Abstract
Background
Talazoparib monotherapy in patients with germline BRCA-mutated, early-stage triple-negative breast cancer (TNBC) showed activity in the neoadjuvant setting in the phase II NEOTALA study (NCT03499353). These biomarker analyses further assessed the mutational landscape of the patients enrolled in the NEOTALA study.
Methods
Baseline tumor tissue from the NEOTALA study was tested retrospectively using FoundationOne®CDx. To further hypothesis-driven correlative analyses, agnostic heat-map visualizations of the FoundationOne®CDx tumor dataset were used to assess overall mutational landscape and identify additional candidate predictive biomarkers of response.
Results
All patients enrolled (N = 61) had TNBC. In the biomarker analysis population, 75.0% (39/52) and 25.0% (13/52) of patients exhibited BRCA1 and BRCA2 mutations, respectively. Strong concordance (97.8%) was observed between tumor BRCA and germline BRCA mutations, and 90.5% (38/42) of patients with tumor BRCA mutations evaluable for somatic-germline-zygosity were predicted to exhibit BRCA loss of heterozygosity (LOH). No patients had non-BRCA germline DNA damage response (DDR) gene variants with known/likely pathogenicity, based on a panel of 14 non-BRCA DDR genes. Ninety-eight percent of patients had TP53 mutations. Genomic LOH, assessed continuously or categorically, was not associated with response.
Conclusion
The results from this exploratory biomarker analysis support the central role of BRCA and TP53 mutations in tumor pathobiology. Furthermore, these data support assessing germline BRCA mutational status for molecular eligibility for talazoparib in patients with TNBC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Germline pathogenic variants in BRCA1 and BRCA2 account for approximately 5% of all breast cancers and up to 30% of hereditary breast cancer. Germline BRCA1/2 mutation carriers have a high incidence of breast cancer in their lifetime [1]; the lifetime probability is approximately 57–65% for those with BRCA1 mutations and 45–49% for those with BRCA2 mutations [2]. BRCA1/2 mutations can undergo loss of heterozygosity (LOH), wherein the non-mutated allele loses function in parallel with the original identified mutation [3]. Women with germline BRCA mutations are more prone to developing breast cancer at a younger age and with more aggressive disease than those with somatic mutations [2].
Approximately 70% of breast tumors containing germline BRCA1 mutations are diagnosed as triple-negative breast cancer (TNBC), a highly aggressive subtype of breast cancer associated with poor outcomes due to limited treatment options [2, 4]. A standard treatment approach for patients with early-stage TNBC is neoadjuvant chemotherapy, including patients with BRCA1/2 mutations [4, 5]. The FDA has approved pembrolizumab plus neoadjuvant chemotherapy as standard of care for patients with high-risk early-stage TNBC [6]. In addition, several further trials have demonstrated that platinum-containing regimens were effective for TNBC in the neoadjuvant setting; however, the efficacy of neoadjuvant platinum for patients with BRCA mutations remains unclear [5, 7,8,9,10].
BRCA1/2 mediate the repair of DNA double-strand breaks (DSBs) by homologous recombination repair (HRR) [11]. Cancer cells with germline BRCA1/2 mutations rely on poly(ADP-ribose) polymerase (PARP) enzymes 1 and 2 for DNA repair [12]. Inhibition of single-strand DNA repair mechanisms mediated by PARP1/2 results in increased single-strand DNA breaks, which eventually culminate in the accumulation of DNA DSBs that are normally repaired via HRR [11, 13]. Hence, PARP inhibition is synthetically lethal in BRCA-mutated cancer cells, which are deficient in HRR [11]. Moreover, PARP inhibitors trap PARP to sites of DNA damage and prevent replication fork progression, resulting in irreparable DNA DSBs [11, 14]. Synthetic lethality therefore provides a mechanistic rationale for the treatment of BRCA-mutated breast cancer, including TNBC, with PARP inhibitors [15, 16]. In addition, pre-clinical models suggest that BRCA LOH may increase sensitivity to PARP inhibition [17].
Talazoparib is an oral PARP inhibitor approved for use in the US, EU, and multiple other countries as a monotherapy for the treatment of patients with deleterious or suspected deleterious germline BRCA-mutated human epidermal growth factor receptor 2 (HER2)-negative locally advanced or metastatic breast cancer [18, 19]. In the phase II NEOTALA study, talazoparib monotherapy in patients with germline BRCA-mutated, HER2-negative early breast cancer (Stage I–III; disease confined to the breast) including TNBC was active in the neoadjuvant setting (pathologic complete response [pCR] rate by independent central review [ICR] was 45.8% [95% confidence interval (CI), 32.0–60.6%] in the population evaluable for primary analysis on pCR) and showed a safety profile consistent with previous findings [20, 21].
The aim of the biomarker analyses reported here was to assess the mutational landscape of patients in the NEOTALA study and to explore potential associations of these factors with patient outcomes [22]. This was achieved by evaluating tumor tissue as well as blood and saliva samples from patients enrolled in the NEOTALA study with exploration of tumor BRCA mutations including germline-somatic concordance and zygosity; the non-BRCA germline DNA damage response (DDR) mutational landscape; homologous recombination deficiency (HRD) as assessed via genomic LOH; and the non-BRCA tumor mutational landscape. In addition, TP53, MYC, and RAD21 genes were selected for exploratory analysis on pCR based on mutational prevalence, potential for crosstalk with DDR pathways [23, 24], and potential associations with outcome previously detected in the phase III EMBRACA study [25, 26].
Patients and methods
Study design and patients
NEOTALA (NCT03499353) was a phase II, non-randomized, single-arm, open-label study evaluating the efficacy and safety of talazoparib in the neoadjuvant setting for patients with early-stage germline BRCA1/2-mutated TNBC. Details of the primary study have been published [20, 21].
Briefly, eligible patients were ≥18 years old with histologically confirmed early HER2-negative carcinoma of the breast, germline BRCA1/2 mutations, and no evidence of distant metastases. The primary endpoint was pCR by ICR after 24 weeks of neoadjuvant talazoparib treatment followed by surgery. Residual cancer burden (RCB) by ICR was a key secondary endpoint. RCB 0 refers to no residual invasive cancer, or pCR [27]. RCB I refers to minimal RCB, RCB II to moderate RCB, and RCB III to extensive RCB (progressive disease) [28].
Mutational analysis
To assess molecular eligibility for the NEOTALA study, peripheral blood was tested using BRACAnalysis CDx® (Myriad Genetics Laboratories, Inc., Salt Lake City, UT, USA) [20, 21]. In the analyses reported here, baseline tumor tissue was tested retrospectively using FoundationOne®CDx (Foundation Medicine, Inc., Cambridge, MA, USA), which is a qualitative next-generation sequencing (NGS)-based in vitro diagnostic test that uses targeted high throughput hybridization-based capture technology for detection of substitutions, insertion, and deletion alterations (indels), and copy number alterations (CNAs) in 324 genes and select gene rearrangements. BRCA1/2 zygosity was assessed using somatic-germline-zygosity (SGZ) [29], and germline versus tumor sequence comparisons were used to determine whether tumor BRCA mutations were of germline or somatic origin. Tumor genomic LOH (gLOH) was assessed as an exploratory metric of HRD as previously described [30,31,32].
Germline mutational status of 14 non-BRCA DDR genes (ATM, BAP1, BARD1, BRIP1, CHEK2, FANCC, MLH1, MRE11A, NBN, PALB2, RAD50, RAD51C, RAD51D, and XRCC2) was retrospectively assessed in baseline saliva samples using CustomNext®-Cancer (Ambry Genetics, CA, USA). For exploratory tumor and germline analyses, mutations were defined as known/likely pathogenic/deleterious single-nucleotide variants, insertions, deletions, rearrangements, or amplifications, unless otherwise stated. For analysis of prevalence of non-BRCA tumor mutations, CNAs were analyzed both collectively with other mutations and separately as these alterations often reflect larger genomic changes that are not necessarily associated with a specific gene [33, 34].
Patient population and biomarker analysis
The biomarker analysis populations included all patients from the NEOTALA study intent-to-treat (ITT) population with tumor samples suitable for analysis by NGS. The total number of patients evaluable for exploratory biomarker analyses varied depending on the assay (i.e., mutation detection, zygosity prediction for detected mutations [limited to short variants in tumors of sufficient quality and purity], or gLOH) and matrix (tumor or saliva). Agnostic heat-map visualizations of the FoundationOne®CDx tumor dataset were used to assess the overall mutational landscape and to identity candidate predictive biomarkers.
Efficacy endpoints
The association between the mutational status of TP53, MYC, RAD21, or RB1 and the primary endpoint pCR as per ICR was investigated using logistic regression. In addition, the association between the mutational status of these genes and RCB was explored. RCB index was assessed as a continuous variable [28] in this case rather than categorically, to facilitate this correlative analysis.
Results
Patients
A total of 61 female patients with TNBC and an average age of 44.6 years were enrolled in the ITT analysis and safety population.
Non-BRCA DDR germline landscape
Forty-nine patients were included in the population evaluable for non-BRCA germline DDR mutations. Variants of ambiguous or unknown significance were detected in the saliva of 3 (6.1%) patients, each for PALB2 and RAD50; 2 (4.1%) patients for ATM; and 1 (2.0%) patient, each for BAP1, BARD1, BRIP1, FANCC, and RAD51C (Supplementary Table S1). No patients had known/likely pathogenic non-BRCA germline DDR variants.
BRCA mutations and zygosity
Of 52 tumor samples from the population evaluable by FoundationOne®CDx, 39 (75.0%) and 13 (25.0%) patients exhibited BRCA1 and BRCA2 mutations, respectively; 1 (1.9%) patient exhibited mutations in both genes; and 1 (1.9%) patient had mutations in neither (Supplementary Table S2). Among the 45 patients evaluable for both germline and tumor analyses, 44 (97.8%) exhibited the same BRCA mutation in tumor as originally detected in germline. The remaining patient exhibited a germline BRCA1 rearrangement but lacked a tumor BRCA (tBRCA) mutation (Fig. 1). In the population evaluable for SGZ, 38/42 (90.5%) patients with tBRCA mutations were predicted to exhibit BRCA LOH (Table 1). Twenty-nine (69.0%) patients were predicted to have one or more BRCA1 mutation with LOH and 9 (21.4%) were predicted to have one or more BRCA2 mutation with LOH. Of the four patients not exhibiting BRCA LOH, two exhibited heterozygous tBRCA2 mutations, one exhibited a heterozygous tBRCA1 mutation, and one exhibited a heterozygous tBRCA2 mutation and a tBRCA1 rearrangement not amenable to SGZ prediction.
Percentage of tumors concordant with germline BRCA1 or BRCA2 mutational status. The proportion of patients with a known germline BRCA1 mutation per Myriad Genetics laboratory results who had a BRCA1 mutation (defined as known or likely pathogenic variant) detected in tumor using FoundationOne®CDx is shown, and likewise for BRCA2. One patient was discordant for BRCA mutational status (germline BRCA1 exon 13 ins 6 kb [large rearrangement]) but exhibited no BRCA1/2 mutation in tumor. All patients showing concordant BRCA1 or BRCA2 mutational status exhibited the same mutation in tumor as originally detected in germline
Non-BRCA tumor mutational landscape
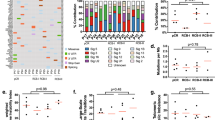
The overall tumor mutational landscape in the evaluable population is shown in Fig. 2. Based on variants with known or likely pathogenic impact, TP53 mutations were near universal across all patients with one exception (n = 51; 98.1%). Genes with mutations having ≥10% prevalence included BRCA1 (n = 39; 75.0%), MYC (n = 14; 26.9%), RAD21 (n = 14; 26.9%), BRCA2 (n = 13; 25.0%), RB1 (n = 11; 21.2%), PTEN (n = 10; 19.2%), and PIK3CA (n = 9; 17.3%) (Table 2). Excluding CNAs, non-BRCA genes with a mutational prevalence ≥10% included RB1 (n = 11; 21.2%), PIK3CA (n = 7; 13.5%), and PTEN (n = 7; 13.5%) (Table 2). In this TNBC subpopulation, MYC (14 [26.9%] patients) and RAD21 (14 [26.9%] patients) were the most frequently mutated genes based on copy number changes with known or likely pathogenic impact (Table 2). In this study, despite the higher prevalence of BRCA1 versus BRCA2 mutations, PIK3CA tended to be more frequently co-mutated with BRCA2 (n = 5/13) than with BRCA1 (n = 4/39).
Tumor mutational landscape – population evaluable by FoundationOne®CDx. The figure displays only those genes altered in two or more patients. For rearrangements, if a partner gene was present, both genes were labeled. If a patient has multiple alterations in a gene, this is indicated by the addition of a black square on the tile. The tile background color depends on the SV-LOH status determined in the following order: SV-LOH, SV-not under LOH, SV NA, or if neither of the alterations are SVs, the tile will be colored as a CN or RE. For example, (i) a patient with one SVLOH and one SV-not under LOH will be colored light blue, (ii) two SVs not under LOH will be colored light green, (iii) a patient with one SV and one CN will be colored as determined by its SV LOH status. CN copy number, LOH loss of heterozygosity, NA not available, RE rearrangement, SV short variant
An exploratory analysis of the association between the tumor mutational status of TP53, MYC, and RAD21 and pCR (ICR: breast/axilla dataset) is presented in Supplementary Table S3. The mutational status of MYC or RAD21 had no observed association with pCR (for each MYC and RAD21: odds ratio = 0.52; 95% CI, 0.12–2.30; p = 0.3893). The mutational status of TP53 had no observed association with pCR (p = 0.9629), although this analysis was confounded by the numerical imbalance favoring TP53 mutations (Supplementary Table S3). Likewise, there was no apparent association between the mutational status of these genes and RCB scores (Supplementary Fig. S1).
A joint visualization of tumor gLOH status and pCR (ICR: breast/axilla dataset) is provided in Fig. 3 showing gLOH distribution ranges from 1.5 to 42.2% for evaluable patients, with no apparent relationship of gLOH score to response, and no signs of any potential threshold gLOH score distinguishing responders from non-responders. An analysis of response employing a gLOH threshold of ≥16% (which is predictive of biallelic BRCA loss in ovarian and breast cancer [30]) found that gLOH was elevated in 24/27 (88.9%) tumors evaluable for both gLOH and pCR. Consistent with the results displayed in Fig. 3, no association between tumor gLOH status and pCR was observed but this was confounded by the imbalance in prevalence favoring gLOH-high (odds ratio of gLOH-high vs. gLOH-low = not evaluable; p = 0.9447) (Supplementary Table S4).
Exploration of potential associations of tumor mutations in other genes with response
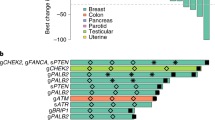
To explore additional potential associations of tumor mutational status with outcome, an agnostic heat-map visualization of response as a function of mutational status was performed in the evaluable biomarker population (Fig. 4). In general, despite a low number of patients, mutations in most genes appeared to exhibit a similar distribution across response categories. For example, no association was apparent between PIK3CA and/or PTEN mutational status and response category. One potential exception was RB1, in which 6/11 (54.5%) patients with alterations had pCRs by ICR versus 1/11 (9.1%) patients with a pathologic partial response by ICR (Fig. 4), although the mutational status of RB1 was not found to be significantly associated with pCR (odds ratio = 5.67; 95% CI, 0.62–52.09; p = 0.1254) (Supplementary Table S3). Similarly, RCB scores were typically lower for RB1-mutant patients in the population with tumors evaluable for FoundationOne®CDx and RCB analysis (Supplementary Fig. S1).
Discussion
In this biomarker analysis of the NEOTALA study, tumor BRCA mutations were evident in all but one patient. Strong concordance (97.8%) was observed between tumor BRCA and germline BRCA mutations analyzed by FoundationOne®CDx and BRACAnalysis CDx®, respectively, except for one patient who exhibited a germline BRCA1 mutation but lacked a tumor BRCA mutation. This exception could reflect genomic instability in that the mutation may have been lost [35], or alternatively, this germline rearrangement not being detectable in the tumor by FoundationOne®CDx. The strong concordance observed in this study is consistent with the findings in the phase III EMBRACA, phase II ABRAZO, and phase III OlympiAD studies in which 95.3%, 96.4%, and 99.3% of patients, respectively, exhibited concordance between tumor and germline BRCA mutational status [26, 36, 37].
In this TNBC population, a total of 38/42 (90.5%) patients with tumor BRCA mutations evaluable for SGZ were predicted to exhibit BRCA LOH. Previous studies support a high prevalence of BRCA1/2 mutational LOH in breast cancer [30, 38]. This high incidence of BRCA LOH coupled with a low overall population size precludes meaningful analysis of tumor BRCA zygosity on response. This finding contrasts with another study analyzing germline BRCA1/2-mutated breast cancer tumors [17]; a greater proportion of patients in this study had one or more BRCA1 mutation with LOH (69.0%) than patients with one or more BRCA2 mutation with LOH (21.4%). In this previous analysis, LOH was seen in 90.2% of BRCA1 carriers and 54.3% of BRCA2 carriers [17].
No patients had non-BRCA germline DDR gene variants with known/likely pathogenicity, which may reflect the small patient numbers in this study and indicate that patients were enrolled based on the presence of germline BRCA mutations. PALB2 and RAD50 were the most commonly occurring variants of ambiguous or unknown significance (each 6.1%). In this TNBC population, tumor PIK3CA mutations, excluding CNAs, had a prevalence of 13.5%, which aligns with previous findings that show PIK3CA is frequently mutated in breast cancer [39, 40] and is potentially an actionable target in the advanced setting [41].
In this study, tumor TP53 mutations, based on variants with known or likely pathogenic impact, excluding CNAs, were near-universal (n = 51; 98.1%). This finding is consistent with the anticipated high prevalence of TP53 mutations in TNBC [15, 42]. In the NEOTALA study, MYC and RAD21 both exhibited CNAs in 26.9% of tumors, with no association with pCR or RCB. This joint high prevalence is consistent with MYC and RAD21 being closely linked on amplicon 8q24 and frequently coamplified in BRCA1-mutated breast cancer [43]. In the best versus worst responder visualization followed by Cox regression analysis of the EMBRACA study, an association between MYC amplification and short overall survival (OS) was observed in patients with TNBC receiving talazoparib (hazard ratio [HR] = 1.88; 95% CI, 1.10–3.20); this association was not evident in patients with TNBC receiving chemotherapy (HR = 0.71; 95% CI, 0.30–1.64) [25]. This finding was of interest considering that MYC has previously been linked to DDR gene regulation [24, 44]. Carey et al. demonstrate that MYC expression directly activates homologous recombination in response to DNA damage and could be used to predict response of patients with TNBC to therapy with PARP inhibitor combinations [24]. No such association with OS was seen in EMBRACA for RAD21 [25], a structural component of the cohesin complex, despite being closely linked chromosomally with MYC [43].
In the NEOTALA study, an agnostic heat-map visualization of response as a function of mutational status revealed that mutations in most genes appeared to exhibit a similar distribution across response categories. A weak association of RB1 alterations with response was observed, although this did not reach statistical significance (p = 0.1254). This finding is interesting given that a previous study demonstrated that RB1 can localize to DNA DSBs, thus promoting DNA end resection and homologous recombination [45]. Notably, all four short variants of RB1 under LOH exhibited pCR, which is intriguing given a report demonstrating that RB1 loss is associated with tumors that are extremely sensitive to platinum retreatment in ovarian cancer [46]. Further exploration of the findings from this ad hoc exploratory analysis is warranted.
Genomic LOH (an exploratory metric of HRD), assessed continuously or categorically, was not associated with response. However, it should be noted that this lack of association is in a germline BRCA-mutated context, with all but three evaluable patients gLOH-high (i.e., gLOH ≥ 16%), and therefore does not address the free-standing predictive utility of gLOH in TNBC. For example, a composite HRD score comprising gLOH, telomeric allelic imbalance, and large-scale state transitions identified TNBC tumors, including BRCA1/2-non-mutated, as being more likely to respond to platinum-containing therapy [47].
The limitations of the NEOTALA study have previously been reported [21]; however, the limitations of the current study include the following: some of the retrospective exploratory analyses involved small sample sizes and certain analyses were confounded due to imbalances in the presence or absence of particular biomarkers.
Overall, the results from this exploratory biomarker analysis of the NEOTALA study support the central role of BRCA mutations in tumor pathobiology in patients with germline BRCA1/2-mutated TNBC and the value of assessing germline BRCA mutational status for molecular eligibility for talazoparib in this indication.
Data availability
Upon request and subject to review, Pfizer will provide the data that support the findings of this study. Subject to certain criteria, conditions and exceptions, Pfizer may also provide access to the related individual de-identified participant data. See https://www.pfizer.com/science/clinical-trials/trial-data-and-results for more information.
Abbreviations
- CI:
-
Confidence interval
- CN:
-
Copy number
- CNA:
-
Copy number alteration
- DDR:
-
DNA damage response
- DSB:
-
Double-strand break
- EU:
-
European Union
- FDA:
-
US Food and Drug Administration
- gLOH:
-
Genomic LOH
- HR:
-
Hazard ratio
- HER2:
-
Human epidermal growth factor receptor 2
- HRD:
-
Homologous recombination deficiency
- HRR:
-
Homologous recombination repair
- ICR:
-
Independent central review
- ITT:
-
Intent-to-treat
- LOH:
-
Loss of heterozygosity
- NA:
-
Not available
- ND:
-
Not determined
- NR:
-
No response
- NGS:
-
Next-generation sequencing
- OS:
-
Overall survival
- PARP:
-
Poly(ADP-ribose) polymerase
- pCR:
-
Pathologic complete response
- pPR:
-
Pathologic partial response
- RCB:
-
Residual cancer burden
- RE:
-
Rearrangement
- SGZ:
-
Somatic-germline-zygosity
- SV:
-
Short variant
- TNBC:
-
Triple-negative breast cancer
- US:
-
United States
References
Gonçalves A, Bertucci A, Bertucci F. PARP inhibitors in the treatment of early breast cancer: the step beyond? Cancers. 2020;12:1378. https://doi.org/10.3390/cancers12061378.
Mylavarapu S, Das A, Roy M. Role of BRCA mutations in the modulation of response to platinum therapy. Front Oncol. 2018;8:16. https://doi.org/10.3389/fonc.2018.00016.
Zhang X, Sjoblom T. Targeting loss of heterozygosity: a novel paradigm for cancer therapy. Pharmaceuticals (Basel). 2021;14:57. https://doi.org/10.3390/ph14010057.
Lee JS, Yost SE, Yuan Y. Neoadjuvant treatment for triple negative breast cancer: recent progresses and challenges. Cancers. 2020;12:1404. https://doi.org/10.3390/cancers12061404.
Loibl S, O’Shaughnessy J, Untch M, Sikov WM, Rugo HS, McKee MD, et al. Addition of the PARP inhibitor veliparib plus carboplatin or carboplatin alone to standard neoadjuvant chemotherapy in triple-negative breast cancer (BrighTNess): a randomised, phase 3 trial. Lancet Oncol. 2018;19:497–509. https://doi.org/10.1016/S1470-2045(18)30111-6.
U.S. Food and Drug Administration. KEYTRUDA® (pembrolizumab) prescribing information. https://www.accessdata.fda.gov/drugsatfda_docs/label/2024/125514s160lbl.pdf (March 2024). Accessed 23 May 2024.
Tung N, Arun B, Hacker MR, Hofstatter E, Toppmeyer DL, Isakoff SJ, et al. TBCRC 031: randomized phase II study of neoadjuvant cisplatin versus doxorubicin-cyclophosphamide in germline BRCA carriers with HER2-negative breast cancer (the INFORM trial). J Clin Oncol. 2020;38:1539–48. https://doi.org/10.1200/JCO.19.03292.
von Minckwitz G, Schneeweiss A, Loibl S, Salat C, Denkert C, Rezai M, et al. Neoadjuvant carboplatin in patients with triple-negative and HER2-positive early breast cancer (GeparSixto; GBG 66): a randomised phase 2 trial. Lancet Oncol. 2014;15:747–56. https://doi.org/10.1016/S1470-2045(14)70160-3.
Sikov WM, Berry DA, Perou CM, Singh B, Cirrincione CT, Tolaney SM, et al. Impact of the addition of carboplatin and/or bevacizumab to neoadjuvant once-per-week paclitaxel followed by dose-dense doxorubicin and cyclophosphamide on pathologic complete response rates in stage II to III triple-negative breast cancer: CALGB 40603 (Alliance). J Clin Oncol. 2015;33:13–21. https://doi.org/10.1200/JCO.2014.57.0572.
Wang C-J, Xu Y, Lin Y, Zhu H-J, Zhou Y-D, Mao F, et al. Platinum-based neoadjuvant chemotherapy for breast cancer with BRCA mutations: a meta-analysis. Front Oncol. 2020;10:592998. https://doi.org/10.3389/fonc.2020.592998.
Lord CJ, Ashworth A. PARP inhibitors: synthetic lethality in the clinic. Science. 2017;355:1152–8. https://doi.org/10.1126/science.aam7344.
Javle M, Curtin NJ. The potential for poly (ADP-ribose) polymerase inhibitors in cancer therapy. Ther Adv Med Oncol. 2011;3:257–67. https://doi.org/10.1177/1758834011417039.
Morales J, Li L, Fattah FJ, Dong Y, Bey EA, Patel M, et al. Review of poly (ADP-ribose) polymerase (PARP) mechanisms of action and rationale for targeting in cancer and other diseases. Crit Rev Eukaryot Gene Expr. 2014;24:15–28. https://doi.org/10.1615/critreveukaryotgeneexpr.2013006875.
Murai J, Huang SY, Renaud A, Zhang Y, Ji J, Takeda S, et al. Stereospecific PARP trapping by BMN 673 and comparison with olaparib and rucaparib. Mol Cancer Ther. 2014;13:433–43. https://doi.org/10.1158/1535-7163.Mct-13-0803.
Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490:61–70. https://doi.org/10.1038/nature11412.
Helleday T. The underlying mechanism for the PARP and BRCA synthetic lethality: clearing up the misunderstandings. Mol Oncol. 2011;5:387–93. https://doi.org/10.1016/j.molonc.2011.07.001.
Maxwell KN, Wubbenhorst B, Wenz BM, De Sloover D, Pluta J, Emery L, et al. BRCA locus-specific loss of heterozygosity in germline BRCA1 and BRCA2 carriers. Nat Commun. 2017;8:319. https://doi.org/10.1038/s41467-017-00388-9.
U.S. Food and Drug Administration. TALZENNA® (talazoparib) prescribing information. https://labeling.pfizer.com/ShowLabeling.aspx?id=11046 (February 2024). Accessed 23 May 2024.
European Medicines Agency. TALZENNA® (talazoparib) Summary of product characteristics. https://www.ema.europa.eu/en/documents/product-information/talzenna-epar-product-information_en.pdf (May 2024). Accessed 23 May 2024.
Litton JK, Beck JT, Jones JM, Andersen J, Blum JL, Mina LA, et al. Neoadjuvant talazoparib in patients with germline BRCA1/2 (gBRCA1/2) mutation-positive, early HER2-negative breast cancer (BC): results of a phase 2 study. J Clin Oncol. 2021;39:505. https://doi.org/10.1200/JCO.2021.39.15_suppl.505.
Litton JK, Beck JT, Jones JM, Andersen J, Blum JL, Mina LA, et al. Neoadjuvant talazoparib in patients with germline BRCA1/2 mutation-positive, early-stage triple-negative breast cancer: results of a phase II study. Oncologist. 2022;28:845–55. https://doi.org/10.1093/oncolo/oyad139.
Telli ML, Litton JK, Beck JT, Jones JM, Andersen J, Mina LA, et al. Neoadjuvant talazoparib (TALA) in patients (pts) with germline BRCA1/2 (gBRCA1/2) mutation-positive, early HER2-negative breast cancer (BC): exploration of tumor BRCA mutational status and zygosity and overall mutational landscape in a phase 2 study. J Clin Oncol. 2021;39(suppl 15):Abstract 554. https://doi.org/10.1200/JCO.2021.39.15_suppl.554.
Bassi C, Fortin J, Snow BE, Wakeham A, Ho J, Haight J, et al. The PTEN and ATM axis controls the G1/S cell cycle checkpoint and tumorigenesis in HER2-positive breast cancer. Cell Death Differ. 2021;28:3036–51. https://doi.org/10.1038/s41418-021-00799-8.
Carey JPW, Karakas C, Bui T, Chen X, Vijayaraghavan S, Zhao Y, et al. Synthetic lethality of PARP inhibitors in combination with MYC blockade is independent of BRCA status in triple-negative breast cancer. Cancer Res. 2018;78:742–57. https://doi.org/10.1158/0008-5472.CAN-17-1494.
Ettl J, Laird AD, Blum JL, Rugo HS, Hurvitz SA, Martín M, et al. A retrospective analysis of association of MYC and RAD21 amplification with final overall survival (OS) data in the phase 3 EMBRACA study with talazoparib. In: Virtual SABCS, 8–12 December 2020.
Blum JL, Laird AD, Litton JK, Rugo HS, Ettl J, Hurvitz SA, et al. Determinants of response to talazoparib in patients with HER2-negative, germline BRCA1/2-mutated breast cancer. Clin Cancer Res. 2022;28:1383–90. https://doi.org/10.1158/1078-0432.CCR-21-2080.
Symmans WF, Wei C, Gould R, Yu X, Zhang Y, Liu M, et al. Long-term prognostic risk after neoadjuvant chemotherapy associated with residual cancer burden and breast cancer subtype. J Clin Oncol. 2017;35:1049–60. https://doi.org/10.1200/JCO.2015.63.1010.
Symmans WF, Peintinger F, Hatzis C, Rajan R, Kuerer H, Valero V, et al. Measurement of residual breast cancer burden to predict survival after neoadjuvant chemotherapy. J Clin Oncol. 2007;25:4414–22. https://doi.org/10.1200/JCO.2007.10.6823.
Sun JX, He Y, Sanford E, Montesion M, Frampton GM, Vignot S, et al. A computational approach to distinguish somatic vs. germline origin of genomic alterations from deep sequencing of cancer specimens without a matched normal. PLoS Comput Biol. 2018;14:e1005965. https://doi.org/10.1371/journal.pcbi.1005965.
Sokol ES, Pavlick D, Khiabanian H, Frampton GM, Ross JS, Gregg JP, et al. Pan-cancer analysis of BRCA1 and BRCA2 genomic alterations and their association with genomic instability as measured by genome-wide loss of heterozygosity. JCO Precis Oncol. 2020;4:442–65. https://doi.org/10.1200/po.19.00345.
Swisher EM, Lin KK, Oza AM, Scott CL, Giordano H, Sun J, et al. Rucaparib in relapsed, platinum-sensitive high-grade ovarian carcinoma (ARIEL2 Part 1): an international, multicentre, open-label, phase 2 trial. Lancet Oncol. 2017;18:75–87. https://doi.org/10.1016/S1470-2045(16)30559-9.
Coleman RL, Oza AM, Lorusso D, Aghajanian C, Oaknin A, Dean A, et al. Rucaparib maintenance treatment for recurrent ovarian carcinoma after response to platinum therapy (ARIEL3): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet. 2017;390:1949–61. https://doi.org/10.1016/S0140-6736(17)32440-6.
Ooi A, Inokuchi M, Horike SI, Kawashima H, Ishikawa S, Ikeda H, et al. Amplicons in breast cancers analyzed by multiplex ligation-dependent probe amplification and fluorescence in situ hybridization. Hum Pathol. 2019;85:33–43. https://doi.org/10.1016/j.humpath.2018.10.017.
Kneissig M, Bernhard S, Storchova Z. Modelling chromosome structural and copy number changes to understand cancer genomes. Curr Opin Genet Dev. 2019;54:25–32. https://doi.org/10.1016/j.gde.2019.02.005.
Ganesan S. Tumor suppressor tolerance: reversion mutations in BRCA1 and BRCA2 and resistance to PARP inhibitors and platinum. JCO Precis Oncol. 2018;2:1–4. https://doi.org/10.1200/po.18.00001.
Turner NC, Laird AD, Telli ML, Rugo HS, Mailliez A, Ettl J, et al. Genomic analysis of tumors from patients with advanced breast cancer and germline BRCA1/2 mutations treated with talazoparib in the phase 2 ABRAZO study. NPJ Breast Cancer. 2023;9:81. https://doi.org/10.1038/s41523-023-00561-y.
Hodgson D, Lai Z, Dearden S, Barrett JC, Harrington EA, Timms K, et al. Analysis of mutation status and homologous recombination deficiency in tumors of patients with germline BRCA1 or BRCA2 mutations and metastatic breast cancer: OlympiAD. Ann Oncol. 2021;32:1582–9. https://doi.org/10.1016/j.annonc.2021.08.2154.
Nik-Zainal S, Davies H, Staaf J, Ramakrishna M, Glodzik D, Zou X, et al. Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature. 2016;534:47–54. https://doi.org/10.1038/nature17676.
Mosele F, Stefanovska B, Lusque A, Tran Dien A, Garberis I, Droin N, et al. Outcome and molecular landscape of patients with PIK3CA-mutated metastatic breast cancer. Ann Oncol. 2020;31:377–86. https://doi.org/10.1016/j.annonc.2019.11.006.
Avivar-Valderas A, McEwen R, Taheri-Ghahfarokhi A, Carnevalli LS, Hardaker EL, Maresca M, et al. Functional significance of co-occurring mutations in PIK3CA and MAP3K1 in breast cancer. Oncotarget. 2018;9:21444–58. https://doi.org/10.18632/oncotarget.25118.
Pascual J, Turner NC. Targeting the PI3-kinase pathway in triple-negative breast cancer. Ann Oncol. 2019;30:1051–60. https://doi.org/10.1093/annonc/mdz133.
Darb-Esfahani S, Denkert C, Stenzinger A, Salat C, Sinn B, Schem C, et al. Role of TP53 mutations in triple negative and HER2-positive breast cancer treated with neoadjuvant anthracycline/taxane-based chemotherapy. Oncotarget. 2016;7:67686–98. https://doi.org/10.18632/oncotarget.11891.
Annunziato S, de Ruiter JR, Henneman L, Brambillasca CS, Lutz C, Vaillant F, et al. Comparative oncogenomics identifies combinations of driver genes and drug targets in BRCA1-mutated breast cancer. Nat Commun. 2019;10:397. https://doi.org/10.1038/s41467-019-08301-2.
Curti L, Campaner S. MYC-induced replicative stress: a double-edged sword for cancer development and treatment. Int J Mol Sci. 2021;22:6168. https://doi.org/10.3390/ijms22126168.
Vélez-Cruz R, Manickavinayaham S, Biswas AK, Clary RW, Premkumar T, Cole F, et al. RB localizes to DNA double-strand breaks and promotes DNA end resection and homologous recombination through the recruitment of BRG1. Genes Dev. 2016;30:2500–12. https://doi.org/10.1101/gad.288282.116.
da Costa A, do Canto LM, Larsen SJ, Ribeiro ARG, Stecca CE, Petersen AH, et al. Genomic profiling in ovarian cancer retreated with platinum-based chemotherapy presented homologous recombination deficiency and copy number imbalances of CCNE1 and RB1 genes. BMC Cancer. 2019;19:422. https://doi.org/10.1186/s12885-019-5622-4.
Telli ML, Timms KM, Reid J, Hennessy B, Mills GB, Jensen KC, et al. Homologous recombination deficiency (HRD) score predicts response to platinum-containing neoadjuvant chemotherapy in patients with triple-negative breast cancer. Clin Cancer Res. 2016;22:3764–73. https://doi.org/10.1158/1078-0432.CCR-15-2477.
Acknowledgments
Medical writing support was provided by Maddie Higgins, MBiolSci, of CMC AFFINITY, a division of IPG Health Medical Communications, and was funded by Pfizer.
Funding
This study was sponsored by Pfizer.
Author information
Authors and Affiliations
Contributions
Conceptualization: MLT, JKL, MD. Data curation: MLT, JKL, JA, MD, YY, WFS, MM, JLB. Formal analysis: MLT, JKL, JTB, YY, JFH, LAA, AA, MM, ADL. Investigation: JTB, JMJ, LAM, MD, YY. Methodology: MLT, JKL, WFS, MM, ADL. Project administration: JTB, YY, KN, MM, ADL. Resources: JTB, JMJ. Supervision: JTB, LAM, YY, ADL. Validation: YY. Visualization: LAM, YY.
Corresponding author
Ethics declarations
Competing interests
MLT reports research support (to her institution) from Arvinas, AstraZeneca, Bayer, Blueprint Medicines, Genentech, GlaxoSmithKline, Hummingbird Biosciences, Merck, Novartis, OncoSec Medical, Pfizer, RefleXion, and Replicate; and consulting/advisory fees from AstraZeneca, Daiichi Sankyo, G1 Therapeutics, Genentech/Roche, Gilead, GlaxoSmithKline, Guardant, Merck, Natera, Novartis, OncoSec Medical, Pfizer, and Sanofi-Aventis. JKL reports research support (to her institution) from AstraZeneca, EMD Serono, Genentech, GlaxoSmithKline, Medivation/Pfizer, Merck, Novartis, Pfizer, and Zenith Epigenetics; fees for participation on the speakers’ bureaus for Clinical Care Options, Med Learning Group, Medpage, Medscape, Physicians’ Education Resource, PrIME Oncology, and UpToDate; honoraria and patent/royalty payments from UpToDate; travel fees from Clinical Care Options, Med Learning Group, Medscape, and Physicians’ Education Resource; and consulting/advisory from AstraZeneca, Ayala Pharmaceuticals, Medivation/Pfizer, and Pfizer—all uncompensated. JTB reports research support (to his institution) from AbbVie, Amgen, Ascentage Pharma Group, AstraZeneca, Bayer, Boston Biomedical/Bristol Myers Squibb, Celgene, Daiichi Sankyo, EMD Serono, Evelo Biosciences, Genentech/Roche, GlaxoSmithKline, Hutchison, Immunomedics, Laekna Therapeutics, Lilly, Mirati Therapeutics, Nektar, Novartis, Pfizer, Polynoma, and Seattle Genetics. JMJ has nothing to disclose. JA reports fees for participation on the speakers’ bureaus for AstraZeneca/Daiichi Sankyo, Genentech, Genomic Health, Immunomedics, Novartis, Puma Biotechnology, and Seattle Genetics; and consulting/advisory fees from AstraZeneca/Daiichi Sankyo, Athenex, Biotheranostics, Myriad, Novartis, Pfizer, Puma Biotechnology, and Seattle Genetics. LAM was an employee at Banner MD Anderson Cancer Center at the time these analyses were performed and has nothing to disclose. RB has nothing to disclose. MD reports consulting/advisory fees from Immunomedics, Novartis, Pfizer, and Seattle Genetics; and honoraria from Amgen. YY was an employee of the City of Hope Comprehensive Cancer Center and Beckman Research Institute at the time of these analyses and reports research support from Eisai, Genentech, and Merck; consulting/advisory fees from BCI Pharma, Daiichi Sankyo, Eisai, Genentech, Gilead, Novartis, Pfizer, and Puma Biotechnology; fees for participation on the speakers’ bureaus for AstraZeneca, Daiichi Sankyo/Lilly, Eisai, Genentech, Gilead, Merck, and Pfizer; and fees for expert testimony for Novartis. WFS reports consulting/advisory fees from Almac Diagnostics and Merck; support for travel from Luminex and Merck; stock ownership in Delphi Diagnostics, Eiger BioPharmaceuticals, ISIS Pharmaceuticals, and Nuvera Biosciences; and an uncompensated relationship with Delphi Diagnostics. JFH was an employee of Foundation Medicine, Inc. at the time of these analyses. LAA is an employee of Foundation Medicine, Inc., and a stockholder of Roche Holdings AG. ADL is an employee of Pfizer and may hold Pfizer stock. AA, MM and KN were employees at Pfizer at the time of these analyses. JLB was a physician at Texas Oncology-Baylor Charles A. Sammons Cancer Center, US Oncology Network at the time these analyses were performed and reports consulting/advisory fees and honoraria from AstraZeneca, Athenex, Inc., Biotheranostics, Inc., Immunomedics, Inc., OncLive, Pfizer, Inc., Puma Biotechnology, Research to Practice, Sanofi, Tempus, and TD2 and participation on the speakers’ bureaus for Pfizer, Inc., and Tempus.
Ethical approval
This study was conducted according to the International Ethical Guidelines for Biomedical Research Involving Human Patients, the International Council for Harmonisation Guidelines for Good Clinical Practice, the 1964 Declaration of Helsinki and its later amendments, and other applicable national and local regulations. The study protocol was reviewed and approved by an institutional review board, ethics committee and research ethics board. This article does not contain any studies with animals performed by any of the authors.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Clinical Trial and registration date: April 17, 2018. NCT03499353. https://clinicaltrials.gov/study/NCT03499353.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Telli, M.L., Litton, J.K., Beck, J.T. et al. Neoadjuvant talazoparib in patients with germline BRCA1/2 mutation-positive, early-stage triple-negative breast cancer: exploration of tumor BRCA mutational status. Breast Cancer 31, 886–897 (2024). https://doi.org/10.1007/s12282-024-01603-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12282-024-01603-4