Abstract
Ovarian cancer (OV) is a malignant tumor that ranks first among gynecological cancers, thus posing a significant threat to women's health. Immunogenic cell death (ICD) can regulate cell death by activating the adaptive immune system. Here, we aimed to comprehensively characterize the features of ICD-associated genes in ovarian cancer, and to investigate their prognostic value and role in the response to immunotherapy. After analyzing datasets from The Cancer Genome Atlas, we utilized weighted gene coexpression network analysis to screen for hub genes strongly correlated with ICD genes in OV, which was subsequently validated with OV samples from the Gene Expression Omnibus (GEO) database. A prognostic risk model was then constructed after combining univariate, multivariate Cox regression and LASSO regression analysis to recognize nine ICD-associated molecules. Next, we stratified all OV patients into two subgroups according to the median value. The multivariate Cox regression analysis showed that the risk model could predict OV patient survival with good accuracy. The same results were also found in the validation set from GEO. We then compared the degree of immune cell infiltration in the tumor microenvironment between the two subgroups of OV patients, and revealed that the high-risk subtype had a higher degree of immune infiltration than the low-risk subtype. Additionally, in contrast to patients in the high-risk subgroup, those in the low-risk subgroup were more susceptible to chemotherapy. In conclusion, our research offers an independent and validated model concerning ICD-related molecules to estimate the prognosis, degree of immune infiltration, and chemotherapy susceptibility in patients with OV.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ovarian cancer, an aggressive disease among gynecological cancers, comprises approximately 90%-95% of primary tumors of the ovary, with the remaining 5%-10% arising as metastases from primary cancers elsewhere [1]. According to the latest cancer statistics for 2024, 19,680 new OV cases and 12,740 OV deaths are expected to occur in the United States [2]. However, the initial symptoms of ovarian cancer show no distinctive clinical features [3], and the screening role of medical institutions for patients with this type of cancer is limited. Most patients are diagnosed at advanced stages, and treatment options are limited [4]. Therefore, although the occurrence of ovarian cancer is less frequent than that of other gynecologic malignancies, including cervical cancer and endometrial cancer, the mortality rate of OV is the highest [1]. Thus, finding new biomarkers for early diagnosis as soon as possible will help improve the outcome of OV patients.
Different stimuli usually determine whether tumor cell death is immunogenic or nonimmunogenic. In recent years, due to the development of the definition of immunogenic cell death, more and more studies have explored its impact on cancer therapy [5, 6]. As a novel subtype of cell death, ICD induces cell death by altering the cell surface composition [7] and releasing soluble mediators [8] such as high mobility group protein (HMGB1). These two signals act on the dendritic cell-expressed receptors CD91 and TLR4. In addition, the ATP released by cellular autophagy can bind to the surface of dendritic cell (DCs) via P2RX7, thus stimulating DCs to present tumor antigens to T cells and triggering chemotherapy-resistant tumor cell death via an IFN-γ-mediated immune response [9, 10]. Recently, many researches highlighted the potential value of ICD in antitumor immune responses [11,12,13]. Previous studies identified a subset of ICD gene signatures that are closely related to ovarian cancer and validated their differential expression and prognostic role in OV [14, 15]. However, the expression of those ICD-associated factors in OV prognosis is still unknown. Therefore, the impact of ICD-related genes on ovarian cancer should be further investigated based on a large amount of transcriptomic data, which will contribute to identifying more valuable biomarkers and therapeutic targets for OV.
In our study, we first performed WGCNA using OV data from the TCGA and GEO databases to identify hub genes highly correlated with ICDs. Then, GO and KEGG analyses were conducted to determine enrichment pathways for these genes. Importantly, the integration of univariate and multivariate Cox regression and LASSO analyses helped us create an OV risk model containing nine ICD-related factors. Finally, immune infiltration, mutation, and chemosensitivity analyses were utilized to elucidate the potential underlying mechanisms involved. Taken together, these results allowed us to identify candidate genes for ovarian cancer.
Materials and Methods
Training Set
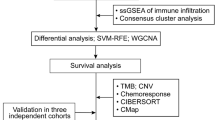
Figure 1 shows the procedures used in this study. Both clinical and transcriptic data for OV samples were downloaded from the TCGA-OV cohort. A set of 34 ICD genes used for analysis were obtained based on the previous study [14]. All samples were cancerous, with 352 cases in total. Clinical information on TCGA molecular typing and immunophenotyping data were downloaded based on the previous study (Supplementary Table 1) [16].
Validation Set
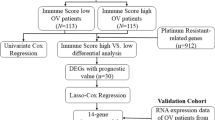
The risk model was then verified with GEO dataset GSE30161 from the GPL570 platform, which contains 58 cancer samples with transcriptome profiles and clinical data. Clinical information is shown in Supplementary Table 2. The criteria for data processing were as follows: (1) The pairs of null probes were removed. (2) A probe who corresponded to more than one gene was deleted. (3) The median expression level of one gene who corresponded to more than one probe was defined as the final expression value of the gene.
Identifying ICD-Related Genes in Ovarian Cancer
The enrichment scores of ICD gene sets calculated by ssGSEA were utilized to determine the levels of ICD genes in OV, followed by WGCNA of transcription profiles to determine the distribution of the soft threshold versus the mean connectivity [17]. To ensure the network adhere being scale-free, we applied 0.85 as the for the scale-free fit index. By using the mean-linked hierarchical technique, genes were clustered according to the standard that the smallest amount of genes per module was 30. Based on the ssGSEA enrichment scores of ICD genes and the top 75% of the variance in the ovarian cancer expression profile, we constructed the weighted gene coexpression network. Ultimately, employing the standard of gene correlation MM > 0.7 and trait significance GS > 0.3, the modules significantly related to the enrichment scores of ICD genes were identified as the core set of ICD-associated genes for subsequent studies.
Functional Enrichment Analysis
To discover functional pathways associated with these ICD-related factors, Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses were performed using the “clusterProfiler” R package, during which the pathway terms with p < 0.05 were considered significantly.
Establishment and Validation of the Risk Model for Ovarian Cancer
Univariate Cox analysis was performed to identify those prognostically relevant genes that met a p-value of less than 0.05. LASSO analysis was then used to determine these important components, and each hub gene's expression weight was taken into account while building a risk model. Next, based on whether their risk score was high or low, all OV patients were separated into two subtypes. We utilized the Kaplan–Meier method to compare the overall survival (OS) curves of the various groups in order to gain a deeper understanding of the predictive value for prognosis. Finally, the GEO dataset GSE30161 was selected as the validation set and scores were calculated using the same formula to verify the generalizability of the model.
Analysis of Molecular Features and Treatment Sensitivity in High- and Low-Risk Subgroups of Patients with Ovarian Cancer
Molecular Features and Pathway Alterations
With the data from the training dataset, the differences in single nucleotide variants (SNV) frequency were investigated through the R package “maftools”. Using Fisher’s test, mutation profiles were calculated and visually depicted using waterfall plots. Subsequently, we plotted the distribution of CNV using the R package “ggplot2” pair. Furthermore, we acquired the hallmark (v7.5.1) gene set and TCGA expression data as input data. Through the ssGSEA analyses within the R package “GSVA”, we estimated the scores of the samples in each pathway. Then, we plotted the distribution of different pathways within the two subgroups according to their enrichment scores based on the R package “pheatmap”.
Comparison of Immune Infiltration Between Two Subgroups of Patients with Ovarian Cancer
In fact, immune cells and stromal cells—two nontumor cells that reside in the tumor microenvironment—play indispensable roles in tumor diagnosis and prognostic prediction. Based on the scores of immune and stromal cells obtained through ESTIMATE analysis, the extent of immune infiltration was quantified in the tumor microenvironment of OV patients..
Chemoresistance Analysis Within the Two Subgroups of Ovarian Cancer Patients
Using the R package “oncoPredict”, we analyzed the correlations between log2(IC50) and the risk score for each chemotherapeutic agent across multiple cell lines from the GDSC2 database. This prediction relied on cell line expression data and information regarding chemotherapeutic agent responses in the GDSC2 database (http://www.cancerrxgene.org) and expression profiles in the training set.
Immunotherapy Response in the Two Subgroups of OV Patients
To test whether the ICD-related gene signature predicts immunotherapy prognosis, we selected data related to immunotherapy response. Subsequently, we performed statistical tests to compare the distributions of complete response (CR) or partial response (PR) and progressive disease (PD) or stable disease (SD) in different subgroups. In addition, statistical tests between subgroups with various levels of response were also performed.
Statistical Analysis
Differences between the two subgroups were verified by the Student’s t-test. Wilcoxon analysis was used to compare differences among multiple groups. To represent significance, we defined ns as p > 0.05, * as p < 0.05, ** as p < 0.01, *** as p < 0.001, and **** as p < 0.0001.
Results
The Landscape of ICD Genetic Alterations in Ovarian Cancer
The 34 ICD genes were differentially expressed according to age, stage, grade, lymphatic infiltration status, TCGA subtype, immune subtype and tumor diameter groups via rank sum test difference analysis and the Kruskal‒Wallis test (Fig. 2A–D). As shown in Fig. 2B, genes such as ENTPD1 were significantly upregulated in the Grade 1/2 subgroup, and PDIA3 was more highly expressed in the Grade 3/4 subgroup. As shown in Fig. 2D, genes such as FOXP3, CASP8, IFNG, CASP1, IFNB1, CD8B, MYD88, PRF1 and CXCR3 were present in the three subgroups C1, C2 and C3 with very significant differences in expression. In the lymphatic infiltration group, the expression of CD4, FOXP3, IL6, IL10, TLR4, CASP1, LY96, NLRP3, PRF1 and CXCR3 were increased in the YES subgroup, while the expression of ATG5 and HMGB1 were increased in the NO subgroup (Fig. 2E). As shown in Fig. 2F, different TCGA-OV subgroups exhibited varying levels of expression for CD4, IFNGR1, FOXP3, ATG5, BAX, P2RX7, IFNG, IL1B, NT5E, IL6, IL10, TLR4, CASP1, CD8A, LY96, CD8B, PRF1, CXCR3, and TNF. The expression of genes such as IL6, IFNB1 and TNF significantly differed among the subgroups with different sizes of residual tumor foci, as Fig. 2G illustrates. Furthermore, we analyzed overall mutations in ICD genes. As shown in Fig. 2H, the majority of the ICD genes in the training set had low mutation frequencies. However, NLRP3 mutations were notably more frequent than mutations in other genes, predominantly missense mutations. The distribution of all ICD genes on chromosome is shown in Fig. 2I. As shown in Fig. 2J, we also performed CNV analysis and found that CNVs were frequent, with amplification being the predominant alteration observed.
Clustering of ICD-Related Modules in Ovarian Cancer
Based on the the optimal soft threshold 3, the enrichment score of the ICD gene set in ovarian cancer was calculated by ssGSEA (Fig. 3A). Subsequently, cluster analysis was used to construct modules, and the closely related modules were merged into new modules according to clinical characteristics and the expression of ICD genes (Fig. 3B). Next, module eigengenes (MEs) were determined based on the first principal component of each module within the gene expression profile, followed by Pearson analysis between MEs in each module and clinical characteristics (Fig. 3C). Finally, we identified blue (cor = 0.915, p = 9.526e-141), red (cor = 0.509, p = 8.713e-25), purple (cor = − 0.341, p = 4.335e-11), black (cor = − 0.444, p = 1.579e-18) and greenyellow (cor = -0.337, p = 7.659e-11) modules with strong correlations with ICD gene expression levels (Fig. 3D), and screened 274 ICD-related genes under other criteria of MM > 0.7 and GS > 0.3.
Identification of the ICD-associated molecules in OV. A Network topology analysis under the various soft thresholds. B Gene tree and color of each module. C Correlation between modules and ICD score. D Point graph showing the identification of the ICD genes in ovarian cancer under MM > 0.7 and GS > 0.3. MM module membership, GS gene significant
Biological Function Analysis of ICD-Associated Genes in Ovarian Cancer
After then, 274 genes linked to ICDs underwent GO and KEGG functional enrichment analysis. According to GO analysis, these factors were more abundant in biological processes, such as T cell activation and positive regulation of cytokine production; cellular components, such as external side of plasma membrane and secretory granule membrane; and molecular functions, such as peptide binding and immune receptor activity (Supplementary Fig. 1). Additionally, the KEGG pathway enrichment of these genes was associated with phagosomes and tuberculosis. Together, these genes were associated with immunogenic cell death process.
Establishment of an ICD-Associated Model for OV
To determine which ICD-associated molecules are closely associated with the prognosis of patients with OV, univariate Cox regression analysis of the above 274 genes was first performed. Of the 14 identified prognostic factors, the top8 are displayed in Supplementary Fig. 2, of which CLEC5A was a factor related to a better prognosis in patients with low CLEC5A expression. These genes were subsequently included in further LASSO analysis. Finally, we identified nine primary ICD-related genes associated with survival based on the default penalty parameter (λ) (Fig. 4A, B), including CXCL11, BATF2, STAT1, CLEC5A, TAP1, VSIG4, TMIGD3, GIMAP7, and ANKRD22 (Fig. 4C). Moreover, we modeled the risk of these nine genes in patients with ovarian cancer based on the LASSO regression coefficients (Table 1). Afterward, using multivariate analysis, we assessed the risk of this model in all OV patients from TCGA and classified the patients into subgroups based on risk: high-risk and low-risk. According to the findings, individuals with a high-risk score might have a shorter overall survival time (Fig. 5A). We further evaluated the precision of this risk model for forecasting the various survival periods of OV patients by creating ROC curves. The area under the curve (AUC) at one year, three years and five years for predicting OS in patients with OV were 0.598, 0.630, and 0.669, respectively, as indicated in Fig. 5B. Moreover, we discovered that the high-risk subgroup had a higher likelihood of mortality (Fig. 5C–D). Again, the difference in gene expression between the two subgroups is proven in Fig. 5E.
Validation of the ICD-Associated Model Using the GEO-OV Dataset
With the prognostic information from the GEO-OV dataset GSE30161, we validated the findings in the OV cohort in TCGA. Similarly, we first separated patients into two risk groups, after which prognostic values and ROC curves were obtained. As shown by K–M curves in Supplementary Fig. 3A, low-risk patients had a higher probability of having improved overall survival. Additionally, based on ROC curves at various survival times, the AUCs for 1-, 3- and 5-year OS were shown to be 0.613, 0.586 and 0.702, respectively (Supplementary Fig. 3B). In the high-risk subtype, likewise, there were higher deaths (Supplementary Fig. 3C–D). Furthermore, we verified that the trend of gene expression was similar in the datasets of GEO and TCGA, except for the gene not detected by sequencing (Supplementary Fig. 3E). Collectively, these results validated the robustness of the nine ICD-associated genes prognostic signature constructed above.
Association Between the Prognostic Model and Clinical Features in OV Patients
First, through univariate and multivariate Cox regression analyses, this nine ICD-related genes signature was proven to be an independent prognostic factor (Fig. 6 and Supplementary Fig. 4). Furthermore, the distribution of risk scores was compared between different clinical characteristic subgroups (Fig. 7A–H). We found that the OI subgroup in the TCGA cohort had lower risk score compared to other subgroups. And there were significant differences between every two subgroups (Fig. 7D). In addition, we found that despite a difference in the risk score between the YES subgroup of patients with lymphatic invasion and the NO subgroup, there was no statistically significant difference (Fig. 7E). Moreover, each of the two tumor-residual subgroup (Fig. 7F) and immunological subgroups (Fig. 7H) showed notable differences in risk scoring.
Mutation and Pathway Enrichment in Two Subgroups of OV
Due to the potential role of inherited mutations in disease progression, we compared differences in mutation and pathway enrichment between OV patients with two different subtypes. After sorting by the mutation frequency of each gene, we obtained the top 20 genes displayed in Supplementary Fig. 5A. Among them, TP53 and TTN had higher mutation frequencies of 91% and 43%, respectively. And missense mutations accounted for the majority of TP53 mutations. Supplementary Fig. 5B further displays the top 10 molecules with the highest CNV frequency. The findings showed that patients in the high-risk group had higher degrees of CNV in all genes and that amplification was the most common form of variation. To further evaluate the pathway enrichment in the two subgroups, we calculate enrichment scores for every patient through ssGSEA (Supplementary Fig. 6A). Our results suggested significant differences in functional pathways between the two subgroups. For example, E2F targets occurred more frequently in the low-risk subtype than in the high-risk subtype, while TGF-β signaling, angiogenesis pathway and Wnt-β catenin signaling were more enriched in the high-risk subgroup. However, the two groups did not show significant differences in each clinical characteristics (Supplementary Fig. 6B).
Analysis of Tumor Immune Microenvironment in the Two Subgroups
As a method widely used to assess the status of immune infiltration, ESTIMATE analysis integrates the stromal score and immune score obtained by ssGSEA based on gene expression signatures to analyze tumor purity [18]. Our study revealed that the ultimate ESTIMATE value was slightly greater in the high-risk subgroup than in the low-risk subgroup, although no significant difference existed (Fig. 8A). Similarly, the ESTIMATE score was shown to be positively related to tumor purity (Fig. 8B). Additionally, we plotted 28 immune cell infiltration profiles in the two subgroups, and found that 5 types of immune cells, namely, plasmacytoid dendritic cells, memory B cells, central memory CD8+ T cells, type 2 T helper cells, and eosinophil, showed significant differences in infiltration between the two subgroups (Fig. 8C).
Analysis of Chemotherapy Resistance in the Two Subgroups of Patients with OV
We examined the connection between nine ICD-associated molecules and chemotherapy resistance in OV patients using the GDSC2 database. Spearman’s test was then utilized to uncover the association between the log2(IC50) of each chemotherapeutic agent and the ICD-related model. Figures 9A and B show the top 6 chemotherapeutic agents that were positively and negatively associated with the signature, respectively, based on the order of p-values from smallest to largest. It was clear that the sensitivity to each chemotherapeutic agent in the low-risk subgroup was greater than that in the high-risk subgroup (Fig. 9A).
Prediction of Immunotherapy Response in the Two Subgroups of Patients with Bladder Cancer
Since publicly published response data for ovarian cancer immunotherapy do not exist, we used bladder cancer immunotherapy response data to test the significance of the constructed gene signature for guiding immunotherapy. First, K–M curves showed that following immunotherapy treatments, individuals with OV in the high-risk subtype were more likely to live shorter lives than those in the low-risk subtype (p < 0.001) (Fig. 10A). In addition, patients who responded to immunotherapy were more likely to have the low-risk subtype (30.87%) than the high-risk subgroup (14.77%) (Fig. 10B). Moreover, the high-risk subgroup (55.22%) had a higher probability of patients who did not react to immunotherapy than the low-risk subgroup (44.78%). In immunotherapy-responsive patients, the proportion of patients who had high-risk score was lower (32.35%) than those had low-risk score (67.65%) (Fig. 10C).
Immunotherapy response in the two groups in bladder cancer samples A K–M analysis show the significant OS with a p < 0.001. B The distribution of immunotherapy response ways in the two groups in bladder cancer patients. C The distribution of the two groups in bladder cancer patients in NoR and R subtypes after the immunotherapy
Discussion
Ovarian cancer is a malignant disease in females and has a poor prognosis [19]. Meanwhile, many new treatment strategies nowadays, including targeted therapy [20], biological therapy [21], and immunotherapy [22] showed unsatisfactory effects. Immune checkpoint blockers [23] have long been highlighted as promising breakthroughs in cancer treatment. However, the efficacy of immunotherapy in tumors classified as immunologically “cold” types, including OV, is largely limited by various factors [24]. In recent years, immunogenic cell death has shown promise, bringing new hope to immunotherapy-resistant patients. Immunogenic cell death is a special form of regulated cell death. Universally, a variety of chemotherapeutic agents, such as anthracyclines and oxaliplatin, not only induce apoptosis but also prompt dying cells to express or release damage-associated molecular patterns (DAMPs), thereby stimulating the immune response to tumors and consequently transforming tumor cells from nonimmunogenic to immunogenic. This has far-reaching significance for antitumour therapy [25, 26]. Previous studies have shown that cancer vaccines targeting ICDs can render “cold” tumors immunogenic, thus increasing their susceptibility to immunotherapy [27]. Therefore, understanding the genetic signature associated with ICDs in OV will help us to determine the potential value of ICDs in OV treatment at a deep level.
Our study explored for the first time the comprehensive role of ICD-related gene signatures in ovarian cancer and demonstrated new ideas for prognostic prediction and individualized treatment of OV. First, we identified 14 ICD-associated genes that are valuable for OV prognosis. To clarify the specific roles these genes play in OV, we performed functional analysis and investigated their effects on immune-related pathways. Subsequently, KEGG pathway analysis revealed that molecules associated with ICDs were mainly enriched in the cytokine production signaling pathway and T-cell activation. Indeed, previous researches have indicated that BAK1, an ICD-associated molecule, can regulate multiple cellular phenotypes by inducing cytokine production [28]. Additionally, Minute et al. detected ICD markers when coculturing tumor cells with cytotoxic immune cells, which were able to trigger DAMPs and further activate the immunotoxic response [29]. As a result, ICDs and ICD-related factors play active roles in the activation of T cells, which leads to better treatment results for ovarian cancer patients.
At the core, by combining univariate, multivariate Cox and LASSO regression methods, we created a predictive model with nine ICD-related genes—CXCL11, BATF2, STAT1, CLEC5A, TAP1, VSIG4, TMIGD3, GIMAP7, and ANKRD22. CXCL11, a protein-coding gene located on chromosome 4, was demonstrated to regulate the circUBAP2/miR-4756/IFIT1/3 axis, thus modulating hepatocellular cell migration and tumor metastasis [30]. In addition, elevated expression of CXCL11 in colon cancer was proven to contribute to an unfavorable clinical outcome [31]. What’s more, CXCL11 was found to be related to antitumor immunity. In NSCLC, docetaxel-stimulated CXCL11 induced CD8+ T-cell infiltration by promoting HMGB1 release [32]. Moreover, BATF2 has been shown to participate in the progression and drug susceptibility of many tumors. For example, it was proposed that BATF2 blocks ERK signaling to suppress gastric cancer [33], and that overexpressing BATF2 increases drug sensitivity by inhibiting Wnt/β-catenin signaling in gastric cancer [34]. The protein-coding gene STAT1 plays an extensive role in cancer biology [35, 36]. A previous study revealed that the overexpression of STAT1 could downregulate PD-L1 expression and therefore facilitate tumor immunotherapy [37]. CLEC5A had been reported to trigger a variety of immune-inflammatory reactions by activating macrophages [38]. TAP1 was found to be involved in multidrug resistance, biological progression, and immune evasion [39,40,41]. VSIG4 had been shown to be related to poor OS and increased immunosuppressive cell infiltration in several types of cancer [42, 43]. TMIGD3 was reported to play roles in tumor proliferation, formation, and metastasis in osteosarcoma malignancy, esophageal cancer and osteosarcoma [44,45,46]. GIMAP7 had been identified as a candidate marker for tumor prognosis prediction and immunotherapy response [47]. ANKRD22 was indicated to be related to prognosis of various cancers [48,49,50]. In conclusion, these nine ICD-related genes can affect tumors through different mechanisms and more basical experiments are needed to uncover their role in the control of tumor immunotherapy of OV patients.
Recently, based on the impacts of ICDs on promoting the immune response, researchers have explored immunotherapy strategies targeting ICDs to complement conventional treatments for various tumors. Nevertheless, the potential impact of ICDs on ovarian cancer needs in-depth exploration. On the basis of the model involving nine ICD-associated genes, we classified OV patients in our study into matching high-risk and low-risk subtypes. We then evaluated the model’s capacity to predict survival time using datasets from the TCGA-OV and GEO-OV. The efficiency of this model to predict OS was demonstrated by the AUC for the 1-year, 3-year and 5-year survival times of OV patients. Next, we investigated the potential association between this model and clinical data. The results revealed a markedly elevated risk score in the YES subgroup of patients with lymphatic invasion compared with that in the NO subgroup. At the same time, CNV and SNV were more common in the high-risk subgroup. The immune microenvironment was also compared between the two subgroups. It was suggested that both stromal cells and immune cells infiltration were elevated in the low-risk subtype compared to the high-risk subtype. In addition, the log2(IC50) value of each chemotherapeutic agent was greater in the high-risk subgroup. Most importantly, it was indicated that a better prognosis was shown in the low-risk subgroup after immunotherapy treatments, and a significantly greater proportion of patients with OV in the low-risk subtype respond to treatment.
In conclusion, we comprehensively described the associations between ICD-associated factors and ovarian cancer. The nine ICD-related genes model we constructed here had high sensitivity and specificity for the prognostic prediction of OV. Moreover, these ICD-associated genes might also become candidate genes for basic research on OV, potentially developed new strategies for mechanistic research and immunotherapy. However, there were still some shortcomings in our study. This ICD-related prognostic model was only validated with data from public databases. Additional fundamental experiments are required to confirm the certain roles of these molecules and their regulatory mechanism.
Data Availability
All analysis in this article were performed according to transcriptomic profiles and clinical features obtained from public databases, mainly the TCGA and the GEO and databases. For further enquiries, please contact the corresponding author.
Abbreviations
- OV:
-
Ovarian cancer
- ICD:
-
Immunogenic cell death
- TCGA:
-
The Cancer Genome Atlas
- GEO:
-
Gene Expression Omnibus
- WGCNA:
-
Weighted gene co-expression network analysis
- GO:
-
Gene ontology
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- HR:
-
Hazard ratio
- CRT:
-
Calreticulin
- HMGB1:
-
High mobility group protein
- OS:
-
Overall survival
- SNV:
-
Single nucleotide variants
- CNV:
-
Copy number variation
- ME:
-
Module eigengenes
- CI:
-
Confidence interval
References:
McGuire, S. (2016). World Cancer Report 2014. Geneva, Switzerland: World Health Organization, International Agency for Research on Cancer, WHO Press, 2015. Advances in Nutrition, 7(2), 418–419. https://doi.org/10.3945/an.116.012211
Siegel, R. L., Giaquinto, A. N., & Jemal, A. (2024). Cancer statistics, 2024. A Cancer Journal for Clinicians, 74(1), 12–49. https://doi.org/10.3322/caac.21820
Stewart, C., Ralyea, C., & Lockwood, S. (2019). Ovarian cancer: An integrated review. Seminars in Oncology Nursing, 35(2), 151–156. https://doi.org/10.1016/j.soncn.2019.02.001
Jayson, G. C., Kohn, E. C., Kitchener, H. C., & Ledermann, J. A. (2014). Ovarian cancer. Lancet, 384(9951), 1376–1388. https://doi.org/10.1016/S0140-6736(13)62146-7
Krysko, D. V., Garg, A. D., Kaczmarek, A., Krysko, O., Agostinis, P., & Vandenabeele, P. (2012). Immunogenic cell death and DAMPs in cancer therapy. Nature Reviews Cancer, 12(12), 860–875. https://doi.org/10.1038/nrc3380
Kroemer, G., Galluzzi, L., Kepp, O., & Zitvogel, L. (2013). Immunogenic cell death in cancer therapy. Annual Review of Immunology, 31, 51–72. https://doi.org/10.1146/annurev-immunol-032712-100008
Obeid, Michel, Tesniere, Antoine, Ghiringhelli, François, Fimia, Gian Maria, Apetoh, Lionel, Perfettini, Jean-Luc., Castedo, Maria, Mignot, Grégoire., Panaretakis, Theoharis, Casares, Noelia, Métivier, Didier, Larochette, Nathanael, van Endert, Peter, Ciccosanti, Fabiola, Piacentini, Mauro, Zitvogel, Laurence, & Kroemer, Guido. (2007). Calreticulin exposure dictates the immunogenicity of cancer cell death. Nature Medicine, 13(1), 54–61. https://doi.org/10.1038/nm1523
Knittle, A. M., Helkkula, M., Johnson, M. S., Sundvall, M., & Elenius, K. (2017). SUMOylation regulates nuclear accumulation and signaling activity of the soluble intracellular domain of the ErbB4 receptor tyrosine kinase. Journal of Biological Chemistry, 292(48), 19890–19904. https://doi.org/10.1074/jbc.M117.794271
Zhao, L., Zhang, S., Chen, H., Kroemer, G., Kepp, O., & Liu, P. (2022). Interference of immunogenic chemotherapy by artificially controlled calreticulin secretion from tumor cells. Methods in Cell Biology, 172, 99–114. https://doi.org/10.1016/bs.mcb.2021.12.027
Lau, Tat San, Chan, Loucia Kit Ying., Man, Gene Chi Wai., Wong, Chi Hang, Lee, Jacqueline Ho Sze., Yim, So Fan, Cheung, Tak Hong, McNeish, Iain A., & Kwong, Joseph. (2020). Paclitaxel induces immunogenic cell death in ovarian cancer via TLR4/IKK2/SNARE-dependent exocytosis. Cancer Immunology Research, 8(8), 1099–1111. https://doi.org/10.1158/2326-6066.CIR-19-0616
Tzogani, Kyriaki, Penttilä, Karri, Lähteenvuo, Johanna, Lapveteläinen, Tuomo, Anglada, Lucía Lopez., Prieto, Carolina, Garcia-Ochoa, Blanca, Enzmann, Harald, Gisselbrecht, Christian, Delgado, Julio, & Pignatti, Francesco. (2021). EMA review of belantamab mafodotin (blenrep) for the treatment of adult patients with relapsed/refractory multiple myeloma. Oncologist, 26(1), 70–76. https://doi.org/10.1002/onco.13592
Markham, A. (2020). Lurbinectedin: First approval. Drugs, 80(13), 1345–1353. https://doi.org/10.1007/s40265-020-01374-0
Jia, Le., Pang, Minghui, Fan, Man, Tan, Xuan, Wang, Yiqian, Huang, Menglin, Liu, Yijing, Wang, Qin, Zhu, Yanhong, & Yang, Xiangliang. (2020). A pH-responsive pickering nanoemulsion for specified spatial delivery of immune checkpoint inhibitor and chemotherapy agent to tumors. Theranostics, 10(22), 9956–9969. https://doi.org/10.7150/thno.46089
Garg, A. D., De Ruysscher, D., & Agostinis, P. (2016). Immunological metagene signatures derived from immunogenic cancer cell death associate with improved survival of patients with lung, breast or ovarian malignancies: A large-scale meta-analysis. Oncoimmunology, 5(2), e1069938. https://doi.org/10.1080/2162402X.2015.1069938
Zhang, W., Liu, T., Jiang, L., Chen, J., Li, Q., & Wang, J. (2022). Immunogenic cell death-related gene landscape predicts the overall survival and immune infiltration status of ovarian cancer. Frontiers in Genetics, 13, 1001239. https://doi.org/10.3389/fgene.2022.1001239
Malta, T. M., Noushmehr, H., Cancer Genome Atlas Research Network. (2018). The immune landscape of cancer. Immunity, 48(4), 812–830. https://doi.org/10.1016/j.immuni.2018.03.023
Langfelder, P., & Horvath, S. (2008). WGCNA: An R package for weighted correlation network analysis. BMC Bioinformatics, 9, 559. https://doi.org/10.1186/1471-2105-9-559
Yoshihara, Kosuke, Shahmoradgoli, Maria, Martínez, Emmanuel, Vegesna, Rahulsimham, Kim, Hoon, Torres-Garcia, Wandaliz, Treviño, Victor, Shen, Hui, Laird, Peter W., Levine, Douglas A., Carter, Scott L., Getz, Gad, Stemke-Hale, Katherine, Mills, Gordon B., & Verhaak, Roel G.W.. (2013). Inferring tumour purity and stromal and immune cell admixture from expression data. Nature Communications, 4, 2612. https://doi.org/10.1038/ncomms3612
Roett, M. A., & Evans, P. (2009). Ovarian cancer: An overview. American Family Physician, 80(6), 609–616.
Kurnit, K. C., Fleming, G. F., & Lengyel, E. (2021). Updates and new options in advanced epithelial ovarian cancer treatment. Obstetrics and Gynecology, 137(1), 108–121. https://doi.org/10.1097/AOG.0000000000004173
Cortez, A. J., Tudrej, P., Kujawa, K. A., & Lisowska, K. M. (2018). Advances in ovarian cancer therapy. Cancer Chemotherapy and Pharmacology, 81(1), 17–38. https://doi.org/10.1007/s00280-017-3501-8
Morand, S., Devanaboyina, M., Staats, H., Stanbery, L., & Nemunaitis, J. (2021). Ovarian cancer immunotherapy and personalized medicine. International Journal of Molecular Sciences. https://doi.org/10.3390/ijms22126532
Lee, E. K., & Konstantinopoulos, P. A. (2019). Combined PARP and immune checkpoint inhibition in ovarian cancer. Trends Cancer, 5(9), 524–528. https://doi.org/10.1016/j.trecan.2019.06.004
Budczies, Jan, Kluck, Klaus, Beck, Susanne, Ourailidis, Iordanis, Allgäuer, Michael, Menzel, Michael, Kazdal, Daniel, Perkhofer, Lukas, Kleger, Alexander, Schirmacher, Peter, Seufferlein, Thomas, & Stenzinger, Albrecht. (2022). Homologous recombination deficiency is inversely correlated with microsatellite instability and identifies immunologically cold tumors in most cancer types. The Journal of Pathology: Clinical Research, 8(4), 371–382. https://doi.org/10.1002/cjp2.271
Yu, Z., Guo, J., Hu, M., Gao, Y., & Huang, L. (2020). Icaritin exacerbates mitophagy and synergizes with doxorubicin to induce immunogenic cell death in hepatocellular carcinoma. ACS Nano, 14(4), 4816–4828. https://doi.org/10.1021/acsnano.0c00708
Zhai, J., Gu, X., Liu, Y., Hu, Y., Jiang, Y., & Zhang, Z. (2023). Chemotherapeutic and targeted drugs-induced immunogenic cell death in cancer models and antitumor therapy: An update review. Frontiers in Pharmacology, 14, 1152934. https://doi.org/10.3389/fphar.2023.1152934
Jin, M. Z., & Wang, X. P. (2021). Immunogenic cell death-based cancer vaccines. Frontiers in Immunology, 12, 697964. https://doi.org/10.3389/fimmu.2021.697964
Garg, Abhishek D., Dudek, Aleksandra M., Ferreira, Gabriela B., Verfaillie, Tom, Vandenabeele, Peter, Krysko, Dmitri V., Mathieu, Chantal, & Agostinis, Patrizia. (2013). ROS-induced autophagy in cancer cells assists in evasion from determinants of immunogenic cell death. Autophagy, 9(9), 1292–1307. https://doi.org/10.4161/auto.25399
He, C., Huang, X., Zhang, Y., Lin, X., & Li, S. (2020). T-cell activation and immune memory enhancement induced by irreversible electroporation in pancreatic cancer. Clinical and Translational Medicine, 10(2), e39. https://doi.org/10.1002/ctm2.39
Liu, Gao, Sun, Jian, Yang, Zhang-Fu., Zhou, Cheng, Zhou, Pei-Yun., Guan, Ruo-Yu., Sun, Bao-Ye., Wang, Zhu-Tao., Zhou, Jian, Fan, Jia, Qiu, Shuang-Jian., & Yi, Yong. (2021). Cancer-associated fibroblast-derived CXCL11 modulates hepatocellular carcinoma cell migration and tumor metastasis through the circUBAP2/miR-4756/IFIT1/3 axis. Cell Death & Disease, 12(3), 260. https://doi.org/10.1038/s41419-021-03545-7
Liu, Meng-jie, Guo, Hui, Jiang, Li.-li, Jiao, Min, Wang, Shu-hong, Tian, Tao, Xiao, Fu., & Wang, Wen-juan. (2021). Elevated RBP-Jκ and CXCL11 expression in colon cancer is associated with an unfavorable clinical outcome. Cancer Management and Research, 13, 3651–3661. https://doi.org/10.2147/CMAR.S298580
Gao, Q., Wang, S., Chen, X., Cheng, S., Zhang, Z., Li, F., Huang, L., Yang, Y., Zhou, B., Yue, D., & Wang, D. (2019). Cancer-cell-secreted CXCL11 promoted CD8(+) T cells infiltration through docetaxel-induced-release of HMGB1 in NSCLC. Journal for ImmunoTherapy of Cancer, 7(1), 42. https://doi.org/10.1186/s40425-019-0511-6
Xie, Jian-Wei., Huang, Xiao-Bo., Chen, Qi-Yue., Ma, Yu-Bin., Zhao, Ya-Jun., Liu, Li-Chao., Wang, Jia-Bin., Lin, Jian-Xian., Jun, Lu., Cao, Long-Long., Lin, Mi., Ru-Hong, Tu., Zheng, Chao-Hui., Huang, Chang-Ming., & Li, Ping. (2020). m(6)A modification-mediated BATF2 acts as a tumor suppressor in gastric cancer through inhibition of ERK signaling. Molecular Cancer, 19(1), 114. https://doi.org/10.1186/s12943-020-01223-4
Yang, Wei, Bian, Wu., Ma, Ning, Wang, Yongfang, Guo, Jianhui, Zhu, Jun, & Zhao, Sihui. (2019). BATF2 reverses multidrug resistance of human gastric cancer cells by suppressing Wnt/β-catenin signaling. In Vitro Cellular & Developmental Biology—Animal, 55(6), 445–452. https://doi.org/10.1007/s11626-019-00360-5
Zhang, Y., & Liu, Z. (2017). STAT1 in cancer: Friend or foe? Discovery Medicine, 24(130), 19–29.
Zhang, Ying, Liu, Zhaoyong, Yang, Xia, Weiqing, Lu., Chen, Yelong, Lin, Youbin, Wang, Jin, Lin, Suxia, & Yun, Jing-Ping. (2021). H3K27 acetylation activated-COL6A1 promotes osteosarcoma lung metastasis by repressing STAT1 and activating pulmonary cancer-associated fibroblasts. Theranostics, 11(3), 1473–1492. https://doi.org/10.7150/thno.51245
Sasidharan, N. V., Toor, S. M., Ali, B. R., & Elkord, E. (2018). Dual inhibition of STAT1 and STAT3 activation downregulates expression of PD-L1 in human breast cancer cells. Expert Opinion on Therapeutic Targets, 22(6), 547–557. https://doi.org/10.1080/14728222.2018.1471137
Chen, R., Wu, W., Chen, S. Y., Liu, Z. Z., Wen, Z. P., Yu, J., et al. (2022). A pan-cancer analysis reveals CLEC5A as a biomarker for cancer immunity and prognosis. Frontiers in immunology., 13, 831542. https://doi.org/10.3389/fimmu.2022.831542
Liverani, Chiara, De Vita, Alessandro, Spadazzi, Chiara, Miserocchi, Giacomo, Cocchi, Claudia, Bongiovanni, Alberto, De Lucia, Anna, La Manna, Federico, Fabbri, Francesco, Tebaldi, Michela, Amadori, Dino, Tasciotti, Ennio, Martinelli, Giovanni, Mercatali, Laura, & Ibrahim, Toni. (2022). Lineage-specific mechanisms and drivers of breast cancer chemoresistance revealed by 3D biomimetic culture. Molecular Oncology, 16(4), 921–939. https://doi.org/10.1002/1878-0261.13037
Dhatchinamoorthy, K., Colbert, J. D., & Rock, K. L. (2021). Cancer immune evasion through loss of MHC class I antigen presentation. Frontiers in Immunology, 12, 636568. https://doi.org/10.3389/fimmu.2021.636568
Anika Tabassum, Md., Samdani, Nazmus, Dhali, Tarak Chandra, Alam, Rahat, Ahammad, Foysal, Samad, Abdus, & Karpiński, Tomasz M. (2021). Transporter associated with antigen processing 1 (TAP1) expression and prognostic analysis in breast, lung, liver, and ovarian cancer. Journal of Molecular Medicine, 99(9), 1293–1309. https://doi.org/10.1007/s00109-021-02088-w
Liao, Yunmei, Guo, Sheng, Chen, Yongwen, Cao, Dayan, Huan, Xu., Yang, Chengying, Fei, Lei, Ni, Bing, & Ruan, Zhihua. (2014). VSIG4 expression on macrophages facilitates lung cancer development. Laboratory Investigation, 94(7), 706–715. https://doi.org/10.1038/labinvest.2014.73
Xu, T., Jiang, Y., Yan, Y., Wang, H., Lu, C., Xu, H., Li, W., Fu, D., Lu, Y., & Chen, J. (2015). VSIG4 is highly expressed and correlated with poor prognosis of high-grade glioma patients. American Journal of Translational Research, 7(6), 1172–1180.
Iyer, Swathi V., Ranjan, Atul, Elias, Harold K., Parrales, Alejandro, Sasaki, Hiromi, Roy, Badal C., Umar, Shahid, Tawfik, Ossama W., & Iwakuma, Tomoo. (2016). Genome-wide RNAi screening identifies TMIGD3 isoform1 as a suppressor of NF-κB and osteosarcoma progression. Nature Communications, 7, 13561. https://doi.org/10.1038/ncomms13561
Zhou, M. H., & Wang, X. K. (2020). Microenvironment-related prognostic genes in esophageal cancer. Translational Cancer Research, 9(12), 7531–7539. https://doi.org/10.21037/tcr-20-2288
Ranjan, A., Iyer, S. V., & Iwakuma, T. (2017). Suppressive roles of A3AR and TMIGD3 i1 in osteosarcoma malignancy. Cell Cycle, 16(10), 903–904. https://doi.org/10.1080/15384101.2017.1308153
Qin, Y., Liu, H., Huang, X., Huang, L., Liao, L., Li, J., et al. (2022). GIMAP7 as a potential predictive marker for pan-cancer prognosis and immunotherapy efficacy. Journal of Inflammation Research, 15, 1047–1061. https://doi.org/10.2147/JIR.S342503
Wu, Y., Liu, H., Gong, Y., Zhang, B., & Chen, W. (2021). ANKRD22 enhances breast cancer cell malignancy by activating the Wnt/β-catenin pathway via modulating NuSAP1 expression. Bosnian Journal of Basic Medical Sciences, 21(3), 294–304. https://doi.org/10.17305/bjbms.2020.4701
Yin, Jun, Wenfan, Fu., Dai, Lu., Jiang, Zeyong, Liao, Hongying, Chen, Wenbin, Pan, Lei, & Zhao, Jian. (2017). ANKRD22 promotes progression of non-small cell lung cancer through transcriptional up-regulation of E2F1. Scientific Reports, 7(1), 4430. https://doi.org/10.1038/s41598-017-04818-y
Wu, M., Li, X., Zhang, T., Liu, Z., & Zhao, Y. (2019). Identification of a nine-gene signature and establishment of a prognostic nomogram predicting overall survival of pancreatic cancer. Frontiers in Oncology, 9, 996. https://doi.org/10.3389/fonc.2019.00996
Funding
This work was funded by Shaanxi Province Natural Science Basic Research Program S2022-JC-QN-1528, Shaanxi Province Natural Science Basic Research Program 2021JQ-397 and Scientific Research Development Fund of the First Affiliated Hospital of Xi’an Jiaotong University 2022QN-10.
Author information
Authors and Affiliations
Contributions
Chen R designed the research, conducted all analysis and organized the paper; Ren J participated in organization of the paper. Wang YF participated in the language checking of the article. Zhang X participated in revising of the paper; Jia YF, Liu C and Qu K revised and edited the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
All authors in this study declare no competing interests.
Ethical Approval
All data are from open databases.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, R., Ren, J., Wang, Y. et al. A Comprehensive Analysis Exploring the Impact of an Immunogenic Cell Death-Related Panel for Ovarian Cancer. Mol Biotechnol (2024). https://doi.org/10.1007/s12033-024-01215-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12033-024-01215-5