Abstract
Background
ZFPM2-AS1, as an oncogenic lncRNA, plays an essential role in the progression of several tumors. However, the prognostic significance, biological function, and molecular mechanism of ZFPM2-AS1 in most tumors have not been fully elucidated.
Methods
We analyzed differentially expressed immune-related lncRNAs (IRlncRNAs) and clustered gastric adenocarcinoma (GAC) samples based on these lncRNAs expression. Then, WGCNA and survival analysis were performed to determine key IRlncRNA (ZFPM2-AS1) in GAC. The comprehensive analysis was performed to evaluate the association between ZFPM2-AS1 expression and survival, tumor microenvironment (TME), immune-related factors, and related signal pathways in pan-cancers. Furthermore, we constructed a co-expression network of ZFPM2-AS1, and NUP107 and C8orf76 were identified as target mRNAs. We further evaluated the role of NUP107 and C8orf76 in the GAC microenvironment. More importantly, real-time polymerase chain reaction (qRT-PCR) was employed to validate ZFPM2-AS1, NUP107 and C8orf76 expression.
Results
ZFPM2-AS1 was remarkably overexpressed and correlated with poor overall survival in most tumors. Further analysis showed that ZFPM2-AS1 was related to various immune cells infiltrated in the microenvironment of most tumors. GSEA revealed that ZFPM2-AS1 in GAC was primarily involved in immune-related pathways. Furthermore, NUP107 and C8orf76 were identified as potential target mRNAs of ZFPM2-AS1, which was related to infiltrating immune cells in the GAC microenvironment. qRT-PCR verified that ZFPM2-AS, NUP107 and C8orf76 were highly expressed in gastric cancer cells.
Conclusion
ZFPM2-AS1 could be a potential biomarker for cancer prognosis, and a promising immune target for cancer therapy. Furthermore, ZFPM2-AS1 might play an immunosuppressive role in the GAC microenvironment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Long non-coding RNAs (lncRNAs) are a heterogeneous class of RNA molecules with longer than 200 nucleotides, which locate within inter-genic region in the genome [1, 2]. LncRNAs are numerous and diverse; according to most estimates, the number of human lncRNA is much higher than that of protein-coding genes [3]. Based on the subcellular localization, lncRNAs can be divided into cytoplasmic lncRNAs and nuclear lncRNAs. Cytoplasmic lncRNAs play a crucial role in post-transcriptional regulation, such as crosstalking with mRNAs via sponging miRNAs. Nuclear lncRNAs mainly participate in the modulation of epigenetic events, transcription or mRNA processing [4]. With the advancement of RNA sequencing technologies and computational prediction techniques, the total number of lncRNAs is continuously increasing [5, 6]. Their growing ranks have attracted the attention of the cancer research community. In the past few decades, accumulating evidence has elucidated the crucial function of lncRNAs in modulating various biological processes of cancer, such as differentiation, apoptosis, DNA damage regulation, and immune response [7,8,9]. LncRNA ZFPM2-AS1, located on chromosome 8q23, has been reported to participate in the occurrence and development of several cancers. For example, ZFPM2-AS1 contributes to lung adenocarcinoma cell growth and epithelial to mesenchymal transition [10]. ZFPM2-AS1 also facilitates hepatocellular carcinoma (HCC) cell cycle progression, cell proliferation, and invasion via miR-653/GOLM1 [11]. In addition, ZFPM2-AS1 promotes the growth, migration, and invasion of thyroid cancer cells through the miR-515-5p/TUSC3 axis [12]. However, the expression pattern, biological function, molecular mechanism, and prognostic significance of ZFPM2-AS1 in most tumor types have not been fully elucidated.
The tumor microenvironment (TME) refers to the physiological and biochemical elements around tumor cells, including the extracellular matrix, the tumor vascular system, stromal cells, immune cells, and the acidic and hypoxic environment [13,14,15]. A growing body of studies has suggested that TME has profound impacts on tumor development and therapeutic responses. For instance, differences of immune cells infiltrated in the TME, including tumor-associated macrophages (TAMs), dendritic cells, and T lymphocytes, have been found in tumor patients [16, 17]. Furthermore, CD8+ T cells, TAMs, and CD4+ T cells in the TME relate to clinical outcomes of multiply tumors, including lung cancer, urothelial cancer, melanoma, and gastric cancer [18,19,20]. Recently, lncRNAs have been studied for their promising role in the regulation of TME. LncRNAs contribute to controlling and mediating the interaction of tumor cells and infiltrating immune cells in the TME, as well as essential mechanisms of immune response [21]. Therefore, the correlation between ZFPM2-AS1 and immune cells infiltrated in the TME is helpful to understand the function of ZFPM2-AS1 in the development of tumors.
Chemotherapy is the first-line therapeutic plan for advanced GC patients, which can significantly improve their life quality and overall survival rate [4]. The widely used chemotherapeutic drugs primarily include platinum drugs, 5-fluorouracil (5-FU), epirubicin, docetaxel, and so on [22]. However, the development of drug resistance becomes a major challenge in clinic oncology, resulting in tumor recurrence and metastasis. Although the detailed mechanism of drug resistance is still inconclusive, lncRNAs have received increased attention to improving the effectiveness of anticancer therapy [23]. Recently, accumulating studies have revealed that lncRNAs are involved in the chemoresistance of GC via various mechanisms, including the induction of autophagy, inhibition of apoptosis, modulation of drug resistance-related genes and signaling pathways, and regulation of cancer stem cells [24]. Therefore, exploring the effect of ZFPM2-AS1 on drug resistance is helpful to enhance therapy efficiency of GC patients. In this study, the non-negative matrix factorization (NMF) algorithm was applied to cluster gastric adenocarcinoma (GAC) samples, and Kaplan–Meier analysis, ESTIMATE, and CIBERSORT algorithms were used to estimate the survival and immune microenvironment differences between clusters. Then, a key immune-related lncRNA (ZFPM2-AS1) was identified associated with clusters and prognosis of GAC patients via Weighted Gene Co-Expression Network Analysis (WGCNA) and survival analysis. We further analyzed the differential expression, clinical prognosis, TME, tumor mutational burden (TMB), microsatellite instability status (MSI), drug sensitivity, and related signal pathways of ZFPM2-AS1 in 33 tumor types. Furthermore, we further constructed a ZFPM2-AS1-mRNA co-expression network and performed differential expression, survival, TMB, MSI, TME, and immune subtype analysis of target mRNAs (NUP107 and C8orf76). Finally, several experiments were carried out to verify the results of bioinformatics analysis. Our study provides a theoretical basis for therapeutic targets and novel mechanisms of ZFPM2-AS1 in GAC and pan-cancers.
2 Materials and methods
2.1 Data collection and differentially expressed immune-related lncRNAs (DE-IRlncRNAs) analysis
The flow diagram of the study is shown in Fig. 1. The lncRNA data and clinical information of 339 GAC and 30 normal samples were extracted from the TCGA. Subsequently, the list of 2498 immune-related genes (IRGs) was obtained from the ImmPort database. To identify the relationship between IRGs and corresponding lncRNAs expresion, Pearson correlation analysis was carried out in the R software. The immune-related lncRNAs (IRlncRNAs) were determined according to the correlation coefficient (|R| > 0.4 and P < 0.001). Furthermore, we identified differentially expressed lncRNAs (DElncRNAs) in GAC samples using the R package “limma”. |Log2FC| > 1 and FDR < 0.05 were regarded as the threshold. Finally, the overlapping lncRNAs between IRlncRNAs and DElncRNAs were determined as DE-IRlncRNAs for further analysis.
2.2 Identification of molecular subtypes and assessment of immune microenvironment
The NMF algorithm is a powerful technique for identifying distinct molecular subtypes and evaluating the optimal number of clusters based on gene expression microarrays. According to the expression of DE-IRlncRNAs, 339 GAC samples were clustered by the NMF algorithm. The optimal number of clusters and the average profile width of the common member matrix were identified using the R package “NMF”. Then, the Kaplan–Meier method was utilized to perform overall survival (OS) in different molecular subtypes of the patients. Furthermore, the ESTIMATE and CIBERSORT algorithms were applied to identify the immune microenvironment differences in different molecular subtypes.
2.3 WGCNA and key DE-IRlncRNAs identification
WGCNA is a system biology method to analyze gene expression patterns of multiple samples. It can divide genes with similar expression patterns into the same modules and identify the association between specific traits and modules. To find the module associated with clinical characteristics and clusters, WGCNA was performed using the R package “WGCNA” based on the expression of DE-IRlncRNAs. Then, we selected DE-IRlncRNAs in the most significant module (the grey module) for survival analysis, and ZFPM2-AS1 was determined as the key lncRNA for pan-cancer analysis.
2.4 Differential expression analysis and survival analysis of ZFPM2-AS1 in pan-cancers
To further explore the role of ZFPM2-AS1 in pan-cancers, we used the UCSC Xena database to download the transcription expression data, clinical information, stemness score, and immune subtype data of 33 TCGA GDC tumors. We obtained the gene symbol of transcription expression data from the Ensembl and extracted ZFPM2-AS1 expression levels of 33 cancer types. Then, the comparison of ZFPM2-AS1 between tumor and normal tissues was performed in 18 cancer types (BLCA, BRCA, COAD, CHOL, GBM, ESCA, KIRP, KICH, HNSC, KIRC, LIHC, LUSC, GAC, LUAD, PRAD, THCA, READ, and UCEC), which had more than five associated adjacent noncancerous tissues. Furthermore, based on the median expression level of ZFPM2-AS1, 33 cancer samples were classified into low-expression and high-expression groups. To identify the role of ZFPM2-AS1 in the cancer patients’ survival, the Kaplan–Meier method was utilized to perform OS, progression-free interval (PFI), and disease-specific survival (DSS) analyses in 33 cancer types. P < 0.05 was regarded as the threshold.
2.5 Tumor immune microenvironment (TIME), TMB, and MSI analysis
The ESTIMATE algorithm was applied to estimate the stromal score and immune score of tumor samples. We examined the correlation between the expression of ZFPM2-AS1 and the stromal/immune score using the Spearman rank correlation coefficient (P < 0.05). Subsequently, the CIBERSORT algorithm was utilized to evaluate the composition ratio of infiltrating immune cells in the TME. We performed correlation analysis to investigate the relationship between ZFPM2-AS1 expression and the proportion of immune cells (P < 0.05).
TMB, which refers to the number of somatic mutations contained in a tumor cell, is a promising biomarker for immunotherapy response in patients with tumors. MSI, a predictor to guide immunotherapy, represents a hypermutator phenotype in tumors caused by impaired DNA mismatch repair. We further assessed the relationship between ZFPM2-AS1 expression and TMB, and MSI to investigate whether ZFPM2-AS1 could be a promising target for tumor immunotherapy (P < 0.05).
2.6 Drug sensitivity analysis and GSEA analysis
We downloaded gene expression and drug sensitivity data of cancer samples from the CellMiner database. We filtered drug sensitivity data with FDA standard certification and clinical laboratory verification. Then, we identified the correlation between drug sensitivity and ZFPM2-AS1 expression. Besides, to explore the role of ZFPM2-AS1 in pan-cancers, GSEA was carried out for the KEGG pathway using the R package “clusterprofiler”. P < 0.05 was regarded as the cutoff.
2.7 The construction of ZFPM2-AS1 co-expression network in GAC
To explore potential mechanisms of ZFPM2-AS1 in GAC, target mRNAs of ZFPM2-AS1 were identified based on correlation coefficient |R| > 0.3 and P < 0.001. We further visualized the ZFPM2-AS1-mRNA co-expression network using the Cytoscape software. In addition, we performed the differential analysis and survival analysis on target mRNAs of ZFPM2-AS1 in GAC. NUP107 and C8orf76 were selected for differential expression analysis, survival analysis, TMB, and MSI analysis in pan-cancers.
2.8 TME and immune subtype analysis of NUP107 and C8orf76 in GAC
We evaluated the association between NUP107 and C8orf76 expression and the stromal/immune score of the TME (P < 0.05). Then, we intersected transcription expression data with stemness score (DNA methylation-based) (DNAss) and stemness score (RNA expression-based) (RNAss). The relationship between NUP107 and C8orf76 expression and DNAss and RNAss of GAC was identified using the Spearman correlation test. Furthermore, six immune subtypes (C1, wound healing; C2, INF-r dominant; C3, inflammatory; C4, lymphocyte depleted; C5, immunological quiet; and C6, TGF-β dominant) were obtained from the UCSC Xena database. We performed the immune subtype analysis of NUP107 and C8orf76 in GAC using R language loaded with packages “limma,” “ggplot2,” and “reshape2”.
2.9 Cell culture, transfection and real-time polymerase chain reaction (qRT-PCR)
The normal gastric cell line (GES-1) and GC cell lines (AGS, MKN-28, HGC-27, and MKN-45) were obtained from ATCC (Manassas, VA, USA). GES-1 cells were cultured in DMEM medium (Gibco). MKN-28, HGC-27, and MKN-45 were grown in RPMI-1640 medium (Gibco). AGS were cultured in F-12 K medium (Gibco). All mediums were supplemented with 10% fetal bovine serum (Gibco), and 10 mg/mL streptomycin, and 100 U/mL penicillin. The plasmids of lncRNA ZFPM2-AS1 and negative control were purchased from GeneCreate (Wuhan, China). Then, they were transfected into AGS via lipofectamine 2000 (Invitrogen, USA). After 48 h, the transfected cell line was applied for further experiments, and the transfection efficiency was evaluated by qRT-PCR.
Total RNAs in cell lines were extracted using TRIzol (Invitrogen) based on the manufacturer’s protocol. Then, PrimeScript RT Reagent Kit was utilized to synthesize cDNA (Thermo Scientific). qRT-PCR was carried out using SYBR Green Master Mix (Thermo Scientific). The 2−ΔΔCt method was utilized to calculate mRNA expression levels, and GAPDH was used as an internal control. The primer sequences were as follows: ZFPM2-AS1 forward primer: CAATGGGACTAAGCCAGGCA; reverse primer: GGGCTCCACCAACAACCATA; C8orf76 forward primer: 5′-TTATACGAACCAGGCTTCTGC-3′; reverse primer: 5′-GCCAACACACTTCACCTCTG-3′; NUP107 forward primer: 5′-GTCTGTGACACCTGGGAAGA-3′; reverse primer: 5′-CTTGATTCTCTTCCAGAACTCTCT-3′; GAPDH forward primer: 5′-CTCGCTTCGGCAGCACA-3′; reverse primer: 5′-AACGCTTCACGAATTTGCGT-3′.
2.10 Cell counting kit-8 (CCK-8) assay
The proliferative capacity of GC cells was measured using CCK-8 assay. Transfected AGS cells were seeded into 96-well plates at a density of 3 × 104 cells/mL. After incubation for 24 h, 48 h, 72 h, and 96 h, 10 μL CCK-8 (Vazyme, Nanjing, China) was added to each well and absorbance was measured at 450 nm. CCK8 assay also performed to detect the cisplatin sensitivity of GC cells. Transfected AGS cells were inoculated onto 96-well plates overnight and then different concentrations of cisplatin (1, 2, 3, 4, 5, 6, 7, 8 μM) were added to each well for continued culture. After 48 h, cells were incubated with 10 μL CCK-8 for 1 h before the detection of absorbance.
2.11 Statistical analyses
Statistical analyses were performed using Perl (version 5.30.1), GraphPad Prism (version 7.0), and R (version 4.0.2). P < 0.05 was defined as statistically significant.
3 Results
3.1 DE-IRlncRNAs-based molecular subtypes and TIME analysis
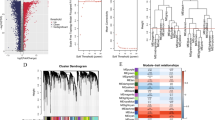
The list of 2498 IRGs was obtained from the ImmPort database, and 1035 IRlncRNAs were identified according to the Pearson correlation analysis (|R| > 0.3 and P < 0.001). Among them, 406 IRlncRNAs were differentially expressed between the GAC and normal tissues (|Log2FC| > 1 and FDR < 0.05) (Fig. 2). Then, these 406 DE-IRlncRNAs were used for NMF cluster analysis, and the optimal number of clusters was determined as 2 via the comprehensive correlation coefficient (Fig. 3a and b). All GAC patients were divided into two clusters, including Cluster 1 (n = 141) and Cluster 2 (n = 198). The Kaplan–Meier curve showed that Cluster 1 patients had a worse OS rate than Cluster 2 (P = 0.017) (Fig. 3c). We further investigated the differences in the immune microenvironment between the two clusters. As shown in Fig. 3d, the immune, estimate, and stromal scores were significantly elevated in Cluster 1 compared with Cluster 2. Furthermore, CIBERSORT revealed that the abundance of T cells CD8, T cells CD4 memory activated, Dendritic cells resting, and B cells naive were significantly different in Cluster 1 and Cluster 2 (P < 0.05) (Fig. 3e).
Identification of DE-IRlncRNAs-based molecular subtypes. a The cophenetic, rss, and silhouette distributions when the number of clusters was set as 2–10. b Consensus map of NMF clustering. c The survival curves of 2 molecular subtypes. d Comparisons of immune, estimate, and stroma scores between 2 molecular subtypes. e Comparisons of infiltrating immune cells between 2 molecular subtypes
3.2 WGCNA analysis and identification of key DE-IRlncRNAs
To determine the crucial modules related to GAC, WGCNA was carried out based on the expression data of 406 DE-IRlncRNAs. The results of WGCNA revealed that the grey module had the highest correlation with Cluster (r = − 0.3, P = 2e−08) and grade (r = 0.25, P = 3e−06) (Fig. S1). Then, the Kaplan–Meier method was utilized to evaluate the prognostic value of 125 DE-IRlncRNAs in the grey module. The results showed that 12 DE-IRlncRNAs (AP001107.5, AL365181.3, TNFRSF10A-AS1, AP000695.2, A2M-AS1, AL023803.1, AP000695.1, AC136475.3, AL590666.2, AP003071.4, AC005586.1, and ZFPM2-AS1) in the grey module were associated with the survival of GAC patients (Table 1 and Fig. S2). Finally, ZFPM2-AS1 with the most significant prognostic value was selected for pan-cancer analysis.
3.3 ZFPM2-AS1 expression analysis and survival analysis in pan-cancers
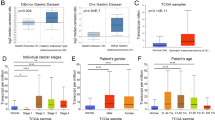
We assessed the lncRNA levels of ZFPM2-AS1 in 33 cancer types from TCGA. As shown in Fig. 4a and b, ZFPM2-AS1 levels were significantly increased in BLCA, BRCA, COAD, ESCA, KICH, HNSC, KIRP, KIRC, LIHC, LUSC, GAC, LUAD, THCA, and READ. Furthermore, to evaluate the impact of ZFPM2-AS1 in cancer patients’ survival, we divided all samples into high-expression and low-expression groups and compared the OS, DSS, and PFI rates of the 33 cancer types. The Kaplan–Meier curves revealed that ZFPM2-AS1 high expression had a worse OS rate than those with ZFPM2-AS1 low expression in KIRC, LGG, LIHC, MESO, SARC, GAC, UCEC, and UVM (P < 0.05) (Fig. 4c). Higher expression levels of ZFPM2-AS1 also indicated worse DSS in BRCA, KIRC, LGG, LIHC, MESO, SARC, GAC, and UCEC, and worse PFI in BLCA, COAD, LGG, SARC, GAC, THYM, UCEC, UVM (P < 0.05) (Fig. 5a and b).
3.4 The correlation of ZFPM2-AS1 expression with TIME
We obtained the immune and stromal scores of all tumor samples and assessed the correlation between ZFPM2-AS1 expression and both scores of the TME. We visualized the results at P < 0.001. As shown in Fig. 6, the high expression of ZFPM2-AS1 was negatively correlated with the stromal score in BLCA but positively associated with COAD, BRCA, DLBC, KIRC, GBM, KICH, READ, LGG, PRAD, TGCT, PCPG, UCEC, and THCA. The expression of ZFPM2-AS was positively related to the immune score in KICH, GBM, LIHC, KIRC, LGG, PRAD, PCPG, and READ, while negatively associated with MESO, HNSC, GAC, LUSC, SKCM, and TGCT.
We further analyzed the relationship between the level of 22 infiltrating immune cells and ZFPM2-AS1 expression. We visualized the results at P < 0.001. The result showed that highly expressed ZFPM2-AS1 was negatively related to B cells naïve in BLCA but positively related to Dendritic cells resting. In BRCA, ZFPM2-AS1 expression was negatively associated with Macrophages M1, T cells CD8, and T cells follicular helper, but positively associated with Mast cells resting and T cells CD4 memory resting. In COAD, the expression of ZFPM2-AS1 was negatively associated with T cells CD8 and NK cells activated but positively associated with Neutrophils. In GBM, the level of ZFPM2-AS expression was positively correlated with Mast cells activated but negatively correlated with Mast cells resting. In HNSC, the expression of ZFPM2-AS1 was negatively associated with B cells naïve, Mast cells resting, T cells CD8 and T cells regulatory (Tregs), but positively associated with Dendritic cells activated, Dendritic cells resting, Macrophages M0, Mast cells activated, and Neutrophils. Besides, high level of ZFPM2-AS1 was positively related to Monocytes in KIRP and Neutrophils in LGG but negatively related to NK cells resting in LIHC and T cells CD8 in LUSC. In SKCM, ZFPM2-AS expression was positively correlated with Macrophages M0, Mast cells activated, T cells CD4 memory resting, T cells gamma delta, but negatively correlated with B cells naïve, Plasma cells, T cells CD8, T cells follicular helper, and Tregs. In TGCT, highly expressed ZFPM2-AS1 was positively associated with Macrophages M2 and Tregs but negatively associated with B cells naïve and T cells CD4 memory activated. Furthermore, the expression of ZFPM2-AS1 had a significant positive relationship with T cells follicular helper in GAC and Macrophages M0 in SARC, while had a significant negative relationship with NK cells activated in PCPG (Fig. S3).
3.5 Immune-related factors, TMB, and MSI, drug sensitivity and GSEA analysis of ZFPM2-AS1 in pan-cancers
To explore the immune-related role of ZFPM2-AS in tumors, we analyzed the relationship between ZFPM2-AS1 expression and immune-related factors. Heatmaps showed that ZFPM2-AS1 was correlated with several known immune checkpoints, including CTLA4, PDCD1 (PD-1), CD274 (PD-L1), TIGIT, LAG-3, HAVCR2 (TIM-3), and IDO1, in most tumors (Fig. 7a). We further evaluated the correlation of ZFPM2-AS1 expression with TMB and MSI. As shown in Fig. 7b, ZFPM2-AS1 expression was correlated with TMB in COAD, BRCA, LGG, BLCA, LIHC, SKCM, ACC, PAAD, UCEC, GAC, and SARC. ZFPM2-AS1 expression was correlated with MSI in COAD, UCEC, KICH, UVM, GAC, LUAD, and SKCM (Fig. 7c).
The relationship between ZFPM2-AS1 expression and immune-related factors, TMB, and MSI in pan-cancers. a The relationship between ZFPM2-AS1 expression and immune-related factors. b The relationship between ZFPM2-AS1 expression and TMB. c The relationship between ZFPM2-AS1 expression and MSI. *P < 0.05, **P < 0.01, ***P < 0.001
Then, we explored the association between ZFPM2-AS1 and drug sensitivity. The result revealed that ZFPM2-AS1 expression had a positive relationship with the sensitivity of cobimetinib, and trametinib. The expression of ZFPM2-AS1 had a negative relationship with the sensitivity of idarubicin, cyclophosphamide, 5-FU, and oxaliplatin (Fig. 8). Furthermore, to explore the role of ZFPM2-AS1 in various types of cancer, GSEA was performed according to the expression of ZFPM2-AS1. We found that ZFPM2-AS1 in pan-cancer tissues was mainly involved in antigen processing and presentation, cytokine-cytokine receptor interaction, focal adhesion, and cytosolic DNA sensing pathway. Importantly, we found ZFPM2-AS1 in GAC primarily involved in immune-related pathways, including antigen processing and presentation, Toll-like receptor signaling pathway, and RIG-I-like receptor signaling (Fig. S4).
3.6 Construction of ZFPM2-AS1 co-expression network
To investigate the underlying mechanisms of ZFPM2-AS1 in GAC, we constructed a ZFPM2-AS1-mRNA co-expression network. As shown in Fig. 9a, a total of 48 genes were positively correlated with ZFPM2-AS1. Among them, NUP107 and C8orf76 were remarkably upregulated in GC samples (Fig. 9b). Similar results were observed in the pairing analysis between GC tissues and adjacent noncancerous tissues from the same patient (Fig. S5). Furthermore, the survival analysis showed that GC patients with NUP107 low expression had a higher survival rate than those with NUP107 high expression. GC patients with C8orf76 low expression also had a higher survival rate than those with C8orf76 high expression (Fig. S6).
3.7 Differential expression analysis, survival analysis, TMB, and MSI analysis of NUP107 and C8orf76 in pan-cancers
We further evaluated the difference of NUP107 and C8orf76 expression in the other 32 types of tumors profiled in TCGA. The result revealed that NUP107 was significantly differentially expressed in 15 cancer types. Upregulated expression of NUP107 was observed in 13 types of cancer (BLCA, BRCA, CHOL, COAD, ESCA, GBM, HNSC, KIRC, KIRP, LIHC, LUAD, LUSC, PRAD, READ, and UCEC), and downregulated expression of NUP107 was observed in two types of cancer (KICH and THCA) (Fig. 10a). The expression of C8orf76 was remarkably upregulated in 16 types of cancer (BLCA, BRCA, CHOL, COAD, ESCA, GBM, HNSC, KICH, KIRC, KIRP, LIHC, LUAD, LUSC, PRAD, READ, and UCEC) (Fig. 10a).
Expression analysis, survival analysis, TMB, and MSI analysis of NUP107 and C8orf76 in pan-cancers. a Expression level of NUP107 and C8orf76 in pan-cancers. *P < 0.05, **P < 0.01, ***P < 0.001. b The survival curve of NUP107 and C8orf76 for OS in pan-cancers. c The correlation between NUP107 and C8orf76 expression and TMB, and MSI in pan-cancers
Subsequently, to probe the effect of NUP107 and C8orf76 in cancer patients’ survival, we divided all samples into high-expression and low-expression groups and compared the OS rate of the other 32 cancer types. As shown in Fig. 10b, the high expression of NUP107 had a worse OS rate than the low expression of NUP107 in KICH, KIRP, LGG, LIHC, SARC, MESO, and THYM. The high expression of C8orf76 had a worse OS rate than the low expression of C8orf76 in BRCA, KICH, KIRC, LGG, LIHC, LUAD, MESO, THYM, and UVM. Furthermore, TMB and MSI analyses were carried out to investigate whether NUP107 and C8orf76 could be promising immunotherapy targets. As shown in Fig. 10c, NUP107 and C8orf76 expression was related to TMB in GAC, THYM, LUSC, BLCA, LUAD, PRAD, BRCA, KICH, PAAD, and LGG. NUP107 and C8orf76 expression was correlated with TMB in UCEC, READ, GAC, and TGCT.
3.8 The correlation of NUP107 and C8orf76 expression with TME and immune subtype of GAC
We performed correlation analysis to determine the association between NUP107 and C8orf76 expression and DNAss, RNAss, stromal score, and immune score of GAC. As shown in Fig. 11a, NUP107 expression was positively associated with RNAss of GAC, while C8orf76 expression was significantly associated with RNAss and DNAss of GAC. Figure 11a also showed that the expression of NUP107 and C8orf76 had a negative relationship with the stromal score and immune score of GAC. We also conducted immune subtype analysis to investigate the relationship between NUP107 and C8orf76 expression and immune infiltrate types in the TME of GAC. The result revealed that the expression of NUP107 and C8orf76 was significantly different in the immune subtype (P < 0.001) (Fig. 11b).
Correlation analysis of NUP107 and C8orf76 expression and GAC microenvironment, stem cell scores, and immune subtype. a The correlation between NUP107 and C8orf76 expression and RNAss, DNAss, immune score, estimate score, and stroma score. b The correlation between NUP107 and C8orf76 expression and immune subtype. *P < 0.05, **P < 0.01, ***P < 0.001
3.9 Validation of ZFPM2-AS1, NUP107 and C8orf76 by qRT-PCR
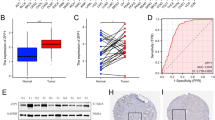
We performed qRT-PCR in GC cells to validate ZFPM2-AS1, NUP107 and C8orf76 expression levels. The results showed that ZFPM2-AS1, NUP107 and C8orf76 expression were significantly highly expressed in GC cells compared with normal gastric cells (Fig. 12a–c). The expression levels of NUP107 and C8orf76 were consistent with the results of our bioinformatics analysis. Furthermore, we detected NUP107 and C8orf76 expression in AGS cells after transfecting the overexpression plasmid of ZFPM2-AS1. The result of qRT-PCR showed that ZFPM2-AS1 positively regulated NUP107 and C8orf76 expression in AGS cells (Fig. 12d–f). These results suggested that NUP107 and C8orf76 might be potential targets of ZFPM2-AS1. Collectively, these findings further demonstrated that the bioinformatics analysis was reliable.
The expression of ZFPM2-AS1, NUP107, and C8orf76 in GC cells. a qRT-PCR validation of ZFPM2-AS1 expression in GC cells. b qRT-PCR validation of NUP107 expression in GC cells. c qRT-PCR validation of C8orf76 expression in GC cells. d ZFPM2-AS1 expression in AGS cells after transfecting ZFPM2-AS1 plasmid. e NUP107 expression in AGS cells after overexpressing ZFPM2-AS1. f C8orf76 expression in AGS cells after overexpressing ZFPM2-AS1. g The proliferation capacity of AGS cells after ZFPM2-AS1 overexpression. h The sensitivity to cisplatin in AGS cells after ZFPM2-AS1 overexpression. *P < 0.05, **P < 0.01, ***P < 0.001
3.10 The effect of ZFPM2-AS1 on the proliferation and cisplatin sensitivity of GC cells
To explore the role of ZFPM2-AS1 in GC cell proliferation and cisplatin sensitivity, ZFPM2-AS1 overexpression plasmid was transfected into AGS cells. Then, the result of CCK-8 assay revealed that highly expressed ZFPM2-AS1 significantly promoted the proliferation of AGS cells (Fig. 12g). Furthermore, we investigated whether cisplatin sensitivity of AGS cells was regulated by ZFPM2-AS1. The results indicated that overexpression of ZFPM2-AS1 reduced the sensitivity to cisplatin in AGS cells. Our finding suggested that high expression levels of ZFPM2-AS1 may be resistant to cisplatin treatment (Fig. 12h).
4 Discussion
LncRNAs are multifunctional molecules that play indispensable roles in the pathogenesis of numerous human genetic diseases, including cancer. In the past few decades, various lncRNAs have been found to be abnormally expressed in different tumor cells. Their role in modulating tumors also has been studied to varying degrees [25, 26]. ZFPM2-AS1, located in 8q23.1, plays an oncogenic role in several tumors. For example, Han et al. detected that ZFPM2-AS1 is highly expressed in lung adenocarcinoma (LUAD) cells. Mechanistically, ZFPM2-AS1 destabilizes ZFPM2 via interacting with UPF1 to facilitate proliferation, and invasion of LUAD cells [10]. Yan et al. also found that ZFPM2-AS1 is significantly increased in small cell lung cancer (SCLC) cells and tissues. Downregulation of ZFPM2‑AS1 attenuates the migration, proliferation, and invasion viability of SCLC cells via the miR-3612/TRAF4 axis [27]. Based on the data from qRT-PCR analysis and TCGA database, ZFPM2-AS1 is recognized as an oncogenic lncRNA in HCC. High levels of ZFPM2-AS1 promote HCC cell cycle progression, cell proliferation, and invasion through miR-653/GOLM1 [11]. Furthemore, a recent study indicated that ZFPM2-AS1 promote thyroid cancer progression via sponging miR-515-5p to modulate TUSC3 expression [12]. However, the underlying molecular mechanisms of dysregulated ZFPM2-AS1 in most tumors remain unclear. In this study, we applied the NMF algorithm to identify a new cluster of GAC, which was correlated with survival and had significant differences in the TIME. Then, ZFPM2-AS1 was identified associated with clusters and prognosis of GAC patients via Weighted Gene Co-Expression Network Analysis (WGCNA) and survival analysis. Furthermore, we first revealed that ZFPM2-AS1 was upregulated in 14 types of cancer, suggesting that ZFPM2-AS1 might be an oncogenic lncRNA in pan-cancers. According to Kaplan–Meier survival curves, ZFPM2-AS1 was also found to be associated with the survival of patients in most tumor types. These results implied that ZFPM2-AS1 might be a promising biomarker for cancer diagnosis and prognosis.
Recently, TME is emerging as a crucial factor in multiple stages of cancer progression, specifically distant metastasis, immune escape, and local resistance, thus affecting the development of frontline interventions in clinical oncology [28]. Importantly, immune cells, as indispensable non-tumor cells in the TME, are considered to be of great value in the diagnosis, prognosis, and treatment of tumors [17, 29, 30]. Our results showed that the level of ZFPM2-AS expression was significantly associated with multiply infiltrating immune cells in the microenvironment of most tumors. Based on these results, we collected 47 common immune-related genes further to explore the immune-related role of ZFPM2-AS in tumors. PDCD1 (PD-1)/CD274 (PD-L1) and CTLA-4 are the hottest immune checkpoints in the current studies. PD-1 is an inhibitory cell surface receptor, which can recognize and bind to PD- L1. The binding of PD-1 to its ligands PD-L1 inhibits the activation of T cells and interrupts the immune surveillance of the TME, leading to tumor immune escape [31, 32]. According to our findings, the expression of ZFPMA-AS1 was highly correlated with PD-1 and PD-L1 in several tumor types. As an inhibitory receptor expressed mainly by T cells, CTLA-4 functions to impair the activation of T cells and is overexpressed upon T cell activation [33]. CTLA-4 is also constitutively expressed on regulatory T cells, which facilitates immune suppression in the TME [34, 35]. Here, we found that the expression of ZFPMA-AS1 was remarkably associated with CTLA-4 in most type of tumors. Furthermore, TIGIT, LAG-3, HAVCR2 (TIM-3), and IDO1 are known as novel immune checkpoint targets that can suppress the activation of effector T lymphocytes, hence mediating a tumor immune escape mechanism [32, 36, 37]. In our study, these novel immune checkpoints were also highly correlated with ZFPMA-AS1expression in most tumors. These findings further implied that ZFPMA-AS1 might affect the progression of cancer by regulating immune-active status in the TME and could be a promising immune target for cancer therapy.
Chemotherapy can improve the survival of patients with locally advanced metastatic or unresectable GC, which plays a vital role in the comprehensive treatment of tumors [22]. However, the development of chemoresistance becomes a major challenge, resulting in the failure of clinical treatment. In recent studies, lncRNAs play an essential role in regulating the drug sensitivity of GC cells. For example, Zhang et al. discovered that CRNDE is associated with the chemosensitivity of GC patients. Mechanistically, CRNDE reduces chemoresistance of GC cells to 5-FU through SRSF6-modulated selective splicing of PICALM [38]. Wu et al. found that FOXD1-AS1 induces cisplatin resistance in GC via activating PIK3CA/PI3K/AKT/mTOR signaling [39]. In addition, Lin et al. revealed that upregulated LINC00200 enhances the chemoresistance of GC cells to oxaliplatin via the E2F1/RAD51 axis [40]. To investigate the indicator function of ZFPMA-AS1 in anticancer drug selection, we analyzed the relationship between anticancer drug sensitivity and ZFPMA-AS1 expression. We found that the expression of ZFPMA-AS was significantly related to the first-line chemotherapeutic drugs, such as 5-FU and oxaliplatin. Our study provided a novel insight for exploring the treatment of tumors and avoiding the resistance of tumors. Furthermore, GSEA further determined that ZFPM2-AS1 in GAC is primarily involved in immune-related pathways, including antigen processing and presentation, Toll-like receptor signaling pathway, and RIG-I-like receptor signaling.
To further explore potential mechanisms of ZFPM2-AS1 in GAC, We constructed a co-expression network to identify target mRNAs. NUP107 and C8orf76 were identified as target mRNAs. NUP107, located on the nuclear rim, is a crucial component of the nuclear pore complex [41, 42]. Recently, NUP107 was reported to participate in the progression of several cancers. For example, Shi et al. demonstrated that the expression of NUP107 is significantly upregulated in cervical cancer samples and cell lines. Besides, overexpression of NUP107 promotes the sensitivity of cervical cancer cells to oxidative insults [43]. Alanee et al. discovered that a single nucleotide variant in NUP107 might be predictive of chemosensitivity in ovarian cancer patients [44]. Based on the bioinformatics analysis, NUP107 is identified as a pivotal gene involved in pancreatic ductal adenocarcinoma (PDAC), which may be a molecular marker for the early diagnosis [45]. Recently, Nong et al. discovered that NUP107, as abridge genes of hepatitis B virus infection-related HCC, is significantly associated with low survival rates of patients [46]. C8orf76, located on chromosome 8q24.13, is a nuclear protein-coding gene [47]. Data from the Human Protein Atlas reveal that upregulated C8orf76 is correlated with the poor prognosis in endometrial cancer, breast cancer, and renal cancer (https://www.proteinatlas.org/ENSG00000189376-C8orf76/pathology). Recent studies reported C8orf76 as a tumor-promoting factor that promotes cell proliferation and metastasis of GC cells. Mechanistic evidence revealed that C8orf76 directly bind to DUSP5P1 promoter region to activate its transcription, therefore inducing the MAPK/ERK signaling pathway [48]. In our study, NUP107 and C8orf76 were found to be overexpressed in GAC and significantly associated with the survival of GAC patients. These findings further implied that NUP107 and C8orf76 might be involved in the development and prognosis of GAC.
To further identify the role of NUP107 and C8orf76 in the TME of GAC, we explore the relationship between NUP107 and C8orf76 expression and DNAss, RNAss, stromal score, and immune score of GAC. The results revealed that NUP107 and C8orf76 expression was significantly associated with immune cells in the GAC microenvironment. Immune type analysis also proved that NUP107 and C8orf76 expression was significantly different in the immune subtype. More importantly, qRT-PCR revealed that NUP107 and C8orf76 expression levels were consistent with the results of bioinformatics analysis. Based on these results, we speculated that ZFPM2-AS1 might play an immunosuppressive role in the microenvironment of GAC by regulating the expression of NUP107 and C8orf76. However, more experiments are needed to confirm our prediction.
5 Conclusions
In conclusion, we applied the NMF algorithm to identify a new cluster of GAC, which was correlated with survival and had significant differences in the TIME. Then, ZFPM2-AS1 was identified associated with clusters and prognosis of GAC patients via WGCNA and survival analysis. We further performed a comprehensive analysis of ZFPM2-AS1, revealing its promising role as an indicator of cancer patients’ survival. Meanwhile, ZFPMA-AS1 was found to be involved in regulating the TIME and associated with anticancer drug sensitivity. We speculated that ZFPMA-AS1 might affect the progression of cancer by regulating immune-active status in the TME and could be an immune target for cancer therapy. Furthermore, we constructed a co-expression network and identified potential target mRNAs (NUP107 and C8orf76) of ZFPM2-AS1 in GAC. More interestingly, NUP107 and C8orf76 might play a significant role in the TIME of GAC and could be a potential biomarker for cancer prognosis. Further experiments verified that ZFPM2-AS1, NUP107 and C8orf76 are highly expressed in GC cells and high levels of ZFPM2-AS1 promote the proliferation of GC cells and reduce the sensitivity to cisplatin.
Data availability
All data in our study were available from the TCGA database (http://cancergenome.nih.gov/).
Abbreviations
- LncRNAs:
-
Long non-coding RNAs
- NMF:
-
Non-negative matrix factorization
- TME:
-
Tumor microenvironment
- TAMs:
-
Tumor-associated macrophages
- GAC:
-
Gastric adenocarcinoma
- WGCNA:
-
Weighted gene co-expression network analysis
- TMB:
-
Tumor mutational burden
- MSI:
-
Microsatellite instability status
- DE-IRlncRNAs:
-
Differentially expressed immune-related lncRNAs
- IRGs:
-
Immune-related genes
- IRlncRNAs:
-
Immune-related lncRNAs
- DElncRNAs:
-
Differentially expressed lncRNAs
- 5-FU:
-
5-Fluorouracil
- CCK-8:
-
Cell counting kit-8
- BLCA:
-
Bladder urothelial carcinoma
- BRCA:
-
Breast invasive carcinoma
- COAD:
-
Colon adenocarcinoma
- CHOL:
-
Colon adenocarcinoma
- GBM:
-
Glioblastoma multiforme
- ESCA:
-
Esophageal carcinoma
- KIRP:
-
Kidney renal papillary cell carcinoma
- KICH:
-
Kidney chromophobe
- HNSC:
-
Head and neck squamous cell carcinoma
- KIRC:
-
Kidney renal clear cell carcinoma
- LIHC:
-
Liver hepatocellular carcinoma
- LUSC:
-
Lung squamous cell carcinoma
- LUAD:
-
Lung adenocarcinoma
- PRAD:
-
Prostate adenocarcinoma
- THCA:
-
Thyroid carcinoma
- READ:
-
Rectum adenocarcinoma
- UCEC:
-
Uterine corpus endometrial carcinoma
- ACC:
-
Adrenocortical carcinoma
- CESC:
-
Cervical squamous cell carcinoma and endocervical adenocarcinoma
- ESCA:
-
Esophageal carcinoma
- DLBC:
-
Lymphoid neoplasm diffuse large B-cell lymphoma
- LAML:
-
Acute myeloid leukemia
- LGG:
-
Brain lower grade glioma
- PCPG:
-
Pheochromocytoma and paraganglioma
- MESO:
-
Mesothelioma
- PAAD:
-
Pancreatic adenocarcinoma
- OV:
-
Ovarian serous cystadenocarcinoma
- SARC:
-
Sarcoma
- UVM:
-
Uveal melanoma
- SKCM:
-
Skin cutaneous melanoma
- TGCT:
-
Testicular germ cell tumors
- THYM:
-
Thymoma
- UCS:
-
Uterine carcinosarcoma
- OS:
-
Overall survival
- PFI:
-
Progression-free interval
- DSS:
-
Disease-specific survival
- TIME:
-
Tumor immune microenvironment
References
Kopp F, Mendell JT. Functional classification and experimental dissection of long noncoding RNAs. Cell. 2018;172(3):393–407. https://doi.org/10.1016/j.cell.2018.01.011.
Chen L, He M, Zhang M, Sun Q, Zeng S, Zhao H, Yang H, Liu M, Ren S, Meng X, et al. The Role of non-coding RNAs in colorectal cancer, with a focus on its autophagy. Pharmacol Ther. 2021;226:107868. https://doi.org/10.1016/j.pharmthera.2021.107868.
Djebali S, Davis CA, Merkel A, Dobin A, Lassmann T, Mortazavi A, Tanzer A, Lagarde J, Lin W, Schlesinger F, et al. Landscape of transcription in human cells. Nature. 2012;489(7414):101–8. https://doi.org/10.1038/nature11233.
Yuan L, Xu ZY, Ruan SM, Mo S, Qin JJ, Cheng XD. Long non-coding RNAs towards precision medicine in gastric cancer: early diagnosis, treatment, and drug resistance. Mol Cancer. 2020;19(1):96. https://doi.org/10.1186/s12943-020-01219-0.
Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP, et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature. 2009;458(7235):223–7. https://doi.org/10.1038/nature07672.
Clark MB, Mercer TR, Bussotti G, Leonardi T, Haynes KR, Crawford J, Brunck ME, Cao KA, Thomas GP, Chen WY, et al. Quantitative gene profiling of long noncoding RNAs with targeted RNA sequencing. Nat Methods. 2015;12(4):339–42. https://doi.org/10.1038/nmeth.3321.
Huarte M. The emerging role of lncRNAs in cancer. Nat Med. 2015;21(11):1253–61. https://doi.org/10.1038/nm.3981.
Hu Q, Egranov SD, Lin C, Yang L. Long noncoding RNA loss in immune suppression in cancer. Pharmacol Ther. 2020;213:107591. https://doi.org/10.1016/j.pharmthera.2020.107591.
Lin C, Yang L. Long noncoding RNA in cancer: wiring signaling circuitry. Trends Cell Biol. 2018;28(4):287–301. https://doi.org/10.1016/j.tcb.2017.11.008.
Han S, Cao D, Sha J, Zhu X, Chen D. LncRNA ZFPM2-AS1 promotes lung adenocarcinoma progression by interacting with UPF1 to destabilize ZFPM2. Mol Oncol. 2020;14(5):1074–88. https://doi.org/10.1002/1878-0261.12631.
Zhang XW, Li QH, Xu ZD, Dou JJ. STAT1-induced regulation of lncRNA ZFPM2-AS1 predicts poor prognosis and contributes to hepatocellular carcinoma progression via the miR-653/GOLM1 axis. Cell Death Dis. 2021;12(1):31. https://doi.org/10.1038/s41419-020-03300-4.
Ren R, Du Y, Niu X, Zang R. ZFPM2-AS1 transcriptionally mediated by STAT1 regulates thyroid cancer cell growth, migration and invasion via miR-515–5p/TUSC3. J Cancer. 2021;12(11):3393–406. https://doi.org/10.7150/jca.51437.
Singh SR, Rameshwar P, Siegel P. Targeting tumor microenvironment in cancer therapy. Cancer Lett. 2016;380(1):203–4. https://doi.org/10.1016/j.canlet.2016.04.009.
Chen Q, Liu G, Liu S, Su H, Wang Y, Li J, Luo C. Remodeling the tumor microenvironment with emerging nanotherapeutics. Trends Pharmacol Sci. 2018;39(1):59–74. https://doi.org/10.1016/j.tips.2017.10.009.
Balkwill FR, Capasso M, Hagemann T. The tumor microenvironment at a glance. J Cell Sci. 2012;125(Pt 23):5591–6. https://doi.org/10.1242/jcs.116392.
Mantovani A, Marchesi F, Malesci A, Laghi L, Allavena P. Tumour-associated macrophages as treatment targets in oncology. Nat Rev Clin Oncol. 2017;14(7):399–416. https://doi.org/10.1038/nrclinonc.2016.217.
Fridman WH, Zitvogel L, Sautes-Fridman C, Kroemer G. The immune contexture in cancer prognosis and treatment. Nat Rev Clin Oncol. 2017;14(12):717–34. https://doi.org/10.1038/nrclinonc.2017.101.
Turley SJ, Cremasco V, Astarita JL. Immunological hallmarks of stromal cells in the tumour microenvironment. Nat Rev Immunol. 2015;15(11):669–82. https://doi.org/10.1038/nri3902.
Nishino M, Ramaiya NH, Hatabu H, Hodi FS. Monitoring immune-checkpoint blockade: response evaluation and biomarker development. Nat Rev Clin Oncol. 2017;14(11):655–68. https://doi.org/10.1038/nrclinonc.2017.88.
Mariathasan S, Turley SJ, Nickles D, Castiglioni A, Yuen K, Wang Y, Kadel EE III, Koeppen H, Astarita JL, Cubas R, et al. TGFbeta attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature. 2018;554(7693):544–8. https://doi.org/10.1038/nature25501.
Zhou Y, Zhu Y, Xie Y, Ma X. The role of long non-coding RNAs in immunotherapy resistance. Front Oncol. 2019;9:1292. https://doi.org/10.3389/fonc.2019.01292.
Smyth EC, Nilsson M, Grabsch HI, van Grieken NC, Lordick F. Gastric cancer. Lancet. 2020;396(10251):635–48. https://doi.org/10.1016/S0140-6736(20)31288-5.
Jiang W, Xia J, Xie S, Zou R, Pan S, Wang ZW, Assaraf YG, Zhu X. Long non-coding RNAs as a determinant of cancer drug resistance: towards the overcoming of chemoresistance via modulation of lncRNAs. Drug Resist Updat. 2020;50:100683. https://doi.org/10.1016/j.drup.2020.100683.
Liu Y, Ao X, Wang Y, Li X, Wang J. Long non-coding RNA in gastric cancer: mechanisms and clinical implications for drug resistance. Front Oncol. 2022;12:841411. https://doi.org/10.3389/fonc.2022.841411.
Iyer MK, Niknafs YS, Malik R, Singhal U, Sahu A, Hosono Y, Barrette TR, Prensner JR, Evans JR, Zhao S, et al. The landscape of long noncoding RNAs in the human transcriptome. Nat Genet. 2015;47(3):199–208. https://doi.org/10.1038/ng.3192.
Goodall GJ, Wickramasinghe VO. RNA in cancer. Nat Rev Cancer. 2021;21(1):22–36. https://doi.org/10.1038/s41568-020-00306-0.
Yan Z, Yang Q, Xue M, Wang S, Hong W, Gao X. YY1-induced lncRNA ZFPM2-AS1 facilitates cell proliferation and invasion in small cell lung cancer via upregulating of TRAF4. Cancer Cell Int. 2020;20:108. https://doi.org/10.1186/s12935-020-1157-7.
Chen F, Zhuang X, Lin L, Yu P, Wang Y, Shi Y, Hu G, Sun Y. New horizons in tumor microenvironment biology: challenges and opportunities. BMC Med. 2015;13:45. https://doi.org/10.1186/s12916-015-0278-7.
Zeng D, Zhou R, Yu Y, Luo Y, Zhang J, Sun H, Bin J, Liao Y, Rao J, Zhang Y, et al. Gene expression profiles for a prognostic immunoscore in gastric cancer. Br J Surg. 2018;105(10):1338–48. https://doi.org/10.1002/bjs.10871.
Jiang Y, Zhang Q, Hu Y, Li T, Yu J, Zhao L, Ye G, Deng H, Mou T, Cai S, et al. ImmunoScore signature: a prognostic and predictive tool in gastric cancer. Ann Surg. 2018;267(3):504–13. https://doi.org/10.1097/SLA.0000000000002116.
Ribas A, Wolchok JD. Cancer immunotherapy using checkpoint blockade. Science. 2018;359(6382):1350–5. https://doi.org/10.1126/science.aar4060.
Jiang X, Wang J, Deng X, Xiong F, Ge J, Xiang B, Wu X, Ma J, Zhou M, Li X, et al. Role of the tumor microenvironment in PD-L1/PD-1-mediated tumor immune escape. Mol Cancer. 2019;18(1):10. https://doi.org/10.1186/s12943-018-0928-4.
Valk E, Rudd CE, Schneider H. CTLA-4 trafficking and surface expression. Trends Immunol. 2008;29(6):272–9. https://doi.org/10.1016/j.it.2008.02.011.
Takahashi T, Tagami T, Yamazaki S, Uede T, Shimizu J, Sakaguchi N, Mak TW, Sakaguchi S. Immunologic self-tolerance maintained by CD25(+)CD4(+) regulatory T cells constitutively expressing cytotoxic T lymphocyte-associated antigen 4. J Exp Med. 2000;192(2):303–10. https://doi.org/10.1084/jem.192.2.303.
Bagchi S, Yuan R, Engleman EG. Immune checkpoint inhibitors for the treatment of cancer: clinical impact and mechanisms of response and resistance. Annu Rev Pathol. 2021;16:223–49. https://doi.org/10.1146/annurev-pathol-042020-042741.
Qin S, Xu L, Yi M, Yu S, Wu K, Luo S. Novel immune checkpoint targets: moving beyond PD-1 and CTLA-4. Mol Cancer. 2019;18(1):155. https://doi.org/10.1186/s12943-019-1091-2.
Gur C, Ibrahim Y, Isaacson B, Yamin R, Abed J, Gamliel M, Enk J, Bar-On Y, Stanietsky-Kaynan N, Coppenhagen-Glazer S, et al. Binding of the Fap2 protein of Fusobacterium nucleatum to human inhibitory receptor TIGIT protects tumors from immune cell attack. Immunity. 2015;42(2):344–55. https://doi.org/10.1016/j.immuni.2015.01.010.
Zhang F, Wang H, Yu J, Yao X, Yang S, Li W, Xu L, Zhao L. LncRNA CRNDE attenuates chemoresistance in gastric cancer via SRSF6-regulated alternative splicing of PICALM. Mol Cancer. 2021;20(1):6. https://doi.org/10.1186/s12943-020-01299-y.
Wu Q, Ma J, Wei J, Meng W, Wang Y, Shi M. FOXD1-AS1 regulates FOXD1 translation and promotes gastric cancer progression and chemoresistance by activating the PI3K/AKT/mTOR pathway. Mol Oncol. 2021;15(1):299–316. https://doi.org/10.1002/1878-0261.12728.
Lin M, Xu M, Xu Z, Weng Z, Lin B, Lan Y, Liu Q, Lin X, Pan J. LINC00200 contributes to the chemoresistance to oxaliplatin of gastric cancer cells via regulating E2F1/RAD51 axis. Hum Cell. 2021;34(4):1163–73. https://doi.org/10.1007/s13577-021-00523-1.
Walther TC, Alves A, Pickersgill H, Loiodice I, Hetzer M, Galy V, Hulsmann BB, Kocher T, Wilm M, Allen T, et al. The conserved Nup107–160 complex is critical for nuclear pore complex assembly. Cell. 2003;113(2):195–206. https://doi.org/10.1016/s0092-8674(03)00235-6.
Loiodice I, Alves A, Rabut G, Van Overbeek M, Ellenberg J, Sibarita JB, Doye V. The entire Nup107-160 complex, including three new members, is targeted as one entity to kinetochores in mitosis. Mol Biol Cell. 2004;15(7):3333–44. https://doi.org/10.1091/mbc.e03-12-0878.
Shi R, Xu L, Huang L, Cheng JX. Nucleoporin 107 promotes the survival of tumor cells in cervical cancers. Gynecol Obstet Invest. 2020;85(1):41–52. https://doi.org/10.1159/000502788.
Alanee S, Delfino K, Wilber A, Robinson K, Brard L, Semaan A. Single nucleotide variant in nucleoporin 107 may be predictive of sensitivity to chemotherapy in patients with ovarian cancer. Pharmacogenet Genomics. 2017;27(7):264–9. https://doi.org/10.1097/FPC.0000000000000288.
Shen Q, Yu M, Jia JK, Li WX, Tian YW, Xue HZ. Possible molecular markers for the diagnosis of pancreatic ductal adenocarcinoma. Med Sci Monit. 2018;24:2368–76. https://doi.org/10.12659/msm.906313.
Nong W, Ma L, Lan B, Liu N, Yang H, Lao X, Deng Q, Huang Z. Comprehensive identification of bridge genes to explain the progression from chronic hepatitis B virus infection to hepatocellular carcinoma. J Inflamm Res. 2021;14:1613–24. https://doi.org/10.2147/JIR.S298977.
Jin DH, Park SE, Lee J, Kim KM, Kim S, Kim DH, Park J. Copy number gains at 8q24 and 20q11-q13 in gastric cancer are more common in intestinal-type than diffuse-type. PLoS ONE. 2015;10(9):e0137657. https://doi.org/10.1371/journal.pone.0137657.
Wang X, Liang Q, Zhang L, Gou H, Li Z, Chen H, Dong Y, Ji J, Yu J. C8orf76 promotes gastric tumorigenicity and metastasis by directly inducing lncRNA DUSP5P1 and associates with patient outcomes. Clin Cancer Res. 2019;25(10):3128–40. https://doi.org/10.1158/1078-0432.CCR-18-2804.
Acknowledgements
The authors gratefully acknowledge the TCGA Research Network for the availability of data.
Funding
This study was supported by the National Key Research and Development Program of China (2018YFC2002000).
Author information
Authors and Affiliations
Contributions
DC, and ZFX designed the study, analyzed the data, and wrote the manuscript. MMW, and XJ generated figures and tables, and revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, D., Wang, M., Jiang, X. et al. Comprehensive analysis of ZFPM2-AS1 prognostic value, immune microenvironment, drug sensitivity, and co-expression network: from gastric adenocarcinoma to pan-cancers. Discov Onc 13, 24 (2022). https://doi.org/10.1007/s12672-022-00487-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-022-00487-0