Abstract
Papillary thyroid carcinoma (PTC) is considered an indolent neoplasm but it may demonstrate aggressive behavior. We aimed to identify clinical and pathological characteristics and molecular signatures associated with aggressive forms of PTCs. We selected 43 aggressive PTC cases based on the presence of metastases at the time of diagnosis, the development of distant metastasis during follow-up, and/or biochemical recurrence, and 43 PTC patients that were disease-free upon follow-up, matching them according to age, sex, pT, and pN parameters. Twenty-four pairs (a total of 48 cases) and 6 normal thyroid tissues were studied using targeted mRNA screening of cancer-associated genes employing NanoString nCounter® technology. In general, aggressive PTCs showed distinctive clinical and morphological features. Among adverse prognostic parameters, the presence of necrosis and an increased mitotic index were associated with shorter disease-free and overall survivals. Other parameters associated with shorter disease-free or overall survivals include a lack of tumor capsule, the presence of vascular invasion, tumor-infiltrating lymphocytes, fibrosclerotic changes, age > 55 years, and a high pTN stage. Various pathways were differentially regulated in non-aggressive as compared to aggressive PTC, including the DNA damage repair, the MAPK, and the RAS pathways. In particular, the hedgehog pathway was differentially de-regulated in aggressive PTC as compared to non-aggressive PTC cases, being WNT10A and GLI3 genes significantly up- and down-regulated in aggressive PTC and GSK3B up-regulated in non-aggressive PTC cases. In conclusion, our study revealed specific molecular signatures and morphological features in aggressive PTC that may be useful to predict more aggressive behavior in a subset of PTC patients. These findings may be useful when developing novel, tailored treatment options for these patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Papillary thyroid carcinoma (PTC) derives from follicular cells and is the most common thyroid malignancy, comprising more than 80% of all malignant thyroid tumors [1,2,3]. PTC is generally considered a tumor with indolent behavior, slow progression, and low mortality rates [4]. However, a subset of cases may present an aggressive clinical course. Clinical and morphological features and gene alterations allow us to predict the aggressive behavior of this tumor, but only up to a certain extent [5]. Compared to classical PTCs, the hobnail, tall cell, columnar, and solid subtypes predominate in aggressive cases [6, 7]. Since several years, it has been ascertained that prognosis is worsened by higher disease stages, tumor size above 3–4 cm, presence of distant metastases, extrathyroidal extension, neoplastic infiltration of the resection margins, and/or older age at diagnosis [8, 9].
In terms of the impact of the molecular signature on prognosis, various studies have attempted to better characterize the genetic make-up of these neoplasms. The BRAF p.V600E mutation is the most common genomic alteration in PTC (found in approximately 70% of cases). It is currently considered that the BRAF mutation by itself is not a high-risk factor, but it has a strong adverse prognostic impact in association with TERT promoter mutations [10, 11]. Mutations of NRAS, KRAS, and HRAS are also common [12], especially in the follicular variant [13], but lack a significant and independent prognostic role. Other molecular alterations accumulate in PTC during cancer progression, such as those in TP53, PIK3CA, and AKT1 [14, 15]. In this context, TERT promoter mutations (C228T and C250T) are present in more advanced PTCs and are strongly associated with a higher risk of distant metastases, recurrence, and cancer-related mortality [10, 16].
Apart from genomic alterations, some studies investigated gene or microRNA expression profiling in PTC. However, only a few studies were designed with the specific purpose of characterizing PTCs with aggressive clinical features. The most recent data are obtained by analysis of publicly available datasets (i.e., TCGA) but are affected by the lack of a detailed characterization of cases and by an unbalanced selection of cases with or without aggressive features [17,18,19,20,21,22].
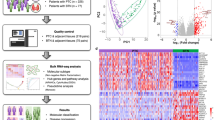
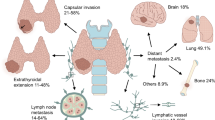
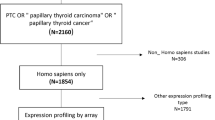
We therefore aimed to test the profiles of gene expression through the NanoString nCounter® technology in a series of well-characterized clinically aggressive PTC, matched with non-aggressive PTC cases, to identify molecular biomarkers potentially predictive of an aggressive clinical course that might be integrated with the current pathological characterization. A summary of the study design and main results is illustrated in Fig. 1.
Material and Methods
Case Selection
A group of 43 cases with aggressive clinical behavior, defined based on the presence of metastasis at diagnosis, development of distant metastases during follow-up, and/or biochemical recurrence (> 2 ng/ml levels of thyroglobulin after TSH stimulation), was selected from a large series of PTC undergoing thyroidectomy from 2002 to 2017 at “Città della Salute e della Scienza” Hospital (Turin, Italy), San Luigi Gonzaga Hospital (Orbassano, Turin, Italy), and Mauriziano Hospital (Turin, Italy). Six cases had a biochemical recurrence, only. Thirty-seven cases had metachronous metastases. These latter cases included one case with synchronous lung metastases, only, four cases with metachronous multiorgan dissemination (including lymph nodes lung, brain, bone), and 32 cases with metachronous lymph node involvement, with or without infiltration of soft tissues in the cervical neck. These cases were matched for age, sex, and T and N parameters with 43 controls, undergoing surgery in the same period, which resulted in free of disease during the clinical follow-up (median follow-up 12.1 years, range 4.4–19 years). Characteristics of lymph node metastases in both aggressive and non-aggressive cases are reported in Supplementary Table 1.
Representative hematoxylin and eosin (H&E) stained slides were re-evaluated in all cases enrolled in the study by three of us (FM, MV, MP). The following parameters were re-assessed and recorded: histological subtype, TNM stage according to AJCC 8th edition, tumor diameter, uni- or multifocal presentation (including bilaterality), presence of tumor capsule, presence and extent of microscopic extrathyroidal invasion, presence of angioinvasion, status of surgical margins, presence of necrosis, presence of tumor-infiltrating lymphocytes (TILs), and presence of fibro-sclerotic changes. TILs were coded as absent, present “non-brisk,” and present “brisk” according to standard evaluation methods applied in breast cancer [23]. For statistical purposes, cases were grouped as TILs absent or present, in this latter case including both non-brisk and brisk patterns. Fibro-sclerotic changes were coded as mild (in the presence of thin and rare bands of collagen deposition) or extensive (in the presence of more abundant and thick fibrous bands). Representative images of the different patterns of distribution of TILs and fibro-sclerotic changes are illustrated in Fig. 2. Mitotic count was determined as the number of mitotic figures in 2 mm2. Five cases (four in the aggressive PTC group and one in the non-aggressive PTC group) met the criteria for high-grade differentiated thyroid carcinomas because of the presence of coagulative necrosis. One case had also a mitotic index of 5 in 2 mm2. However, due to their limited number, these cases were not further considered as a separate group but necrosis and mitotic index were considered individually for the sake of statistical analyses.
Representative images of the distribution of tumor-infiltrating lymphocytes and fibro-sclerotic changes in papillary thyroid carcinoma cases. Top left: tumor-infiltrating lymphocytes and fibro-sclerotic changes absent; top right: tumor-infiltrating lymphocytes present, brisk, and fibro-sclerotic changes absent; bottom left: tumor-infiltrating lymphocytes present, brisk, and fibro-sclerotic changes present mild; bottom right: tumor-infiltrating lymphocytes present, non-brisk, and fibro-sclerotic changes present extensively
A pool of thyroid tissues devoid of macroscopically recognizable alterations from patients operated on due to follicular nodular disease was used as the baseline for gene expression testing and consisted of 3 female (aged 33, 50, and 71) and 3 male patients (aged 40, 50, and 69).
Before the study started, all cases were de-identified and coded by a pathology staff member not involved in the study, and all data were accessed anonymously. The study was approved by the local Ethical Committee (#610, date December 20th, 2017) and conducted in accordance with the principles set out in the Declaration of Helsinki. Considering the retrospective nature of this research protocol and that it had no impact on patients’ care, no specific written informed consent was required.
Molecular Analyses
For molecular testing, 24 pairs of aggressive and their non-aggressive match were analyzed by means of gene expression profiling (for a total of 48 cases tested) along with the pool of 6 samples of non-neoplastic thyroid tissue. In this sub-series, the aggressive cases were also matched with non-aggressive cases for the histological subtype.
For RNA extraction, two 10-μm-thick formalin-fixed and paraffin-embedded (FFPE) tissue sections were obtained from each tissue block and collected in sterile Eppendorf tubes by means of manual microdissection. The same procedure was adopted for normal thyroid tissues, making sure that only healthy/unaltered parenchymal areas were dissected. RNA isolation was performed using the FFPE RNA Isolation Kit (Roche Diagnostics GmbH, Mannheim, Germany), according to the manufacturer’s protocols. Total RNA concentration was assessed using a NanoDrop spectrophotometer (Thermo Fisher Scientific, Inc., Wilmington, DE, USA).
NanoString nCounter® technology was used to detail the transcriptomic cancer-associated landscape in PTC. Three hundred nanograms of total RNA from each sample were hybridized to the nCounter PanCancer Pathways Panel, according to the manufacturer’s instructions (NanoString Technologies, Seattle, WA, USA). The nCounter PanCancer Pathways panel includes 730 genes from 13 canonical pathways and 40 selected reference genes. The analyses were set up according to the protocol provided by the manufacturer. Expression data were normalized and analyzed with the nSolver Analysis Software (version 4.0.62). For background correction, the mean count of negative controls plus two times the standard deviation was subtracted from the counts of each gene. The means of the supplied positive controls and the geometric mean of the housekeeping genes were used to normalize the measured expression values. Both positive and negative controls were included in the panel, according to the manufacturer’s instructions. Additionally, the advanced analysis module (version 2.0.134) was used to perform differential expression analyses. Briefly, by the differential expression analysis tool (nSolver Advanced Analysis module), for each gene, a single linear regression was fit using all selected covariates to predict expression. Volcano plots were generated to display each gene’s -log10 (p-value) and log2 fold change with the selected covariate. Highly statistically significant genes fell at the top of the plot above the horizontal lines, and highly differentially expressed genes fell to either side. Horizontal lines indicated various p-value thresholds.
Statistical Analyses
All analyses were performed using Stata/MP 15.0 Statistical Software (STATA, College Station, TX). The differences in the distribution of clinical-pathological variables were analyzed using parametric and non-parametric tests (student’s t test, Pearson’s chi-square test, Bonferroni’s correction, Wilcoxon’s rank test). Overall survival (OS) was calculated from the surgical excision date of the primary tumor to the date of death or last check-up. The disease-free survival (DFS) was calculated from the date of surgical excision of the primary tumor to the date of the first relapse or last check-up. Survival curves were plotted using the Kaplan-Meier method, and the statistical comparisons were performed using the log-rank test. The Cox regression analyses were carried out on DFS and OS to calculate HRs and 95% CIs for the different study groups. Cox proportional hazards regression model was used in multivariate survival analysis to assess the independent role of parameters significant at univariate analysis. All statistical tests were two sided. A p-value < 0.05 was considered significant.
Results
Clinical and Pathological Characteristics
As shown in Table 1, aggressive PTC was more frequently multifocal (26/43, 60.5%) as compared to non-aggressive cases (16/43, 37.2%) (p = 0.031). The tumor capsule was absent in more than half of aggressive cases (29/43, 67.4%) and in only 17/43 (39.5%) of non-aggressive ones (p = 0.015). Furthermore, aggressive PTCs had more frequent TILs (26/43, 60.5%) as compared to the non-aggressive group (11/43, 25.6%) (p = 0.001). Only 10/43 (23.3%) non-aggressive PTCs had extensive fibro-sclerotic changes, as compared to 21/43 (48.8%) aggressive cases (p = 0.047). Although not statistically significant, angioinvasion was observed more commonly in aggressive (30/43, 69.8%) as compared to non-aggressive cases (21/43, 48.8%) (p = 0.077). Of note, at follow-up, 11/43 (25.6%) patients with aggressive PTC died (8 of their PTC and 3 due to other causes), as compared to 4/43 (9.3%) patients with non-aggressive PTC who died of other causes, all not related to their PTC (p = 0.047).
By means of Cox univariate analyses (Table 2), in terms of DFS, the presence of angioinvasion (p = 0.019), TILs (p = 0.001), necrosis (p = 0.050), and higher mitotic counts (p = 0.001) were all correlated with an adverse prognosis. Extensive fibro-sclerotic changes were also associated with shorter DFS (p = 0.038 and p = 0.021 in the log rank test, Fig. 3). On the other hand, the presence of a complete tumor capsule was a favorable morphologic feature (p = 0.029). In terms of OS, tumor diameter larger than 4 cm (p = 0.004), presence of necrosis (p < 0.001), and higher mitotic index (p = 0.001) were parameters correlated with adverse prognosis in terms of OS. However, none of the parameters significant in univariate analyses showed independent statistical significance in multivariate analyses.
OS in patients with aggressive PTC was significantly shorter than in patients with non-aggressive PTC (p = 0.038) (Fig. 4).
Gene Expression Profile of Aggressive and Non-Aggressive PTC Cases
The clinical and pathological characteristics of the 48 patients analyzed for gene expression profiling are summarized in Supplementary Table 2.
In aggressive PTC, 38 genes were significantly upregulated and 3 were significantly downregulated as compared to the non-neoplastic tissue (Supplementary Table 3; volcano plot image in Fig. 5a). In non-aggressive PTC, 80 genes were significantly upregulated and 9 were significantly downregulated as compared to non-neoplastic thyroid tissue (Supplementary Table 4; volcano plot image in Fig. 5b). Three genes, only, were significantly de-regulated in aggressive but not in non-aggressive PTC, namely two upregulated (PKMYT1 and WNT10A) and one downregulated (GLI3) (Table 3). By contrast, 45 genes were significantly upregulated and seven were downregulated in non-aggressive, but not in aggressive PTC. Thirty-six upregulated and two downregulated genes were in common between aggressive and non-aggressive PTC cases. However, even in this subset, four genes (DUSP4, INHBB, JAG2, and MLF1) were significantly upregulated in the group of non-aggressive cases as compared to the aggressive ones (Fig. 6).
The differential gene expression between aggressive and non-aggressive cases according to different pathways is presented in Supplementary Fig. 1. None of the genes related to DNA damage repair (POLD4, H2AFX, POLR2J, ATM, MAD2L2, PRKDC) that were upregulated in non-aggressive PTC cases were found differentially expressed in aggressive PTC (Supplementary Fig. 1a1/2).
Two genes related to the NOTCH pathway, DTX2 and JAG2, were upregulated (Supplementary Fig. 1b1/2) in both groups. All genes de-regulated in the aggressive PTC group and belonging to the transcriptional misregulation pathway were also de-regulated in the non-aggressive PTC group (Supplementary Fig. 1c1/2).
Deregulation of the hedgehog pathway involved different genes in aggressive and non-aggressive PTC. In fact, WNT10A and GLI3 were found up- and downregulated in aggressive PTCs, whereas GSK3B was found upregulated in non-aggressive PTCs (Supplementary Fig. 1d1/2).
The analyses of the WNT pathway revealed overlap in only one gene between the two groups (PPP3CB), whereas several other genes were found to be de-regulated in non-aggressive cases (CCND3, GSK3B, SFRP4, SMAD3, AXIN1, RAC2, JUN) (Supplementary Fig. 1e1/2).
Regarding the JAK-STAT pathway, upregulation of EPOR, BCL2L1, and CBL genes was found upregulated in both groups, while PIK3R2, CCND3, SOS2, OSM, JAK3, PIM1, and IL2RB genes were up- and TPO down-regulated in the non-aggressive group, only (Supplementary Fig. 1f1/2).
Considering the RAS pathway, PDGFC, KRAS, RAC2, SOS2, and PIK3R2 genes were upregulated in non-aggressive cases, only, whereas RASA4, MET, PDGFA, RIN1, TIAM1, LAT, and BCL2L1 were upregulated in both groups (Supplementary Fig. 1g1/2).
Regarding the genes labeled as “driver genes,” there was an overlap for ATRX, TRAF7, STK11, MET, SMAD2, CBL, and RUNX1, with deregulation of several other genes in non-aggressive PTC, only (Supplementary Fig. 1h1/2). A similar picture was observed for the sub-analysis of the MAPK pathway, with upregulation of TGFB1, PPP3CB, CACNB3, DUSP4, HSB1, and PDGFA in both groups, but upregulation of KRAS, MAP3K1, SOS2, DUSP5, RAC2, and CDC25B and downregulation of CACNA2D1 and JUN in the non-aggressive PTC group, only (Supplementary Fig. 1k1/2).
Finally, the PI3K pathway was significantly de-regulated in both groups, with 11 genes upregulated in both groups and several others de-regulated in non-aggressive cases (Supplementary Fig. 1l1/2).
Discussion
In the present study, we reported that clinically aggressive PTC is associated with peculiar pathological features and gene expression profiles.
PTCs in general show indolent behavior with localized disease and typically do not recur or metastasize beyond locoregional lymph nodes. A recent meta-analysis, using univariate comparisons, has identified male sex, advanced age, tumor size, multifocality, vascular invasion, extra-thyroidal extension, and lymph node metastasis as risk factors for distant metastasis in well-differentiated thyroid cancers [24]. In our series, we could not confirm sex, age, and TNM stage as parameters of aggressiveness in the aggressive vs non-aggressive PTC group comparison, since these parameters were balanced in the case match procedure. Moreover, the relatively small sample size for each group—designed to be balanced for molecular analyses—and the not complete homogeneity of compositions of the two groups (i.e., RAI treatment) limit the statistical impact of clinical pathological correlates, as demonstrated by the low level of statistical significance in DFS and OS of non-aggressive and aggressive PTC cases. We could confirm multifocality and angioinvasion as parameters significantly more prevalent in aggressive PTC, the latter also showing a significant association with shorter disease-free survival.
Concerning additional pathological parameters, we found that the presence of TILs was more frequent in aggressive PTCs as compared to non-aggressive cases. This finding is in contrast with previous data on the association of immune cell infiltrate (including B and T cells) and a more favorable outcome [25,26,27]. As a limitation of our study, we did not perform subtyping of the tumor-associated immune compartment, but it is conceivable that our case series selection approach influenced our results as compared to literature studies. Moreover, our data are strengthened by the significant association of the presence of TILs with shorter disease-free survival (with a trend to significance also in terms of shorter overall survival).
Another feature associated with an aggressive clinical course in PTC was the presence of fibro-sclerotic tissue within the tumor. Our findings are supported by a few previous studies [28, 29] but are in contrast with a recent report on a large unselected series [30]. However, the value of our results is supported by a disease-free survival analysis that showed a significant adverse impact of the presence of extensive fibrotic changes. These fibrous tissue modifications were appearing as trabecular, net-like structures permeating tumor architecture. In a setting of parathyroid tumors, the presence of thick fibrous bands is commonly referred to as a sign of malignancy [31, 32].
Finally, increased mitotic activity and the presence of necrosis were identified as strong adverse prognostic indicators, both in disease-free and overall survival analyses, although their distribution was not statistically different comparing aggressive and non-aggressive PTC cases. This latter finding is strongly supportive of the concept of the high-grade differentiated thyroid cancer group, as a specific prognostic category within differentiated thyroid cancers and is in line with the novel category proposed by the fifth edition of the WHO book on endocrine tumors, which embeds aggressive differentiated carcinomas and poorly differentiated carcinomas in a single group named high-grade thyroid carcinomas [33, 34].
A second part of our study was focused on the definition of gene expression profiles in a subset of aggressive and non-aggressive PTC cases, selected from the pathological series and matched also for the histological subtype. Despite other studies employed NanoString nCounter technology to explore the molecular make-up of thyroid malignances [35,36,37], to the best of our knowledge, this is the first study using the nCounter® PanCancer Pathways Panel to explore different canonical pathways in correlation with PTC aggressiveness. Our study falls in the scarce literature aimed at testing gene expression profiles as determinants of aggressiveness in PTC. The weakness of our approach is mostly based on the targeted approach (although covering over 700 genes), the lack of genomic data of the cases investigated, and the lack of RNA validation using an alternative approach or protein analysis by means of immunohistochemistry. By contrast, the strengths of our study are the robustness of the nanostring technology and the study design in terms of case selection. In fact, as compared to similar studies, we designed a comparative study on aggressive and non-aggressive PTC cases based on a strongly balanced and matched selection. A recent study based on TCGA data identified a specific signature of immune-related genes (namely HSPA1A, NOX5, and FGF23) associated with prognosis in PTC, but using data from a series of nearly 500 PTC cases with 3.2% of patients, only, died of disease and less than 2% of cases with distant metastases [38]. Another very recent study, also based on TCGA data, compared gene expression profiles in 455 non recurrent vs 46 recurrent PTC tumors specimens, identifying differential expression profiles in 40 genes with a significant influence of the tumor genotype [19]. In this study, recurrent PTC cases with BRAF-like signature significantly overexpressed—among other genes—FN1, ITGA3, and MET and downregulated TPO and TG.
Our study clearly shows that clinically aggressive and non-aggressive PTC cases in our series consistently share a gene expression signature as compared to normal thyroid tissue. In terms of pathway analysis, up-regulation of the RAS, MAPK, and PI3K pathways was consistent in both groups, although with an enrichment of de-regulated genes in the non-aggressive PTC group. However, a larger number of genes was deregulated in non-aggressive PTC as compared to normal tissues than those deregulated in aggressive PTC cases. The larger set of de-regulated genes depicted in non-aggressive PTC cases shows a more significant upregulation of genes belonging to the DNA damage repair and JAK/STAT pathways and a downregulation of genes belonging to the WNT pathway. Interestingly, the hedgehog pathway was differentially de-regulated in aggressive PTC as compared to non-aggressive PTC cases. In fact, whereas WNT10A and GLI3 were significantly up- and down-regulated in aggressive PTC, GSK3B was upregulated in non-aggressive PTC cases.
These findings suggest that these genes are likely involved in molecular mechanisms of aggressiveness. In fact, WNT10A/beta-catenin pathway activation has been demonstrated to promote cellular characteristics of aggressiveness, such as proliferation and migration in thyroid cancer [39]. Data on GLI3-driven molecular mechanisms in thyroid cancer are not available. GLI3 activation/overexpression has been shown to promote tumor progression in colon cancer [40] but not in other models such as gallbladder or cervical cancer [41, 42] thus supporting tumor type-specific activities. On the other hand, GSK3B expression has been described to inhibit the progression of several cancer models, such as, among others, lung [43], gastric [44], and hepatocellular [45] cancers. Finally, PKMYT1 was significantly upregulated in aggressive PTC cases, only. Although no data are available on the role of this gene in thyroid cancer, PKMYT1 has been shown to exert an important effect on tumor immunity and progression in several cancer models [46].
In conclusion, we demonstrate that clinically aggressive PTC is characterized by peculiar pathological features, including an increased TILs infiltration and a more prevalent occurrence of intratumoral fibrosis, supporting the value of a careful reporting of these parameters in the PTC diagnostic work-up. Moreover, although non-aggressive PTC in our series showed a larger number of deregulated genes as compared to aggressive PTC cases, the existence of a set of differentially regulated genes in the two groups suggests that different molecular mechanisms are active in promoting clinical aggressiveness in PTC with special reference to the impairment of the hedgehog pathway. Although limited by the relative small sample size and biases in case selection, these results have a prime impact highlighting the need to evaluate the stromal component as part of the diagnostic workup for PTC, with special reference to the presence of fibrosclerotic changes, a pathological characteristic that is currently not considered in standardized pathology reporting for thyroid cancer [47]. Moreover, gene expression data pave the way for molecular validation studies targeting regulators of the hedgehog pathway to identify novel biomarkers of aggressiveness.
Availability of Data and Material
Not applicable.
Code Availability
Not applicable.
References
Haugen BR, Alexander EK, Bible KC, Doherty GM, Mandel SJ, Nikiforov YE, Pacini F, Randolph GW, Sawka AM, Schlumberger M, Schuff KG, Sherman SI, Sosa JA, Steward DL, Tuttle RM, Wartofsky L (2016) 2015 American Thyroid Association Management Guidelines for Adult Patients with Thyroid Nodules and Differentiated Thyroid Cancer: The American Thyroid Association Guidelines Task Force on Thyroid Nodules and Differentiated Thyroid Cancer. Thyroid 26:1–133. https://doi.org/10.1089/thy.2015.0020
Genere N, El Kawkgi OM, Giblon RE, Vaccarella S, Morris JC, Hay ID, Brito JP (2021) Incidence of Clinically Relevant Thyroid Cancers Remains Stable for Almost a Century: A Population-Based Study. Mayo Clin Proc 96:2823–2830. https://doi.org/10.1016/j.mayocp.2021.04.028
Li M, Delafosse P, Meheus F, Borson-Chazot F, Lifante JC, Simon R, Groclaude P, Combes JD, Dal Maso L, Polazzi S, Duclos A, Colonna M, Vaccarella S; Thyroid Cancer Group FRANCIM (2021) Temporal and geographical variations of thyroid cancer incidence and mortality in France during 1986–2015: The impact of overdiagnosis. Cancer Epidemiol 75:102051. https://doi.org/10.1016/j.canep.2021.102051
Roman BR, Morris LG, Davies L (2017) The thyroid cancer epidemic, 2017 perspective. Curr Opin Endocrinol Diabetes Obes 24:332–336. https://doi.org/10.1097/MED.0000000000000359
Papp S, Asa SL (2015) When thyroid carcinoma goes bad: a morphological and molecular analysis. Head Neck Pathol 9:16–23. https://doi.org/10.1007/s12105-015-0619-z
Nath MC, Erickson LA (2018) Aggressive Variants of Papillary Thyroid Carcinoma: Hobnail, Tall Cell, Columnar, and Solid. Adv Anat Pathol 25:172–179. https://doi.org/10.1097/PAP.0000000000000184
Baloch ZW, Asa SL, Barletta JA, Ghossein RA, Juhlin CC, Jung CK, LiVolsi VA, Papotti MG, Sobrinho-Simões M, Tallini G, Mete O (2022) Overview of the 2022 WHO Classification of Thyroid Neoplasms. Endocr Pathol 33:27–63. https://doi.org/10.1007/s12022-022-09707-3
Shaha AR, Shah JP, Loree TR (1996) Risk group stratification and prognostic factors in papillary carcinoma of thyroid. Ann Surg Oncol 3:534–538. https://doi.org/10.1007/BF02306085
Voutilainen PE, Multanen MM, Leppäniemi AK, Haglund CH, Haapiainen RK, Franssila KO (2001) Prognosis after lymph node recurrence in papillary thyroid carcinoma depends on age. Thyroid 11:953–957. https://doi.org/10.1089/105072501753211028
Xing M, Liu R, Liu X, Murugan AK, Zhu G, Zeiger MA, Pai S, Bishop J (2014) BRAF V600E and TERT promoter mutations cooperatively identify the most aggressive papillary thyroid cancer with highest recurrence. J Clin Oncol 32:2718–2726. https://doi.org/10.1200/JCO.2014.55.5094
Zhao L, Wang L, Jia X, Hu X, Pang P, Zhao S, Wang Y, Wang J, Zhang Y, Lyu Z (2020) The Coexistence of Genetic Mutations in Thyroid Carcinoma Predicts Histopathological Factors Associated With a Poor Prognosis: A Systematic Review and Network Meta-Analysis. Front Oncol 10:540238. https://doi.org/10.3389/fonc.2020.540238.
Cancer Genome Atlas Research Network (2014) Integrated genomic characterization of papillary thyroid carcinoma. Cell 159:676–690. https://doi.org/10.1016/j.cell.2014.09.050
Zhu Z, Gandhi M, Nikiforova MN, Fischer AH, Nikiforov YE (2003) Molecular profile and clinical-pathologic features of the follicular variant of papillary thyroid carcinoma. An unusually high prevalence of ras mutations. Am J Clin Pathol 120:71–77. https://doi.org/10.1309/ND8D-9LAJ-TRCT-G6QD
Ricarte-Filho JC, Ryder M, Chitale DA, Rivera M, Heguy A, Ladanyi M, Janakiraman M, Solit D, Knauf JA, Tuttle RM, Ghossein RA, Fagin JA (2009) Mutational profile of advanced primary and metastatic radioactive iodine-refractory thyroid cancers reveals distinct pathogenetic roles for BRAF, PIK3CA, and AKT1. Cancer Res 69:4885–4893. https://doi.org/10.1158/0008-5472.CAN-09-0727
Nikiforova MN, Wald AI, Roy S, Durso MB, Nikiforov YE (2013) Targeted next-generation sequencing panel (ThyroSeq) for detection of mutations in thyroid cancer. J Clin Endocrinol Metab 98:E1852–60. https://doi.org/10.1210/jc.2013-2292
Bullock M, Ren Y, O'Neill C, Gill A, Aniss A, Sywak M, Sidhu S, Delbridge L, Learoyd D, de Vathaire F, Robinson BG, Clifton-Bligh RJ (2016) TERT promoter mutations are a major indicator of recurrence and death due to papillary thyroid carcinomas. Clin Endocrinol (Oxf) 85:283–290. https://doi.org/10.1111/cen.12999
Chen Z, Wang Y, Li D, Le Y, Han Y, Jia L, Yan C, Tian Z, Song W, Li F, Zhao K, He X (2022) Single-Cell RNA Sequencing Revealed a 3-Gene Panel Predicted the Diagnosis and Prognosis of Thyroid Papillary Carcinoma and Associated With Tumor Immune Microenvironment. Front Oncol 12:862313. https://doi.org/10.3389/fonc.2022.862313
Luo Y, Chen R, Ning Z, Fu N, Xie M (2022) Identification of a Four-Gene Signature for Determining the Prognosis of Papillary Thyroid Carcinoma by Integrated Bioinformatics Analysis. Int J Gen Med 15:1147–1160. https://doi.org/10.2147/IJGM.S346058
Nieto HR, Thornton CEM, Brookes K, Nobre de Menezes A, Fletcher A, Alshahrani M, Kocbiyik M, Sharma N, Boelaert K, Cazier JB, Mehanna H, Smith VE, Read ML, McCabe CJ (2022) Recurrence of Papillary Thyroid Cancer: A Systematic Appraisal of Risk Factors. J Clin Endocrinol Metab 107:1392–1406. https://doi.org/10.1210/clinem/dgab836
Shi J, Wu P, Sheng L, Sun W, Zhang H (2021) Ferroptosis-related gene signature predicts the prognosis of papillary thyroid carcinoma. Cancer Cell Int 21:669. https://doi.org/10.1186/s12935-021-02389-7
Wang T, Tian K, Ji X, Song F (2021) A 6 transcription factors-associated nomogram predicts the recurrence-free survival of thyroid papillary carcinoma. Medicine (Baltimore) 100:e27308. https://doi.org/10.1097/MD.0000000000027308
Ma J, Han W, Lu K (2021) Comprehensive Pan-Cancer Analysis and the Regulatory Mechanism of ASF1B, a Gene Associated With Thyroid Cancer Prognosis in the Tumor Micro-Environment. Front Oncol 11:711756. https://doi.org/10.3389/fonc.2021.711756
Hendry S, Salgado R, Gevaert T, Russell PA, John T, Thapa B, Christie M, van de Vijver K, Estrada MV, Gonzalez-Ericsson PI, Sanders M, Solomon B, Solinas C, Van den Eynden GGGM, Allory Y, Preusser M, Hainfellner J, Pruneri G, Vingiani A, Demaria S, Symmans F, Nuciforo P, Comerma L, Thompson EA, Lakhani S, Kim SR, Schnitt S, Colpaert C, Sotiriou C, Scherer SJ, Ignatiadis M, Badve S, Pierce RH, Viale G, Sirtaine N, Penault-Llorca F, Sugie T, Fineberg S, Paik S, Srinivasan A, Richardson A, Wang Y, Chmielik E, Brock J, Johnson DB, Balko J, Wienert S, Bossuyt V, Michiels S, Ternes N, Burchardi N, Luen SJ, Savas P, Klauschen F, Watson PH, Nelson BH, Criscitiello C, O'Toole S, Larsimont D, de Wind R, Curigliano G, André F, Lacroix-Triki M, van de Vijver M, Rojo F, Floris G, Bedri S, Sparano J, Rimm D, Nielsen T, Kos Z, Hewitt S, Singh B, Farshid G, Loibl S, Allison KH, Tung N, Adams S, Willard-Gallo K, Horlings HM, Gandhi L, Moreira A, Hirsch F, Dieci MV, Urbanowicz M, Brcic I, Korski K, Gaire F, Koeppen H, Lo A, Giltnane J, Rebelatto MC, Steele KE, Zha J, Emancipator K, Juco JW, Denkert C, Reis-Filho J, Loi S, Fox SB (2017) Assessing Tumor-infiltrating Lymphocytes in Solid Tumors: A Practical Review for Pathologists and Proposal for a Standardized Method From the International Immunooncology Biomarkers Working Group: Part 1: Assessing the Host Immune Response, TILs in Invasive Breast Carcinoma and Ductal Carcinoma In Situ, Metastatic Tumor Deposits and Areas for Further Research. Adv Anat Pathol 24:235–251. https://doi.org/10.1097/PAP.0000000000000162
Vuong HG, Duong UNP, Pham TQ, Tran HM, Oishi N, Mochizuki K, Nakazawa T, Hassell L, Katoh R, Kondo T (2018) Clinicopathological Risk Factors for Distant Metastasis in Differentiated Thyroid Carcinoma: A Meta-analysis. World J Surg 42:1005–1017. https://doi.org/10.1007/s00268-017-4206-1
Yang Z, Yin L, Zeng Y, Li Y, Chen H, Yin S, Zhang F, Yang W (2021) Diagnostic and prognostic value of tumor-infiltrating B cells in lymph node metastases of papillary thyroid carcinoma. Virchows Arch 479:947–959. https://doi.org/10.1007/s00428-021-03137-y
Cunha LL, Morari EC, Guihen AC, Razolli D, Gerhard R, Nonogaki S, Soares FA, Vassallo J, Ward LS (2012) Infiltration of a mixture of immune cells may be related to good prognosis in patients with differentiated thyroid carcinoma. Clin Endocrinol (Oxf) 77:918–925. https://doi.org/10.1111/j.1365-2265.2012.04482.x
Villagelin DG, Santos RB, Romaldini JH (2011) Is diffuse and peritumoral lymphocyte infiltration in papillary thyroid cancer a marker of good prognosis? J Endocrinol Invest 34:e403–8. https://doi.org/10.3275/7870
Liu X, Zhang S, Gang Q, Shen S, Zhang J, Lun Y, Xu D, Duan Z, Xin S (2018) Interstitial fibrosis in papillary thyroid microcarcinoma and its association with biological behavior. Oncol Lett 15:4937–4943. https://doi.org/10.3892/ol.2018.7928
Takeda M, Mikami T, Numata Y, Okamoto M, Okayasu I (2013) Papillary thyroid carcinoma with heterotopic ossification is a special subtype with extensive progression. Am J Clin Pathol 139:587–598. https://doi.org/10.1309/AJCPQZQN50HKIAHA
Wang HQ, Li Y, Song X, Ma YQ, Li JL, Li YX, Wang GF, Liu P, Liu PL, Shen C, Shi HY (2022) Significance of interstitial fibrosis and p16 in papillary thyroid carcinoma. Endocr J. Jun 18. https://doi.org/10.1507/endocrj.EJ22-0010
Schantz A, Castleman B (1973) Parathyroid carcinoma. A study of 70 cases. Cancer 31:600–605. https://doi.org/10.1002/1097-0142(197303)31:3%3C600::AID-CNCR2820310316%3E3.0.CO;2-0
McClenaghan F, Qureshi YA (2015) Parathyroid cancer. Gland Surg 4:329–338. https://doi.org/10.3978/j.issn.2227-684X.2015.05.09
Xu B, David J, Dogan S, Landa I, Katabi N, Saliba M, Khimraj A, Sherman EJ, Tuttle RM, Tallini G, Ganly I, Fagin JA, Ghossein RA (2022) Primary high-grade non-anaplastic thyroid carcinoma: a retrospective study of 364 cases. Histopathology 80:322–337. https://doi.org/10.1111/his.14550
Wong KS, Dong F, Telatar M, Lorch JH, Alexander EK, Marqusee E, Cho NL, Nehs MA, Doherty GM, Afkhami M, Barletta JA (2021) Papillary Thyroid Carcinoma with High-Grade Features Versus Poorly Differentiated Thyroid Carcinoma: An Analysis of Clinicopathologic and Molecular Features and Outcome. Thyroid 31:933–940. https://doi.org/10.1089/thy.2020.0668
Giannini R, Moretti S, Ugolini C, Macerola E, Menicali E, Nucci N, Morelli S, Colella R, Mandarano M, Sidoni A, Panfili M, Basolo F, Puxeddu E (2019) Immune Profiling of Thyroid Carcinomas Suggests the Existence of Two Major Phenotypes: An ATC-Like and a PDTC-Like. J Clin Endocrinol Metab 104:3557–3575. https://doi.org/10.1210/jc.2018-01167
Chitikova Z, Pusztaszeri M, Makhlouf AM, Berczy M, Delucinge-Vivier C, Triponez F, Meyer P, Philippe J, Dibner C (2015) Identification of new biomarkers for human papillary thyroid carcinoma employing NanoString analysis. Oncotarget 6:10978–10993. https://doi.org/10.18632/oncotarget.3452
Armanious H, Adam B, Meunier D, Formenti K, Izevbaye I (2020) Digital gene expression analysis might aid in the diagnosis of thyroid cancer. Curr Oncol 27:e93-e99. https://doi.org/10.3747/co.27.5533
Gan X, Guo M, Chen Z, Li Y, Shen F, Feng J, Cai W, Xu B (2021) Development and validation of a three-immune-related gene signature prognostic risk model in papillary thyroid carcinoma. J Endocrinol Invest 44:2153–2163. https://doi.org/10.1007/s40618-021-01514-7
Dong T, Zhang Z, Zhou W, Zhou X, Geng C, Chang LK, Tian X, Liu S (2017) WNT10A/β-catenin pathway in tumorigenesis of papillary thyroid carcinoma. Oncol Rep 38:1287–1294. https://doi.org/10.3892/or.2017.5777
Shen M, Zhang Z, Wang P (2021) GLI3 Promotes Invasion and Predicts Poor Prognosis in Colorectal Cancer. Biomed Res Int 2021:8889986. https://doi.org/10.1155/2021/8889986
Ichimiya S, Onishi H, Nagao S, Koga S, Sakihama K, Nakayama K, Fujimura A, Oyama Y, Imaizumi A, Oda Y, Nakamura M (2021) GLI2 but not GLI1/GLI3 plays a central role in the induction of malignant phenotype of gallbladder cancer. Oncol Rep 45:997–1010. https://doi.org/10.3892/or.2021.7947
Zhu H, Xia L, Shen Q, Zhao M, Gu X, Bouamar H, Wang B, Sun LZ, Zhu X (2018) Differential effects of GLI2 and GLI3 in regulating cervical cancer malignancy in vitro and in vivo. Lab Invest 98:1384–1396. https://doi.org/10.1038/s41374-018-0089-5
Zhu MC, Zhang YH, Xiong P, Fan XW, Li GL, Zhu M (2022) Circ-GSK3B up-regulates GSK3B to suppress the progression of lung adenocarcinoma. Cancer Gene Ther. https://doi.org/10.1038/s41417-022-00489-8
Ma X, Chen H, Li L, Yang F, Wu C, Tao K (2021) CircGSK3B promotes RORA expression and suppresses gastric cancer progression through the prevention of EZH2 trans-inhibition. J Exp Clin Cancer Res 40:330. https://doi.org/10.1186/s13046-021-02136-w
Li Y, Zhou Y, Ma L, Liu D, Dai Z, Shen J (2020) miR-3677–3p promotes hepatocellular carcinoma progression via inhibiting GSK3β. Acta Biochim Biophys Sin (Shanghai) 52:1404–1412. https://doi.org/10.1093/abbs/gmaa125
Shao C, Wang Y, Pan M, Guo K, Molnar TF, Kocher F, Seeber A, Barr MP, Navarro A, Han J, Ma Z, Yan X (2021) The DNA damage repair-related gene PKMYT1 is a potential biomarker in various malignancies. Transl Lung Cancer Res 10:4600–4616. https://doi.org/10.21037/tlcr-21-973
Ghossein R, Barletta JA, Bullock M, Johnson SJ, Kakudo K, Lam AK, Moonim MT, Poller DN, Tallini G, Tuttle RM, Xu B, Gill AJ (2021) Data set for reporting carcinoma of the thyroid: recommendations from the International Collaboration on Cancer Reporting. Hum Pathol 110:62–72. https://doi.org/10.1016/j.humpath.2020.08.009
Funding
Open access funding provided by Università degli Studi di Torino within the CRUI-CARE Agreement. This study was partially supported by a grant from the Italian Association for Cancer Research (AIRC; IG 20100 year 2017 to MP). CM is supported by a grant from the Department of Medical Sciences, 19B/22/DSM.
Author information
Authors and Affiliations
Contributions
MP designed the study. JM and FC collected the clinico-pathological data and prepared the database. FM, MV, and MP revised the cases. JM and GO performed gene expression analysis using the NanoString nCounter® technology and elaborated the data. SOA performed the statistical analysis. MV, MP, and JM interpreted the obtained results. MP, MV, and JM wrote the first draft of the manuscript, and all authors critically revised it. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Ethics Approval
The study was approved by the local Ethical Committee (#610, date December 20th, 2017) and conducted in accordance with the principles set out in the Declaration of Helsinki. Considering the retrospective nature of this research protocol and that it had no impact on patients’ care, no specific written informed consent was required. All cases were de-identified, and all clinical-pathological data were accessed anonymously.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Metovic, J., Cabutti, F., Osella-Abate, S. et al. Clinical and Pathological Features and Gene Expression Profiles of Clinically Aggressive Papillary Thyroid Carcinomas. Endocr Pathol 34, 298–310 (2023). https://doi.org/10.1007/s12022-023-09769-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12022-023-09769-x