Abstract
Human milk is the best nutrition for infants, providing optimal support for the developing immune system and gut microbiota. Hence, it has been used as source for probiotic strain isolation, including members of the genus Bifidobacterium, in an effort to provide beneficial effects to infants who cannot be exclusively breastfed. However, not all supplemented bifidobacteria can effectively colonise the infant gut, nor confer health benefits to the individual infant host; therefore, new isolates are needed to develop a range of dietary products for this specific age group. Here, we investigated the beneficial potential of Bifidobacterium breve DSM 32583 isolated from human milk. We show that in vitro B. breve DSM 32583 exhibited several characteristics considered fundamental for beneficial bacteria, including survival in conditions simulating those present in the digestive tract, adherence to human epithelial cell lines, and inhibition of growth of potentially pathogenic microorganisms. Its antibiotic resistance patterns were comparable to those of known beneficial bifidobacterial strains, and its genome did not contain plasmids nor virulence-associated genes. These results suggest that B. breve DSM 32583 is a potential probiotic candidate.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In recent years, there has been a significant increase in the number of food and dietary supplement products containing probiotic microorganisms, with specific strains of the genus Bifidobacterium being used worldwide [1, 2]. Probiotics are defined as ‘live microorganisms which when administered in adequate amounts confer a health benefit on the host’ [3]. Previous studies have reported various beneficial effects that Bifidobacterium strains exert on human health, such as regulation of intestinal microbial homeostasis [4], production of vitamins [5], modulation of local and systemic immune responses [6], and the hydrolysis of bile salts [7].
Bifidobacteria are among the first gut colonisers and the most ubiquitous inhabitants of the gastrointestinal tract, especially at early life stages. They can be regarded as founding microbiota members that exert crucial influence on the intestinal environment, the structure of the early-life microbial communities, and ultimately host development [8]. Bifidobacteria dominate the microbiota of vaginally delivered breast-fed infants [9]. Numerous studies have reported isolation of genomically identical Bifidobacterium strains from faecal samples of breastfeeding mother-infant pairs and the corresponding human milk, suggesting vertical transmission [10, 11]. The high Bifidobacterium abundance in breast-fed infants has been associated with their carbohydrate metabolism capabilities and linked to the presence of genes involved in the degradation of human milk oligosaccharides (HMOs) in their genomes [12, 13].
The ability to exert health benefits makes members of Bifidobacterium excellent potential probiotic candidates. However, studies have shown that not all probiotic-containing formulas or supplements ensure successful colonisation of the infant gut or facilitate an increase in the abundance of bifidobacteria in this environment [14, 15]. In addition, significant strain-specific differences in probiotic performance in vitro were described [16]. Therefore, ongoing research in this field is necessary to develop new and safe strains for formula and food production.
The initial screening and selection of beneficial strains involve experimental assessments covering a number of important criteria, for example, phenotypic and genotypic stability of the candidate strain, carbohydrate fermentation patterns, production of growth-inhibiting metabolites and other antimicrobial substances, and subsequent ability to inhibit potential pathogens [17, 18]. The probiotic strain must be able to tolerate acidic and bile-rich conditions in the gastrointestinal tract in order to survive passage through the digestive system, making resistance to gastric acid and bile salts one of the fundamental criteria in the selection of bacterial candidates for commercial use [19]. The ability of probiotic strains to adhere to the epithelium is another important selection criterion, as adhesion to the intestinal mucosa is regarded as a prerequisite for colonisation and can facilitate displacement of potentially pathogenic microorganisms from host cells [20].
A significant aspect of the initial probiotic strain selection is the assessment of antibiotic resistance patterns. The European Food Safety Authority (EFSA) states that bacterial strains used in commercial products should not harbour transferable antibiotic resistance and strongly recommends that minimum inhibitory concentrations (MICs) of the most important antimicrobial agents used in human medicine are evaluated at the probiotic strain selection stage [21]. Given that antibiotic resistance transfer can occur via a number of different mechanisms, it is recommended that whole genome sequences of bacterial candidate strains are screened for their mobilome, i.e. the presence of insertion elements, transposases, bacteriophages, and plasmids, in addition to phenotypic antimicrobial resistance assessment [18].
Here, we describe the functional properties of the novel strain Bifidobacterium breve DSM 32583, isolated from human milk and subjected to genotypic and phenotypic evaluation, to assess its beneficial or ‘probiotic’ potential. Genomic analysis and experimental approaches were used to investigate its antibiotic resistance patterns, carbohydrate fermentation profile, resistance to gastric acid, adherence to human epithelial cell lines, and its antimicrobial activity. Our results indicate that B. breve DSM 32583 possesses several promising characteristics of a probiotic candidate.
Materials and Methods
The strain isolation and the majority of phenotypic characterisation experiments listed in the ‘Materials and Methods’ section were carried out by ProbiSearch SLU, Madrid, Spain, unless indicated otherwise.
Common Name and Systematic Identification
The isolate has been deposited as DSM 32583 and WS 5622 in the German Collection of Microorganisms (DSMZ, Braunschweig) and the Weihenstephan Strain Collection (WS, Freising), respectively.
Isolation of B. breve DSM 32583
The strain was isolated as described previously [22]. Healthy women, after normal full-term pregnancy, without mastitis and other perinatal problems were enrolled to the study aiming at evaluating the diversity of lactobacilli and bifidobacteria in human milk (study B-06/262, Ethical Committee on Clinical Research of Hospital Clínico, Madrid).
Genotypic Identification
The whole genome of B. breve DSM 32583 has been sequenced and described [22]. Previously, we employed average nucleotide identity (ANI) analysis to confirm the affiliation of DSM 32583 to the B. breve taxon, with the 98.1% ANI value to the type strain B. breve DSM 20213T [22]. Here, complete genome sequences for Bifidobacterium species associated with human hosts—infants in particular (Bifidobacterium longum subsp. longum and Bifidobacterium longum subsp. infantis, B. breve, Bifidobacterium bifidum, Bifidobacterium catenulatum, Bifidobacterium pseudocatenulatum)—were downloaded from NCBI and used in comparative analysis employing Mashtree v.1.2.0 (Supplementary Table 1) [23].
The presence of mobile insertion elements in the genome of B. breve DSM 32583 was investigated using ISfinder (transposases) [24], PHASTER (phage-related sequences) [25], and PlasmidFinder (the presence of plasmids) [26]. Virulence genes and pathogenicity islands were searched for using VirulenceFinder v.2.0 [27] and PAIDB v.2.0 [28]. The presence of putative antimicrobial resistance genes was assessed using the CARD database (v.3.2.4) [29]. Additionally, regions flanking the detected tet(O) gene homologue were screened against the NCBI database [30] using the online blastn suite [31], as well as aligned to respective genomic regions of B. longum strains H34, F313, and Y1 (accession numbers: KY697301.1, KY697302.1, KY697303.1) [32] using MAFFT v.7 [33]. The presence of putative bacteriocin gene clusters was investigated using Bactibase and BAGEL4 [34, 35].
Prediction of HMO clusters was performed by comparing known bifidobacterial protein sequences to the genome of B. breve DSM 32583 using local blastp (e-value < 1 × 10−50, percentage identity > 70%). HMO clusters were annotated ‘present’ if all cluster components were identified at the above homology level. Incomplete clusters (more than 3 locally clustered genes) were annotated as ‘partially present’.
Phenotypic Identification
Gram-staining and catalase assays were conducted as described previously [36]. The activity of fructose-6-phosphate phosphoketolase (F6PPK) was confirmed through the enzymatic-colorimetric assay [37].
Antimicrobial Susceptibility Testing
Minimum inhibitory concentrations (MICs) were determined according to recommendations of the (EFSA) using a micro-dilution method within the ISO 10932 IDF 223 International Standard by Probisearch SL, Madrid, Spain, and Dairy Research Institute of Asturias-Spanish National Research Council (IPLA-CSIC), Villaviciosa, Spain. Additionally, MICs were also determined by E-test strips by Probisearch SL, Madrid, Spain, and LADR Laborverbund Dr. Kramer & Kollegen, Geesthacht, Germany.
Briefly, the MIC of 16 antibiotics, namely ampicillin, vancomycin, gentamycin, kanamycin, streptomycin, clindamycin, tetracycline, chloramphenicol, tylosin, ciprofloxacin, colistin, fosfomycin, neomycin, erythromycin, virginiamycin, linezolid, trimethoprim, and rifampicin was determined. Colonies from overnight culture plates were resuspended in sterile saline solution at turbidity of a McFarland standard of 1 (3 × 108 CFU/ml). The bacterial solution was then used to inoculate at 1:1000 LSM-cys liquid medium, composed of a 90% Iso-Sensitest medium (BD Difco, USA) and a 10% of MRS-cys medium (0.3 g/l) (BD Difco, USA), as indicated in Standard ISO 10932:2010. Next, 100 μl of the diluted culture was distributed into the well of antibiotic-precoated plates VetMIC Lact-1 (version 1) and Lact-2 (SVA National Veterinary Institute, Sweden). The plates were then incubated for 48 h under anaerobic conditions at 37°C, and the minimal inhibitory concentration (MIC), defined as the lowest antibiotic concentration at which there is no visual growth (ISO 10932:2010), was determined.
Additionally, MICs of selected antibiotics were also determined by E-test strips (AB BIODISK, Solna, Sweden) following manufacturer’s instructions. Briefly, individual colonies from overnight culture were resuspended in 5 ml of sterile saline until density corresponding to a McFarland standard of 0.5 was obtained (1.5 × 108 CFU/ml). A sterile cotton swab was dipped into the standardised inoculum and used to inoculate an agar plate. Inoculated plates were allowed to dry for approximately 15 min before application of the E-test strips with preformed antimicrobial gradients. After 24 h of incubation, the MIC was defined as the value corresponding to the first point on the E-test strip where growth did not occur along the inhibition ellipse. For bacteriostatic agents (e.g. tetracycline, erythromycin, and clindamycin), the MIC was read at the point where growth was inhibited by 80% (i.e. the first point of significant inhibition as judged visually).
The Carbohydrate Fermentation and Enzymatic Activity Profiles
The carbohydrate fermentation profile was determined with the API 50 CH system (BioMérieux, France) according to the manufacturer’s instructions by Probisearch SL, Madrid, Spain. The enzymatic activity profile was assayed using the API Zym galleries (BioMérieux, France) following the manufacturer’s instructions.
pH Stress Survival Assay
The survival of the isolate at low pH was tested in an in vitro model of the human stomach and small intestine based on that described by Marteau et al. [38]. Limosilactobacillus fermentum CECT 5716, Lactobacillus johnsonii La1, Lacticaseibacillus rhamnosus GG, and Lacticaseibacillus casei Immunitas were tested as controls. UHT-treated human milk (25 ml) containing approximately 109 CFU/ml of the isolate were diluted in 5 ml of sterile electrolyte solution containing 6.2 g/l of NaCl, 2.2 g/l of KCl, 0.22 g/l of CaCl2, and 1.2 g/l of NaHCO3 to simulate the in vivo dilution by saliva. Then, 5 ml of porcine gastric juice was added, and the mixture was incubated at 37°C on a shaker (paddle speed 200 ± 5 min⁻1). The pH curve in the stomach-resembling compartment was controlled to reproduce the values found in monogastric animals after yoghourt consumption [39]: pH 5.0 at initiation, pH 4.1 at 20 min, pH 3.0 at 40 min, and pH 2.1 at 60 min. Fractions were taken from these compartments at 20, 40, 60, and 80 min, in a manner that simulates the normal gastric emptying [38]. After adjusting their pH to 6.5 ± 0.2 with 1 M NaHCO3, they were mixed with 10 ml of a sterile electrolyte solution containing 5 g/l of NaCl, 0.6g/l of KCl, 0.3 g/l of CaCl2, 4% of porcine bile, and 7% of pancreatin (Sigma-Aldrich, UK), which simulates the content of the duodenal juice. After 120 min of successive exposure to these conditions, bacterial survival was determined by plating the samples onto TOS agar plates (Merck, Germany), which were then incubated anaerobically at 37°C for 48 h. These assays were performed in quadruplicate, and the values were expressed as the mean ± SD. Cumulative percentages of bacteria that survived the passage through the simulated gastric and duodenal compartments for the total collection period were obtained by summing the results of successive sampling periods.
Adhesion Assays to Caco-2 Cells
The adherence of B. breve DSM 32583 to HT-29 and Caco-2 cells was examined as described previously [40]. Additionally, L. fermentum CECT 5716, L. rhamnosus GG, and L. casei Imunitas were tested. Cells were grown in DMEM medium (PAA, Linz, Austria) containing 25 mM glucose and 1 mM sodium pyruvate and supplemented with 10% heat-inactivated (30 min, 56°C) foetal calf serum, 2 mM L-glutamine, 1% non-essential amino acid preparation, 100 U/ml penicillin, and 100 mg/ml streptomycin. For the adherence assays, HT-29 and Caco-2 were cultured to confluence in 2 ml of medium devoid of antibiotics. Approximately 10 days post confluence, 1 ml of the medium was replaced with 1 ml of B. breve DSM 32853 suspension (108 CFU/ml in DMEM). The inoculated cultures were incubated for 1 h at 37°C in 5% CO2. Then, the monolayer was washed five times with sterile PBS, fixed with methanol, stained with Gram stain and examined microscopically. The adherent bacteria in 20 random microscopic fields were counted for each test.
Growth Inhibition of Other Bacterial Species
An overlay method previously described [41, 42] was used to determine the ability of the strain to inhibit the growth of other microorganisms. The following bacteria were employed as indicator organisms: Enterococcus faecium P21, Enterococcus faecalis TAB28, Listeria monocytogenes ScottA, L. monocytogenes Ohio, Listeria innocua RdC, Staphylococcus aureus CECT 5191, Staphylococcus epidermidis CECT 231, Salmonella enterica serovar Choleraesuis CECT 4155, S. Choleraesuis CECT 409, S. Choleraesuis CECT 443, S. enterica serovar Enteritidis 4396, Escherichia coli CECT 4076 (O157:H7), E. coli RJM1, E. coli RJM2, Klebsiella pneumoniae CECT 142, Klebsiella oxytoca CECT 860T, and Proteus vulgaris CECT 484. The plates overlaid with bacterial indicators were incubated at 37°C for 48 h, while those overlaid with yeasts cells or fungal spores were incubated at 30°C for up to 120 h. The plates were examined for zones of inhibition around the strain streaks. All experiments assaying inhibitory activity were performed in triplicate.
Conjugated Linoleic Acid (CLA) and Conjugated Linolenic Acid (CLNA) Production
All reagents used were HPLC grade: hexane and sulphuric acid were obtained from Labscan (Ireland), linoleic acid (LA) (C18:2 cis9 cis12) from Sigma-Aldrich (USA), linolenic acid (ALA) (C18:3 cis9 cis12 cis15) from Nu-Chek Prep, Inc. (USA), and high conjugated linoleic acid (CLA) content oil (Tonalin R) from Cognis (Germany). LA and ALA were prepared as a 30 mg/ml stock solution containing 2% (w/v) Tween-80 and filter sterilised through a 0.45-µm pore size membrane.
B. breve DSM 32583 was grown overnight at 37°C in MRS broth supplemented with 0.05% (w/v) L-cysteine-HCL (Sigma-Aldrich, USA) and 0.2% (w/v) Tween-80 (MRS-Cys broth) under anaerobic conditions. Three percent (v/v) of the culture were transferred to fresh MRS-Cys broth (10 ml) containing free LA (0.5 mg/ml) and/or free ALA (0.5 mg/ml) and incubated at 37°C for 24 h under anaerobic conditions. The strain was also tested for CLA/CLNA production in 10% reconstituted skim milk supplemented with 0.05% (w/v) L-cysteine and 0.8% (w/v) casamino acids (milk-based medium).
Lipid isolation from culture media was carried out using a chloroform/methanol (2:1, v/v) solution according to Folch method modified by Iverson et al. [43]. The lipid residues obtained were subjected to a N2 flow and remained dissolved in chloroform at − 20°C until spectrophotometric analysis. For this analysis, lipid extract (200 μl) was placed on a quartz 96-wells plate, and total CLA was quantified at a wavelength of 233 nm in a spectrophotometer according to Rodriguez-Alcalá et al. [44]. Measurements were obtained in triplicate.
The concentrations of CLA and CLNA in the culture media were determined using a direct methylation method. Heptadecanoic acid (C17:0; Sigma-Aldrich, USA) was added to the samples as an internal standard. The fatty acid methyl esters (FAMEs) were dissolved in n-hexane and determined by gas chromatography (GC) in a chromatograph equipped with a VF-23 column (30 m × 0.25 nm × 0.25 μm; Varian, Netherlands). For gas–liquid (GLC) analysis, the initial temperature was 80°C. Then, the temperature was increased to 170°at 30°C/min, held at 170°C for 3 min, increased to 230°C at 30°C/min, and finally held at 230°C for 7 min. Helium was used as the carrier gas at a pressure of 15 psig and with a split ratio of 1:50. The injection volume was 0.5 μl, and the analysis time was 15 min. Peaks were identified by comparing the retention times of CLA methylated standards (Nu-check, USA) and by gas chromatography-mass spectrometry (GC/MS). CLA and CLNA concentrations were expressed as μg/ml, and their conversion rates from LA and ALA were calculated using the formula (CLA/(CLA + LA)) × 100 and (CLNA/(CLNA + LNA)) × 100, respectively.
Results
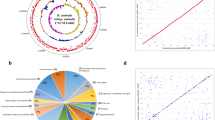
B. breve DSM 32583 was isolated from human milk and is a non-motile, non-spore forming, catalase negative, F6PPK-positive, rod-shaped anaerobic Gram-positive bacterium. The average nucleotide identity (ANI) analysis previously confirmed the affiliation of DSM 32583 to the B. breve taxon (Supplementary Table 2) [22]. Additionally, relatedness analysis of representative infant-associated Bifidobacterium species placed B. breve DSM 32583 within the B. breve cluster (Fig. 1).
Mobilome of B. breve DSM 32583
We assessed the genome of B. breve DSM 32583 for the presence of insertion sequences. This analysis revealed that all long open reading frames (ORFs) (> 50% coverage of known insertion sequence (IS) length) that showed high similarity to known transposases were homologous to IS previously reported for bifidobacteria, in particular the strain B. longum NCC2705 [45] (Supplementary Table 3).
The search for the presence of putative bacteriophage particles in the genome of B. breve DSM 32583 did not identify any complete prophage sequence, but only three incomplete prophage-associated regions were detected. The first was predicted to contain a phage tail protein similar to that from phage Strept phiSASD1, the second a transposase similar to that found in phage Bacter Diva, while the third region was predicted to contain a gene of unknown function similar to that previously identified in phage Entero phiEF24C (Supplementary Table 4). Furthermore, analysis of the genomic data also suggested the absence of plasmids.
Resistome: Virulence Factors and Antimicrobial Resistance
The screening for virulence and pathogenic potential of B. breve DSM 32583 revealed a complete absence of virulence genes.
Using the CARD database [29], the genome of B. breve DSM 32583 was examined for the presence of putative genes associated with antibiotic resistance. This analysis detected homologues of two resistance genes commonly identified in other bifidobacteria, the rifamycin-resistant beta-subunit of RNA polymerase (rpoB) and the tetracycline resistance gene tet(O) (Supplementary Table 5).
The identified homologue of the tet(O) gene in B. breve DSM 32583 was not found to be located in the vicinity of putative transposases, with only short, incomplete sequence regions (less than 80 nt) identified as identical to known insertion sequence elements upstream and downstream (Supplementary Table 3). However, manual comparison against the NCBI database (September 2022) revealed a putative homologue of a protein previously proposed to be transposon-associated (tnpV) directly upstream of the tet(O) gene. The tnpV gene was originally identified in the Clostridium perfringens transposon Tn4451, which is mobilisable but not self-mobile, i.e. in need of a transposase in close proximity. Previous reports suggested that the tnpV sequence showed similarity to viral transcriptional regulators based on PSI-BLAST searches, but its function and relevance remain unknown [46].
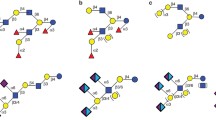
Phenotypic tetracycline resistance (MIC µg/ml ≥ 256) in bifidobacteria harbouring the tet(O) gene has been linked to the presence of a 99-bp gene encoding a putative Cpp-like protein downstream of the tetracycline resistance gene [32]. Sequence comparisons between the genome of B. breve DSM 32583 and nucleotide sequences downstream of the tet(O) gene previously reported for tetracycline-resistant B. longum strains revealed partial homology to these regions (86.3% nucleotide identity over a 150-bp aligned region, on average). This might suggest the presence of the putative and not yet fully characterised cpp2 gene, whose relevance is difficult to ascertain, especially considering the lower MIC levels observed for B. breve DSM 32583, as described below. The proposed genomic architecture of the regions flanking the tet(O) gene homologue in B. breve DSM 32583 is represented in Fig. 2a.
a Schematic representation of putative genomic architecture of the regions flanking the identified tet(O) gene homologue in B. breve DSM 32583. b Diagram depicting presence and absence of homologues of known HMO degradation clusters in B. breve DSM 32583 (blastp, e-value.−50, percentage identify > 50%). HMO clusters were annotated ‘present’ if all cluster components were identified at the above homology level. Incomplete clusters (more than 3 locally clustered genes) were annotated as ‘partially present’
Given the results of the genomic analysis, B. breve DSM 32583 was subjected to antibiotic sensitivity testing according to the recommendations of EFSA using a micro-dilution method, as well as E-test strips, with sensitivity to tetracycline tested by three independent laboratories.
The results of this analysis revealed that B. breve DSM 32583 exhibited antibiotic sensitivities within the recommended EFSA cut-off values (Table 1) and varying results for tetracycline. The MIC values recorded for this antibiotic ranged from 4 to 16 µg/ml between the testing institutions. A discrepancy between the results of genomic and phenotypic analysis was also observed for rifampicin. While genomic screening suggested high protein homology to the known amino acid sequence conveying resistance to rifampicin in Bifidobacterium adolescentis DSM 20083T (92.48% identity), the results of the antimicrobial resistance testing revealed that B. breve DSM 32583 was susceptible to this antibiotic (Supplementary Table 5 and Table 1).
Carbohydrate Fermentation Patterns of B. breve DSM 32583
The carbohydrate fermentation pattern of B. breve DSM 32583 was tested with API Rapid 50 CH fermentation strips by two different laboratories. Overall, profiles were qualitatively similar (Supplementary Table 6), with inter-laboratory variation between data reported for L-arabinose, amygdalin, D-trehalose, gentobiose, and L-fucose.
Members of B. breve, B. longum subsp. infantis, B. longum, and B. pseudocatenulatum have previously been reported to contain genomic clusters required for HMO utilisation [12, 13, 47]. Therefore, we searched for the presence of these clusters in B. breve DSM 32583. Lacto-N-tetraose and lacto-N-neo-tetraose HMO clusters have been well annotated in B. breve UCC2003, lnt cluster (BBR_RS13080-BBR_RS13100), lac cluster (BBR_RS18470-BBR_RS18480), the nah cluster (BBR_RS18490-BBR_RS18520), and lnp/glt cluster (BBR_RS18650-BBR_RS18675) [13], and homologues to all of these clusters were identified in B. breve DSM 32583. This suggests that B. breve DSM 32583 has genomic potential to degrade oligosaccharides present in human milk (Fig. 2b and Supplementary Table 7).
Enzymatic Activity
Enzymatic activity of B. breve DSM 32583 was investigated against 19 different enzymes using the API ZYM assay. Typically, Bifidobacterium strains have high activities of β-galactosidase and α-glucosidase, and moderate-to-high activity of α-galactosidase. Additionally, weak or absent activities of α-mannosidase, and α-fucosidase are characteristic for members of the genus Bifidobacterium, and so are weak protease activities [48, 49].
B. breve DSM 32583 exhibited esterase (C4), esterase lipase (C8), leucine arylamidase, valine arylamidase, cystine arylamidase, acid phosphatase, naftol-AS-BI-phosphohydrolase, α-galactosidase, β-galactosidase, α-glucosidase, β-glucosidase, and N-acetyl-β-glucosaminidase activities among the tested enzymes (Supplementary Table 8).
Resistance to Gastric Acid
The ability of B. breve DSM 32583 to survive conditions simulating those present in the gastrointestinal tract was tested and compared with that of L. fermentum CECT 5716, L. rhamnosus GG, L. johnsonii La1, and L. casei Immunitas. Viability of B. breve DSM 32583 and L. casei Immunitas remained at around 15% on average between the 20 min mark at pH 5 and the 60 min mark at pH 3, while the results recorded for the remaining tested strains in the same time frame showed more variability (Table 2). After 80 min and pH reduction to 2.1, around 7% of B. breve DSM 32583 on average remained viable in the gastric compartment, while the viable counts for L. casei Immunitas dropped to 0.1%. The cumulative survival of B. breve DSM 32583 during passage through the simulated gastric and duodenal compartments (about 52% on average) was comparable to that of L. casei Immunitas and L. fermentum CECT 5716 (about 53% and 51%, respectively), while an overall lower value was recorded for L. rhamnosus GG, L. johnsonii La1 (around 41% on average).
Adhesion Assays to Caco-2 and HT-29 Cells
The adherence of B. breve DSM 32583 was tested and compared with that of selected lactic acid bacteria (Table 3). B. breve DSM 32583 exhibited levels of adherence to HT-29 cells that were comparable to those of L. fermentum CECT 5716 and L. rhamnosus GG, with 862 ± 361, 886 ± 597, 820 ± 350 bacterial cells detected in the assessed microscopic fields, respectively. In contrast, the number of adherent cells recorded for L. casei Immunitas was lower (164 ± 89 cells). Similarly, numbers of bacterial cells that adhered to Caco-2 cells were higher for B. breve DSM 32583, L. fermentum CECT 5716, and L. rhamnosus GG (330 ± 124, 277 ± 185, and 362 ± 139, respectively), while the number of observed adherent L. casei Immunitas cells was considerably lower, with only 17 ± 11 cells recorded. Overall, these results suggest high ability of B. breve DSM 32583 to adhere to HT-29 and Caco-2 cells.
Growth Inhibition of Other Bacterial Species
The potential of probiotics to antagonise potential pathogens is of particular interest; therefore, the ability of B. breve DSM 32583 to inhibit growth of selected strains was evaluated. B. breve DSM 32583 was able to inhibit growth of indicator bacteria (Table 4). In all 17 cases tested, a zone of inhibition with a radius larger than 3 mm was observed after direct contact of B. breve DSM 32583 with the indicator strain cultures, and in 12 out of 17 cases tested, the radius was greater than 6 mm. This inhibition is likely to result from production of organic acids and the decrease in the environmental pH, rather than from the production of compounds with bactericidal properties. This observation could be supported by the results of genomic analysis, which indicated the absence of bacteriocin genes in the genome of B. breve DSM 32583.
Conjugated Linoleic Acid (CLA) and Conjugated Linolenic Acid (CLNA) Production
Lastly, we assessed the potential of B. breve DSM 32583 to produce CLA and CLNA. The concentration of CLA produced by the strain in MRS medium supplemented with L-cysteine (MRS-Cys) reached 171 ± 38 µg/ml, indicating that the minimal conversion rate from the added linoleic acid (LA) was approximately 74%. B. breve DSM 32583 was able to produce different CLA isomers, such as cis 9, trans 11 CLA, trans 10, cis 12 CLA and trans 9, trans 11 CLA (Table 5, Fig. 3a). The conversion rate of linolenic acid (LNA) into CLNA was higher than that of CLA, approximately 99%, with the amount of produced CLNA reaching the concentration of 219 ± 9 µg/ml. Two CLNA isomers (cis 9, trans 11, cis 15 CLNA and trans 9, trans 11, cis 15 CLNA) could be detected in the chromatogram profiles (Table 5, Fig. 3b).
Gas chromatography chromatogram profiles of the fatty acid content present in the culture media obtained from B. breve DSM 32583 grown in MRS broth supplemented with 0.5 mg/ml linoleic (a) and linolenic acid (b). Extensions of the anterior chromatogram indicating the conjugated linoleic (CLA) and linolenic (CLNA) acid isomers are presented in boxed fields
Subsequently, B. breve DSM 32583 was tested for CLA and/or CLNA production in the milk-based medium. (Table 5). CLA production was lower in this medium compared with MRS supplemented with L-cysteine, reaching 75 ± 10 µg/ml at approximately 31% conversion rate, while the CLNA production and conversion rates were comparable to those observed in MRS-Cys medium (Table 5).
Discussion
Human milk and exclusive breastfeeding are the best form of nutrition for infants, providing all essential macro- and micronutrients alongside other health-promoting factors [50]. One of the advantageous properties of human milk is the presence of beneficial bacteria that colonise the infant gut and modulate the developing immune system. Therefore, human milk is nowadays an important source for isolation of probiotic strains [51]. In particular, beneficial properties of members of families Lactobacillaceae and Bifidobacteriaceae are frequently assessed due to their established history of safe use [52]. Each new potentially probiotic strain should be validated by genotypic and phenotypic methods to ensure both its safety and its potential to exert health-promoting effects on the host [53].
In the present work, we describe functional properties of Bifidobacterium breve DSM 32583 isolated from human milk. Previously performed average nucleotide identity analysis identified our strain as a member of the B. breve taxon [22]. Here, additional analysis of relatedness to representative Bifidobacterium strains belonging to several infant-associated species also placed B. breve DSM 32583 within the B. breve cluster, confirming its affiliation to this well-studied species that has previously been associated with a number of health-promoting properties [13, 14, 54].
Genomic analysis of the bacterial mobilome allows to identify potentially harmful mobile elements and becomes a standard in the characterisation of probiotic candidates. Prokaryotic genome plasticity and stability can be impacted by the presence of different mobile elements, such as insertion sequences (IS), transposases, plasmids, and prophages [55]. Transposable elements have been suggested to play a role in environmental adaptation and species diversification through their involvement in chromosomal deletions and/or rearrangements mediated by their ability to catalyse the movement of DNA fragments between different locations in the genome by recognising specific target sequences [56]. For B. breve DSM 32583, all detected ORFs that showed high similarity to known transposases were homologous to IS previously reported for bifidobacteria, in particular the strain B. longum NCC2705 [45].
Prophages have long been known to contribute to horizontal gene transfer events through transduction—a process in which a bacteriophage transfers non-viral DNA from one bacterial cell to another [57]. The B. breve DSM 32583 genome contains no plasmids and no complete prophage sequences, with only three incomplete prophage-associated regions present. Therefore, the probability of horizontal gene transfer in case of B. breve DSM 32583 seems very low.
Genomic resistance to antibiotics has to be assessed for probiotic candidates and does not pose a safety concern, as long as the antibiotic resistance genes are intrinsic and not located on mobile elements like plasmids [58]. The screen for the presence of putative genes associated with antibiotic resistance revealed putative homologues of the rifamycin-resistant beta-subunit of RNA polymerase (rpoB) and the tetracycline resistance gene tet(O). Previous studies have indicated that resistance to rifamycin in bifidobacteria is linked to a number of different mutations observed across the rpoB gene, including regions other than the resistance-conveying sequence ‘hot spots’ [59]. Moreover, the widespread presence of tetracycline resistance genes tet(W/M/O/S) in bifidobacteria is very well documented, with no reports of related adverse observations [60,61,62] and the wide use in the functional food industry, including dairy products and other probiotic formulations [63]. The tet(W) gene seems to be the most prevalent in different bifidobacterial species, as it has been detected at high frequencies in strains belonging to B. longum subsp. longum [61] and B. animalis subsp. lactis [61, 64]. Furthermore, it has been identified in genomes of a number of bifidobacterial strains featured in commercial products with long-term known safety, for example, in B. animalis subsp. lactis BB-12 [65] or B. breve BB02, B. animalis subsp. lactis BL03, and B. animalis subsp. lactis BI04 [66].
Previous literature shows that the tet(W) gene in the genus Bifidobacterium seems to be integrated into the chromosome with its surrounding regions showing strain-specificity and is frequently flanked by transposase target sequences or genes coding for transposases [60, 64]. This suggests that, under appropriate conditions, the gene could be transferred, but this has never been experimentally confirmed [64, 65]. Here, the homologue of the tet(O) gene in B. breve DSM 32583 was not found to be located in the vicinity of putative transposases. Manual sequence searches revealed the presence of a putative homologue of a non-self-mobile protein previously proposed to be transposon-associated (tnpV) upstream of the tet(O) gene. However, it’s function remains unknown [46].
Phenotypic antibiotic sensitivity testing according to the EFSA guidelines revealed antibiotic sensitivities within the recommended EFSA cut-off values. Interestingly, values for tetracycline showed variation ranging from 4 to 16 µg/ml between the testing institutions. While the value of 16 µg/ml is formally above the recommended EFSA cut-off of 8 µg/ml, the results obtained for B. breve DSM 32583 fall within the inter-laboratory variation of MICs previously reported for other commercially used bifidobacteria, such as B. animalis subsp. lactis BB-12 [1], HN019, Bl-04, and B420 [67], B. longum subsp. longum BORI, and B. breve M-16V [1]. Despite the fact that tetracycline-class drugs are broad spectrum antibiotics and have many clinical applications, their utility has been declining in the past 50 years due to an increase in rates of acquired resistance and the availability of other antimicrobials, such as cephalosporins [68]. Nowadays, tetracycline-class antibiotics are mainly used as a second-line treatment option in human medicine due to many side effects, such as ototoxicity [68, 69], but remain among the most frequently used in livestock and poultry worldwide [70].
Many probiotics are known to carry several chromosomally located resistances [71]. Given that bacteraemia caused by probiotics has only been reported in extremely rare cases and susceptible cohorts [72], it has been proposed that probiotic strains should be susceptible to at least two major commonly used antibiotics [73]. Since B. breve DSM 32583 is susceptible to ampicillin, vancomycin, gentamycin, streptomycin, clindamycin, chloramphenicol, and erythromycin, the strain meets this criterion. Moving forward, approaches to reduce antibiotic resistance in probiotic candidates might take advantage of recently proposed genome engineering methods [74, 75]. While current food law regulations and consumer expectations are not yet in line with these pioneering ideas, future dietary product developments may implement such techniques.
Another condition for probiotic strains is the absence of bacterial virulence factors, which have been shown to enable pathogens to replicate and disseminate within a host by negatively modulating or eluding host defences [76]. Screening for virulence and pathogenic potential of B. breve DSM 32583 revealed no virulence genes, as expected for bifidobacteria.
Being saccharolytic, bifidobacteria utilise carbohydrates as their sole source of carbon and energy. As such, they play an important role in shaping the gut environment through their carbohydrate metabolism capabilities [77]. Therefore, we assessed the carbohydrate fermentation pattern of B. breve DSM 32583 at two independent institutions. Overall, the profiles were qualitatively similar, with some inter-laboratory variation, and indicated that B. breve DSM 32583 can metabolise a wide range of carbohydrates, which is consistent with previous reports. Indeed, it has been widely proven that bifidobacteria can metabolise different carbohydrates, including host-derived gastric mucin, and host- and plant-derived oligosaccharides, such as galacto-oligosaccharides and fructo-oligosaccharides or pectin, but the bifidobacterial metabolic capacity for specific carbohydrates is species- and strain-dependent [77].
Bifidobacterial species and strains associated with early life, e.g. B. breve and B. longum subsp. infantis, often contain catabolic genes that specifically target human milk oligosaccharides (HMOs) for degradation and metabolism [78]. The B. breve DSM 32583 genome contains putative homologues to Bifidobacterium gene clusters previously implicated in the degradation of lacto-N-tetraose and lacto-N-neo-tetraose. This suggest the potential of B. breve DSM 32583 to degrade oligosaccharides present in human milk, which will have to be verified in future studies.
An important characteristic of bacterial strains used in probiotic formulations is their ability to survive the passage through the gastrointestinal tract, in particular through the acidic environment of the stomach and the duodenum, where the bile salts are secreted, and retain their optimal functionality [79]. We therefore tested the ability of B. breve DSM 32583 to survive conditions simulating those present in the gastrointestinal tract and compared it with that of L. fermentum CECT 5716, L. rhamnosus GG, L. johnsonii La1, and L. casei Immunitas. B. breve DSM 32583 showed good gastrointestinal survival rates, and we observed strain-specific variability in the viability of bacterial cells, consistent with previous reports. For example, Marteau et al. [38] reported cumulative deliveries of a viable B. bifidum strain, recovered from fermented milk product Ofilus (Yoplait), from the gastric compartment into the duodenal compartment at 67%, while in a study conducted by Venema et al. [80] using a computer-controlled in vitro model of the stomach and small intestine (TIM-1), 5.3% of the viable ingested bifidobacterial dose (B. longum SP07/3 and B. bifidum MF20/5) and 1% of the viable ingested Lactobacillus dose (Lactobacillus gasseri, PA16/8) survived passage through the gastric compartment. Significant variability in the survival of different bifidobacterial strains was previously shown by Zuo et al. [16], who reported gastric juice tolerance of 0.01 and 70.13% for B. bifidum IF3-211 and B. lactis Bb12, respectively.
Probiotic candidates should be metabolically active in the gastrointestinal tract, we therefore assessed the enzymatic activity of B. breve DSM 32583 and its ability to produce health-promoting conjugated acids. The strain exhibited esterase (C4), esterase lipase (C8), leucine arylamidase, valine arylamidase, cystine arylamidase, acid phosphatase, naphthol-AS-BI-phosphohydrolase, α-galactosidase, β-galactosidase, α-glucosidase, β-glucosidase, and N-acetyl-β-glucosaminidase activities. The enzymatic profile of B. breve DSM 32583 was generally similar to those previously reported for members of this species, including B. breve ATCC 15698, B. breve ATCC 15699, B. breve ATCC 15700, B. breve ATCC 15701, and B. breve IDCC4401 [54].
Conjugated linoleic acid (CLA) and conjugated linolenic acid (CLNA) have been reported to exert various physiological benefits to host health, including anti-inflammatory [81, 82], anti-hypertensive [83], and anti-atherosclerotic effects [84]. Since their production by probiotic bacteria has recently become a sought-after characteristics [85], we assessed the potential of B. breve DSM 32583 to produce CLA and CLNA, with the results indicating that our strain is able to produce these compounds in different growth conditions.
Another prerequisite for probiotic characterisation is the adherence to intestinal epithelial cells. Adherence to the intestinal mucosa mediated by a close interaction between bacterial adhesins and the host cell surface is the first step in gut colonisation [86]. Therefore, the ability of a candidate strain to adhere to intestinal cells is another prerequisite for probiotic characterisation [87]. In addition, probiotic bacteria able to form a tight bond with the host epithelial cells may compete with potential pathogens for the same receptors, thus preventing potentially pathogenic microorganisms from colonising the host [20]. In the present work, B. breve DSM 32583 showed good adherence to HT-29 as well as Caco-2, which was comparable to other probiotic strains, namely L. fermentum CECT 5716 and L. rhamnosus GG.
Furthermore, B. breve DSM 32583 inhibited the growth of pathogen strains in vitro. The ability of probiotic bacteria to protect against infectious agents is considered to result from their considerable potential to interact with the host’s immune system and to produce compounds displaying antimicrobial properties, e.g. lactic and acetic acid or bacteriocins [87].
Genotypic and phenotypic characterisation of candidate probiotic strains is the first step in the assessment of their safety and their potential suitability for commercial use. At this early stage of characterisation, we have shown that B. breve DSM 32583 possesses beneficial characteristics required from a potential probiotic candidate. However, while we compared phenotypic traits of B. breve DSM 32583 to those of well-characterised probiotic members of Lactobacillaceae, we have not yet answered questions related to differences in functionality between our strain and other human milk-associated members of the genus Bifidobacterium. This is an important aspect that should be considered in the future, especially given that strain-specific differences have previously been reported for early-life bifidobacteria [16].
Some of the major limitations in probiotic development are associated with the reproducibility of preliminary results in vivo. While the results for bacterial survival and enzymatic activity might be promising in vitro, the use of cell lines and the addition of digestive enzymes do not fully reflect natural conditions in the gastrointestinal tract. Complementary, recently published results strengthen our understanding of the beneficial effects of B. breve DSM 32583 discussed here. We have shown good survival and metabolic activity of the strain in an infant gut model [88] and conducted a small pilot-trial, whose results suggested beneficial effects of the strain on the incidence of respiratory and GI infections in infants [89]. Further assessments in human intervention studies will have to ensure the strain’s suitability and efficacy to infants.
Concluding Remarks
B. breve DSM 32583 described in this report showed several characteristics considered fundamental for probiotic candidates. It survived conditions simulating those present in the digestive tract, was able to adhere to human epithelial cells, and inhibited growth of potentially pathogenic microorganisms. Its antibiotic resistance patterns were comparable to those of known probiotic strains, and its genome did not contain plasmids nor any virulence genes. These findings suggest that B. breve DSM 32583 may potentially be suitable for commercial applications.
Data Availability
No datasets were generated or analysed during the current study.
References
Kim MJ, Ku S, Kim SY, Lee HH, Jin H, Kang S, Li R, Johnston TV, Park MS, Ji GE (2018) Safety Evaluations of Bifidobacterium bifidum BGN4 and Bifidobacterium longum BORI. Int J Mol Sci 19(5):1422
Day RL, Harper AJ, Woods RM, Davies OG, Heaney LM (2019) Probiotics: current landscape and future horizons. Future Sci OA 5(4):FSO391
Hill C, Guarner F, Reid G, Gibson GR, Merenstein DJ, Pot B, Morelli L, Canani RB, Flint HJ, Salminen S, Calder PC, Sanders ME (2014) The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat Rev Gastro Hepat 11(8):506–514
Tojo R, Suarez A, Clemente MG, de los Reyes-Gavilan CG, Margolles A, Gueimonde M, Ruas-Madiedo P (2014) Intestinal microbiota in health and disease: role of bifidobacteria in gut homeostasis. World J Gastroenterol 20(41):15163–15176
LeBlanc JG, Milani C, de Giori GS, Sesma F, van Sinderen D, Ventura M (2013) Bacteria as vitamin suppliers to their host: a gut microbiota perspective. Curr Opin Biotechnol 24(2):160–168
O’Neill I, Schofield Z, Hall LJ (2017) Exploring the role of the microbiota member Bifidobacterium in modulating immune-linked diseases. Emerg Top Life Sci 1(4):333-349. https://doi.org/10.1042/ETLS20170058
Tanaka H, Doesburg K, Iwasaki T, Mierau I (1999) Screening of lactic acid bacteria for bile salt hydrolase activity. J Dairy Sci 82(12):2530–2535
Roger LC, Costabile A, Holland DT, Hoyles L, McCartney AL (2010) Examination of faecal Bifidobacterium populations in breast- and formula-fed infants during the first 18 months of life. Microbiol-Sgm 156:3329–3341
Shao Y, Forster SC, Tsaliki E, Vervier K, Strang A, Simpson N, Kumar N, Stares MD, Rodger A, Brocklehurst P, Field N, Lawley TD (2019) Stunted microbiota and opportunistic pathogen colonization in caesarean-section birth. Nature 574(7776):117–121
Jost T, Lacroix C, Braegger CP, Rochat F, Chassard C (2014) Vertical mother-neonate transfer of maternal gut bacteria via breastfeeding. Environ Microbiol 16(9):2891–2904
Milani C, Mancabelli L, Lugli GA, Duranti S, Turroni F, Ferrario C, Mangifesta M, Viappiani A, Ferretti P, Gorfer V, Tett A, Segata N, van Sinderen D, Ventura M (2015) Exploring vertical transmission of bifidobacteria from mother to child. Appl Environ Microbiol 81(20):7078–7087
Sela DA, Chapman J, Adeuya A, Kim JH, Chen F, Whitehead TR, Lapidus A, Rokhsar DS, Lebrilla CB, German JB, Price NP, Richardson PM, Mills DA (2008) The genome sequence of Bifidobacterium longum subsp infantis reveals adaptations for milk utilization within the infant microbiome, P Natl Acad Sci USA 105(48):18964-18969
James K, Motherway MO, Bottacini F, van Sinderen D (2016) Bifidobacterium breve UCC2003 metabolises the human milk oligosaccharides lacto-N-tetraose and lacto-N-neo-tetraose through overlapping, yet distinct pathways. Sci Rep 6:38560
Maldonado J, Gil-Campos M, Maldonado-Lobón JA, Benavides MR, Flores-Rojas K, Jaldo R, Jiménez Del Barco I, Bolívar V, Valero AD, Prados E, Peñalver I, Olivares M (2019) Evaluation of the safety, tolerance and efficacy of 1-year consumption of infant formula supplemented with Lactobacillus fermentum CECT5716 Lc40 or Bifidobacterium breve CECT7263: a randomized controlled trial. BMC Pediatr 19(1):361
Heppner N, Reitmeier S, Heddes M, Merino MV, Schwartz L, Dietrich A, List M, Gigl M, Meng C, van der Veen DR, Schirmer M, Kleigrewe K, Omer H, Kiessling S, Haller D (2024) Diurnal rhythmicity of infant fecal microbiota and metabolites: a randomized controlled interventional trial with infant formula. Cell Host Microbe 32(4):573-587.e5
Zuo F, Yu R, Feng X, Chen L, Zeng Z, Khaskheli GB, Ma H, Chen S (2016) Characterization and in vitro properties of potential probiotic Bifidobacterium strains isolated from breast-fed infant feces. Ann Microbiol 66(3):1027–1037
Tuomola E, Crittenden R, Playne M, Isolauri E, Salminen S (2001) Quality assurance criteria for probiotic bacteria. Am J Clin Nutr 73(2 Suppl):393S-398S
EFSA (2018) Guidance on the characterisation of microorganisms used as feed additives or as production organisms. EFSA J 16(3):e05206
Ouwehand AC, Salminen S, Isolauri E (2002) Probiotics: an overview of beneficial effects. Antonie Van Leeuwenhoek 82(1–4):279–289
Styriak I, Nemcova R, Chang YH, Ljungh A (2003) Binding of extracellular matrix molecules by probiotic bacteria. Lett Appl Microbiol 37(4):329–333
EFSA (2012) Guidance on the assessment of bacterial susceptibility to antimicrobials of human and veterinary importance. EFSA J 10(6). https://doi.org/10.2903/j.efsa.2012.2740
Kujawska M, Schaubeck M, Hall LJ, Neuhaus K (2023) Draft genome sequence of Bifidobacterium breve DSM 32583, isolated from human milk. Microbiol Resour Announc 12:e0041223
Katz LS, Griswold T, Morrison SS, Caravas JA, Zhang S, den Bakker HC, Deng X, Carleton HA (2019) Mashtree: a rapid comparison of whole genome sequence files. J Open Source Softw 4(44):1762
Siguier P, Varani A, Perochon J, Chandler M (2012) Exploring bacterial insertion sequences with ISfinder: objectives, uses, and future developments. Methods Mol Biol 859:91–103
Arndt D, Grant JR, Marcu A, Sajed T, Pon A, Liang Y, Wishart DS (2016) PHASTER: a better, faster version of the PHAST phage search tool. Nucleic Acids Res 44(W1):W16–W21
Carattoli A, Zankari E, Garcìa-Fernandez A, Larsen MV, Lund O, Villa L, Aarestrup FM, Hasman H (2014) PlasmidFinder and pMLST: in silico detection and typing of plasmids. Antimicrob Agents Chemother. https://doi.org/10.1128/AAC.02412-14
Joensen KG, Scheutz F, Lund O, Hasman H, Kaas RS, Nielsen EM, Aarestrup FM (2014) Real-time whole-genome sequencing for routine typing, surveillance, and outbreak detection of verotoxigenic Escherichia coli. J Clin Microbiol 52(5):1501–1510
Yoon SH, Park Y-K, Kim JF (2015) PAIDB v2. 0: exploration and analysis of pathogenicity and resistance islands. Nucleic Acids Res 43(D1):D624-D630
Alcock BP, Raphenya AR, Lau TTY, Tsang KK, Bouchard M, Edalatmand A, Huynh W, Nguyen AV, Cheng AA, Liu S, Min SY, Miroshnichenko A, Tran HK, Werfalli RE, Nasir JA, Oloni M, Speicher DJ, Florescu A, Singh B, Faltyn M, Hernandez-Koutoucheva A, Sharma AN, Bordeleau E, Pawlowski AC, Zubyk HL, Dooley D, Griffiths E, Maguire F, Winsor GL, Beiko RG, Brinkman FSL, Hsiao WWL, Domselaar GV, McArthur AG (2020) CARD 2020: antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res 48(D1):D517–D525
Sayers EW, Beck J, Brister JR, Bolton EE, Canese K, Comeau DC, Funk K, Ketter A, Kim S, Kimchi A, Kitts PA, Kuznetsov A, Lathrop S, Lu Z, McGarvey K, Madden TL, Murphy TD, O’Leary N, Phan L, Schneider VA, Thibaud-Nissen F, Trawick BW, Pruitt KD, Ostell J (2020) Database resources of the National Center for Biotechnology Information. Nucleic Acids Res 48(D1):D9–D16
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinformatics 10:421
Wang N, Hang X, Zhang M, Liu X, Yang H (2017) Analysis of newly detected tetracycline resistance genes and their flanking sequences in human intestinal bifidobacteria. Sci Rep 7(1):6267
Rozewicki J, Li S, Amada KM, Standley DM, Katoh K (2019) MAFFT-DASH: integrated protein sequence and structural alignment. Nucleic Acids Res 47(W1):W5–W10
Hammami R, Zouhir A, Le Lay C, Hamida JB, Fliss I (2010) BACTIBASE second release: a database and tool platform for bacteriocin characterization. BMC Microbiol 10(1):1–5
van Heel AJ, de Jong A, Song C, Viel JH, Kok J, Kuipers OP (2018) BAGEL4: a user-friendly web server to thoroughly mine RiPPs and bacteriocins. Nucleic Acids Res 46(W1):W278–W281
Coico R (2005) Gram staining. Curr Protoc Microbiol Appendix 3. Appendix 3C. https://doi.org/10.1002/9780471729259.mca03cs00
Scardovi V (1986) Genus Bifidobacterium Orla-Jensen 1924. In: Sneath PHA, Mair NS, Sharpe ME, Holt JG (eds) Bergey’smanual of systematic bacteriology. Williams and Wilkins, Baltimore, pp 1418–1434
Marteau P, Minekus M, Havenaar R, Huis in't Veld JH (1997) Survival of lactic acid bacteria in a dynamic model of the stomach and small intestine: validation and the effects of bile. J Dairy Sci 80(6):1031–1037
Conway P, Gorbach S, Goldin B (1987) Survival of lactic acid bacteria in the human stomach and adhesion to intestinal cells. J Dairy Sci 70(1):1–12
Coconnier MH, Klaenhammer TR, Kerneis S, Bernet MF, Servin AL (1992) Protein-mediated adhesion of Lactobacillus acidophilus BG2FO4 on human enterocyte and mucus-secreting cell lines in culture. Appl Environ Microbiol 58(6):2034–2039
Magnusson J, Schnurer J (2001) Lactobacillus coryniformis subsp. coryniformis strain Si3 produces a broad-spectrum proteinaceous antifungal compound. Appl Environ Microbiol 67(1):1–5
Martin R, Olivares M, Marin ML, Xaus J, Fernandez L, Rodriguez JM (2005) Characterization of a reuterin-producing Lactobacillus coryniformis strain isolated from a goat’s milk cheese. Int J Food Microbiol 104(3):267–277
Iverson SJ, Lang SL, Cooper MH (2001) Comparison of the Bligh and Dyer and Folch methods for total lipid determination in a broad range of marine tissue. Lipids 36(11):1283–1287
Rodríguez-Alcalá LM, Braga T, Malcata FX, Gomes A, Fontecha J (2011) Quantitative and qualitative determination of CLA produced by Bifidobacterium and lactic acid bacteria by combining spectrophotometric and Ag+-HPLC techniques. Food Chem 125:1373–1378
Schell MA, Karmirantzou M, Snel B, Vilanova D, Berger B, Pessi G, Zwahlen MC, Desiere F, Bork P, Delley M, Pridmore RD, Arigoni F (2022) The genome sequence of Bifidobacterium longum reflects its adaptation to the human gastrointestinal tract. P Natl Acad Sci USA 99(22):14422-14427
Adams V, Lyras D, Farrow KA, Rood JI (2002) The clostridial mobilisable transposons. Cell Mol Life Sci 59(12):2033–2043
Garrido D, Ruiz-Moyano S, Lemay DG, Sela DA, German JB, Mills DA (2015) Comparative transcriptomics reveals key differences in the response to milk oligosaccharides of infant gut-associated bifidobacteria. Sci Rep 5:13517
Delgado S, O’Sullivan E, Fitzgerald G, Mayo B (2008) In vitro evaluation of the probiotic properties of human intestinal Bifidobacterium species and selection of new probiotic candidates. J Appl Microbiol 104(4):1119–1127
Arboleya S, Ruas-Madiedo P, Margolles A, Solis G, Salminen S, de los Reyes-Gavilan CG, Gueimonde M (2011) Characterization and in vitro properties of potentially probiotic Bifidobacterium strains isolated from breast-milk. Int J Food Microbiol 149(1):28–36
Habicht JP (2004) Expert consultation on the optimal duration of exclusive breastfeeding: the process, recommendations, and challenges for the future. Adv Exp Med Biol 554:79–87
Damaceno QS, Gallotti B, Reis IMM, Totte YCP, Assis GB, Figueiredo HC, Silva TF, Azevedo V, Nicoli JR, Martins FS (2023) Isolation and identification of potential probiotic bacteria from human milk. Probiotics Antimicrob Prot 15(3):491–501
Lyons KE, Ryan CA, Dempsey EM, Ross RP, Stanton C (2020) Breast milk, a source of beneficial microbes and associated benefits for infant health. Nutrients 12(4):1039
da Silva TF, Glória RA, Americo MF, Freitas ADS, de Jesus LCL, Barroso FAL, Laguna JG, Coelho-Rocha ND, Tavares LM, le Loir Y, Jan G, Guédon É, Azevedo VAC (2024) Unlocking the potential of probiotics: a comprehensive review on research, production, and regulation of probiotics. Probiotics Antimicrob Proteins. https://doi.org/10.1007/s12602-024-10247-x
Choi IY, Kim J, Kim SH, Ban OH, Yang J, Park MK (2021) Safety evaluation of Bifidobacterium breve IDCC4401 isolated from infant feces for use as a commercial probiotic. J Microbiol Biotechnol 31(7):949–955
Siefert JL (2009) Defining the mobilome. Methods Mol Biol 532:13–27
Serrato-Capuchina A, Matute DR (2018) The role of transposable elements in speciation. Genes (Basel) 9(5):254
Borodovich T, Shkoporov AN, Ross RP, Hill C (2022) Phage-mediated horizontal gene transfer and its implications for the human gut microbiome. Gastroenterol Rep (Oxf) 10:goac012
Gueimonde M, Sánchez B, de los Reyes-Gavilán CG, Margolles A (2013) Antibiotic resistance in probiotic bacteria. Front Microbiol 4. https://doi.org/10.3389/fmicb.2013.00202
Lokesh D, Parkesh R, Kammara R (2018) Bifidobacterium adolescentis is intrinsically resistant to antitubercular drugs. Sci Rep 8(1):11897
Kazimierczak KA, Flint HJ, Scott KP (2006) Comparative analysis of sequences flanking tet(W) resistance genes in multiple species of gut bacteria. Antimicrob Agents Chemother 50(8):2632–2639
Aires J, Doucet-Populaire F, Butel MJ (2007) Tetracycline resistance mediated by tet(W), tet(M), and tet(O) genes of Bifidobacterium isolates from humans. Appl Environ Microbiol 73(8):2751–2754
Gueimonde M, Sánchez B, de los Reyes-Gavilán CG, Margolles A (2013) Antibiotic resistance in probiotic bacteria. Front Microbiol 4:202
Masco L, Huys G, De Brandt E, Temmerman R, Swings J (2005) Culture-dependent and culture-independent qualitative analysis of probiotic products claimed to contain bifidobacteria. Int J Food Microbiol 102(2):221–230
Gueimonde M, Florez AB, van Hoek AH, Stuer-Lauridsen B, Stroman P, de los Reyes-Gavilan CG, Margolles A (2010) Genetic basis of tetracycline resistance in Bifidobacterium animalis subsp. lactis. Appl Environ Microbiol 76(10):3364–3369
Polit A, Yang H, Amund D (2018) Investigating the transmissibility of tet(W) in bifidobacteria exposed to acid and bile stress. Biosci Microbiota Food Health 37(2):39–43
Douillard FP, Mora D, Eijlander RT, Wels M, de Vos WM (2018) Comparative genomic analysis of the multispecies probiotic-marketed product VSL#3. PLoS ONE 13(2):e0192452
FDA (2013) GRN No. 445 Bifidobacterium animalis subsp. lactis strains HN019, Bi-07, Bl-04 and B420. https://www.cfsanappsexternal.fda.gov/scripts/fdcc/?set=GRASNotices
LaPlante KL, Dhand A, Wright K, Lauterio M (2022) Re-establishing the utility of tetracycline-class antibiotics for current challenges with antibiotic resistance. Ann Med 54(1):1686–1700
Bisht M, Bist SS (2011) Ototoxicity: the hidden menace. Indian J Otolaryngol Head Neck Surg 63(3):255–259
Granados-Chinchilla F, Rodriguez C (2017) Tetracyclines in food and feedingstuffs: from regulation to analytical methods, bacterial resistance, and environmental and health implications. J Anal Methods Chem 2017:1315497
Toth AG, Judge MF, Nagy SA, Papp M, Solymosi N (2023) A survey on antimicrobial resistance genes of frequently used probiotic bacteria, 1901 to 2022. Euro Surveill 28(14). https://doi.org/10.2807/1560-7917.ES.2023.28.14.2200272
Acuna-Gonzalez A, Kujawska M, Youssif M, Atkinson T, Grundy S, Hutchison A, Tremlett C, Clarke P, Hall LJ (2023) Bifidobacterium bacteraemia is rare with routine probiotics use in preterm infants: a further case report with literature review. Anaerobe 80:102713
Borriello SP, Hammes WP, Holzapfel W, Marteau P, Schrezenmeir J, Vaara M, Valtonen V (2003) Safety of probiotics that contain lactobacilli or bifidobacteria. Clin Infect Dis 36(6):775–780
Zuo F, Chen S, Marcotte H (2020) Engineer probiotic bifidobacteria for food and biomedical applications - current status and future prospective. Biotechnol Adv 45:107654
Zuo F, Zeng Z, Hammarström L, Marcotte H (2019) Inducible plasmid self-destruction (IPSD) assisted genome engineering in lactobacilli and bifidobacteria. ACS Synth Biol 8(8):1723–1729
Cross AS (2008) What is a virulence factor? Crit Care 12(6):196
Pokusaeva K, Fitzgerald GF, van Sinderen D (2011) Carbohydrate metabolism in bifidobacteria. Genes Nutr 6(3):285–306
Lordan C, Roche AK, Delsing D, Nauta A, Groeneveld A, MacSharry J, Cotter PD, van Sinderen D (2024) Linking human milk oligosaccharide metabolism and early life gut microbiota: bifidobacteria and beyond. Microbiol Mol Biol Rev 88(1):e0009423
Collado MC, Sanz Y (2007) Induction of acid resistance in Bifidobacterium: a mechanism for improving desirable traits of potentially probiotic strains. J Appl Microbiol 103(4):1147–1157
Venema K, Verhoeven J, Verbruggen S, Espinosa L, Courau S (2019) Probiotic survival during a multi-layered tablet development as tested in a dynamic, computer-controlled in vitro model of the stomach and small intestine (TIM-1). Lett Appl Microbiol 69(5):325–332
Moraes C, Oliveira CA, Amaral ME, Landini GA, Catisti R (2017) Liver metabolic changes induced by conjugated linoleic acid in calorie-restricted rats. Arch Endocrinol Metab 61(1):45–53
Boussetta T, Raad H, Letteron P, Gougerot-Pocidalo MA, Marie JC, Driss F, El-Benna J (2009) Punicic acid a conjugated linolenic acid inhibits TNFalpha-induced neutrophil hyperactivation and protects from experimental colon inflammation in rats. PLoS ONE 4(7):e6458
Zhao WS, Zhai JJ, Wang YH, Xie PS, Yin XJ, Li LX, Cheng KL (2009) Conjugated linoleic acid supplementation enhances antihypertensive effect of ramipril in Chinese patients with obesity-related hypertension. Am J Hypertens 22(6):680–686
Zlatanos SN, Laskaridis K, Sagredos A (2008) Conjugated linoleic acid content of human plasma. Lipids Health Dis 7:34
Mei Y, Chen H, Yang B, Zhao J, Zhang H, Chen W (2022) Research progress on conjugated linoleic acid bio-conversion in Bifidobacterium. Int J Food Microbiol 369:109593
Westerlund B, Korhonen TK (1993) Bacterial proteins binding to the mammalian extracellular matrix. Mol Microbiol 9(4):687–694
Toscano M, De Vecchi E, Gabrieli A, Zucotti GV, Drago L (2015) Probiotic characteristics and in vitro compatibility of a combination of Bifidobacterium breve M-16 V, Bifidobacterium longum subsp. infantis M-63 and Bifidobacterium longum subsp. longum BB536. Ann Microbiol 65:1079–1086
Klaassens ES, Baak ML, Mekkes NJ, Bongoni R, Schaubeck M (2024) Effect of protein modification in synbiotic infant formula on probiotic metabolic activity and bacterial composition in an infant gut-model. Microbiome Res Rep 3:38. https://doi.org/10.20517/mrr.2024.13
Alba C, Carrera M, Álvarez-Calatayud G, Arroyo R, Fernández L, Rodríguez JM (2024) Evaluation of safety and beneficial health effects of the human-milk strain Bifidobacterium breve DSM32583: an infant pilot trial. Nutrients 16(8):1134
Acknowledgements
We are very grateful to all mothers for donating milk for isolation of probiotic strains. We also want to thank Juan Miguel Rodriguez Gomez for his advice and scientific support.
Author information
Authors and Affiliations
Contributions
MK performed bioinformatics analyses except for mobilome and virulence factors analysis, which was performed by KN with the help of CH. SA carried out in vitro analyses. MK and KN interpreted the data. EJ contributed and analysed in vitro data. MK wrote the paper and visualised the data. All authors have read and approved the final manuscript as submitted and agreed to be accountable for all aspects of the work. MS and LJH coordinated and supervised the project and downstream data analysis, drafted and reviewed initial drafts and also finalised the final draft prior to submission.
Corresponding author
Ethics declarations
Competing Interests
Financial interests: Lindsay Hall and Klaus Neuhaus report financial support provided by HiPP GmbH und Co Vertrieb KG. Monika Schaubeck reports a relationship with HiPP GmbH und Co Vertrieb KG that includes employment. Monika Schaubeck has patent pending to HiPP & Co. Esther Jiménez is employee of Probisearch, SLU, a company that received funding from HiPP GmbH & Co. Vertrieb KG, Pfaffenhofen, Germany, to perform phenotypic characterisation experiments and data analysis. If there are other authors, they declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kujawska, M., Neuhaus, K., Huptas, C. et al. Exploring the Potential Probiotic Properties of Bifidobacterium breve DSM 32583—A Novel Strain Isolated from Human Milk. Probiotics & Antimicro. Prot. (2024). https://doi.org/10.1007/s12602-024-10346-9
Accepted:
Published:
DOI: https://doi.org/10.1007/s12602-024-10346-9