Abstract
Purpose of the Review
To evaluate recent studies related to the paradox of high HDL-C with mortality and atherosclerotic cardiovascular disease (ASCVD) risk.
Recent Findings
Two observational studies (Cardiovascular Health in Ambulatory Care Research Team [CANHEART] and Copenhagen City Heart Study and the Copenhagen General Population Study [Copenhagen Heart Studies]) of adults without pre-existing ASCVD have shown a significant U-shaped association of HDL-C with all-cause and cause-specific mortality. Both studies showed that low HDL-C levels consistently increased hazard risk (HR) for all-cause and cause-specific mortality. In the CANHEART study, high HDL-C levels, HDL-C > 90 mg/dL, were associated with increased HR for non-CVD/non-cancer mortality. In the Copenhagen Heart Studies, women with HDL-C ≥ 135 mg/dL showed increased HR for all-cause and CVD mortality, while men with HDL-C > 97 mg/dL showed increased HR for all-cause and CVD mortality. Genetic association studies failed to show that genetic etiologies of high HDL-C significantly reduced risk for myocardial infarction (MI), while hepatocyte nuclear factor-4 (HNF4A) was significantly associated with high HDL-C and increased MI risk. Candidate gene studies have identified scavenger receptor B class I (SCARB1) and lymphocyte activation gene-3 (LAG3) as genes significantly associated with high HDL-C and increased MI risk.
Summary
Low HDL-C remains as a significant factor for increased disease risk while high HDL-C levels are not associated with cardioprotection. Clinical CVD risk calculators need revision.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
HDL-cholesterol (HDL-C) and its use in atherosclerotic cardiovascular disease (ASCVD) risk calculators is well-established, and its measurement remains a cornerstone in medical education and clinical practice [1]. As an example, in the Framingham Risk Calculator, HDL-C levels ≥ 60 mg/dL inform the health care provider that ASCVD risk factor calculations can be reduced by 1, concluding that HDL-C levels above this cut point are cardioprotective [2]. However, there is now an abundance of data from randomized clinical trials (RCTs), observational, genetic association and candidate gene studies that challenge this paradigm of high HDL-C levels and cardioprotection [3].
RCTs are the gold standard by which safety and efficacy of potential therapeutics for human use are adjudicated. Within the past 15 years, a number of pharmaceutical companies have conducted RCTs to raise HDL-C levels as a means to further reduce CVD events, with the majority targeting inhibition of cholesteryl ester transfer protein (CETP) to achieve their stated goals [4,5,6]. These trials resulted in failures or underwhelming findings to support their primary objectives, leading most of these companies to abandon pursuit of further studies related to HDL metabolism.
This review will not address the exhaustive literature surrounding the composition, subfractions, and functional assays attributed to HDL (such as cholesterol efflux) but will instead focus on the recent literature related to high HDL-C levels and the paradox of increased all-cause, CVD, and non-CVD mortality.
Observational Studies of High HDL-C
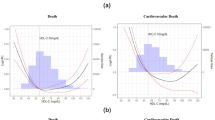
Results from studies like the Cardiovascular Health in Ambulatory Care Research Team Study (CANHEART) and the Copenhagen Heart Studies came to the same conclusion that HDL-C levels demonstrated a U-shaped association with all-cause and cause-specific mortality (Table 1) [7••, 8••]. In the CANHEART Study, investigators examined the association of fasting cholesterol measurements with cause-specific mortality in a large observational cohort (n = 631,762 men and women) of subjects living in Ontario, Canada, who were between the ages of 40 and 105 years of age without pre-existing CVD conditions. These investigators observed a U-shaped association of HDL-C with CVD and non-CVD mortality. Specifically, men with low HDL-C (≤ 30 mg/dL) had an increased CVD mortality (hazard ratio [HR] 1.81; 95th percentile confidence intervals [CI] 1.45–2.25), cancer mortality (HR 1.61, CI 1.32–1.97), and other mortality HR (HR 2.01, CI 1.63–2.47) compared with the reference group (HDL-C 41–50 mg/dL), while men with HDL-C > 90 mg/dL also showed significantly increased risk (HR 1.72, CI 1.103–2.682) for non-CVD death. In women, those with low HDL-C (≤ 30 mg/dL) had an increased CVD (HR 2.26, CI 1.56–3.29), cancer mortality (HR 1.96, CI 1.43–2.69), and other mortality risk (HR 2.86, CI 2.17–3.76) compared with the reference group (HDL-C 51–60 mg/dL), while women with HDL-C > 90 mg/dL had an increased risk for non-CVD death (HR 1.32, CI 1.01–1.71). In both men and women, the CANHEART study found that high HDL-C levels did not significantly reduce CVD mortality, indicating that high HDL-C levels did not confer cardioprotection.
In the Copenhagen Heart Studies [8••], investigators examined the association of HDL-C with all-cause, CVD, and non-CVD mortality in two large prospective studies, the Copenhagen City Heart Study and the Copenhagen General Population Study (combined total of 52,268 white men and 64,240 white women of Danish ancestry). As observed in the CANHEART study, these investigators observed a similar pattern of a U-shaped association of HDL-C to mortality. Specifically, the adjusted HR for all-cause mortality was 1.27 (CI 1.12–1.45) in men with HDL-C < 39 mg/dL, 1.36 (CI 1.09–1.70) in men with HDL-C between 97 and 115 mg/dL, and 2.06 (CI 1.44–2.95) for men with HDL-C ≥ 116 mg/dL. In women, the adjusted HR for all-cause mortality was 1.78 (CI 1.43–2.22) for those with HDL-C < 39 mg/dL, 1.10 (CI 0.83–1.46) for those with HDL-C between 116 and 134 mg/dL, and 1.68 (CI 1.09–2.58) in those with HDL-C ≥ 135 mg/dL. These investigators also examined cause-specific mortality rates in subjects with extreme HDL-C, such that in men and women, the HR for CVD mortality was 2.53 and 2.89, respectively. The HR for cancer mortality was 1.76 in men and 1.33 in women. The HR for other mortality was 1.90 in men and 1.51 in women. No significant increase or decrease in risk for CVD outcomes (myocardial infarction [MI], ischemic CVD, and ischemic stroke) was found in subjects with extreme HDL-C levels.
Additionally, investigators from the Copenhagen Heart Studies also examined the association of HDL-C with hospitalizations due to infectious diseases [9]. These investigators again observed a U-shaped association in that subjects with low HDL-C (31 mg/dL), and those with high HDL-C (100 mg/dL) had HRs of 1.75 (CI 1.31–2.34) and 1.43 (CI 1.16–1.76), respectively, compared with the reference group. On the other hand, in participants of the UK Biobank study, Trinder et al. [10] observed that genetic etiologies underlying high HDL-C (HDL polygenic score) were significantly associated with lower infectious disease hospitalizations. Altogether, the results are intriguing and should provide an impetus for further mechanistic and causal studies related to immunological functions of HDL and risk for disease [11, 12].
Genetic Association Studies of High HDL-C
One of the largest-to-date of the genetic association studies related to HDL-C and CVD risk was conducted by Voight et al. [3]. These investigators conducted two Mendelian randomization analyses in order to determine if genetic etiologies of HDL-C were a causal factor in MI risk. In the first analysis, they examined whether a loss-of-function coding single nucleotide polymorphism (SNP) in the endothelial lipase gene (LIPG rs61755018, Asn396Ser) would predict MI risk in a combined population of subjects with MI (n = 20,913 cases) or controls (n = 95,407). In the second analysis (n = 12,482 MI cases and n = 41,331 controls), they examined whether a genetic score encompassing fourteen common SNPs associated with HDL-C levels would predict lower MI risk. The results showed that neither the LIPG SNP nor the genetic score were significantly associated with lower MI risk, concluding that HDL-C per se was not a causal factor in MI risk. Of note, the investigators did identify HNF4A (hepatocyte nuclear factor 4A) as being associated with increased HDL-C and MI risk, but this association was not explored further but could be a candidate gene for the high HDL-C paradox. Others have noted that HNF4A is a key nuclear receptor transcription factor regulating bile acid synthesis and the final steps of reverse cholesterol transport [13, 14].
These investigators subsequently used molecular inversion probes (MIP) to re-sequence coding and regulatory regions of GALNT2, APOA5, APOC3, SCARB1, CCDC92, ZNF664, CETP, and LIPG in 797 subjects with extremely high HDL-C levels compared with 735 subjects with low-to-normal HDL-C [15]. The main objective of this re-sequencing effort was to identify variations within noncoding regulatory regions within these genes significantly associated with HDL-C and to perform this re-sequencing effort in a smaller sample size. Using the MIP methodology, the investigators were able to replicate their previous findings of certain SNPs associated with HDL-C but also identified a rare noncoding variant within CETP that was significantly associated with HDL-C.
Candidate Gene Studies Associated with High HDL-C
We and others have used a candidate gene approach, with a particular focus on scavenger receptor class B type I (SCARB1 gene and SR-B1 protein) gene due to its physiological role in HDL metabolism and lipid uptake in mice and in man [16,17,18,19,20,21,22,23]. Zanoni et al. [24] had identified a rare coding variant (P376L) within the SCARB1 gene that was significantly associated with increased HDL-C and MI risk, providing more evidence for the role of SCARB1 in the paradox of HDL-C.
In the Multi-Ethnic Study of Atherosclerosis (MESA), we showed that the common noncoding rs10846744 SNP within SCARB1 was significantly associated with increased subclinical atherosclerosis and incident MI risk [19,20,21]. In MESA and as reported in other databases, the frequency of carriers homozygous for the risk C allele was 7.9% in Whites, 55% in African-Americans, 21.6% in Chinese-Americans, and 15.4% in Hispanics. Carriers of the C allele were found to have an odds ratio of 1.45 for MI compared with carriers of the G allele; henceforth, we refer to the C allele as the MI risk allele and the G allele as the reference allele. Our observations in MESA were also reported in CARDIoGRAM in that the rs10846744 risk allele was significantly associated with increased prevalent CVD [25].
We were intrigued about the mechanism by which the rs10846744 SNP increased MI risk given its location within a regulatory enhancer region of SCARB1. In unpublished work, we did not observe significant changes in SR-B1 RNA or protein expression in either peripheral monocyte-derived macrophages or EBV-transformed B lymphoblasts derived from risk C carriers. Using bioinformatics data from the UCSC browser and our own ChIP-Seq data (GEO accession GSE110761), we confirmed transcription factors were binding at this site, which is characteristic of a regulatory or enhancer region. With this observation, we subjected EBV-transformed B lymphoblasts from carriers homozygous for the reference or MI risk allele to RNA-Seq, which revealed that carriers of the risk C allele had fivefold less of the immune checkpoint inhibitor lymphocyte activation gene-3 (LAG3) RNA expression compared with carriers of the reference G allele [26•]. We then asked the question of whether from this regulatory region there contained significant chromatin gene-gene interactions between SCARB1 and LAG3. We used a number of chromatin capture assays (Hi-C, 4C-Seq) to examine chromatin gene-gene interactions between these loci on chromosome 12 (cis) and with other chromosomes (trans or long distance interactions) (unpublished data). Pioneers in the field of 3D chromatin architecture have developed and continue to refine methodologies that evaluate the effect of cis and trans gene-gene interactions on RNA transcriptional regulation [27,28,29,30]. Briefly, Hi-C assays evaluate unbiased global chromatin interactions (many-to-many) without immunoprecipitation followed by high depth next-gen sequencing (NGS). Bioinformatics analysis can be performed using programs such as Hi-C Pro and Juicer, and Juicebox and the WashU Epigenome Browser can be used for visualization of the chromatin contacts [31,32,33]. Circularized chromatin conformation capture (4C-Seq) assays are very informative in evaluating chromatin interactions from a specified gene locus based on inverse PCR reactions (one-to-many). Bioinformatics analysis can be performed using 4Cker software with readouts described as near bait (NB), cis, and trans [34]. RNA-Seq is then performed to determine the correlation between 4C-Seq chromatin contacts and gene expression. In this manner, the chromatin capture assays provide a tool to integrate changes in gene-gene interaction networks with differentially expressed RNA transcription, such as what we observed between SCARB1 and LAG3.
When we were examining the associations of SCARB1 rs10846744 with CVD risk, no matter the extent of adjusting the multivariable regression models with traditional CVD risk factors (including HDL-C), we could not identify a factor that attenuated the significant association of rs10846744 with CVD risk [19, 20]. Thus, when we first identified LAG3 via RNA-Seq analyses, we were intrigued by the fact that LAG3 is an immune checkpoint inhibitor. LAG3 is a gene that belongs to the Ig superfamily and is a significant suppressor, or immune checkpoint inhibitor, in the expansion of activated T effector (Teff) cells [35]. In humans, the LAG3 gene resides on the short arm of chromosome 12 (12p13.32) and is within 8.4 kB of CD4 [36]. LAG3 protein is a ligand to MHC class II molecules on antigen-presenting cells (APCs), and through this interaction operates to limit effector T cell expansion and homeostasis [37]. LAG3 is expressed in B cells, T cells, and NK lymphocytes, monocytes, and dendritic cells [DCs] [38, 39], and its distribution is approximately 50% intracellular and 50% cell surface [40]. Activation of cells promotes transit of intracellular LAG3 to the cell surface, where LAG3 is then subject to cleavage by ADAM10 and ADAM17 metalloproteases, which results in soluble LAG3 (sLAG3) [41]. Of the two metalloproteases, ADAM17 is considered the one to primarily affect cleavage of transmembrane LAG3 to release sLAG3. In addition to transmembrane LAG3 binding to MHC class II to limit effector T cell expansion, in vitro studies show that sLAG3 also binds to MHC class II and regulates CD4-driven signaling pathways [42].
We were the first to report in MESA that plasma sLAG3 protein levels were an independent predictor of HDL-C and increased CVD risk [26•]. In subjects with hyperalphalipoproteinemia (HDL-C ≥ 60 mg/dL), low plasma LAG3 protein levels significantly increased MI risk (odds ratio 1.45) and plasma LAG3 added predictive value to the Framingham risk score. Plasma LAG3 was inversely associated with HDL-C levels (p = 0.007), IL-10 levels (p < 0.0001), and with hs-CRP (p = 0.087), although the latter was of borderline significance. In a multivariate regression analysis, rs10846744 remained as an independent predictor of sLAG3, demonstrating the significant influence of this noncoding SCARB1 variant on LAG3 expression. Using a number of in vitro approaches, we showed that cellular LAG3 protein deficiency in EBV-transformed human B lymphoblasts reduced lipid raft formation and phosphosignaling, increased secretion of pro-inflammatory cytokines such as TNFα, and exhibited lower secretion of IL-10 [26•].
The association of LAG3 deficiency with increased CVD risk was also observed in a recent clinical study. Zhu et al. [43] observed that Chinese patients with documented coronary artery disease demonstrated significantly lower expression by flow cytometry of CD49b+ LAG3+ Tregs (Tr1) cells compared with control subjects. This immunosuppressive Treg subset has been characterized by the surface expression of LAG3 and a source of IL-10 secretion [44,45,46,47]. Our results and those of Zhu et al. [43] are consistent that humans with LAG3 deficiency are at increased risk for CVD and have lower IL-10 levels.
Conclusions
We and others are challenging the paradigm that high HDL-C levels are cardioprotective. Since the Framingham Risk Calculator was first developed and to the present time, HDL-C ≥ 60 mg/dL levels can be used to reduce risk factors in clinical decision-making for therapeutic intervention. The now large body of evidence from RCTs, large observational genetic association studies along with candidate gene studies like SCARB1 and LAG3 suggest that there is no clinical utility in the use of high HDL-C levels alone in CVD risk assessments. This should prompt a reassessment of high HDL-C cutoffs in CVD risk calculations, and clinical health care providers should be duly cautioned in advising patients that high HDL-C levels are cardioprotective. To be clear, observational studies are uniform in that low HDL-C levels are significantly associated with increased risk for CVD and all-cause mortality. In the meantime, more financial resources from federal and non-federal sources are needed to support biomedical research in identifying the causal role of HDL in health and disease states.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Grundy SM, Stone NJ, Bailey AL, Beam C, Birtcher KK, Blumenthal RS, et al. 2018 AHA/ACC/AACVPR/AAPA/ABC/ACPM/ADA/AGS/APhA/ASPC/NLA/PCNA guideline on the management of blood cholesterol: a report of the American College of Cardiology/American Heart Association task force on clinical practice guidelines. Circulation. 2019;139(25):e1082–e143. https://doi.org/10.1161/CIR.0000000000000625.
Bosomworth NJ. Practical use of the Framingham risk score in primary prevention: Canadian perspective. Can Fam Physician. 2011;57(4):417–23.
Voight BF, Peloso GM, Orho-Melander M, Frikke-Schmidt R, Barbalic M, Jensen MK, et al. Plasma HDL cholesterol and risk of myocardial infarction: a Mendelian randomisation study. Lancet. 2012;380(9841):572–80. https://doi.org/10.1016/s0140-6736(12)60312-2.
Barter PJ, Caulfield M, Eriksson M, Grundy SM, Kastelein JJ, Komajda M, et al. Effects of torcetrapib in patients at high risk for coronary events. N Engl J Med. 2007;357(21):2109–22. https://doi.org/10.1056/NEJMoa0706628.
Schwartz GG, Olsson AG, Abt M, Ballantyne CM, Barter PJ, Brumm J, et al. Effects of dalcetrapib in patients with a recent acute coronary syndrome. N Engl J Med. 2012;367(22):2089–99. https://doi.org/10.1056/NEJMoa1206797.
Group HTRC, Bowman L, Hopewell JC, Chen F, Wallendszus K, Stevens W, et al. Effects of anacetrapib in patients with atherosclerotic vascular disease. N Engl J Med. 2017;377(13):1217–27. https://doi.org/10.1056/NEJMoa1706444.
•• Ko DT, Alter DA, Guo H, Koh M, Lau G, Austin PC, et al. High-density lipoprotein cholesterol and cause-specific mortality in individuals without previous cardiovascular conditions: the CANHEART study. J Am Coll Cardiol. 2016;68(19):2073–83. https://doi.org/10.1016/j.jacc.2016.08.038This large observational study from Canada showed a U-shaped assocation of HDL-cholesterol with increased mortality.
•• Madsen CM, Varbo A, Nordestgaard BG. Extreme high high-density lipoprotein cholesterol is paradoxically associated with high mortality in men and women: two prospective cohort studies. Eur Heart J. 2017;38(32):2478–86. https://doi.org/10.1093/eurheartj/ehx163This large observational study from Denmark showed a U-shaped association of HDL-cholesterol with increased cardiovascular and other mortality.
Madsen CM, Varbo A, Tybjaerg-Hansen A, Frikke-Schmidt R, Nordestgaard BG. U-shaped relationship of HDL and risk of infectious disease: two prospective population-based cohort studies. Eur Heart J. 2018;39(14):1181–90. https://doi.org/10.1093/eurheartj/ehx665.
Trinder M, Walley KR, Boyd JH, Brunham LR. Causal inference for genetically determined levels of high-density lipoprotein cholesterol and risk of infectious disease. Arterioscler Thromb Vasc Biol. 2020;40(1):267–78. https://doi.org/10.1161/ATVBAHA.119.313381.
Guo L, Ai J, Zheng Z, Howatt DA, Daugherty A, Huang B, et al. High density lipoprotein protects against polymicrobe-induced sepsis in mice. J Biol Chem. 2013;288(25):17947–53. https://doi.org/10.1074/jbc.M112.442699.
Catapano AL, Pirillo A, Bonacina F, Norata GD. HDL in innate and adaptive immunity. Cardiovasc Res. 2014;103(3):372–83. https://doi.org/10.1093/cvr/cvu150.
Ohoka N, Okuhira K, Cui H, Wu W, Sato R, Naito M, et al. HNF4alpha increases liver-specific human ATP-binding cassette transporter A1 expression and cholesterol efflux to apolipoprotein A-I in response to cholesterol depletion. Arterioscler Thromb Vasc Biol. 2012;32(4):1005–14. https://doi.org/10.1161/ATVBAHA.111.238360.
Crestani MEDF, Caruso D, Mitro N, Gilardi F, Chacon ABV, Patelli R, et al. LXR (liver X receptor) and HNF-4 (hepatocyte nuclear factor-4): key regulators in reverse cholesterol transport. Biochem Soc Trans. 2001;32:92–6.
Khetarpal SA, Babb PL, Zhao W, Hancock-Cerutti WF, Brown CD, Rader DJ, et al. Multiplexed targeted resequencing identifies coding and regulatory variation underlying phenotypic extremes of high-density lipoprotein cholesterol in humans. Circ Genom Precis Med. 2018;11(7):e002070. https://doi.org/10.1161/CIRCGEN.117.002070.
Acton SRA, Landschulz KT, Xu S, Hobbs HH, Krieger M. Identification of scavenger receptor SR-BI as a high density lipoprotein receptor. Science. 1996;271:518–20.
Rigotti A, Trigatti BL, Penman M, Rayburn H, Herz J, Krieger M. A targeted mutation in the murine gene encoding the high density lipoprotein (HDL) receptor scavenger receptor class B type I reveals its key role in HDL metabolism. Proc Natl Acad Sci U S A. 1997;94(23):12610–5. https://doi.org/10.1073/pnas.94.23.12610.
Trigatti B, Rayburn H, Vinals M, Braun A, Miettinen H, Penman M, et al. Influence of the high density lipoprotein receptor SR-BI on reproductive and cardiovascular pathophysiology. Proc Natl Acad Sci U S A. 1999;96(16):9322–7. https://doi.org/10.1073/pnas.96.16.9322.
Naj AC, West M, Rich SS, Post W, Kao WH, Wasserman BA, et al. Association of scavenger receptor class B type I polymorphisms with subclinical atherosclerosis: the multi-ethnic study of atherosclerosis. Circ Cardiovasc Genet. 2010;3(1):47–52. https://doi.org/10.1161/CIRCGENETICS.109.903195.
Manichaikul A, Naj AC, Herrington D, Post W, Rich SS, Rodriguez A. Association of SCARB1 variants with subclinical atherosclerosis and incident cardiovascular disease: the multi-ethnic study of atherosclerosis. Arterioscler Thromb Vasc Biol. 2012;32(8):1991–9. https://doi.org/10.1161/ATVBAHA.112.249714.
Manichaikul A, Wang XQ, Musani SK, Herrington DM, Post WS, Wilson JG, et al. Association of the lipoprotein receptor SCARB1 common missense variant rs4238001 with incident coronary heart disease. PLoS One. 2015;10(5):e0125497. https://doi.org/10.1371/journal.pone.0125497.
Prabhudas M, Bowdish D, Drickamer K, Febbraio M, Herz J, Kobzik L, et al. Standardizing scavenger receptor nomenclature. J Immunol. 2014;192(5):1997–2006. https://doi.org/10.4049/jimmunol.1490003.
Thuahnai ST, Lund-Katz S, Williams DL, Phillips MC. Scavenger receptor class B, type I-mediated uptake of various lipids into cells. Influence of the nature of the donor particle interaction with the receptor. J Biol Chem. 2001;276(47):43801–8. https://doi.org/10.1074/jbc.M106695200.
Zanoni P, Khetarpal SA, Larach DB, Hancock-Cerutti WF, Millar JS, Cuchel M, et al. Rare variant in scavenger receptor BI raises HDL cholesterol and increases risk of coronary heart disease. Science. 2016;351(6278):1166–71. https://doi.org/10.1126/science.aad3517.
Grallert H, Dupuis J, Bis JC, Dehghan A, Barbalic M, Baumert J, et al. Eight genetic loci associated with variation in lipoprotein-associated phospholipase A2 mass and activity and coronary heart disease: meta-analysis of genome-wide association studies from five community-based studies. Eur Heart J. 2012;33(2):238–51. https://doi.org/10.1093/eurheartj/ehr372.
• Golden D, Kolmakova A, Sura S, Vella AT, Manichaikul A, Wang XQ, et al. Lymphocyte activation gene 3 and coronary artery disease. JCI Insight. 2016;1(17):e88628. https://doi.org/10.1172/jci.insight.88628This study was the first to report that deficiency of the immune checkpoint molecule, lymphocyte activation gene-3, was significantly associated with increased HDL-cholestersol and increased risk of myocardial infarction.
Cope NF, Fraser P. Chromosome conformation capture. Cold Spring Harb Protoc. 2009;2009(2):pdb prot5137. https://doi.org/10.1101/pdb.prot5137.
Nagano T, Lubling Y, Stevens TJ, Schoenfelder S, Yaffe E, Dean W, et al. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature. 2013;502(7469):59–64. https://doi.org/10.1038/nature12593.
Nagano T, Lubling Y, Yaffe E, Wingett SW, Dean W, Tanay A, et al. Single-cell Hi-C for genome-wide detection of chromatin interactions that occur simultaneously in a single cell. Nat Protoc. 2015;10(12):1986–2003. https://doi.org/10.1038/nprot.2015.127.
Lieberman-Aiden E, van Berkum NL, Williams L, Imakaev M, Ragoczy T, Telling A, et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science. 2009;326(5950):289–93. https://doi.org/10.1126/science.1181369.
Durand NC, Robinson JT, Shamim MS, Machol I, Mesirov JP, Lander ES, et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 2016;3(1):99–101. https://doi.org/10.1016/j.cels.2015.07.012.
Durand NC, Shamim MS, Machol I, Rao SS, Huntley MH, Lander ES, et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 2016;3(1):95–8. https://doi.org/10.1016/j.cels.2016.07.002.
Li D, Hsu S, Purushotham D, Sears RL, Wang T. WashU Epigenome browser update 2019. Nucleic Acids Res. 2019;47(W1):W158–W65. https://doi.org/10.1093/nar/gkz348.
Raviram R, Rocha PP, Muller CL, Miraldi ER, Badri S, Fu Y, et al. 4C-ker: a method to reproducibly identify genome-wide interactions captured by 4C-Seq experiments. PLoS Comput Biol. 2016;12(3):e1004780. https://doi.org/10.1371/journal.pcbi.1004780.
Huard B, Tournier M, Hercend T, Triebel F, Faure F. Lymphocyte-activation gene 3/major histocompatibility complex class II interaction modulates the antigenic response of CD4+ T lymphocytes. Eur J Immunol. 1994;24(12):3216–21. https://doi.org/10.1002/eji.1830241246.
Bruniquel D, Borie N, Hannier S, Triebel F. Regulation of expression of the human lymphocyte activation gene-3 (LAG-3) molecule, a ligand for MHC class II. Immunogenetics. 1998;48(2):116–24. https://doi.org/10.1007/s002510050411.
Triebel F. LAG-3: a regulator of T-cell and DC responses and its use in therapeutic vaccination. Trends Immunol. 2003;24(12):619–22. https://doi.org/10.1016/j.it.2003.10.001.
Workman CJ, Wang Y, El Kasmi KC, Pardoll DM, Murray PJ, Drake CG, et al. LAG-3 regulates plasmacytoid dendritic cell homeostasis. J Immunol. 2009;182(4):1885–91. https://doi.org/10.4049/jimmunol.0800185.
Workman CJ, Cauley LS, Kim IJ, Blackman MA, Woodland DL, Vignali DAA. Lymphocyte activation Gene-3 (CD223) regulates the size of the expanding T cell population following antigen activation in vivo. J Immunol. 2004;172(9):5450–5. https://doi.org/10.4049/jimmunol.172.9.5450.
Woo SR, Li N, Bruno TC, Forbes K, Brown S, Workman C, et al. Differential subcellular localization of the regulatory T-cell protein LAG-3 and the coreceptor CD4. Eur J Immunol. 2010;40(6):1768–77. https://doi.org/10.1002/eji.200939874.
Li N, Wang Y, Forbes K, Vignali KM, Heale BS, Saftig P, et al. Metalloproteases regulate T-cell proliferation and effector function via LAG-3. EMBO J. 2007;26(2):494–504. https://doi.org/10.1038/sj.emboj.7601520.
Subramanyam MWG, Nabioullin R, Tepper MA. Soluble human lymphocyte activation gene-3 modulates allospecific T cell responses. Int Immunol. 1998;10(5):679–89.
Zhu Z, Ye J, Ma Y, Hua P, Huang Y, Fu X, et al. Function of T regulatory type 1 cells is down-regulated and is associated with the clinical presentation of coronary artery disease. Hum Immunol. 2018;79(7):564–70. https://doi.org/10.1016/j.humimm.2018.05.001.
Ng TH, Britton GJ, Hill EV, Verhagen J, Burton BR, Wraith DC. Regulation of adaptive immunity; the role of interleukin-10. Front Immunol. 2013;4:129. https://doi.org/10.3389/fimmu.2013.00129.
Battaglia M, Gregori S, Bacchetta R, Roncarolo MG. Tr1 cells: from discovery to their clinical application. Semin Immunol. 2006;18(2):120–7. https://doi.org/10.1016/j.smim.2006.01.007.
Gagliani N, Magnani CF, Huber S, Gianolini ME, Pala M, Licona-Limon P, et al. Coexpression of CD49b and LAG-3 identifies human and mouse T regulatory type 1 cells. Nat Med. 2013;19(6):739–46. https://doi.org/10.1038/nm.3179.
Asseman CMS, Leach MW, Coffman RL, Powrie F. An essential role for interleukin 10 in the function of regulatory T cells that inhibit intestinal inflammation. J Exp Med. 1999;190:995–1003.
Funding
This work was supported by a NIH grant, NHLBI HL131862, and the Linda and David Roth Chair of Cardiovascular Research endowment to Dr. Rodriguez.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Dr. Rodriguez has inventorship rights to issued patents related to SCARB1 and LAG3, and she is the founder of Lipid Genomics.
Human and Animal Rights and Informed Consent
This article does contain studies with human subjects. All the studies were approved by respective institutional research review boards.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Vascular Biology
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rodriguez, A. High HDL-Cholesterol Paradox: SCARB1-LAG3-HDL Axis. Curr Atheroscler Rep 23, 5 (2021). https://doi.org/10.1007/s11883-020-00902-3
Accepted:
Published:
DOI: https://doi.org/10.1007/s11883-020-00902-3