Abstract
There is a lack of standardised imaging methods for marine zooplankton due to the difficulty of manipulating small and often fragile specimens. Yet, standardised 2D photographs and 3D scans provide important morphological information to accompany DNA-barcoded specimens for reference databases such as the Barcode of Life Data System (BOLD). Shelled pteropods are considered as bio-indicators to study impacts of ocean acidification, and thus, it is especially important to obtain high-quality records of their fragile aragonitic shells. We used alcohol-based hand sanitiser gel as a medium for photographing pteropods of the genus Limacina prior to micro-CT scanning and destructive DNA analysis. The high viscosity and transparency of the hand sanitiser enabled easy handling of the specimens so that they could be positioned in a standardised orientation and photographed with a stacking microscope. The high-quality photographs provide a record of morphology and allow for subsequent geometric morphometric analyses. This method did not impact the downstream micro-CT and molecular analyses of the same specimens and resulted in publicly available 2D and 3D digital vouchers as well as ten reference DNA barcodes (partial Cytochrome Oxidase I gene sequences). While alcohol-based hand sanitiser entered our daily lives due to a distressing pandemic, we could make use of it as a cheap and easily available resource to make high quality voucher photographs of shelled pteropods. Digital vouchers serve as a record of their morphology for further taxonomic analyses and facilitate studies assessing shell growth and impacts of ocean acidification.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Shelled pteropods are holoplanktonic gastropods that are a common subject of global change research, due to their important ecological and biogeochemical roles and susceptibility to ocean acidification. Shelled pteropods have been regarded as bio-indicators of ocean acidification, because their thin aragonitic shell is sensitive to dissolution under acidified conditions (Orr et al. 2005; Bednaršek et al. 2017). Pteropods contribute to the oceanic carbon flux by producing biomass and sequestering carbon through high phytoplankton grazing (Hunt et al. 2008) and large downward fluxes of faecal pellets (Manno et al. 2010), mucus nets (Noji et al. 1997; Conley et al. 2018), and shells (Tsurumi et al. 2005; Fabry et al. 2009). Pteropods are also prey for heteropods (Böer et al. 2005), amphipods (Bernard 2006), cephalopods (Hanlon and Messenger 1998), and, in polar systems, fishes, seabirds (Hunt et al. 2008), and marine mammals (Lalli and Gilmer 1989). Their species diversity and population structure are commonly assessed with DNA barcodes (e.g. Hunt et al. 2010; Jennings et al. 2010; Burridge et al. 2017; Kohnert et al. 2020; Choo et al. 2021), which contributes to a global zooplankton DNA barcoding reference database.
There is an urgent need for more high-quality voucher photographs of specimens to complement DNA barcode reference databases of marine zooplankton (Bucklin et al. 2021). This allows for the morphological identification of specimens, and validation of their DNA barcodes (Bucklin et al. 2016; Laakmann et al. 2020). The preservation of morphological characters of the specimen, in addition to sequencing of DNA barcodes, and collection of associated georeferencing, environmental, and ecological metadata, are important components of an integrative taxonomy pipeline (Padial et al. 2010). With the aid of these multiple sources of information, we can successfully assess species boundaries within the zooplankton assemblage (Hirai et al. 2015; Bode et al. 2017; Burridge et al. 2019), and in some cases, resolve (pseudo-)cryptic species identified through DNA barcodes with more thorough inspection of their morphology (e.g. Burridge et al. 2015; Wall-Palmer et al. 2018). While non-destructive DNA extraction is possible in some zooplankton taxa by subsampling (e.g. Choquet et al. 2018), or by leaching DNA without damage to chitinous exoskeletons or calcite shells (e.g. Cornils 2015; Weiner et al. 2016), we have had no success in preserving the highly soluble aragonitic shells of pteropods using different DNA extraction protocols. In such cases, high-quality voucher photographs play a critical role as the only remaining record of the morphological traits of the DNA barcoded specimen.
There is a lack of methodological detail as to how zooplankton can be prepared for taking a high-quality stacking photograph. Ideally, these specimens should be positioned in a standardised orientation to facilitate measurements of body parts, or to highlight a particular aspect of their anatomy. In coiled gastropods especially, the positioning of the specimen is important to standardise the turn and tilt of the shell (Callomon 2019), which directly impacts the suitability of the photographs in 2D geometric morphometrics analyses. Known methods for positioning small invertebrates for photography vary from malleable tack for dry specimens to glass slides, wax cradles, and stainless steel nuts for wet specimens (Geiger et al. 2007). Micro-computed tomography (micro-CT) scanning is a method where specimens do not need to be placed in a standardised orientation because 3D reconstructions of their shell morphology are obtained (Shimizu et al. 2018, 2021), but it is expensive and time-consuming to have 3D scans for all specimens that are included in a DNA barcoding pipeline.
As part of our integrative taxonomy pipeline for planktonic gastropods, we needed to find a reproducible way to image specimens in a standardised orientation before destructive DNA extraction. Alcohol-based hand sanitiser gel became increasingly more common since the start of the COVID-19 pandemic as a convenient replacement of soap and water to maintain hand hygiene. Because of the availability of alcohol-based hand gel, we were inspired to include it as part of our integrative taxonomy workflow to facilitate the photography of small pteropods. Alcohol-based hand sanitiser has been commonly used in positioning and stabilising spiders (e.g. Valdez-Mondragón 2010; Bilton 2018; LeMay and Agnarsson 2020) and other macroinvertebrates (http://www.gigamacro.com/blog/wet-specimen-holder-macroinvertebrates) for photography; however, it had not been applied to planktonic gastropods or other marine zooplankton. Here, we explored the use of alcohol-based hand sanitiser to obtain high quality 2D images of shelled pteropods, and subsequently 3D micro-CT scans prior to destructive DNA analyses.
Material and methods
We used alcohol-based hand sanitiser in the photography of ten shelled pteropods belonging to the Limacina genus, representing five nominal species: L. helicina s.l., L. retroversa s.l., L. bulimoides, L. trochiformis, and L. lesueurii. Previously, fine black sand was used to position pteropods in a standardised orientation for photography (e.g. Burridge et al. 2017; Choo et al. 2020), although the noise in the background of images meant that they were unsuitable for downstream image recognition or machine learning applications. Here, alcohol-based hand gel was used as a positioning medium for photographing the specimen in a standardised apertural orientation. The store-bought hand sanitiser we used (“Dr. Original” brand) contained the following ingredients: denatured alcohol, water, polyacrylamide, C13-14 Isoparaffin, and Laureth-7.
Under the microscope, we prepared the setup for imaging, which consisted of a watch glass with a small volume of hand sanitiser, atop on a black velvet cloth which provided a dark background (Fig. 1a). Each specimen was gently placed on the blob of hand sanitiser in a watch glass. The specimen was then manipulated carefully into a standard orientation with a thin brush, to ensure the shell axis was parallel to the plane of the camera (Fig. 1a). The specimen and hand sanitiser were then covered with a thin layer of 96% ethanol to reduce glare (Fig. 1b). Subsequently, the shells were photographed using a Zeiss V20 stacking stereomicroscope with Axiovision software (Zeiss, Germany). Once photographed, each specimen was rinsed in 96% ethanol to wash off remaining gel and stored in 96% ethanol at −20 °C.
Imaging setup for Limacina specimens. a A watch glass with hand sanitiser added was placed on a sheet on black velvet under the microscope. The specimen was placed within the hand sanitiser and then manipulated into position. b A small volume of 96% ethanol was added to completely cover the hand sanitiser.
After the photograph was obtained for each specimen, a subset of five individuals was selected for micro-CT scanning. Specimens were prepared and scanned according to the protocol described in (Mekkes et al. 2021). We also obtained the mitochondrial cytochrome oxidase subunit I (COI) barcode of the 10 individuals. DNA was extracted and barcoded according to the protocol in (Choo et al. 2021). The shells were washed in 96% ethanol before micro-CT scanning and in MilliQ water before being placed in lysis buffer for DNA extraction to prevent possible inhibition of the polymerase chain reaction (PCR) due to any remaining (denatured) alcohol. Sequences were checked and edited in Geneious Prime 2021.1.1 (https://www.geneious.com) and submitted to the BOLD database (Ratnasingham and Hebert 2007, 2013).
Results and discussion
Alcohol-based hand sanitiser facilitated the imaging process of ethanol-preserved shelled pteropods and did not interfere with their downstream micro-CT scanning, DNA extraction, and molecular analyses. We experienced that orienting the specimens for stacking photography was efficient with alcohol gel and the transparent shells contrasted well against the clean dark background provided by the black velvet cloth. These high-quality images can be used for publication and geometric morphometric analyses and were included as digital vouchers in the BOLD reference database with minimal editing needed (Fig. 2).
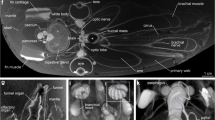
Stacking photographs (top row) and micro-CT 3D reconstructions (bottom row) of one specimen from each of the five nominal Limacina species. (a, f) L. retroversa s.l., (b, g) L. bulimoides, (c, h) L. trochiformis, (d, i) L. lesueurii, and (e, j) L. helicina s.l.. The stacking photographs and micro-CT reconstructions for each species are of the same individual, and species name and collection locality are given below each individual. Sensu lato (s.l.) is included for the species requiring taxonomic revision.
The DNA extraction and COI barcoding of the 10 specimens were successful, including for those five specimens (one for each species) that had the additional step of micro-CT scanning prior to DNA extraction. This adds further evidence to initial exploratory experiments (Hall et al. 2014) that DNA barcoding is compatible with micro-CT scanning and barcodes can be reliably amplified despite the X-ray radiation from the scanning. The DNA vouchers of these specimens are stored and accessible through Naturalis Biodiversity Center in Leiden, the Netherlands (Museum accessions: RMNH.MOL.347126, 347129, 347279-280, 347361, 347363, 347367-68, 347375, 347382), and their 2D photographs and COI barcodes were submitted to BOLD (BOLD accessions: LPCOI001-21-LPCOI010-21). Their micro-CT reconstructions were uploaded onto MorphoSource with digital object identifiers (DOIs) and can be accessed via https://www.morphosource.org/projects/000539891 with their respective DOIs. We note that two of the taxa L. retroversa s.l. and L. helicina s.l. require taxonomic revision (WoRMS Editorial Board 2021).
Our method of photography with alcohol-based hand sanitiser allowed for easy manipulation of small pteropods while providing enough viscosity to stabilise specimens during stacking photography. To support heavier specimens, we cooled the gel in the fridge to increase its viscosity. Different store-bought hand sanitiser brands have varying ingredient compositions that affect their viscosity and transparency (Berardi et al. 2020); therefore, this may require some testing for their suitability before use. Alcohol-based hand sanitisers also vary in terms of their alcohol concentrations, between the range of 60 and 95% (Boyce and Pittet 2002; Edmonds et al. 2012), though we expect that the initial fixation quality of the specimen (ideally in >95% ethanol) matters more than the ethanol concentration of the alcohol gel when it comes to subsequent DNA extraction and PCR (Stein et al. 2013). Alternative media used in positioning specimens for photodocumentation include glycerol, glycerine-gelatine jelly, and water-based lubricating jelly (Evitt 1984; Sasso Porto et al. 2016) although these do not contain ethanol and may affect downstream DNA analyses.
We suggest that the use of alcohol gel for photography can be integrated in the digitisation and barcoding pipelines of other marine zooplankton taxa for reference databases. High-quality voucher photographs of specimens are important to link genetic information with phenotypic characters through high-quality photographs and 3D reconstructions, especially if specimens have to be destroyed for DNA extraction (Bucklin et al. 2021). This is a useful approach for other calcifying zooplankton such as foraminifers and other pelagic gastropods, as well as for chaetognaths and polychaetes (personal observation). The increased flexibility in positioning specimens could also allow for better observation of diagnostic traits and more accurate measurements of appendages in e.g., crustaceans, and could facilitate re-assembly of fragile species that fall apart upon collection as for gelatinous groups such as siphonophores. Furthermore, the transparency of the gel allows to switch between different colours of background by simply using a different coloured cloth under the watchglass. While a dark background is commonly used in publications and typically allows for better thresholding (e.g. for machine learning or image recognition applications), a light background is better suited for analyses of colour variation, which may be indicative of species boundaries in e.g. Limacina bulimoides (Choo et al. 2023).
While alcohol-based hand sanitiser entered our daily lives due to a distressing pandemic, we learned that we could make use of it as a cheap and easily available resource as part of an integrative taxonomy pipeline for shelled pteropods. The ease of positioning specimens in a standard orientation also facilitates the challenging photography of younger life stages, such as veliger larvae in gastropods and nauplii in crustaceans, prior to DNA extraction. This is necessary to provide a complete overview of morphology across the life history of the species, facilitate species identifications, and resolve (pseudo-) cryptic species complexes. We hope that the described methodology can increase the efficiency of collecting morphological information and encourage the inclusion of high quality 2D and 3D digital vouchers in DNA reference databases for marine zooplankton.
References
Bednaršek N, Klinger T, Harvey CJ et al (2017) New ocean, new needs: application of pteropod shell dissolution as a biological indicator for marine resource management. Ecol Indic 76:240–244. https://doi.org/10.1016/j.ecolind.2017.01.025
Berardi A, Perinelli DR, Merchant HA et al (2020) Hand sanitisers amid CoViD-19: a critical review of alcohol-based products on the market and formulation approaches to respond to increasing demand. Int J Pharm 584:119431. https://doi.org/10.1016/j.ijpharm.2020.119431
Bernard KS (2006) The role of the euthesocome pteropod, Limacina retroversa, in the Polar Frontal Zone. Rhodes University, Southern Zone
Bilton DT (2018) Scotolemon doriae Pavesi, 1878, a soil-dwelling harvestman new to Britain (Opiliones: Phalangodidae). Arachnology 17:361–363. https://doi.org/10.13156/arac.2017.17.8.361
Bode M, Laakmann S, Kaiser P et al (2017) Unravelling diversity of deep-sea copepods using integrated morphological and molecular techniques. J Plankton Res 39:600–617. https://doi.org/10.1093/plankt/fbx031
Böer M, Gannefors C, Kattner G et al (2005) The Arctic pteropod Clione limacina: seasonal lipid dynamics and life-strategy. Mar Biol 147:707–717. https://doi.org/10.1007/s00227-005-1607-8
Boyce JM, Pittet D (2002) Guideline for hand hygiene in health-care settings: recommendations of the Healthcare Infection Control Practices Advisory Committee and the HICPAC/SHEA/APIC/IDSA Hand Hygiene Task Force. Am J Infect Control 30:S1–S46. https://doi.org/10.1067/mic.2002.130391
Bucklin A, Lindeque PK, Rodriguez-Ezpeleta N et al (2016) Metabarcoding of marine zooplankton: prospects, progress and pitfalls. J Plankton Res 38:393–400. https://doi.org/10.1093/plankt/fbw023
Bucklin A, Peijnenburg KTCA, Kosobokova KN et al (2021) Toward a global reference database of COI barcodes for marine zooplankton. Mar Biol 168:1–26. https://doi.org/10.1007/s00227-021-03887-y
Burridge AK, Goetze E, Raes N et al (2015) Global biogeography and evolution of Cuvierina pteropods. BMC Evol Biol 15:39. https://doi.org/10.1186/s12862-015-0310-8
Burridge AK, Hörnlein C, Janssen AW et al (2017) Time-calibrated molecular phylogeny of pteropods. PLoS One 12:e0177325. https://doi.org/10.1371/journal.pone.0177325
Burridge AK, Van Der Hulst R, Goetze E, Peijnenburg KTCA (2019) Assessing species boundaries in the open sea: an integrative taxonomic approach to the pteropod genus Diacavolinia. Zool J Linn Soc 187:1016–1040. https://doi.org/10.1093/zoolinnean/zlz049
Callomon P (2019) Standard views for imaging mollusk shells. Am. Malacol. Soc. 1–19
Choo LQ, Bal TMP, Choquet M et al (2020) Novel genomic resources for shelled pteropods: a draft genome and target capture probes for Limacina bulimoides, tested for cross-species relevance. BMC Genomics 21:11. https://doi.org/10.1186/s12864-019-6372-z
Choo LQ, Bal TMP, Goetze E, Peijnenburg KTCA (2021) Oceanic dispersal barriers in a holoplanktonic gastropod. J Evol Biol 34:224–240. https://doi.org/10.1111/jeb.13735
Choo LQ, Spagliardi G, Malinsky M et al (2023) Genome-wide phylogeography reveals cryptic speciation in the circumglobal planktonic calcifier Limacina bulimoides. Mol Ecol 32:3200–3219. https://doi.org/10.1111/mec.16931
Choquet M, Kosobokova K, Kwaśniewski S et al (2018) Can morphology reliably distinguish between the copepods Calanus finmarchicus and C. glacialis, or is DNA the only way? Limnol Oceanogr Methods 16:237–252. https://doi.org/10.1002/lom3.10240
Conley KR, Lombard F, Sutherland KR (2018) Mammoth grazers on the ocean’s minuteness: a review of selective feeding using mucous meshes. Proc R Soc B Biol Sci 285:20180056. https://doi.org/10.1098/rspb.2018.0056
Cornils A (2015) Non-destructive DNA extraction for small pelagic copepods to perform integrative taxonomy. J Plankton Res 37:6–10. https://doi.org/10.1093/plankt/fbu105
Edmonds SL, MacInga DR, Mays-Suko P et al (2012) Comparative efficacy of commercially available alcohol-based hand rubs and World Health Organization-recommended hand rubs: formulation matters. Am J Infect Control 40:521–525. https://doi.org/10.1016/j.ajic.2011.08.016
Evitt WR (1984) Some techniques for preparing, manipulating and mounting dinoflagellates. J Micropalaeontology 3:11–18. https://doi.org/10.1144/jm.3.2.11
Fabry V, McClintock J, Mathis J, Grebmeier J (2009) Ocean acidification at high latitudes: the bellwether. Oceanography 22:160–171. https://doi.org/10.5670/oceanog.2009.105
Geiger DL, Marshall BA, Ponder WF et al (2007) Techniques for collecting, handling, preparing, storing and examining small molluscan specimens. Molluscan Res 27:1–50
Hall AC, Sherlock E, Sykes D (2014) Does micro-CT scanning damage DNA in museum specimens? J Nat Sci Collect 2:22–29
Hanlon RT, Messenger JB (1998) Cephalopod behaviour. Cambridge University Press, Cambridge
Hirai J, Tsuda A, Goetze E (2015) Extensive genetic diversity and endemism across the global range of the oceanic copepod Pleuromamma abdominalis. Prog Oceanogr 138:77–90. https://doi.org/10.1016/j.pocean.2015.09.002
Hunt B, Strugnell J, Bednarsek N et al (2010) Poles apart: the “bipolar” pteropod species Limacina helicina is genetically distinct between the Arctic and Antarctic oceans. PLoS One 5:e9835. https://doi.org/10.1371/journal.pone.0009835
Hunt BPV, Pakhomov EA, Hosie GW et al (2008) Pteropods in Southern Ocean ecosystems. Prog Oceanogr 78:193–221. https://doi.org/10.1016/j.pocean.2008.06.001
Jennings RM, Bucklin A, Ossenbrügger H, Hopcroft RR (2010) Species diversity of planktonic gastropods (Pteropoda and Heteropoda) from six ocean regions based on DNA barcode analysis. Deep Sea Res Part II Top Stud Oceanogr 57:2199–2210. https://doi.org/10.1016/j.dsr2.2010.09.022
Kohnert PC, Cerwenka AF, Brandt A, Schrödl M (2020) Pteropods from the Kuril-Kamchatka Trench and the sea of Okhotsk (Euopisthobranchia; Gastropoda). Prog Oceanogr 181:102259. https://doi.org/10.1016/j.pocean.2019.102259
Laakmann S, Blanco-Bercial L, Cornils A (2020) The crossover from microscopy to genes in marine diversity: from species to assemblages in marine pelagic copepods. Philos Trans R Soc B Biol Sci 375:20190446. https://doi.org/10.1098/rstb.2019.0446
Lalli CM, Gilmer RW (1989) Pelagic snails: the biology of holoplanktonic gastropod molluscs. Stanford University Press, California
LeMay GA, Agnarsson I (2020) New species of smiley-faced spider Spintharus (Araneae, Theridiidae) from Brazil, and comments on unobserved diversity in South America. Zookeys 915:17–24. https://doi.org/10.3897/zookeys.915.47563
Manno C, Tirelli V, Accornero A, Fonda Umani S (2010) Importance of the contribution of Limacina helicina faecal pellets to the carbon pump in Terra Nova Bay (Antarctica). J Plankton Res 32:145–152. https://doi.org/10.1093/plankt/fbp108
Mekkes L, Renema W, Bednaršek N et al (2021) Pteropods make thinner shells in the upwelling region of the California Current Ecosystem. Sci Rep 11:1731. https://doi.org/10.1038/s41598-021-81131-9
Noji TT, Bathmann UV, Von Bodungen B et al (1997) Clearance of picoplankton-sized particles and formation of rapidly sinking aggregates by the pteropod, Limacina retroversa. J Plankton Res 19:863–875. https://doi.org/10.1093/plankt/19.7.863
Orr JC, Fabry VJ, Aumont O et al (2005) Anthropogenic decline in high-latitude ocean carbonate by 2100. Nature 437:681–686. https://doi.org/10.1038/nature04095
Padial JM, Miralles A, De la Riva I, Vences M (2010) The integrative future of taxonomy. Front Zool 7:16. https://doi.org/10.1186/1742-9994-7-16
Ratnasingham S, Hebert PDN (2013) A DNA-based registry for all animal species: the Barcode Index Number (BIN) system. PLoS One 8:e66213. https://doi.org/10.1371/journal.pone.0066213
Ratnasingham S, Hebert PDN (2007) BOLD: The Barcode Of Life Data system: barcoding. Mol Ecol Notes 7:355–364. https://doi.org/10.1111/j.1471-8286.2007.01678.x
Sasso Porto D, Melo GAR, Almeida EAB (2016) Clearing and dissecting insects for internal skeletal morphological research with particular reference to bees. Rev Bras Entomol 60:109–113. https://doi.org/10.1016/j.rbe.2015.11.007
Shimizu K, Kimoto K, Noshita K et al (2018) Phylogeography of the pelagic snail Limacina helicina (Gastropoda: Thecosomata) in the subarctic western North Pacific. J Molluscan Stud 84:30–37. https://doi.org/10.1093/mollus/eyx040
Shimizu K, Noshita K, Kimoto K, Sasaki T (2021) Phylogeography and shell morphology of the pelagic snail Limacina helicina in the Okhotsk Sea and western North Pacific. Mar Biodivers 51:22. https://doi.org/10.1007/s12526-021-01166-z
Stein ED, White BP, Mazor RD et al (2013) Evaluating ethanol-based sample preservation to facilitate use of DNA barcoding in routine freshwater biomonitoring programs using benthic macroinvertebrates. PLoS One 8:e51273. https://doi.org/10.1371/journal.pone.0051273
Tsurumi M, Mackas DL, Whitney FA et al (2005) Pteropods, eddies, carbon flux, and climate variability in the Alaska Gyre. Deep Res Part II Top Stud Oceanogr 52:1037–1053. https://doi.org/10.1016/j.dsr2.2005.02.005
Valdez-Mondragón A (2010) Zootaxa, Two new species of spiders of the genus Selenops Latreille, 1819 (Araneae: Selenopidae) and redescription of Selenops scitus Muma, 1953 from Mexico. Zootaxa 58:47–58
Wall-Palmer D, Burridge AK, Goetze E et al (2018) Biogeography and genetic diversity of the atlantid heteropods. Prog Oceanogr 160:1–25. https://doi.org/10.1016/j.pocean.2017.11.004
Weiner AKM, Morard R, Weinkauf MFG et al (2016) Methodology for single-cell genetic analysis of planktonic foraminifera for studies of protist diversity and evolution. Front Mar Sci 3:255. https://doi.org/10.3389/fmars.2016.00255
WoRMS Editorial Board (2021) World Register of Marine Species. https://www.marinespecies.org. Accessed 3 Nov 2021
Acknowledgements
We wish to warmly thank Deborah Wall-Palmer, Anders Illum, and Nikolaj Scharff for their insights leading to our use of alcohol-based hand sanitiser in pteropod photography, as well as Luis Martell for discussion about its use for hydrozoan imaging. We are also grateful to Jef Huisman and Galice Hoarau for comments on the manuscript and Frank Stokvis and Rob Langelaan at Naturalis Biodiversity Center for their help and expertise in implementing the DNA barcoding and micro-CT scanning pipelines. We would also like to thank the reviewers and editors whose comments have helped to improve our manuscript.
Funding
This study was supported by a Netherlands Organisation for Scientific Research (NWO) Vidi grant 016.161.351 to K. T. C. A. P. Nederlandse Organisatie voor Wetenschappelijk Onderzoek,016.161.351,Katja T.C.A Peijnenburg.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
No animal testing was performed during this study.
Sampling and field studies
All necessary permits for sampling have been obtained by the authors from the competent authorities.
Data availability
The data underlying this article are available in BOLD at http://www.boldsystems.org/, and can be accessed with BOLD Process IDs LPCOI001-21 to LPCOI010-21. The DNA vouchers (RMNH.MOL.347126, 347129, 347279-280, 347361, 347363, 347367-68, 347375, 347382) are accessioned at Naturalis Biodiversity Center, the Netherlands, and are available upon request. The 3D scans are archived on MorphoSource and can be accessed at https://www.morphosource.org/projects/000539891 or via the following DOIs: doi:https://doi.org/10.17602/M2/M486506, doi:https://doi.org/10.17602/M2/M486512, doi:https://doi.org/10.17602/M2/M486521, doi:https://doi.org/10.17602/M2/M486529, doi:https://doi.org/10.17602/M2/M495278, doi:https://doi.org/10.17602/M2/M495286, doi:https://doi.org/10.17602/M2/M495296, doi:https://doi.org/10.17602/M2/M495315, doi:https://doi.org/10.17602/M2/M495327, doi:https://doi.org/10.17602/M2/M495345.
Author contribution
KTCAP, GS, and LQC designed the research. KTCAP and LQC contributed samples used for the study. GS performed sample processing and analyses. LQC wrote the manuscript. All authors reviewed and approved of the manuscript.
Additional information
Communicated by E. Fileman
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is a contribution to the Topical Collection Discovering the water column: integrated taxonomic approaches for measuring marine pelagic biodiversity
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Choo, L.Q., Spagliardi, G. & Peijnenburg, K.T.C.A. The use of hand-sanitiser gel facilitates combined morphological and genetic analysis of shelled pteropods. Mar. Biodivers. 53, 77 (2023). https://doi.org/10.1007/s12526-023-01384-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12526-023-01384-7