic analysis of HD-Zip genes
The identification of full-length protein sequences from pear was first elucidated in conjunction with HD-Zip genes from Arabidopsis and apple using MUSCLE v3.8.31 (Edgar 2004). Consequently, gaps were deleted by GENEDOC (Nicholas et al. 1997). Then, aligned sequences were constructed using Clustal W of MEGA5.0 software (Tamura et al. 2011) with the neighbor-jointing method. A bootstrap of 1000 replicates with pairwise deletion option was performed (Chen et al. 2014). Subfamilies of the HD-Zip proteins of pear and apple were classified according to their phylogenetic relationships with their corresponding Arabidopsis HD-Zip proteins.
Investigation of cis-elements in the promoter region
The 2000 bp genomic sequences upstream of the transcription start site (ATG) were obtained from pear genomic database. PLACE (http://www.dna.affrc.go.ip/PLACE/signalscan.html), an online database of plant cis-acting regulatory DNA elements (cis-elements) (Higo et al. 1999), was used to analyze putative cis-elements in the promoter regions of the HD-Zip I and II genes, as described previously (Zhao et al. 2011).
Gene structure and conserved motif analysis
Gene Structure Display server (GSDS, http://gsds.cbi.pku.edu.cn/) was employed to analyze the exon/intron organization of the HD-Zip genes by alignment of the cDNAs with their corresponding genomic DNA sequences (Guo et al. 2007; Zhao et al. 2011). The MEME program (version 4.9.1, http://meme.nbcr.net/memecgibin/meme.cgi) with the following variables—number of repetitions, if any; maximum number of motifs, 20; and optimum motif width, between six and 200 residues—was used for the annotation of conserved motif (Bailey and Elkan 1995; Zhao et al. 2011).
Plant material and growth conditions
Tissue culture seedlings of pear (P. betulifolia Bunge) were used to study gene expression levels in all experiments. Pear seedlings were maintained on 1/3 MS medium (Murashige and Skoog 1962) under a 16:8 h light:dark photoperiod (cool white fluorescent tubes, 40–60 µmol m−2 s−1) at 25 °C in the culture room (Righetti et al. 2014). For NaCl and/or PEG6000 stress treatment, rooted seedlings were acquired according to the following procedures: 1-month-old tissue culture seedlings were first moved to 1/4 Lepoivre’s medium (Leblay et al. 1991) supplemented with IAA 0.5 mg L−1, BA 1.0 mg L−1, and NAA 0.1 mg L−1 at 22–24 °C in the dark for 7–12 days; subsequently, all tissues were moved in 1/4 Lepoivre’s medium for 1.5–2 months at 25 °C with a 16:8 h light:dark photoperiod (cool white fluorescent tubes, 40–60 μmol m−2 s−1); finally, all tissues with roots were grown in 1/4 Lepoivre’s liquid medium for 2 weeks at 25 °C with a 16:8 h light:dark photoperiod (cool white fluorescent tubes, 40–60 µmol m−2 s−1). To mimic salt- and drought-induced stress, 1/4 Lepoivre’s liquid medium with 150 mmol L−1 NaCl or 20 % PEG6000 (w/v) were employed to maintain rooted seedling growth. Seedlings in 1/4 Lepoivre’s liquid medium were used as control. After 0, 6, 12, and 24 h, young leaves from six seedlings were collected, then flash frozen in liquid nitrogen, and stored at −80 °C for further analysis (Hu et al. 2012).
For the artificial infection experiments of pathogen, A. alternata Japanese pear pathotype L-12 (Wang et al. 2006), stored in our lab, was grown on potato dextrose agar plates (Sigma–Aldrich) for 1 week (Suzuki et al. 2003). The conidial suspension was filtered through a cloth mesh (Suzuki et al. 2003; Wang et al. 2013a). After removal of the supernatant and re-suspension of the conidia in sterile distilled water, an inoculation suspension was prepared containing 1.0 × 106 conidia mL−1. For intact plant assays, 30 mL of the conidial suspension was sprayed on the leaves. Inoculated plants were grown in a sealed light transmittance bottle in a 16/8 h (~40 mmol m−2 s−1) photoperiod growth chamber (25/22 °C) (Wang et al. 2013a). Seedlings were sprayed with H2O as a negative control. After 0, 6, and 9 days, leaves were collected, flash frozen in liquid nitrogen, and stored at −80 °C for further analysis. At least ten plants were assessed for each treatment.
RNA isolation and qRT-PCR
Total RNA from leaves was extracted using TRIzol reagent (Invitrogen, Carlsbad, CA) according to manufacturer’s instructions. RNA was treated with Turbo DNA-free kit (Applied Biosystems, Foster City, CA) according to manufacturer’s instructions to ensure no DNA contamination. The RNA concentration and purity were measured using a NanoDrop TM 3100 Spectrophotometer (Thermo Scientific, Wilmington, DE) (Wang et al. 2013b). The RNA integrity was checked by agarose gel electrophoresis (Wang et al. 2013b). cDNA synthesis was performed with approximately 2 μg total RNA using the superscript III first-strand synthesis cDNA kit according to the manufacturer’s instructions (Wang et al. 2013a, b).
Gene-specific primers for the pear HD-Zip were assigned according to the non-conserved regions at the 3′-end (Table S1). To obtain unique cDNA segments, blasting of primers against pear genome was performed to estimate primer specificity, and then a single band of correct size was produced by real-time PCR. Final single sharp peaks were generated by the corresponding melting curves (Shao et al. 2014).
Amplifications were performed on an Applied Biosystems 7300 system (Applied Biosystems, Shanghai, China) to monitor the expression profiles of HD-Zip I and II genes in response to various abiotic and biotic stress conditions. Reactions were carried out with the following procedure: 2 min at 95 °C, followed by 40 cycles of 45 s at 95 °C, 30 s at 58 °C, and 45 s at 72 °C (Wang et al. 2013b). Fluorescence was measured at the end of each cycle (Wang et al. 2013b). Analysis of the melting curves provided evidence for the absence of nonspecific products and primer dimers (Wang et al. 2013b). For data analysis, on the basis of three independent biological samples as indicated, average threshold cycle (CT) values were calculated for each gene of interest (Wang et al. 2013b) and were normalized and used to calculate the relative transcript levels as described by Livak and Schmittgen (2001). The pear gene Actin (accession number: AF38514) was used as an internal standard for normalization.
Results
Identification of HD-Zip gene family in the pear and apple genome
To obtain comprehensive information of PbHD-Zip genes in the pear genome, the Hidden Markov Model (HMM) profile of the homeobox and HD-Zip domain (PF00046 and PF04618) were used as a query to search the database using the program HMMER3.0 (Liu et al. 2014) with default E value; 118 HD-containing sequences were found in the pear genome. The online program SMART and Pfam with an E value <0.1 were employed to evaluate the integrity of the HD-Zip domain. As a result, 52 non-redundant HD-Zip genes called PbHB1 to PbHB52 were identified. IPR001356, as a corresponding functional domain of PF00046, was employed to search HD-Zip protein sequence from the apple genome, which found 138 HD-containing sequences. All these sequences were searched for the integrity of the HD-Zip domain by SMART and Pfam program. Seventy non-redundant HD-Zip genes were identified in the apple genome, and 80 sequences of HD-Zip in apple (Malus × domestica) were provided by plant transcription factor database (http://planttfdb.cbi.pku.edu.cn, release 3.0). By further searching using Pfam and START, 10 sequences that had only an HD domain but no Zip element were removed. The rest which contained both HD domain and Zip elements were consistent with that of our identification. As shown in Table 1, the total number of HD-Zip genes identified in pear was smaller than that in grape and Populus, while greater than that in peach. Compared with herbaceous plants, the number of HD-Zip genes identified in pear was close to that in maize but much smaller than that in soybean, and greater than that in both Arabidopsis, O. sativa, M. truncatula, B. distachyon, and S. bicolor (Table 1). The total number of HD-Zip genes in the apple genome was next only to that in soybean (Table 1). Based on their phylogenetic relationship with Arabidopsis HD-Zip genes, the HD-Zip family in pear contained 18 genes in subfamily I, 13 in subfamily II, 5 in subfamily III, and 16 in subfamily IV. However, the apple genome contained 24 in subfamily I, 18 in subfamily II, 9 in subfamily III, and 19 in subfamily IV (Table 1). Among these genes, the genes in subfamily III and subfamily IV encoded general longer proteins (average 600–950 amino acids); however, genes in subfamily I and II encoded shorter proteins (average 200–400 amino acids). Table 2 shows the nomenclature and corresponding information.
Phylogenetic and structural analysis of PbHB genes
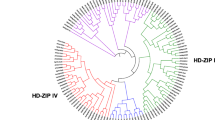
The putative 52 HD-Zip protein sequences were used to generate unrooted phylogenetic tree with 1000 bootstrap replicates. Based on phylogenetic trees, the pear HD-Zip family was classified into four major subfamilies (I–IV) with >50 % bootstrap values (Fig. 1). The number of genes was the greatest in subfamily I, followed by subfamilies II and IV, and the least in subfamily III, which is similar to the distribution of HD-Zip genes in other species (Table 1; Fig. 1).
Phylogenetic relationship and gene structure of pear HD-Zip genes. The unrooted tree of HD-Zip was constructed with the MEGA5.0 program using the full-length amino acid sequences of the 52 pear HD-Zip proteins by the Neighbor-Joining (NJ) method with 1000 bootstrap replicates. The bootstrap values are showed at the branches. The trees contain four subfamilies marked by different colored backgrounds. The proteins were named according to their location in the phylogenetic trees (see Fig. S1). Gene structure is indicated on the right panel. Exons are marked by pink boxes and introns are shown by black lines. The untranslated regions (UTRs) are indicated by gray boxes. The length of protein can be estimated using the scale at the bottom
To gain further assessment about the structural diversity of HD-Zip genes, we subsequently compared the exon/intron organizations and exon types and numbers of individual HD-Zip genes in pear. The results showed that the PbHBs were categorized into four groups. In each group, exon/intron structure of PbHBs was highly conserved; for instance, the HD-Zip genes in the same group exhibited similar exon/intron structures and intron numbers in the coding sequences. Together, the HD-Zip genes in subfamilies I and II have 1–3 exons, with the exception of PbHB13 and 16, which separately had 10 and 7 exons, respectively. However, the HD-Zip genes in subfamilies III contained 17–19 exons and subfamily IV contained 9–14 exons, with the exception of PbHB46, which contained 7 exons (Fig. 1).
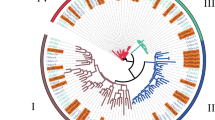
To evaluate the phylogenetic relationship of HD-Zip genes from different species, 52 full-length proteins sequences from pear, 70 from apple, and 47 from Arabidopsis were used to construct phylogenetic trees by three methods, namely minimal evolution (ME), maximum parsimony (MP), and neighbor-joining (NJ). The three phylogenetic trees produced by these methods were closely similar; therefore, the NJ tree was employed for further analyzes (Fig. 2). The results suggested that both PbHBs and MdHBs were divided into four subfamilies (I–IV) as those in Arabidopsis. Subfamily I was divided into six subclades, α, β, γ, δ, ε, and φ, which were consistent with the results in Arabidopsis (Agalou et al. 2008). Subfamily II was divided into six subclades (α–ζ). The HD-Zip subfamily III contained three subclades (α, β, and γ). The HD-Zip subfamily IV was divided into six subclades (α, β, γ, δ, ε, and φ).
Phylogenetic trees of Arabidopsis, apple and pear HD-Zip proteins. The full-length amino acid sequences from apple and pear were in conjunction with that of Arabidopsis using MUSCLE program, then the tree was generated with MEGA 5.0 program by the NJ method. 1000 replicates of the bootstrap were adopted, and the values were indicated at the branches. The unrooted tree generated by the same method were shown on the lower-left corner
Investigation of conserved motifs in PbHB genes
We identified 20 conserved motifs in the pear HD-Zip proteins by MEME web server (Fig. 3). Each of the putative motifs from MEME was annotated by searching Pfam and SMART. The categorization based on the distribution of motifs was consistent with the classification of the phylogenetic tree of HD-Zip gene family. Among 20 conserved motifs, motif 1 and motif 2 existing in all 52 PbHBs encoded the HD domain; motif 9 containing in all 52 PbHBs encoded the Zip domain; motifs 4, 5, 6, 7, 15, and 17 encoded the START domain, which were found in subfamily III and IV, while motif 12 encoding the MEKHLA domain was only observed in subfamily III. Table (S2) lists the complete information on conserved motifs.
Distribution of conserved motifs in pear HD-Zip proteins. Schematic representation of the conserved motifs in pear HD-ZIP proteins were identified by MEME web serve. Each colored box represents a motif in the protein. The conserved amino acid sequences and length of each motif are listed in Table S1. Detailed distribution of 20 motifs were shown in Table S2 and Fig. S1
Identification of putative stress-related and pathogen-related cis-element in the promoters of PbHB family I and II genes
Previous evidence showed that most members of HD-Zip subfamilies I and II are involved in abiotic stress, i.e., salt- and drought-induced stress (Harris et al. 2011). To illustrate the possible response of PbHB I and II genes in various stress responses, we analyzed putative abiotic stress-related cis-elements in the 2 kb promoter regions upstream of the transcription start site (ATG) of PbHB I and II genes. ABA-responsive elements, low temperature-responsive element (LTRE) and dehydration-responsive element (DRE) are major transcription factors that regulate ABA signaling involved in salt- and drought-induced stress (Yoshida et al. 2010). In this study, by searching the promoter sequence against PLACE database, ABRE was detected in most of the genes in subfamily I including PbHB1-4, PbHB17, PbHB1-9, PbHB1-10, PbHB1-11, PbHB1-13 and most of the genes in HD-Zip subfamily II with the exception of PbHB19. DRE which was detected in PbHB6, PbHB8, PbHB10, PbHB11, PbHB12, PbHB14, PbHB15, PbHB17, and PbHB18 of subfamily I and most of the genes in subfamily II with the exception of PbHB23, PbHB31, and PbHB32. LTREs which were detected in most of the genes including PbHB6-15, PbHB6-17, and PbHB6-18 of subfamily I and most of the genes in subfamily II with the exception of PbHB24 and PbHB32. In addition, MYCATRD22 and MYBCORE, which were involved in regulating drought stress, ABA signaling, and water stress (Urao et al. 1993; Abe et al. 1997), were also detected in most of the genes in subfamilies I and II with the exception of PbHB1 (Table 3).
Previous evidence also suggested that members of HD-Zip genes families I and II such as HaHB10 and HaHB4 were involved in response to biotic stress such as P. syringae (Dezar et al. 2011) and S. littoralis or S. frugiperda larvae (Manavella et al. 2008). Pathogen-related cis-elements including ELRECOREPCRP1, AGCBOXNPGLB, WBBOXPCWRKY1, WBOXATNPR1, WBOXNTERF3, GCCCORE, GT1GMSCAM4, and WRKY710S were detected by searching the promoter sequence against PLACE database. Notably, WRKY710S was found in almost all of HD-Zip genes in subfamilies I and II with the exception of PbHB1. WBOXATNPR1, GT1GMSCAM4, and WBOXNTERF3 were detected in the majority of HD HD-Zip genes in subfamilies I and II; however, AGCNOXNPGLB was detected only in PbHB7 and GCCCORE in PbHB7 and PbHB17. In addition, ELRECOREPCRP1 and WBBOXPCWRKY1 were observed in most genes in subfamilies I and II. Tables 3 and S3 provide detailed information about stress-related cis-elements.
Expression patterns of PbHB family I and II genes under abiotic and biotic stress
To obtain more comprehensive information about the regulation roles of pear HD-Zip genes in abiotic stress, i.e., salt- and drought-induced stress, we monitored the expression patterns of 20 genes in HD-Zip subfamilies I and II in 3-month-old tissue culture seedlings under salinity and drought stress. As shown in Fig. 4a, of the 20 HD-Zip genes, the expression level of 15 PbHB genes were up-regulated in response to drought stress; however, those of the other five were down-regulated. The expression level of PbHB2 was only up at 6 h after PEG6000 treatments, that of PbHB1 and PbHB20 was up only at 12 h, and that of PbHB4 and PbHB25 was activated only at 24 h. The transcription abundance of PbHB5, PbHB13, PbHB17, PbHB21, PbHB27, and PbHB30 was observed at all three time points (6–24 h), although the highest expression value of these genes was obtained at different time points. For example, the peak of PbHB13 and PbHB27 was observed at 6 h while the maximum of PbHB22 was obtained at 12 h and the summit of PbHB5 and PbHB17 was obtained at 24 h. The drought stress also induced a slight expression of PbHB7 and PbHB19.
Expression analysis of 20 PbHB I and II genes under PEG-6000, salinity and Alternaria alternata stress. Twenty genes were selected and their time-course expression profiles were evaluated under drought, salinity and pathogen stress treatment by quantitative real-time. a Fold changes were obtained from leaves under drought and salinity stress treatment. b Fold changes were obtained from seedlings infected by A. alternata. Relative expression was calculated using PbActin gene as an internal control. Data represent three independent experiments (±SE, n = 10)
As shown in Fig. 4b, of the 20 HD-Zip genes, nine PbHB genes showed response to salt-induced stress, whereas, the expression of the other 11 PbHB genes, including PbHB1, PbHB6, PbHB7, PbHB10, PbHB14, PbHB15, PbHB16, PbHB20, PbHB23, and PbHB25, was down-regulated. Of the nine up-regulated PbHB genes, the expression of PbHB2, PbHB21, and PbHB27 reached in the peaks at 6 h after NaCl treatment; however, the expression level of PbHB4, PbHB5, and PbHB13 exhibited the highest levels at 12 h and those of PbHB17, PbHB19, and PbHB29 was at 24 h. Notably, the expression of PbHB6, PbHB10, PbHB14, PbHB15, and PbHB23 was down-regulated under both drought- and salt-induced stress. PbHB17, PbHB19, and PbHB27 were up-regulated and displayed comparable expression patterns under both stress treatments.
Plant pathogen fungi A. alternata (Fr) Keissler cause black spot disease on Japanese pear cultivar and are major causes of yield loss (Suzuki et al. 2003). To identify the key regulators involved in pathogen–host interaction, 20 HD-Zip genes regulated by the infection of A. alternata Japanese pear pathotype were screened. It was noted that the primary necrotic lesion formed on leaves was at 6 days post-inoculation, subsequently by expanding lesions after 3 days (data not shown). Therefore, the leaves at 6- and 9-days post-inoculation were selected for monitoring the expression of HD-Zip genes. As shown in Fig. 4c, all 20 HD-Zip genes were responsive to the infection of A. alternata. The expression of these 20 HD-Zip genes was up-regulated at 6 days post-inoculation with the exception of PbHB17, PbHB20, and PbHB30, which showed maximum expression at 9 days post-inoculation. Among 20 HD-Zip genes, the expression level of PbHB16 was the highest, followed by the transcription level of PbHB4, PbHB6, PbHB10, PbHB13, PbHB15, PbHB21, PbHB22, PbHB24, and PbHB28; whereas, the expression of PbHB1 and PbHB5 was activated slightly.
Discussion
In this study, we extensively analyzed the HD-Zip genes in pear genome, including identifying of HD-Zip genes, analyzing of phylogenetic tree, gene structure, conserved motif, and monitoring expression patterns of HD-Zip I and II genes under abiotic and biotic stress conditions. The information contributes a comprehensive insight of the HD-Zip gene family in pear and lays the foundation for further functional feature of individual HD-Zip genes.
Distribution of members, exons, and motifs reflected evolutionary conservation of pear HD-Zip genes
In the present study, a total of 52 non-redundant HD-Zip genes were identified in the pear genome. Recent comprehensive analysis of genes encoding HD-Zip proteins from other plant species, such as Populus (Hu et al. 2012), peach (Zhang et al. 2014), soybean (Chen et al. 2014), Arabidopsis (Olsson et al. 2004), maize (Zhao et al. 2011), and rice (Agalou et al. 2008), enabled comparison of numbers and classification of these genes among different species. For some species, the larger the genome size of the species, the more will be the genes encoding HD-Zip proteins. Zhao et al. (2011) reported that the total number of HD-Zip genes is not positively correlated with the genome size of the corresponding species. Recent studies suggested that the tandem and segmental duplications exerted a substantial role in the expansion of gene family during the process of genome evolution (Zhao et al. 2011). However, notably, the classification of genes, the number distribution of members in individual family, and the percentage of each family relative to the total number of HD-Zip genes are conserved.
It is widely known that the intron/exon organization and intron types and numbers are clues of the evolution relationship within some gene families including HD-Zip genes (Zhao et al. 2011). Studies in peach showed one gene of 14 HD-Zip I members without an intron (Zhang et al. 2014). In Populus, 4 of 22 HD-Zip I genes had no introns (Hu et al. 2012). In our studies, 2 of 18 HD-Zip I genes had no intron, and this feature was not observed in other HD-Zip subfamilies in pear, similar to Populus and peach. The number of introns was the highest (17–19) in subfamily III compared with those in the HD-Zip subfamilies I, II, and IV. Most of the genes of subfamilies I and II contain the fewest of introns, while PbHB13 and PbHB16 were exceptions. These observations in pear are consistent with those in peach, maize, soybean, Populus, Arabidopsis, and rice.
Besides the conserved intron/exon organization and intron types and numbers, conserved number and distribution of motifs is another feature of HD-Zip genes in pear. Subfamily III contains the most number of motifs, while subfamilies I and II contain the fewest of motifs. In addition, the position of motifs among members of the individual subfamily is conserved. These two features in pear are consistent with other species, including Populus, peach, soybean, maize, rice, and Arabidopsis.
Diverse PbHB genes are involved in response to drought, salt stress, and pathogenic infection
Previous studies have shown that HD-Zip genes were involved in regulating adaptive response of plants to various stress conditions, such as drought (Zhang et al. 2012), salt (Harris et al. 2011), bacteria (Dezar et al. 2011), and insects (Manavella et al. 2008). In this study, the expression patterns of PbHB genes revealed that pear HD-Zip genes were widely involved in responding to drought- and salt-induced stress as well as pathogen infection. Interestingly, drought-responsive genes were more than salt-responsive genes. In addition, some drought up-regulated genes were down-regulated in salt stress. Most of the genes were co-expressed in response to both drought- and salt-induced stress. Notably, nearly all of detected genes were responsive to the infection of the pathogen. Moreover, the maximum expression level of most of the genes showed coincidence with the occurrence of symptoms. The result suggests HD-Zip I and II genes may play a vital role in regulating plant defense response to the pathogenic infection.
In conclusion, this study has provided us an insight about the information of HD-Zip family in pear. Identified and phylogenetic analysis of HD-Zip gene family is the first step toward a comprehensive functional feature. Expression patterns of HD-Zip genes under various stress conditions provided us with a benchmark to go a step further for functional analysis. Based on findings from this study, we will be able to narrow down candidate genes for detailed characterization and breeding of new cultivars that can adapt to various stress conditions. Future studies are warranted to identify the HD-Zip genes that are involved in the adaptive development of pear in unfavorable environmental conditions.
Author contribution statement
HW, JL, YC conceived and designed the experiments. HW performed the experiments. HW analyzed the data. JL, YC, XGL contributed reagents/materials/analysis tools. HW wrote the paper.
References
Abe H, Yamaguchi-Shinozaki K, Urao T, Iwasaki T, Hosokawa D, Shinozaki K (1997) Role of Arabidopsis MYC and MYB homologs in drought- and abscisic acid-regulated gene expression. Plant Cell 9:1859–1868
Agalou A et al (2008) A genome-wide survey of HD-Zip genes in rice and analysis of drought-responsive family members. Plant Mol Biol 66:87–103
Ariel F, Diet A, Verdenaud M, Gruber V, Frugier F, Chan R, Crespi M (2010) Environmental regulation of lateral root emergence in Medicago truncatula requires the HD-Zip I transcription factor HB1. Plant Cell 22:2171–2183
Bailey TL, Elkan C (1995) The value of prior knowledge in discovering motifs with MEME. Proc Int Conf Intell Syst Mol Biol 3:21–29
Chen X, Chen Z, Zhao H, Zhao Y, Cheng B, Xiang Y (2014) Genome-wide analysis of soybean HD-Zip gene family and expression profiling under salinity and drought treatments. PLoS One 9:e87156
Deng X et al (2006) A homeodomain leucine zipper gene from Craterostigma plantagineum regulates abscisic acid responsive gene expression and physiological responses. Plant Mol Biol 61:469–489
Dezar CA, Gago GM, Gonzalez DH, Chan RL (2005) Hahb-4, a sunflower homeobox-leucine zipper gene, is a developmental regulator and confers drought tolerance to Arabidopsis thaliana plants. Transgenic Res 14:429–440
Dezar CA et al (2011) HAHB10, a sunflower HD-Zip II transcription factor, participates in the induction of flowering and in the control of phytohormone-mediated responses to biotic stress. J Exp Bot 62:1061–1076
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Finn RD et al (2006) Pfam: clans, web tools and services. Nucleic Acids Res 34:D247–D251
Frank W, Phillips J, Salamini F, Bartels D (1998) Two dehydration-inducible transcripts from the resurrection plant Craterostigma plantagineum encode interacting homeodomain-leucine zipper proteins. Plant J 15:413–421
Guo AY, Zhu QH, Chen X, Luo JC (2007) GSDS: a gene structure display server. Hereditas 29:1023–1026
Guo Y et al (2014) Comparative genomic analysis of Klebsiella pneumonia (LCT-KP214) and a mutant strain (LCT-KP289) obtained after spaceflight. BMC Genom 15:589
Harris JC, Hrmova M, Lopato S, Langridge P (2011) Modulation of plant growth by HD-Zip class I and II transcription factors in response to environmental stimuli. New Phytol 190:823–837
Henriksson E, Olsson A, Johannesson H, Johansson H, Hanson J, Engstrom P, Soderman E (2005) Homeodomain leucine zipper class I genes in Arabidopsis. Expression patterns and phylogenetic relationships. Plant Physiol 139:509–518
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database. Nucl Acids Res 27:297–300
Hu R, Chi X, Chai G et al (2012) Genome-wide identification, evolutionary expansion, and expression profile of homeodomain-leucine zipper gene family in poplar (Populus trichocarpa). PLoS One 7:e31149. doi:10.1371/journal.pone.0031149
Johannesson H, Wang Y, Engstrom P (2001) DNA-binding and dimerization preferences of Arabidopsis homeodomain-leucine zipper transcription factors in vitro. Plant Mol Biol 45:63–73
Johannesson H, Wang Y, Hanson J, Engstrom P (2003) The Arabidopsis thaliana homeobox gene ATHB5 is a potential regulator of abscisic acid responsiveness in developing seedlings. Plant Mol Biol 51:719–729
Leblay C, Chevreau E, Raboin LM (1991) Adventitious shoot regeneration from in vitro leaves of several pear cultivars (Pyrus communis L.). Plant Cell Rep 25:99–105
Letunic I, Doerks T, Bork P (2009) SMART 6: recent updates and new developments. Nucleic Acids Res 37:D229–D232
Liu W, Fu R, Li Q, Li J, Wang L, Ren Z (2013) Genome-wide identification and expression profile of homeodomain-leucine zipper Class I gene family in Cucumis sativus. Gene 531:279–287
Liu J et al (2014) Genome-wide analysis and expression profile of the bZIP transcription factor gene family in grapevine (Vitis vinifera). BMC Genom 15:281
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408
Manavella PA, Dezar CA, Bonaventure G, Baldwin IT, Chan RL (2008) HAHB4, a sunflower HD-Zip protein, integrates signals from the jasmonic acid and ethylene pathways during wounding and biotic stress responses. Plant J 56:376–388
Moens CB, Selleri L (2006) Hox cofactors in vertebrate development. Dev Biol 291:193–206
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue culture. Physiol Plant 15:473–497
Nakashima K, Ito Y, Yamaguchi-Shinozaki K (2009) Transcriptional regulatory networks in response to abiotic stresses in Arabidopsis and grasses. Plant Physiol 149:88–95
Nicholas K, Nicholas H, Deerfield D (1997) GeneDoc: analysis and visualization of genetic variation. EMBNEW.NEWS 4:14
Olsson AS, Engstrom P, Soderman E (2004) The homeobox genes ATHB12 and ATHB7 encode potential regulators of growth in response to water deficit in Arabidopsis. Plant Mol Biol 55:663–677
Righetti L et al (2014) Elimination of the nptII marker gene in transgenic apple and pear with a chemically inducible R/Rs recombinase. Plant Cell Tissue Org 117:335–348
Schena M, Davis RW (1992) HD-Zip proteins: members of an Arabidopsis homeodomain protein superfamily. Proc Natl Acad Sci USA 89:3894–3898
Sessa G, Morelli G, Ruberti I (1993) The Athb-1 and -2 HD-Zip domains homodimerize forming complexes of different DNA binding specificities. EMBO J 12:3507–3517
Shao Y, Qin Y, Zou Y, Ma F (2014) Genome-wide identification and expression profiling of the SnRK2 gene family in Malus prunifolia. Gene 552:87–97
Skinner D, Gasser C (2009) Expression-based discovery of candidate ovule development regulators through transcriptional profiling of ovule mutants. BMC Plant Biol 9:29
Soderman E, Hjellstrom M, Fahleson J, Engstrom P (1999) The HD-Zip gene ATHB6 in Arabidopsis is expressed in developing leaves, roots and carpels and up-regulated by water deficit conditions. Plant Mol Biol 40:1073–1083
Son O et al (2010) ATHB12, an ABA-inducible Homeodomain-Leucine Zipper (HD-Zip) protein of Arabidopsis, negatively regulates the growth of the inflorescence stem by decreasing the expression of a gibberellin 20-oxidase gene. Plant Cell Physiol 51:1537–1547
Sultan SE (2010) Plant developmental responses to the environment: eco-devo insights. Curr Opin Plant Biol 13:96–101
Suzuki T, Shinogi T, Narusaka Y, Park P (2003) Infection behavior of Alternaria alternata Japanese pear pathotype and localization of 1,3-β-d-glucan in compatible and incompatible interactions between the pathogen and host plants. J Gen Plant Pathol 69:91–100
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tron AE, Bertoncini CW, Chan RL, Gonzalez DH (2002) Redox regulation of plant homeodomain transcription factors. J Biol Chem 277:34800–34807
Urao T, Yamaguchi-Shinozaki K, Urao S, Shinozaki K (1993) An Arabidopsis myb homolog is induced by dehydration stress and its gene product binds to the conserved MYB recognition sequence. Plant Cell 5:1529–1539
Wang H, Chang YH, Chen ZY (2006) Study on biological characteristics of Alternaria alternate caused occurrence of pear black spot. J Fruit Sci 23:247–251
Wang H, Liu G, Li C, Powell AL, Reid MS, Zhang Z, Jiang CZ (2013a) Defence responses regulated by jasmonate and delayed senescence caused by ethylene receptor mutation contribute to the tolerance of petunia to Botrytis cinerea. Mol Plant Pathol 14:453–469
Wang H, Stier G, Lin J, Liu G, Zhang Z, Chang YH, Reid MS, Jiang CZ (2013b) Transcriptome changes associated with delayed flower senescence on transgenic petunia by inducing expression of etr1-1, a mutant ethylene receptor. PLoS One 8:e65800
Wang H, Li GB, Zhang DY, Lin J, Sheng BL, Han JL, Chang YH (2013c) Biological functions of HD-Zip transcription factors. Yi Chuan 35(10):1179–1188
Yoshida T, Fujita Y, Sayama H, Kidokoro S, Maruyama K, Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2010) AREB1, AREB2, and ABF3 are master transcription factors that cooperatively regulate ABRE-dependent ABA signaling involved in drought stress tolerance and require ABA for full activation. Plant J 61:672–685
Zhang S, Haider I, Kohlen W, Jiang L, Bouwmeester H, Meijer AH, Schluepmann H, Liu CM, Ouwerkerk PB (2012) Function of the HD-Zip I gene Oshox22 in ABA-mediated drought and salt tolerances in rice. Plant Mol Biol 80:571–585
Zhang CH, Ma RJ, Shen ZJ, Sun X, Korir NK, Yu ML (2014) Genome-wide analysis of the Homeodomain-Leucine Zipper (HD-ZIP) gene family in peach (Prunus persica). Genet Mol Res 13:2654–2668
Zhao Y, Zhou Y, Jiang H, Li X, Gan D, Peng X, Zhu S, Cheng B (2011) Systematic analysis of sequences and expression patterns of drought-responsive members of the HD-Zip gene family in maize. PLoS One 6:e28488
Acknowledgments
This work was supported by the National Natural Science Foundation of China (Grant No. 31300469), the Jiangsu Natural Science Foundation of China (Grant No. BK 20141383) and the Jiangsu Agricultural Science and Technology Independent Innovation Fund of China [Grant No. CX(13)5015, CX(14)5016]. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by G. K. Pandey.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11738_2015_1933_MOESM1_ESM.pptx
Supplementary material 1 (PPTX 153 kb). Fig. S1 Motifs of 52 pear HD-Zip proteins. 20 motifs were identified through MEME (http://meme.nbcr.net/meme/). Motif organization of 52 pear HD-Zip proteins was investigated through MAST (http://meme.nbcr.net/meme/)
11738_2015_1933_MOESM2_ESM.docx
Supplementary material 2 (DOCX 15 kb). Table S1. A list of primer sequences of the 20 selected PbHB I and II genes for qRT-PCR analysis
11738_2015_1933_MOESM5_ESM.txt
Supplementary material 5 (TXT 111 kb). Text S1. A complete list of HD-Zip gene sequences identified from the pear genome in the present study. The list contains 52 PbHB sequences in this study. Amino acid sequences were deduced from their corresponding coding sequences and genomic DNA sequences were obtained from Pear Genome Project and Genome Database for Rosaceae
11738_2015_1933_MOESM6_ESM.txt
Supplementary material 6 (TXT 52 kb). Text S2. A list of HD-Zip protein sequences identified from the apple genome and the Arabidopsis genome in this study
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Wang, H., Lin, J., Li, X.G. et al. Genome-wide identification of pear HD-Zip gene family and expression patterns under stress induced by drought, salinity, and pathogen. Acta Physiol Plant 37, 189 (2015). https://doi.org/10.1007/s11738-015-1933-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-015-1933-5