Abstract
The symbiotic interaction between gut microbiota and the digestive tract is an important factor in maintaining the intestinal environment balance. Colorectal cancer (CRC) is a complex disease involving the interaction between tumour cells and a large number of microorganisms. The microbiota is involved in the occurrence, development and prognosis of colorectal cancer. Several microbiota species have been studied, such as Fusobacterium nucleatum (F. nucleatum), Enterotoxigenic Bacteroides fragilis (ETBF), Streptococcus bovis (S. bovis), Lactobacillus, and Bifidobacterium. Studies about the interaction between microbiota and CRC were retrieved from Embase, PubMed, Ovid and Web of Science up to 21 Oct 2021. This review expounded on the effect of microbiota on CRC, especially the dysregulation of bacteria and carcinogenicity. The methods of gut microbiota modifications representing novel prognostic markers and innovative therapeutic strategies were also described.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
CRC ranks third in the incidence rate and second in the mortality rate among all types of malignant tumours worldwide [1]. CRC is a heterogeneous cancer arising from different genetic and epigenetic events. Generally, CRC is considered a multifactorial disease caused by a high-fat diet, obesity, smoking, drinking, inflammatory processes, environmental factors and genetic alterations. CRC has a trend of increasing morbidity and mortality in developing countries. Recently, several studies revealed that CRC is significantly associated with microbiota features and diet [2,3,4,5,6,7,8,9]. The mode of life and diet seem to be the most important natural factors impacting the gut microbiota. Diet varies among ethnicities, nationalities and regions (rural or urban) [10]. The morbidity of CRC in different populations worldwide may be related to gut microbiota disorders in different regions. Evidence shows that the gut microbiota has a causal relationship with colorectal carcinogenesis [11]. Gut barrier dysfunction and increased tight junction permeability precede the development of colon tumors [12].
2 General overview of the gut microbiota
The human gut microbiota is a complex community of bacteria, archaea, viruses and eukaryotes that is subject-specific and stable in healthy people [13]. Patients with CRC harbour a distinct microbiota. Scanlan et al. analyzed the fecal flora with CRC and colorectal polyps separately and observed an increased number of Clostridium globosa and Clostridium tenderis [14]. Dejea et al. observed that precancerous polyps from patients with familial adenomatous polyposis possessed biofilms with E. coli and B. fragilis, which can have a role in tumour progression [15]. These pathogenic bacteria were increased in patients and contributed to CRC by inducing tumour proliferation, promoting inflammation and DNA damage and protecting cancer from immune attack [16, 17]. Dai and Yachida found that probiotics, including Bifidobacterium and Lactobacillus, decreased, which exerted a protective effect against CRC [14, 16, 18]. With the advent of DNA sequencing technology, the capability to study microbiota composition has increased, allowing for more accurate and rapid identification and classification of microbiota in individuals. Metagenomic sequencing for large-scale fecal samples and mucosa indicated that gut microbiota diversity in CRC patients decreased [19, 20]. This phenomenon was also found in the case of intestinal adenoma [21]. Therefore, the relative abundance and proportion of gut microbiota are crucial for maintaining the intestinal microecosystem, which is a system with complex interactions among gut microbiota species, metabolites and intestinal tissues. The microbiome imbalance is associated with various diseases, including obesity, Crohn’s disease and gastrointestinal malignancies. A growing amount of evidence has supported that the environment and microbiome are risk factors for several malignancies, including CRC [22, 23]. Pathogenic bacteria can also cause tumours under specific conditions in mouse studies [24, 25]. Other studies revealed that bacterial dysbiosis was potentially associated with CRC and could increase CRC occurrence risk [26, 27]. In addition, Lanka et al. pointed out that gut microbiota could be a potential predictor for early postoperative complications and local recurrence [28].
3 The direct effect of pathogenic bacteria
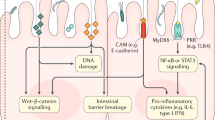
The occurrence of CRC is a comprehensive process influenced by genetic factors. It has different pathogenic mechanisms in the following process: 1. The gut microbiota adheres to intestinal epithelial cells. 2. The gut microbiota binds to Toll-like receptors (TLRs), which are located on the membrane of epithelial cells and endosomes [29]. 3. The gut microbiota synthesizes and secretes cytotoxic biomolecules or metabolites [30]. 4. Cytotoxic biomolecules or metabolites can affect DNA structure and repair sufficiency, pericellular stagnation and chromophore malformation [31, 32]. Experimental results have shown that S. bovis could increase cell proliferation and the secretion of cytokines and transcription factors, such as interleukin-1 (IL-1), interferon-G (IFN-G), IL-8 and nuclear factor kappa-B (NF-κB), which could promote human carcinoma [33]. Gram-negative bacteria could increase the expression of TLR4 [34]. Activated TLR4 may promote CRC occurrence via the Cox-2 or epidermal growth factor receptor (EGFR) signalling pathway and other mechanisms [35]. Li et al. found that the intestinal flora in CRC patients promoted the progression of adenoma, damaged intestinal barrier function, induced chronic low-grade inflammation and activated the Wnt signaling pathway [36].
In healthy people, the intestinal microbiota inhabits the colon mucus without triggering an inflammatory response, which could prevent direct contact with gut microbiota, while Dejea et al. considered that the mucosa-associated microbial community is an important factor in CRC pathogenesis, particularly in the proximal colon [37, 38]. Invasive bacteria can disrupt biofilms and cause a series of carcinogenic effects [39]. A study showed that bacteria were involved in the formation of proximal tumours and not in distal cases, suggesting that CRC could play different roles between the proximal colon and distal colon [40]. The reason could be as follows: first, proximal CRC is more hypermethylated and has a higher mutation rate [41]. Second, the proximal colon has a high level of fermentable carbohydrate substrates, while the distal colon has a thickened mucus layer and increased bacteria, which could attenuate immune activity [42,43,44]. When the local environment changes, for example, in cases of operation and infection, bacteria are more likely to accumulate in the distal colon [20, 43, 44]. Nevertheless, a recent metagenomic study revealed that distal CRC patients had more abundant carcinoma-associated bacterial genes in the feces than proximal CRC patients [45]. The proximal colon contains more abundant F. nucleatum, and their metabolism can induce carcinogens and an inflammatory tumour microenvironment and increase the mutation rate [46]. This phenomenon indicated that site-specific bacteria could cause CRC, and some bacteria play different roles between proximal and distal CRC.
Although F. nucletum is an indigenous species in the normal oral cavity, it is associated with pathological changes of the colon and rectum. It has been discovered that higher F. nucleatum is associated with higher T stages, lymph node metastases, tumour invasion, and larger tumour sizes [3, 5, 47]. Studies have revealed that F. nucleatum is higher in patients with adenomas, high-grade dysplasia and CRC than in healthy people [48]. A higher concentration of F. nucleatum has a negative impact on survival outcomes [3, 48,49,50]. The phenomenon that F. nucleatum levels were positively correlated with younger age and MSI-H in South Africa was found by Viljoen et al. [51]. According to a large cohort study, activating autophagy to stimulate tumour growth and promote chemotherapy tolerance can lead to a shorter survival time [42]. Furthermore, it is strongly associated with the CpG island methylator phenotype, the density of CD3 + T cells, microsatellite instability and chromosomal instability [51,52,53,54]. Gur et al. reported that F. nucleatum could inhibit the function of NK cells by binding to the inhibitory receptor TIGIT via the Fap2 protein [55].
ETBF is associated with chronic colorectal diseases, including enteritis, chronic colorectal dysfunction and even CRC [56, 57]. Viljoen et al. revealed that the levels of ETBF and F. nucleatum were significantly higher in advanced CRC (stage III and IV) and had a positive association between regional lymph node metastases and high-level colonization by Fusobacterium [49, 51]. ETBF triggered β-catenin nuclear signaling, induced c-Myc expression and cellular proliferation and increased colitis and tumors in a Min/+ mouse model [58, 59]. ETBF promotes CRC in the following ways. First, NF-κB plays an important role in the host response to microbial infection by coordinating innate and acquired immune functions [60]. ETBF can stimulate intracellular IL-17 secretion, which activates the NF-κB protein, triggering long-term chronic inflammation and finally leading to tumorigenesis [61]. Furthermore, the activated NF-κB signalling pathway could increase chemokine (CXCL1, CXCL2, and CXCL5) and polyamine metabolism and induce DNA damage [62]. Second, the mitogen-activated protein kinase (MAPK) signaling pathway regulates cell inflammation, proliferation and apoptosis [63]. Under bacterial stimulation, intestinal epithelial cells rapidly secrete IL-8 and monocyte chemotactic protein 1 (MCP-1), which activate the MAPK signaling pathway. This pathway induces intercellular adhesion molecule-1 (ICAM-1) and the enhancement of adhesion between inflammatory cells and endothelial cells, which could decrease the permeability of inflammatory cells, aggravate the inflammatory response, inhibit cell apoptosis and further promote the development of CRC [64]. Third, ETBF also increases the expression of cyclooxygenase (COX)-2 and releases prostaglandin E-2 (PGE2), which activate inflammation associated with signal transducers and activators of transcription (STAT)3 and interact with epithelial cells [65]. Toxin can degrade E-cadherin, thus altering signalling pathways, upregulating spermine oxidase and leading to cell morphology, promoting carcinogenesis and irreversible DNA damage [65]. Finally, ETBF can also promote tumor growth via associated lncRNA1 by activating Ras homologs [66].
S. bovis, also named Streptococcus gallolyticus subsp. gallolyticus (SGG) is one of the first bacteria clearly associated with CRC occurrence [67]. It is more common in early-stage adenomas than in later-stage carcinomas, and its concomitant inflammatory factors concentrate in the intestine by recruiting CD11b+TLR-4+cells [68, 69]. Meamwhile, Aymeric et al. concluded that CRC-specific conditions promote S. bovis colonization by the Wnt pathway and the decreased expression of the bile acid apical transporter gene Slc10A2 [67]. Bovis has also been widely studied, and its antibody may serve as a potential marker for CRC at an early stage of disease [70].
4 Inflammatory factor
Numerous inflammatory factor were found to be associated with protumor or antitumor effects in CRC pathogenesis [71, 72]. IL-1 promotes tumorigenesis and tumor metastasis in CRC [73], and the tumor-suppressive effects of IL-1 have also been reported [73, 74]. Tumor necrosis factor (TNF) plays a dual role in cancer. It can induce apoptosis by activating the TNF receptor and promote tumorigenesis and cancer progression by activating TNF receptor 2 [71]. Gut microbiota dysfunction could induce abnormal immune responses and create a special immune microenvironment causing DNA damage and gene mutation. For example, members of Enterobacteriaceae are known to cause inflammation in the gastrointestinal tract and contribute to CRC [75]. IL-8 has a chemotactic effect on neutrophils, recruiting more immature polymorphic cells and further aggravating inflammation and cell damage [76]. Tseng et al. showed that the IL-6 signaling pathway plays an important role in the occurrence and chemoresistance of various cancers, including CRC [73, 77, 78]. Under inflammatory conditions, NF-κB-induced IL-6 can lead to cancer progression and metastasis via the IL-6/STAT3 signaling pathway [77]. Bacteria-derived lipopolysaccharide stimulated IL-1β- and IL-17-producing T helper cell activation to promote inflammation [79]. By using human and mouse models, Sobhani et al. demonstrated that CRC-associated microbiota induced gene methylation. Several gene promoters were hypermethylated in CRC but not in normal tissues [27]. Hypermethylation of the promoter Wif1 could serve as a surrogate diagnostic marker for early CRC [80].
In contrast, some bacterias could enhance the secretion of anti-inflammatory factors, such as IL-10 and TGF-β, which reduce the concentrations of proinflammatory factors, such as IFN-γ, reduce the infiltration of NK cells and the degree of the inflammatory response [81, 82]. By focusing on TGF-β1 signaling, IL-25 is constitutively produced by gut mucosal cells and restrain intestinal inflammation [79], and IL-6R is considered a potential target for CRC treatment. Tocilizumab, an IL-6R antagonist antibody, blocks the IL-6/STAT3 signaling pathway, which could reduce CRC viability and enhance cell apoptosis [83].
5 Effect of bacterial metabolites
The gut microbiota participates in synthesis and metabolism, while the metabolites regulate intestinal micro ecological balance in return. For example, red meat, processed meat and high protein food are digested by enzymes into toxic nitrogen and sulfur-containing substances [84]. Hydrogen sulfide (H2S), generated by sulfate-reducing bacteria (SRB), can promote inflammation of the colon mucosa, induce DNA damage and methylation, release free radicals and simultaneously inhibit the synthesis of cytochrome oxidase butyric acid and mucus [84, 85]. In addition, a high-fat diet can produce secondary cholic acid, affecting mitosis, activating NF-κB and epidermal growth factor receptor (EGFR) and promoting tumour development [69, 86, 87]. Intestinal compounds activated by microorganisms are derived from some vegetable foods, such as intestinal lignans, which may play a role in carcinogenesis [88]. For example, intestinal microbes can produce short-chain fatty acids (SCFAs), such as butyric acid, by fermenting food, which has been proven to inhibit intestinal inflammation, regulate the immune response, maintain barrier function, reduce precancerous lesions and regulate DNA methylation [27, 89]. SCFAs generated by intestinal microorganisms are primary nutritional substrates for colonocytes [90]. By upregulating secretory IgA and cytokines, prebiotics can enhance host immunity [91]. Butyric acid protects against pathogenetic mechanisms mediated by reactive oxygen species and aids in understanding the apparent protection against CRC. Butyrate is important for colon epithelial cells by increasing apoptosis, decreasing cell proliferation and increasing differentiation [92]. Butyrate substrates have been shown to increase the acute apoptotic response and halt the cell cycle through animal studies [93]. Acting as a histone deacetylase inhibitor, butyrate could also regulate the expression of CDKN1A [94]. They can also increase phytochemicals from active microorganisms, such as polyphenols with anti-inflammatory and antioxidant properties [16]. Several species, such as Lactobacillus rhamnosus GG, Lactobacillus, Lactobacillus casei, and Bifidobacterium breve, can also downregulate inflammatory reactions or preserve intestinal barrier function [95, 96]. It has also been reported that the protective protein p40 can inhibit epithelial cell apoptosis and intestinal barrier destruction induced by cytokines and enhance IgA secretion [97]. Lactate dehydrogenase or other molecules from lactic acid bacteria can induce apoptosis or inhibit CRC invasion [87, 98]. However, some reports hold the opposite view: butyrate treatment could promote epithelial cell proliferation and polyp formation and eventually tumor progression [99]. Therefore, butyric acid supplementation does not necessarily benefit the host’s health, which is highly dependent on the somatic genetic background [100].
6 Effect of antibiotics
Antibiotics might be related to inhibiting tumour proliferation, invasion and growth. For patients with colon cancer, antibiotic therapy is even recommended as an immunotherapy strategy [100, 101]. After antibiotic manipulation of the gut microbiota, a reduction in tumor load was detected, which supported the hypothesis of bacterial involvement in carcinogenesis [102]. For example, metronidazole can eradicate Fusobacterium colonization and inhibit the proliferation of CRC [103]. In the future, antibiotic therapy might become a potentially valuable method to prevent cancer. However, accumulating evidence indicates that antibiotics can damage immunotherapy and induce disease progression by further creating microbial disorders. A meta-analysis including 8 studies showed that current research only suggests a weak association between cumulative antibiotic consumption and risk of CRC because of the heterogeneity and quality of the available research [104, 105]. In addition, antibiotic-mediated bacterial clearance may aggravate unexpected adverse reactions; for example, vancomycin and anti-ctla-4 can lead to more severe and fatal colonic inflammation in mice with colitis [106]. Therefore, a large number of high-quality studies are needed to further confirm the relationship between antibiotic exposure and CRC progression.
7 Treatment of CRC by regulating gut microbiota
The theory of the “driver-passenger” model supposes that disease-related microorganisms may cause DNA damage, increase cell proliferation, produce genotoxic substances and lead to adenoma or cancer [75, 107]. The microenvironment of the tumour changes, which may be beneficial to the growth of bacteria that dominate the tumour. In recent years, an increasing number of people have paid attention to the intestinal flora in different stages of CRC to distinguish the initial flora from the late flora [75, 108]. Thousands of bacteria lie in the intestinal tract, but not all of them are carcinogens. Probiotics are active intestinal microorganisms that bring health benefits to hosts [109]. In addition to direct interaction, probiotics also produce metabolites, such as acetic acid or bacteriocin, which can inhibit the growth of pathogens by lowering the pH value and play a direct antibacterial role [110,111,112].
The most widely used clinically adopted probiotics include Lactobacillus, Bifidobacterium and Clostridium butyricum. Zaharuddin et al. showed that probiotics may modify the intestinal microenvironment, resulting in a decline in proinflammatory cytokines [113]. Geier et al. discovered that lactic acid bacteria and Bifidobacterium could decrease CRC incidence by reducing the activities of azo reductase, nitro reductase and β-glucuronidase [109]. The expression of Th17 cells and the secretion of IL-23 and IL-17 were downregulated by lactic acid bacteria by inhibiting the STAT3 and NF-κB signalling pathways [114]. Another study showed that Lactobacillus rhamnosus, Lactobacillus plantarum and Escherichia coli enhanced intestinal barrier function by upregulating the expression of tight junction proteins, stimulating mucin production and promoting epithelial restitution [115,116,117]. Lactic acid bacteria showed antitumour properties in a CRC mouse model induced by dimethylhydrazine (DMH) [105]. By affecting the metabolism of retinoic acid, Bifidobacterium infantis and Bifidobacterium breve activate intestinal dendritic cells (DC) and release regulatory T cells (Treg) expressing Foxp3 + , regulatory T cells of type 1 (Tr1) and il-10, thus exerting an antitumour effect [115, 116]. Oral probiotics (such as Bifidobacterium and Akkermansia muciniphila) and fecal microbial transplantation (FMT) can significantly enhance immune therapy based on PD-1 by enhancing the response of dendritic cells and T cells [118,119,120]. In animal experiments, Sivan et al. found that oral Bifidobacterium in mice had the same efficacy as an anti-PDL1 immunosuppressant and almost completely inhibited the growth of melanoma in mice [120].
The anti-inflammatory effect can also be produced by gene modification and protein expression. For example, efforts are devoted to studying S-layer proteins, lipoteichoic acid and exopolysaccharides, and the deletion of lipoteichoic acid and immunostimulating protein in lactic acid bacteria downregulates the expression of proinflammatory mediators and inhibits the occurrence of colonic inflammation and cancer [110, 121]. Intestinal microbes recovered through whole metagenomic analysis have become a potential strategy for the prevention and treatment of CRC [27, 105]
8 FMT
The gut microbiota creates a complex microenvironment that can influence the tumor in a very heterogeneous way that relies on intrinsic heterogeneity [77]. Studies based on the intestinal flora of healthy people and patients show that FMTs can restore microbial homeostasis and may help to improve various gastrointestinal diseases, including irritable bowel disease and Clostridium difficile infection [122]. FMT seems to bring some benefits. For example, it increases the diversity of microorganisms without destroying the intestinal flora. FMT of A. shahii had a good response in antibiotic-treated colon tumor mice [123]. In addition, FMT led to a reduction in the number of tumors and inflammation, as well as the inhibition of proinflammatory molecules (IL-1β, IL-6, and TNF-α) and an increase in anti-inflammatory cytokines (IL-10 and TGF-β) [79, 124].
However, from colon cancer mice or CRC patients to sterile mice, FMT experiments revealed that intestinal microflora played a key role in the development of CRC [125, 126]. This treatment has the risk of transferring pathogens and antibiotic-resistant genes, and some people worry that the donor’s microflora and its hidden complexity may cause chronic diseases in Shinjuku [127]. Nooij et al. studied the change in the prevalence and abundance of potentially carcinogenic pks + E. coli after FMT [128]. The pks genome island encodes colibactin, which is a polypeptide-polyketone hybrid that can induce double-stranded DNA breakage and chromosome aberration [129]. At present, there is no validated diagnostic test that can accurately assess the carcinogenic potential of microorganisms. Importantly, all medical decisions require systematic risk and benefit assessment [127]. FDA warned of the potential risks of spreading multidrug-resistant bacteria and subsequently developing life-threatening infections. In a case reported by the FDA, two immunocompromised patients were infected by broad-spectrum β-lactamase (ESBL)-producing E. coli, and one patient died [130]. The evidence of using FMT in cancer is limited, and more clinical trials of solid cancer are just beginning to appear. The risk of adverse events should not be underestimated, including potentially fatal systemic inflammatory response syndrome and unintentional transfer of pathogens, including highly resistant microorganisms [123].
In general, the continuous changes after FMT show that the long-term consequences, whether promising or harmful, should be taken into account [131]. Rooks and others emphasized the importance of FMT in preventing, treating and controlling disease progression by changing the intestinal flora [132]. If FMT could be used to predict CRC and adopted in clinical practice, it could be a novel, convenient, efficient, economical and noninvasive method.
9 Conclusion
At present, the microorganisms promoting CRC mainly include nuclear F. nucleatum, ETBF, and S. bovis, and the antitumour microorganisms mainly include Lactobacillus and Bifidobacterium. The increased content of self auto-induced factor 2, extracted from F. nucleatum in CRC patient feces, was associated with tumor immunity through tumor-associated macrophages and the CD4/CD8 ratio. This indicates that it may be a potential marker for clinical screening [133]. Another study pointed out that combining IgA and IgG against Fusobacteria with CEA and CA 19–9 may be a better method for screening CRC. (AUC = 0.743, specificity = 94.22%, sensitivity = 40.00%) [134]. It is challenging to select a single intestinal microorganism as the diagnostic marker, and the combination of multiple microorganism scans, such as Roseburia, Clostridium and Akkermansia, improves the sensitivity and specificity [36]. In the future, it is possible to formulate a new standard for early screening and recurrence by the flora mentioned above [122, 135].
References
Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer statistics, 2021. CA Cancer J Clin. 2021;71(1):7–33.
Baxter BA, Oppel RC, Ryan EP. Navy beans impact the stool metabolome and metabolic pathways for colon health in cancer survivors. Nutrients. 2018. https://doi.org/10.3390/nu11010028.
Yu L, Zhao G, Wang L, Zhou X, Sun J, Li X, et al. A systematic review of microbial markers for risk prediction of colorectal neoplasia. Br J Cancer. 2022;126:1318–28. https://doi.org/10.1038/s41416-022-01740-7.
Nakatsu G, Li X, Zhou H, Sheng J, Wong SH, Wu WK, et al. Gut mucosal microbiome across stages of colorectal carcinogenesis. Nat Commun. 2015;6:8727. https://doi.org/10.1038/ncomms9727.
Castellarin M, Warren RL, Freeman JD, Dreolini L, Krzywinski M, Strauss J, et al. Fusobacterium nucleatum infection is prevalent in human colorectal carcinoma. Genome Res. 2012;22:299–306. https://doi.org/10.1101/gr.126516.111.
Zamani S, Taslimi R, Sarabi A, Jasemi S, Sechi LA, Feizabadi MM. Enterotoxigenic Bacteroides fragilis: a possible etiological candidate for bacterially-induced colorectal precancerous and cancerous lesions. Front Cell Infect Microbiol. 2019;9:449. https://doi.org/10.3389/fcimb.2019.00449.
He Z, Gharaibeh RZ, Newsome RC, Pope JL, Dougherty MW, Tomkovich S, et al. Campylobacter jejuni promotes colorectal tumorigenesis through the action of cytolethal distending toxin. Gut. 2019;68:289–300. https://doi.org/10.1136/gutjnl-2018-317200.
Shield KD, Freisling H, Boutron-Ruault MC, Touvier M, Marant Micallef C, Jenab M, et al. New cancer cases attributable to diet among adults aged 30–84 years in France in 2015. Br J Nutr. 2018;120:1171–80. https://doi.org/10.1017/S0007114518002544.
Ubago-Guisado E, Rodríguez-Barranco M, Ching-López A, Petrova D, Molina-Montes E, Amiano P, et al. Evidence update on the relationship between diet and the most common cancers from the European Prospective Investigation into Cancer and Nutrition (EPIC) Study: a systematic review. Nutrients. 2021. https://doi.org/10.3390/nu13103582.
Ayeni FA, Biagi E, Rampelli S, et al. Infant and adult gut microbiome and metabolome in rural Bassa and urban settlers from Nigeria. Cell Rep. 2018;23(10):3056–67.
Keku TO, Dulal S, Deveaux A, Jovov B, Han X. The gastrointestinal microbiota and colorectal cancer. Am J Physiol Gastrointest Liver Physiol. 2015;308(5):G351–63.
Soler AP, Miller RD, Laughlin KV, Carp NZ, Klurfeld DM, Mullin JM. Increased tight junctional permeability is associated with the development of colon cancer. Carcinogenesis. 1999;20(8):1425–31.
Jalanka-Tuovinen J, Salonen A, Nikkilä J, et al. Intestinal microbiota in healthy adults: temporal analysis reveals individual and common core and relation to intestinal symptoms. PLoS ONE. 2011;6(7): e23035.
Scanlan PD, Shanahan F, Clune Y, et al. Culture-independent analysis of the gut microbiota in colorectal cancer and polyposis. Environ Microbiol. 2008;10(3):789–98.
Dejea CM, Fathi P, Craig JM, Boleij A, Taddese R, Geis AL, et al. Patients with familial adenomatous polyposis harbor colonic biofilms containing tumorigenic bacteria. Science. 2018;359:592–7. https://doi.org/10.1126/science.aah3648.
Dai Z, Coker OO, Nakatsu G, et al. Multi-cohort analysis of colorectal cancer metagenome identified altered bacteria across populations and universal bacterial markers. Microbiome. 2018;6(1):70.
Long X, Wong CC, Tong L, et al. Peptostreptococcus anaerobius promotes colorectal carcinogenesis and modulates tumour immunity. Nat Microbiol. 2019;4(12):2319–30.
Yachida S, Mizutani S, Shiroma H, et al. Metagenomic and metabolomic analyses reveal distinct stage-specific phenotypes of the gut microbiota in colorectal cancer. Nat Med. 2019;25(6):968–76.
Wirbel J, Pyl PT, Kartal E, et al. Meta-analysis of fecal metagenomes reveals global microbial signatures that are specific for colorectal cancer. Nat Med. 2019;25(4):679–89.
Flemer B, Lynch DB, Brown JM, Jeffery IB, Ryan FJ, Claesson MJ, et al. Tumour-associated and non-tumour-associated microbiota in colorectal cancer. Gut. 2017;66:633–43. https://doi.org/10.1136/gutjnl-2015-309595.
Saito K, Koido S, Odamaki T, et al. Metagenomic analyses of the gut microbiota associated with colorectal adenoma. PLoS ONE. 2019;14(2): e0212406.
Song M, Chan AT. Environmental factors, gut microbiota, and colorectal cancer prevention. Clin Gastroenterol Hepatol. 2019;17(2):275–89.
Wong SH, Yu J. Gut microbiota in colorectal cancer: mechanisms of action and clinical applications. Nat Rev Gastroenterol Hepatol. 2019;16(11):690–704.
Arthur JC, Perez-Chanona E, Mühlbauer M, Tomkovich S, Uronis JM, Fan TJ, et al. Intestinal inflammation targets cancer-inducing activity of the microbiota. Science. 2012;338:120–3. https://doi.org/10.1126/science.1224820.
Uronis JM, Mühlbauer M, Herfarth HH, Rubinas TC, Jones GS, Jobin C. Modulation of the intestinal microbiota alters colitis-associated colorectal cancer susceptibility. PLoS ONE. 2009;4: e6026. https://doi.org/10.1371/journal.pone.0006026.
Kwong T, Wang X, Nakatsu G, Chow TC, Tipoe T, Dai R, et al. Association between bacteremia from specific microbes and subsequent diagnosis of colorectal cancer. Gastroenterology. 2018;155:383-90.e8. https://doi.org/10.1053/j.gastro.2018.04.028.
Sobhani I, Bergsten E, Couffin S, et al. Colorectal cancer-associated microbiota contributes to oncogenic epigenetic signatures. Proc Natl Acad Sci USA. 2019;116(48):24285–95.
Lauka L, Reitano E, Carra MC, et al. Role of the intestinal microbiome in colorectal cancer surgery outcomes. World J Surg Oncol. 2019;17(1):204.
Gay NJ, Symmons MF, Gangloff M, Bryant CE. Assembly and localization of Toll-like receptor signalling complexes. Nat Rev Immunol. 2014;14(8):546–58.
Schmitt M, Greten FR. The inflammatory pathogenesis of colorectal cancer. Nat Rev Immunol. 2021;21:653–67. https://doi.org/10.1038/s41577-021-00534-x.
Veziant J, Gagnière J, Jouberton E, et al. Association of colorectal cancer with pathogenic Escherichia coli: focus on mechanisms using optical imaging. World J Clin Oncol. 2016;7(3):293–301.
Cuevas-Ramos G, Petit CR, Marcq I, Boury M, Oswald E, Nougayrède JP. Escherichia coli induces DNA damage in vivo and triggers genomic instability in mammalian cells. Proc Natl Acad Sci USA. 2010;107:11537–42. https://doi.org/10.1073/pnas.1001261107.
Abdulamir AS, Hafidh RR, Abu BF. The association of Streptococcus bovis/gallolyticus with colorectal tumors: the nature and the underlying mechanisms of its etiological role. J Exp Clin Cancer Res. 2011;30(1):11.
Tang AT, Choi JP, Kotzin JJ, Yang Y, Hong CC, Hobson N, et al. Endothelial TLR4 and the microbiome drive cerebral cavernous malformations. Nature. 2017;545:305–10. https://doi.org/10.1038/nature22075.
Fukata M, Chen A, Vamadevan AS, et al. Toll-like receptor-4 promotes the development of colitis-associated colorectal tumors. Gastroenterology. 2007;133(6):1869–81.
Li L, Li X, Zhong W, Yang M, Xu M, Sun Y, Ma J, Liu T, Song X, Dong W, Liu X, Chen Y, Liu Y, Abla Z, Liu W, Wang B, Jiang K, Cao H. Gut microbiota from colorectal cancer patients enhances the progression of intestinal adenoma in Apc(min/+) mice. EBioMedicine. 2019;48:301–15. https://doi.org/10.1016/j.ebiom.2019.09.021.
Dejea CM, Wick EC, Hechenbleikner EM, et al. Microbiota organization is a distinct feature of proximal colorectal cancers. Proc Natl Acad Sci USA. 2014;111(51):18321–6.
Johansson ME, Larsson JM, Hansson GC. The two mucus layers of colon are organized by the MUC2 mucin, whereas the outer layer is a legislator of host-microbial interactions. Proc Natl Acad Sci USA. 2011;108(Suppl 1):4659–65. https://doi.org/10.1073/pnas.1006451107.
Johnson CH, Dejea CM, Edler D, et al. Metabolism links bacterial biofilms and colon carcinogenesis. Cell Metab. 2015;21(6):891–7.
Peters BA, Dominianni C, Shapiro JA, et al. The gut microbiota in conventional and serrated precursors of colorectal cancer. Microbiome. 2016;4(1):69.
The Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012;487(7407):330–7.
Koropatkin NM, Cameron EA, Martens EC. How glycan metabolism shapes the human gut microbiota. Nat Rev Microbiol. 2012;10(5):323–35.
Lee GH, Malietzis G, Askari A, Bernardo D, Al-Hassi HO, Clark SK. Is right-sided colon cancer different to left-sided colorectal cancer?—A systematic review. Eur J Surg Oncol. 2015;41(3):300–8.
Macfarlane GT, Macfarlane S. Bacteria, colonic fermentation, and gastrointestinal health. J AOAC Int. 2012;95(1):50–60.
Feng Q, Liang S, Jia H, et al. Gut microbiome development along the colorectal adenoma-carcinoma sequence. Nat Commun. 2015;6:6528.
Kostic AD, Chun E, Robertson L, et al. Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe. 2013;14(2):207–15.
Li YY, Ge QX, Cao J, et al. Association of Fusobacterium nucleatum infection with colorectal cancer in Chinese patients. World J Gastroenterol. 2016;22(11):3227–33.
Flanagan L, Schmid J, Ebert M, Soucek P, Kunicka T, Liska V, et al. Fusobacterium nucleatum associates with stages of colorectal neoplasia development, colorectal cancer and disease outcome. Eur J Clin Microbiol Infect Dis. 2014;33:1381–90. https://doi.org/10.1007/s10096-014-2081-3.
Yan X, Liu L, Li H, Qin H, Sun Z. Clinical significance of Fusobacterium nucleatum, epithelial-mesenchymal transition, and cancer stem cell markers in stage III/IV colorectal cancer patients. Onco Targets Ther. 2017;10:5031–46.
Mima K, Nishihara R, Qian ZR, Cao Y, Sukawa Y, Nowak JA, et al. Fusobacterium nucleatum in colorectal carcinoma tissue and patient prognosis. Gut. 2016;65:1973–80. https://doi.org/10.1136/gutjnl-2015-310101.
Viljoen KS, Dakshinamurthy A, Goldberg P, Blackburn JM. Quantitative profiling of colorectal cancer-associated bacteria reveals associations between Fusobacterium spp., Enterotoxigenic Bacteroides fragilis (ETBF) and clinicopathological features of colorectal cancer. PLoS ONE. 2015;10(3):e0119462.
Rubinstein MR, Wang X, Liu W, Hao Y, Cai G, Han YW. Fusobacterium nucleatum promotes colorectal carcinogenesis by modulating E-cadherin/β-catenin signaling via its FadA adhesin. Cell Host Microbe. 2013;14(2):195–206.
Tahara T, Yamamoto E, Suzuki H, et al. Fusobacterium in colonic flora and molecular features of colorectal carcinoma. Cancer Res. 2014;74(5):1311–8.
Ito M, Kanno S, Nosho K, et al. Association of Fusobacterium nucleatum with clinical and molecular features in colorectal serrated pathway. Int J Cancer. 2015;137(6):1258–68.
Gur C, Ibrahim Y, Isaacson B, et al. Binding of the Fap2 protein of Fusobacterium nucleatum to human inhibitory receptor TIGIT protects tumors from immune cell attack. Immunity. 2015;42(2):344–55.
Haghi F, Goli E, Mirzaei B, Zeighami H. The association between fecal enterotoxigenic B. fragilis with colorectal cancer. BMC Cancer. 2019;19(1):879.
Jasemi S, Emaneini M, Fazeli MS, et al. Toxigenic and non-toxigenic patterns I, II and III and biofilm-forming ability in Bacteroides fragilis strains isolated from patients diagnosed with colorectal cancer. Gut Pathog. 2020;12:28.
Wu S, Rhee KJ, Albesiano E, Rabizadeh S, Wu X, Yen HR, Huso DL, Brancati FL, Wick E, McAllister F, Housseau F, Pardoll DM, Sears CL. A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nat Med. 2009;15:1016–22. https://doi.org/10.1038/nm.2015.
Wu S, Morin PJ, Maouyo D, Sears CL. Bacteroides fragilis enterotoxin induces c-Myc expression and cellular proliferation. Gastroenterology. 2003;124:392–400. https://doi.org/10.1053/gast.2003.50047.
Peng C, Ouyang Y, Lu N, Li N. The NF-κB signaling pathway, the microbiota, and gastrointestinal tumorigenesis: recent advances. Front Immunol. 2020;11:1387.
Taniguchi K, Karin M. NF-κB, inflammation, immunity and cancer: coming of age. Nat Rev Immunol. 2018;18(5):309–24.
Chung L, Thiele Orberg E, Geis AL, Chan JL, Fu K, DeStefano Shields CE, Dejea CM, Fathi P, Chen J, Finard BB, Tam AJ, McAllister F, Fan H, Wu X, Ganguly S, Lebid A, Metz P, Van Meerbeke SW, Huso DL, Wick EC, Pardoll DM, Wan F, Wu S, Sears CL, Housseau F. Bacteroides fragilis toxin coordinates a pro-carcinogenic inflammatory cascade via targeting of colonic epithelial cells. Cell Host Microbe. 2018;23:203-214.e5. https://doi.org/10.1016/j.chom.2018.01.007.
Wang C, Li P, Xuan J, et al. Cholesterol enhances colorectal cancer progression via ROS elevation and MAPK signaling pathway activation. Cell Physiol Biochem. 2017;42(2):729–42.
Gu T, Li Q, Egilmez NK. IFNβ-producing CX3CR1(+) macrophages promote T-regulatory cell expansion and tumor growth in the APC(min/+)/Bacteroides fragilis colon cancer model. Oncoimmunology. 2019;8(12): e1665975.
Cheng WT, Kantilal HK, Davamani F. The mechanism of Bacteroides fragilis toxin contributes to colon cancer formation. Malays J Med Sci. 2020;27:9–21. https://doi.org/10.21315/mjms2020.27.4.2.
Bao Y, Tang J, Qian Y, Sun T, Chen H, Chen Z, Sun D, Zhong M, Chen H, Hong J, Chen Y, Fang JY. Long noncoding RNA BFAL1 mediates enterotoxigenic Bacteroides fragilis-related carcinogenesis in colorectal cancer via the RHEB/mTOR pathway. Cell Death Dis. 2019;10:675. https://doi.org/10.1038/s41419-019-1925-2.
Marmolin ES, Hartmeyer GN, Christensen JJ, Nielsen XC, Dargis R, Skov MN, et al. Bacteremia with the bovis group Streptococci: species identification and association with infective endocarditis and with gastrointestinal disease. Diagn Microbiol Infect Dis. 2016;85:239–42. https://doi.org/10.1016/j.diagmicrobio.2016.02.019.
Deng Q, Wang C, Yu K, Wang Y, Yang Q, Zhang J, et al. Streptococcus bovis contributes to the development of colorectal cancer via recruiting CD11b+TLR-4+ cells. Med Sci Monit. 2020;26:e921886. https://doi.org/10.12659/MSM.921886.
Yoshimoto S, Loo TM, Atarashi K, et al. Obesity-induced gut microbial metabolite promotes liver cancer through senescence secretome. Nature. 2013;499(7456):97–101.
Butt J, Werner S, Willhauck-Fleckenstein M, Michel A, Waterboer T, Zörnig I, et al. Serology of Streptococcus gallolyticus subspecies gallolyticus and its association with colorectal cancer and precursors. Int J Cancer. 2017;141:897–904. https://doi.org/10.1002/ijc.30765.
Mager LF, Wasmer MH, Rau TT, Krebs P. Cytokine-induced modulation of colorectal cancer. Front Oncol. 2016;6:96. https://doi.org/10.3389/fonc.2016.00096.
Heo G, Lee Y, Im E. Interplay between the gut microbiota and inflammatory mediators in the development of colorectal cancer. Cancers. 2021. https://doi.org/10.3390/cancers13040734.
Li J, Huang L, Zhao H, Yan Y, Lu J. The role of interleukins in colorectal cancer. Int J Biol Sci. 2020;16:2323–39. https://doi.org/10.7150/ijbs.46651.
Isambert N, Hervieu A, Rébé C, Hennequin A, Borg C, Zanetta S, et al. Fluorouracil and bevacizumab plus anakinra for patients with metastatic colorectal cancer refractory to standard therapies (IRAFU): a single-arm phase 2 study. Oncoimmunology. 2018;7: e1474319. https://doi.org/10.1080/2162402X.2018.1474319.
Tjalsma H, Boleij A, Marchesi JR, Dutilh BE. A bacterial driver-passenger model for colorectal cancer: beyond the usual suspects. Nat Rev Microbiol. 2012;10(8):575–82.
Alfaro C, Sanmamed MF, Rodríguez-Ruiz ME, Teijeira Á, Oñate C, González Á, et al. Interleukin-8 in cancer pathogenesis, treatment and follow-up. Cancer Treat Rev. 2017;60:24–31. https://doi.org/10.1016/j.ctrv.2017.08.004.
Abou-Shousha S, Moaaz M, Sheta M, Motawea MA. An approach to breast cancer immunotherapy: the apoptotic activity of recombinant anti-interleukin-6 monoclonal antibodies in intact tumour microenvironment of breast carcinoma. Scand J Immunol. 2016;83:427–37. https://doi.org/10.1111/sji.12426.
Tseng-Rogenski SS, Hamaya Y, Choi DY, Carethers JM. Interleukin 6 alters localization of hMSH3, leading to DNA mismatch repair defects in colorectal cancer cells. Gastroenterology. 2015;148(3):579–89.
Yang Y, Li L, Xu C, Wang Y, Wang Z, Chen M, et al. Cross-talk between the gut microbiota and monocyte-like macrophages mediates an inflammatory response to promote colitis-associated tumourigenesis. Gut. 2020;70:1495–506. https://doi.org/10.1136/gutjnl-2020-320777.
Amiot A, Mansour H, Baumgaertner I, et al. The detection of the methylated Wif-1 gene is more accurate than a fecal occult blood test for colorectal cancer screening. PLoS ONE. 2014;9(7): e99233.
Round JL, Lee SM, Li J, et al. The Toll-like receptor 2 pathway establishes colonization by a commensal of the human microbiota. Science. 2011;332(6032):974–7.
Ip W, Hoshi N, Shouval DS, Snapper S, Medzhitov R. Anti-inflammatory effect of IL-10 mediated by metabolic reprogramming of macrophages. Science. 2017;356(6337):513–9.
Wang J, Zhou J, Jiang C, et al. LNRRIL6, a novel long noncoding RNA, protects colorectal cancer cells by activating the IL-6-STAT3 pathway. Mol Oncol. 2019;13(11):2344–60.
Scanlan PD, Shanahan F, Marchesi JR. Culture-independent analysis of desulfovibrios in the human distal colon of healthy, colorectal cancer and polypectomized individuals. FEMS Microbiol Ecol. 2009;69(2):213–21.
Tjalsma H, Schöller-Guinard M, Lasonder E, Ruers TJ, Willems HL, Swinkels DW. Profiling the humoral immune response in colon cancer patients: diagnostic antigens from Streptococcus bovis. Int J Cancer. 2006;119(9):2127–35.
Goodwin AC, Destefano Shields CE, Wu S, et al. Polyamine catabolism contributes to enterotoxigenic Bacteroides fragilis-induced colon tumorigenesis. Proc Natl Acad Sci USA. 2011;108(37):15354–9.
Chen ZY, Hsieh YM, Huang CC, Tsai CC. inhibitory effects of probiotic Lactobacillus on the growth of human colonic carcinoma cell line HT-29. Molecules. 2017. https://doi.org/10.3390/molecules22010107.
Chong ES. A potential role of probiotics in colorectal cancer prevention: review of possible mechanisms of action. World J Microbiol Biotechnol. 2014;30(2):351–74.
Tong LC, Wang Y, Wang ZB, et al. Propionate ameliorates dextran sodium sulfate-induced colitis by improving intestinal barrier function and reducing inflammation and oxidative stress. Front Pharmacol. 2016;7:253.
Davani-Davari D, Negahdaripour M, Karimzadeh I, Seifan M, Mohkam M, Masoumi SJ, Berenjian A, Ghasemi Y. Prebiotics: definition, types, sources, mechanisms, and clinical applications. 2019. Foods. https://doi.org/10.3390/foods8030092.
Popov J, Caputi V, Nandeesha N, Rodriguez DA, Pai N. Microbiota-immune interactions in ulcerative colitis and colitis associated cancer and emerging microbiota-based therapies. Int J Mol Sci. 2021. https://doi.org/10.3390/ijms222111365.
Winter J, Nyskohus L, Young GP, Hu Y, Conlon MA, Bird AR, Topping DL, Le Leu RK. Inhibition by resistant starch of red meat-induced promutagenic adducts in mouse colon. Cancer Prev Res. 2011;4:1920–8. https://doi.org/10.1158/1940-6207.CAPR-11-0176.
Le Leu RK, Brown IL, Hu Y, Esterman A, Young GP. Suppression of azoxymethane-induced colon cancer development in rats by dietary resistant starch. Cancer Biol Ther. 2007;6:1621–6. https://doi.org/10.4161/cbt.6.10.4764.
Worthley DL, Whitehall VL, Le Leu RK, Irahara N, Buttenshaw RL, Mallitt KA, Greco SA, Ramsnes I, Winter J, Hu Y, Ogino S, Young GP, Leggett BA. DNA methylation in the rectal mucosa is associated with crypt proliferation and fecal short-chain fatty acids. Dig Dis Sci. 2011;56:387–96. https://doi.org/10.1007/s10620-010-1312-4.
De Marco S, Sichetti M, Muradyan D, et al. Probiotic cell-free supernatants exhibited anti-inflammatory and antioxidant activity on human gut epithelial cells and macrophages stimulated with LPS. Evid Based Complement Alternat Med. 2018;2018:1756308.
Gao J, Li Y, Wan Y, et al. A novel postbiotic from Lactobacillus rhamnosus GG with a beneficial effect on intestinal barrier function. Front Microbiol. 2019;10:477.
Wang Y, Liu L, Moore DJ, et al. An LGG-derived protein promotes IgA production through upregulation of APRIL expression in intestinal epithelial cells. Mucosal Immunol. 2017;10(2):373–84.
Escamilla J, Lane MA, Maitin V. Cell-free supernatants from probiotic Lactobacillus casei and Lactobacillus rhamnosus GG decrease colon cancer cell invasion in vitro. Nutr Cancer. 2012;64(6):871–8.
Belcheva A, Irrazabal T, Robertson SJ, et al. Gut microbial metabolism drives transformation of MSH2-deficient colon epithelial cells. Cell. 2014;158(2):288–99.
Sethi V, Kurtom S, Tarique M, Lavania S, Malchiodi Z, Hellmund L, Zhang L, Sharma U, Giri B, Garg B, Ferrantella A, Vickers SM, Banerjee S, Dawra R, Roy S, Ramakrishnan S, Saluja A, Dudeja V. Gut microbiota promotes tumor growth in mice by modulating immune response. Gastroenterology. 2018;155:33-37.e6. https://doi.org/10.1053/j.gastro.2018.04.001.
Park R, Umar S, Kasi A. Immunotherapy in colorectal cancer: potential of fecal transplant and microbiota-augmented clinical trials. Curr Colorectal Cancer Rep. 2020;16:81–8. https://doi.org/10.1007/s11888-020-00456-1.
Parker KD, Maurya AK, Ibrahim H, Rao S, Hove PR, Kumar D, Kant R, Raina B, Agarwal R, Kuhn KA, Raina K, Ryan EP. Dietary rice bran-modified human gut microbial consortia confers protection against colon carcinogenesis following fecal transfaunation. Biomedicines. 2021. https://doi.org/10.3390/biomedicines9020144.
Bullman S, Pedamallu CS, Sicinska E, et al. Analysis of Fusobacterium persistence and antibiotic response in colorectal cancer. Science. 2017;358(6369):1443–8.
Sanyaolu LN, Oakley NJ, Nurmatov U, Dolwani S, Ahmed H. Antibiotic exposure and the risk of colorectal adenoma and carcinoma: a systematic review and meta-analysis of observational studies. Colorectal Dis. 2020;22:858–70. https://doi.org/10.1111/codi.14921.
Fong W, Li Q, Yu J. Gut microbiota modulation: a novel strategy for prevention and treatment of colorectal cancer. Oncogene. 2020;39(26):4925–43.
Wang F, Yin Q, Chen L, Davis MM. Bifidobacterium can mitigate intestinal immunopathology in the context of CTLA-4 blockade. Proc Natl Acad Sci USA. 2018;115(1):157–61.
Hajishengallis G, Darveau RP, Curtis MA. The keystone-pathogen hypothesis. Nat Rev Microbiol. 2012;10(10):717–25.
Bardou M, Barkun AN, Martel M. Obesity and colorectal cancer. Gut. 2013;62(6):933–47.
Geier MS, Butler RN, Howarth GS. Probiotics, prebiotics and synbiotics: a role in chemoprevention for colorectal cancer. Cancer Biol Ther. 2006;5(10):1265–9.
Fayol-Messaoudi D, Berger CN, Coconnier-Polter MH, Liévin-Le Moal V, Servin AL. pH-, lactic acid-, and non-lactic acid-dependent activities of probiotic Lactobacilli against Salmonella enterica Serovar typhimurium. Appl Environ Microbiol. 2005;71(10):6008–13.
Gillor O, Etzion A, Riley MA. The dual role of bacteriocins as anti- and probiotics. Appl Microbiol Biotechnol. 2008;81(4):591–606.
Gao Z, Guo B, Gao R, Zhu Q, Wu W, Qin H. Probiotics modify human intestinal mucosa-associated microbiota in patients with colorectal cancer. Mol Med Rep. 2015;12:6119–27. https://doi.org/10.3892/mmr.2015.4124.
Sofi F, Dinu M, Pagliai G, Pierre F, Gueraud F, Bowman J, et al. Fecal microbiome as determinant of the effect of diet on colorectal cancer risk: comparison of meat-based versus pesco-vegetarian diets (the MeaTIc study). Trials. 2019;20:688. https://doi.org/10.1186/s13063-019-3801-x.
Chen L, Zou Y, Peng J, et al. Lactobacillus acidophilus suppresses colitis-associated activation of the IL-23/Th17 axis. J Immunol Res. 2015;2015: 909514.
Jeon SG, Kayama H, Ueda Y, et al. Probiotic Bifidobacterium breve induces IL-10-producing Tr1 cells in the colon. PLoS Pathog. 2012;8(5): e1002714.
Konieczna P, Groeger D, Ziegler M, et al. Bifidobacterium infantis 35624 administration induces Foxp3 T regulatory cells in human peripheral blood: potential role for myeloid and plasmacytoid dendritic cells. Gut. 2012;61(3):354–66.
Tojo R, Suárez A, Clemente MG, et al. Intestinal microbiota in health and disease: role of bifidobacteria in gut homeostasis. World J Gastroenterol. 2014;20(41):15163–76.
Gopalakrishnan V, Spencer CN, Nezi L, et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science. 2018;359(6371):97–103.
Routy B, Le Chatelier E, Derosa L, et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science. 2018;359(6371):91–7.
Sivan A, Corrales L, Hubert N, et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science. 2015;350(6264):1084–9.
Khazaie K, Zadeh M, Khan MW, et al. Abating colon cancer polyposis by Lactobacillus acidophilus deficient in lipoteichoic acid. Proc Natl Acad Sci USA. 2012;109(26):10462–7.
Smits LP, Bouter KE, de Vos WM, Borody TJ, Nieuwdorp M. Therapeutic potential of fecal microbiota transplantation. Gastroenterology. 2013;145(5):946–53.
Iida N, Dzutsev A, Stewart CA, Smith L, Bouladoux N, Weingarten RA, Molina DA, Salcedo R, Back T, Cramer S, Dai RM, Kiu H, Cardone M, Naik S, Patri AK, Wang E, Marincola FM, Frank KM, Belkaid Y, Trinchieri G, Goldszmid RS. Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science. 2013;342:967–70. https://doi.org/10.1126/science.1240527.
Wang Z, Hua W, Li C, Chang H, Liu R, Ni Y, Sun H, Li Y, Wang X, Hou M, Liu Y, Xu Z, Ji M. Protective role of fecal microbiota transplantation on colitis and colitis-associated colon cancer in mice is associated with Treg cells. Front Microbiol. 2019;10:2498. https://doi.org/10.3389/fmicb.2019.02498.
Baxter NT, Zackular JP, Chen GY, Schloss PD. Structure of the gut microbiome following colonization with human feces determines colonic tumor burden. Microbiome. 2014;2:20.
Zackular JP, Baxter NT, Iverson KD, et al. The gut microbiome modulates colon tumorigenesis. mBio. 2013;4(6):e00692-13.
Khoruts A, Can FMT. Cause or prevent CRC? Maybe, but there is more to consider. Gastroenterology. 2021;161:1103–5. https://doi.org/10.1053/j.gastro.2021.06.074.
Nooij S, Ducarmon QR, Laros J, Zwittink RD, Norman JM, Smits WK, Verspaget HW, Keller JJ, Terveer EM, Kuijper EJ. Fecal microbiota transplantation influences procarcinogenic Escherichia coli in recipient recurrent Clostridioides difficile patients. Gastroenterology. 2021;161:1218-1228.e5. https://doi.org/10.1053/j.gastro.2021.06.009.
Wilson MR, Jiang Y, Villalta PW, Stornetta A, Boudreau PD, Carrá A, Brennan CA, Chun E, Ngo L, Samson LD, Engelward BP, Garrett WS, Balbo S, Balskus EP. The human gut bacterial genotoxin colibactin alkylates DNA. Science. 2019. https://doi.org/10.1126/science.aar7785.
Suehiro Y, Zhang Y, Hashimoto S, et al. Highly sensitive faecal DNA testing of TWIST1 methylation in combination with faecal immunochemical test for haemoglobin is a promising marker for detection of colorectal neoplasia. Ann Clin Biochem. 2018;55(1):59–68.
Drewes JL, Corona A, Sanchez U, Fan Y, Hourigan SK, Weidner M, Sidhu SD, Simner PJ, Wang H, Timp W, Oliva-Hemker M, Sears CL. Transmission and clearance of potential procarcinogenic bacteria during fecal microbiota transplantation for recurrent Clostridioides difficile. JCI Insight. 2019. https://doi.org/10.1172/jci.insight.130848.
Rooks MG, Garrett WS. Gut microbiota, metabolites and host immunity. Nat Rev Immunol. 2016;16:341–52. https://doi.org/10.1038/nri.2016.42.
Li Q, Peng W, Wu J, et al. Autoinducer-2 of gut microbiota, a potential novel marker for human colorectal cancer, is associated with the activation of TNFSF9 signaling in macrophages. Oncoimmunology. 2019;8(10): e1626192.
Wang HF, Li LF, Guo SH, et al. Evaluation of antibody level against Fusobacterium nucleatum in the serological diagnosis of colorectal cancer. Sci Rep. 2016;6:33440.
Purcell RV, Pearson J, Aitchison A, Dixon L, Frizelle FA, Keenan JI. Colonization with enterotoxigenic Bacteroides fragilis is associated with early-stage colorectal neoplasia. PLoS ONE. 2017;12: e0171602. https://doi.org/10.1371/journal.pone.0171602.
Acknowledgements
This work was supported by the Ministry of Science and Technology of the People’s Republic of China (No. 2017YFC0908204), Department of Science and Technology of Sichuan Province (No. 2021YFS0025), 1.3.5 project for disciplines of excellence, West China Hospital, Sichuan University (No. 20HXJS003), 1·3·5 project for disciplines of excellence-Clinical Research Incubation Project, and West China Hospital, Sichuan University (No. 2019HXFH031).
Author information
Authors and Affiliations
Contributions
JL and YZ designed the review and wrote the manuscript. ZW and LY reviewed the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
All authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, J., Zhu, Y., Yang, L. et al. Effect of gut microbiota in the colorectal cancer and potential target therapy. Discov Onc 13, 51 (2022). https://doi.org/10.1007/s12672-022-00517-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-022-00517-x