Abstract
Purpose
MUC1 is a membrane bound protein that can regulate tumor progression but its role in tumor metastasis and the metastatic microenvironment remains unclear.
Methods
We performed differential gene analysis for primary liver cancer (n = 31) and lung metastases (n = 31) using the Gene Expression Omnibus (GEO) dataset (GSE141016) and obtained RNA sequencing data from 374 liver cancer and 50 normal tissues from The Cancer Genome Atlas (TCGA). We analyzed the prognostic value of MUC1 and the relationship between MUC1 and the TME using online databases and a clinical cohort. Immunohistochemistry detected MUC1 in normal liver, liver cancer, and lung metastases. Multiplex immunohistochemistry staining detected immune cells in the metastatic microenvironment.
Results
High MUC1 expression levels in hepatocellular carcinoma are associated with worse clinical prognosis and higher rates of lung metastasis. In addition, we observed a correlation between MUC1 and multiple immune cells in the metastatic microenvironment. In paired primary liver cancer and lung metastatic tumor tissues from the same patient, we observed higher MUC1 protein levels in lung metastases than in primary liver cancer. Furthermore, MUC1 was negatively correlated with CD8+T and Treg cells in the metastatic tumor microenvironment and positively correlated with DC. In addition, we found that MUC1 was associated with CD8+T cell activation and function using flow cytometry in another cohort of patients with liver cancer.
Conclusion
These data confirm the potential of MUC1 as a prognostic marker and therapeutic target.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Hepatocellular carcinoma (HCC) is one of the most malignant cancers with high incidence and morbidity rates [1]. Despite tremendous advances in clinical treatment over the last decades, HCC remains a leading cause of cancer-related mortality worldwide; metastasis, especially lung metastasis, is largely responsible for the poor prognosis. Metastatic TME is important for the initiation of primary tumor metastasis. However, the mechanism by which tumor cells reshape the pre-metastatic microenvironment remains unclear.
The metastatic TME consists of a variety of cellular and extracellular matrix components, of which immune cells are important components [2]. Cytotoxic CD8+tumor-infiltrating lymphocytes (TILs) can effectively kill tumor cells, but inhibitory immune cells such as myeloid-derived suppressor cells (MDSC) and regulatory T cells (Tregs) may contribute to maintaining a suppressive immune microenvironment that protects tumor cells from immune destruction [3, 4]. In addition, activation of immune checkpoints, such as the PD-L1/PD1 axis (PD1 is a T-cell co-suppressor molecule and checkpoint protein), also causes loss of CD8+TILs effector function; this can lead to CD8+T cell depletion and tumor evasion of the immune system, leading to tumor progression and metastasis [5, 6]. However, little is known about how tumor cells influence CD8+TILs function in the metastatic tumor microenvironment.
Mucin 1 (MUC1) is a membrane-bound protein in the mucin family [7]. MUC1 is normally overexpressed in many types of epithelial adenocarcinomas such as lung, colon, breast, and ovarian cancers [8]. MUC1 can regulate tumor progression through a variety of mechanisms, including immune response, tumor invasion, tumor cell proliferation and apoptosis, and angiogenesis [9, 10]. MUC1 induces apoptosis of activated T cells, leading to the inhibition of human T cell proliferation and cytotoxicity [11, 12]. Moreover, MUC1 binds siglecs and blocks toll-like receptors (TLRS) on DC, promoting immature DC and thereby reducing T cell function. MUC1 interacts with neutrophils and macrophages to protect cancer cells during metastasis [13, 14]. Abnormally high expression of MUC1 on cancer cells causes physical barriers, blocking the detection of tumor-associated antigens (TAAs) and preventing immune cells from killing tumor cells [7]. However, the effects of MUC1 on the liver cancer lung metastasis have not yet been investigated.
In this study, we confirmed the clinical significance of MUC1 in patients with HCC and analyzed the relationship between MUC1 and hepatocellular carcinoma lung metastasis. We investigated the regulation of MUC1 in immune cells in the metastatic TME. We also revealed the potential mechanism by which hepatocellular carcinoma cells upregulate MUC1 and affect the function of CD8+TILs to promote hepatocellular carcinoma lung metastasis. Our study provides new therapeutic targets for HCC treatment.
2 Materials and methods
2.1 Human tissues
A tumor microarray (TMA) from HCC patients (n = 80) with follow-up information was used for analysis. Specimens of primary liver tumors and lung metastases from the same patient (n = 17) were obtained from Shanghai Renji Hospital. These were provided with informed patient consent and approval from the ethics review committees of Shanghai Renji Hospital, Shanghai Jiao Tong University.
2.2 Online data acquisition and analyses
We performed differential gene analysis for primary liver cancer (n = 31) and lung metastases (n = 31) using the Gene Expression Omnibus (GEO) dataset (GSE141016). We obtained RNA sequencing data from 374 liver cancer and 50 normal tissues from The Cancer Genome Atlas (TCGA). Based on the expression level of MUC1, hepatocellular carcinoma patients were divided into MUC1 high and low groups, and Overall Survival (OS) and Progression Free Survival (PFS) were analyzed separately. To assess the abundance of immune cell infiltration, gene expression profiles were used to assess the abundance of immune cell infiltration in tumor using ESTIMATE, and the ESTIMATE scores, immune scores, and stromal scores were calculated. The relationship between MUC1 and immune cell gene expression was calculated using the ssGSEA algorithm. As positive gene sets, gene sets with standardized p < 0.05 and false discovery rate p < 0.05 were accepted.
2.3 Immunohistochemistry (IHC) staining
The assay was performed according to the procedure description previously [15]. The primary anti-MUC1 (Abcam, Cambridge, UK) antibody (1:200 dilution) was used for IHC staining.
2.4 Multiple immunohistochemistry (mIHC) staining
We performed the fluorescent dye experiments according to the procedure description previously [16] by using the CD11c, CD86, CD206, CD66b, CD14, CD11b, CD19, CD8, CD4, Foxp3, and CD56 anti-human antibodies (Abcam). Using the PerkinElmer Vectra3® platform, we scanned the slides and quantified the results.
2.5 Flow cytometry
Flow cytometry experiments were performed according to the procedure described earlier [15]. In particular, cells suspensions were incubated with APC-Cy7 live/dead (BD, 1:100), PC5.5 anti-human CD4 (BD, 1:100), BV421 anti-human TNF-α (BD, 1:100), BV605 anti-human CD8 (BD, 1:100), BV650 anti-human IFNγ (BD, 1:100), APC anti-human IL2 (BD, 1:100), AF700 anti-human GZMB (BD, 1:100), PE anti-human CD69 (BD, 1:100), and BV510 anti-human CD3 (BD, 1:100) antibodies. Flow cytometry was performed using a BD Fortessa FACS with Diva software v6.0. and results were quantified using FlowJo V10 software.
2.6 Statistical analyses
Analyses were performed using GraphPad Prism 7.0 (La Jolla, CA, USA). We compared the differences between the two groups using Student's t-tests. Pearson's correlation test was used to analyze correlations, and the t-test was used to perform univariate analysis. The log-rank test was used for follow-up data, and p values < 0.05 were considered statistically significant.
3 Results
3.1 MUC1 is associated with a worse clinical outcome
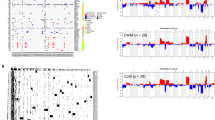
To identify the key genes associated with lung metastases from HCC, we performed differential gene analysis of primary liver cancer and lung metastases using the Gene Expression Omnibus (GEO) dataset (GSE141016) (Fig. 1a). The results revealed that MUC1 was expressed differentially between the two groups and prominently upregulated in lung metastases. To further investigate the reliability of the correlation between MUC1 upregulation and poor prognosis, we explored MUC1 expression levels in the liver hepatocellular carcinoma (LIHC) dataset from TCGA database and found that high MUC1 levels were related to shorter overall survival (OS) and progression-free survival (PFS) (Fig. 1b). We analyzed an HCC cohort and consistently found that patients with high MUC1 levels were prone to worse overall survival (P = 0.0305), metastasis-free survival (P = 0.0349), and lung metastasis-free survival (P = 0.0325) (Fig. 1c). In addition, the expression level of MUC1 in tumor tissue was higher than that in normal liver tissue (P = 0.0462) according to TCGA and GTEx data (Additional file 1: Supplementary Fig. 1a). However, in a clinical cohort of HCC patients, the protein levels of MUC1 were detected using immunohistochemical (IHC) staining and were also not statistically significantly different between HCC and normal liver tissue. (Additional file 1: Supplementary Fig. 1b).
MUC1 associates with a worse clinical outcome. a Volcano plots of differential gene expression in 31 patients with paired primary tumors and lung metastases. Blue and red dots represent downregulated and upregulated genes, respectively. b MUC1 expression analysis of Overall survival and progress free interval of MUC1 high group and MUC1 low group from LIHC of The Cancer Genome Atlas (TCGA). c MUC1 expression analysis of Overall survival, met-free survival and lung met-free survival in a liver cancer cohort from Shanghai Renji Hospital (n = 80). *P < 0.05, **P < 0.01, ***P < 0.001, by log rank test
3.2 MUC1 level is significantly correlated with liver cancer lung metastasis
To investigate the relationship between MUC1 and lung metastasis in HCC, we detected the protein levels of MUC1 in patients with no metastasis, patients with extrahepatic metastasis, and patients with lung metastasis in the HCC cohort by IHC assay. We found that MUC1 was significantly upregulated in patients with extrahepatic metastasis and patients with lung metastasis compared to patients without metastasis (Fig. 2a, b). The percentages of MUC1 high and medium groups were significantly higher in patients with extrahepatic metastasis and lung metastasis than in patients without metastasis (Fig. 2c). We found no statistically significant correlation between MUC1 expression and tumor size and recurrence rate, while extrahepatic metastasis and lung metastasis rates were higher in the MUC1 high group than in the MUC1 low group (Fig. 2d). In addition, we analyzed the MUC1 protein levels in primary neoplasms and lung metastases of HCC from the same patient by immunohistochemical staining and found that the MUC1 protein levels in lung metastases were remarkably higher than those in primary HCC (Fig. 2e, f).
MUC1 expression is significantly correlated with liver cancer lung metastasis. a–c MUC1 expression level in non-metastasis (n = 69), extrahepatic metastasis(n = 11) and lung metastasis (n = 7). (a) Representative pictures of MUC1 immunohistochemistry staining. b Composition of MUC1 expression level in non-metastasis, extrahepatic metastasis and lung metastasis. c Comparison of MUC1 between non-metastasis and extrahepatic metastasis, non-metastasis and lung metastasis, respectively. d Clinical events of MUC1 high group (n = 40) and MUC1 low group (n = 40). e, f Comparison of MUC1 between paired primary tumors and lung metastases. e Representative pictures of MUC1 immunohistochemistry staining and f statistics chart. Scale bars, 50 μm. *P < 0.05, **P < 0.01, ***P < 0.001, by chi-squared test (d) and two-tailed unpaired (c) or paired (f) t test
3.3 MUC1 is associated with the TME
Next, to investigate whether MUC1 regulates tumor metastasis by reprogramming the immune microenvironment, we calculated the ESTIMATE score, immune score, and stromal score according to LIHC from TCGA database, and found that the ESTIMATE score, immune score, and stromal score of the MUC1 high group were higher than those of the MUC1 low group (Fig. 3a). To further explore the effect of MUC1 on the TME, we analyzed the correlation between MUC1 and immune cells using TCGA database; we found that the MUC1 high group had more T cells, NK cells, mast cells, macrophages, CD8 + T, B cells, Tem, TFH, Th1 cells, and Th2 cells and lower Th17 cells compared to the MUC1 low group (Fig. 3b). MUC1 was positively correlated with macrophages, T cells, DC, NK cells, mast cells, CD8 + T cells, B cells, Tem, TFH, Th1, and Th2 cells, and negatively correlated with Tem, Treg, and Th17 cells (Fig. 3c). Furthermore, we analyzed the data using the ssGSEA algorithm which showed that MUC1 was positively correlated with T cells (r = 0.188, p < 0.001), NK cells (r = 0.200, p < 0.001), mast cells(r = 0.124, p < 0.001), macrophages (r = 0.348, p < 0.001), CD8 + T cells (r = 0.144, p = 0.005), B cells (r = 0.199, p < 0.001), Tem (r = 0.296, p < 0.001), TFH (r = 0.339, p < 0.001), Th1 cells (r = 0.285, p < 0.001), and Th2 cells (r = 0.289, p < 0.001), and negatively correlated with Th17 (r = − 0.195, p < 0.001) cells and Tregs (r = − 0.008, p = 0, 090) (Fig. 3d and Additional file 1: Supplementary Fig. 2a).
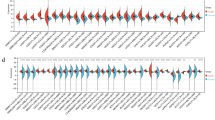
MUC1 associates with tumor immune microenvironment. a Violin plots show ESTIMATE score, Immune score and Stromal score of MUC1 high and low group from TCGA. (b-d) Analyses of MUC1 expression and immune infiltration in LIHC of TCGA. b Enrichment score of infiltration profile of immune subsets based on MUC1 expression levels from TCGA using ssGSEA. c Lollipop plots show correlation between MUC1 and immune infiltration in LIHC of TCGA using ssGSEA. d Correlation analysis of MUC1 and immune cells from TCGA using ssGSEA. *P < 0.05, **P < 0.01, ***P < 0.001, by Pearson correlation analysis
3.4 Association between MUC1 and TILs in lung metastases
To further explore the relationship between MUC1 and immune cells in the TME, we examined metastasis-associated immune cells in paired primary tumor and lung metastasis tissues from the same patient using Fluorescent Multiplex Immunohistochemistry (Fig. 4a–c). In addition, we analyzed the correlation between MUC1 and individual immune cells and showed that MUC1 was positively correlated with DC (r = 0.5439, p = 0.0009) and negatively correlated with Tregs (r = − 0.3464, p = 0.0447) and CD8 + T cells (r = − 0.3757, p = 0.0285) (Fig. 4d and Additional file 1: Supplementary Fig. 3a).
Association of MUC1 and TILs in lung metastasis. a–d Analyses of MUC1 expression and immune infiltration in paired primary tumors and lung metastases form the same patients (n = 17) by multi-immunohistochemistry (mIHC) staining of metastasis-associated lymphatic and myeloid immune cells. Representative mIHC staining images of metastasis-associated a lymphatic and b myeloid immune cells. Scale bars, 50 μm. c Dot plots displaying MUC1 expression level and altered immune cells in paired primary tumors and lung metastases. d Correlation analyses between MUC1 and metastasis-associated immune cells. *P < 0.05, **P < 0.01, ***P < 0.001, by Pearson correlation analysis
3.5 Correlation of MUC1 with immune checkpoints in the TME
To investigate the mechanism of MUC1 effect on T cells in the TME, we analyzed the relationship between MUC1 and immune checkpoints using TCGA database and found that MUC1 was positively correlated with CD274 (PDL1), HAVCR2 (TIM-3), CD226, and CD47. Among them, MUC1 was most strongly correlated with CD274 (PDL1) (r = 0.221, p < 0.001) (Fig. 5a, b). Next, we explored the relationship between MUC1 and PDL1 at the protein level. Immunohistochemical staining showed that PDL1 levels were higher in the lung metastasis group of HCC patients than in HCC patients without extrahepatic metastasis, and MUC1 was positively correlated with PDL1 (r = 0.4077, P = 0.0002) (Fig. 5c-e). In addition, we performed immunohistochemical staining of paired primary tumor and lung metastasis tissue sections from the same patient and found that PDL1 levels were higher in lung metastases than in primary tumors, and that MUC1 was positively correlated with PDL1 (r = 0.3919, P = 0.0219) (Fig. 5f-h).
Correlation of MUC1 with immune checkpoints in tumor immune microenvironment. a, b Correlation analysis of MUC1 expression with immune checkpoints in LIHC of TCGA. c–e Analysis of MUC1 and PDL1 in HCC patients. c Representative pictures of PDL1 IHC staining. d Comparisons of PDL1 level in non-metastatic with metastatic and lung metastatic tumors. e Correlation analysis of MUC1 expression with PDL1 level. f–h Analysis of MUC1 and PDL1 in primary and lung metastatic tumors from the same patient. f Representative pictures of PDL1 IHC staining. g Comparisons of PDL1 level in primary and lung metastatic tumors from the same patient. h Correlation analysis of MUC1 expression with PDL1 level. Scale bars, 50 μm. *P < 0.05, **P < 0.01, ***P < 0.001, by two-tailed unpaired or paired t test (d, g) and Pearson correlation analysis (b, e and h)
3.6 Association between MUC1 expression and CD8 + T cell function and immune checkpoints
To further explore the relationship between MUC1 and T cells in the metastatic TME, we examined CD8+T cell function and immune checkpoints using flow cytometry in a cohort of HCC patients. The patients were divided into MUC1 high and MUC1 low groups based on MUC1 immunohistochemical staining. Figure 6a–c showed that HCC patients in the MUC1 high group had lower levels of the T cell activation marker CD69 in CD8+T cells compared to those in the MUC1 low group. Furthermore, patients in the MUC1 high group had significantly lower levels of the CD8+T function marker granzyme B than those in the MUC1 low group.
Association between MUC1 expression and CD8+T cell function and immune checkpoints. a Representative flow cytometry charts of CD8+T cell function and immune checkpoints based on MUC1 expression. b Statistical map of CD8+T cell immune checkpoints. c Statistical map of CD8+T cell function. *P < 0.05, **P < 0.01, ***P < 0.001, by two-tailed unpaired t test (b, c)
4 Discussion
Tumor metastasis is a complex process that involves multiple cellular and extracellular components [17]. In addition to intrinsic tumor cell characteristics, metastatic TME influences tumor metastasis [18, 19]. Tumor cells and the metastatic TME interact and dynamically change with each other. It has been reported that primary tumors can remodel the pre-metastatic ecological niche before metastasis, making it favorable for tumor cells to metastasize [20]. We identified MUC1 expression levels were increased in hepatocellular carcinoma tissues relative to normal liver tissues from TCGA and GTEx data. However, the protein levels of MUC1 were not detected statistically significantly different in a clinical cohort of HCC patients. This may be because the sample size of the clinical cohort was not large enough. We also found that patients with high MUC1 expression levels were more likely to develop lung metastases and that MUC1 expression levels were significantly higher in lung metastases than in primary liver cancer. The reasons for the increased expression of MUC1 in HCC require further research. We analyzed the correlation between MUC1 and multiple immune cells and found that MUC1 expression was positively correlated with macrophages, T cells, DC, NK, mast cells, CD8+T, B cells, Tem, TFH, Th1 cells and Th2 cells, and negatively correlated with Th17 cells, Tem and Tregs. These findings suggest that HCC may promote lung metastasis by remodeling the metastatic microenvironment through MUC1.
The interaction between immune and tumor cells in the TME plays a key role in controlling tumor progression [21, 22]. CD8+TILs are key effector cells that directly kill tumor cells [23], and the abundance of CD8+TILs is positively correlated with clinical prognosis [24], while the T cell co-suppressor molecule PD-L1 and PD1 receptor are associated with worse clinical prognosis [25]. Recent studies have found that MUC1 is expressed on human T cells and may be involved in the regulation of T cell function [26]. A variety of factors contribute to the loss of TILs effector function, leading to tumor evasion by the immune system and tumor progression and metastasis [27, 28]. To investigate how MUC1 affects T cells in the TME. Using immunofluorescence staining of paired primary liver cancer and lung metastatic tumor tissues from the same patient, we found that MUC1 expression was negatively correlated with CD8+T and Treg cells in the metastatic TME and positively correlated with DC cells. We analyzed the relationship between MUC1 and immune checkpoints using TCGA database and found that MUC1 was positively correlated with CD274. Consistent with this, in another cohort of liver cancer patients, we used flow cytometry and found higher levels of CD69+CD8+T cells and GZMB of CD8+T in the MUC1 high group, but did not find a statistical difference in CD8+T cells which may be related to sample size. According to these results, abnormally high expression level of MUC1 in liver cancer cells may inhibit the function of CD8+TILs through the high expression of PDL1, forming an inhibitory immune microenvironment, resulting in metastasis of cancer cells. Further studies are needed to clarify whether MUC1 affects other immune cells and their roles.
In this study, we reported that high MUC1 expression levels in HCC are associated with worse clinical prognoses and higher rates of lung metastasis. Multiple immune cells were found to be associated with MUC1 expression. In paired primary liver cancer and lung metastatic tumor tissues from the same patient, we observed higher MUC1 protein levels in lung metastases than in primary liver cancer, and MUC1 was negatively correlated with CD8+T and Treg cells in the metastatic TME and positively correlated with DC. In addition, in another cohort of liver cancer patients, we found that MUC1 was associated with CD8+T cell activation and function using flow cytometry. This suggests that tumors may regulate CD8+T cell function in the metastatic TME through MUC1, thereby affecting the HCC lung metastasis process. These data confirm the potential of MUC1 as a prognostic marker and therapeutic target. This will contribute to the development of new strategies for the treatment of tumor metastasis.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Jemal A, et al. Cancer statistics, 2010. CA: a Cancer J Clin. 2010;60(5):277–300.
Quail DF, Joyce JA. Microenvironmental regulation of tumor progression and metastasis. Nat Med. 2013;19(11):1423–37.
Hegde S, Leader AM, Merad M. MDSC: Markers, development, states, and unaddressed complexity. Immunity. 2021;54(5):875–84.
Wing JB, Tanaka A, Sakaguchi S. Human FOXP3 regulatory T cell heterogeneity and function in autoimmunity and cancer. Immunity. 2019;50(2):302–16.
Galluzzi L, et al. Immunostimulation with chemotherapy in the era of immune checkpoint inhibitors. Nat Rev Clin Oncol. 2020;17(12):725–41.
Cervantes-Villagrana RD, et al. Tumor-induced neurogenesis and immune evasion as targets of innovative anti-cancer therapies. Signal Transduct Target Ther. 2020;5(1):99.
Bose M, et al. Targeting tumor-associated MUC1 overcomes anoikis-resistance in pancreatic cancer. Transl Res: the J Lab Clinical Med. 2022. https://doi.org/10.1016/j.trsl.2022.08.010.
Kerckhoffs KGP, et al. Mucin expression in gastric- and gastro-oesophageal signet-ring cell cancer: results from a comprehensive literature review and a large cohort study of Caucasian and Asian gastric cancer. Gastric Cancer: Official J Int Gastric Cancer Association Japanese Gastric Cancer Association. 2020;23(5):765–79.
Maeda T, et al. MUC1-C induces PD-L1 and immune evasion in triple-negative breast cancer. Can Res. 2018;78(1):205–15.
Bose M, et al. Overexpression of MUC1 induces non-canonical TGF-β signaling in pancreatic ductal adenocarcinoma. Front Cell Dev Biol. 2022;10: 821875.
Jiang Z-B, et al. Evodiamine suppresses non-small cell lung cancer by elevating CD8 T cells and downregulating the MUC1-C/PD-L1 axis. J Exp Clin Cancer Res: CR. 2020;39(1):249.
Bouillez A, et al. MUC1-C integrates PD-L1 induction with repression of immune effectors in non-small-cell lung cancer. Oncogene. 2017;36(28):4037–46.
Bhatia R, et al. Cancer-associated mucins: role in immune modulation and metastasis. Cancer Metastasis Rev. 2019;38(1–2):223–36.
Sahraei M, et al. Repression of MUC1 promotes expansion and suppressive function of myeloid-derived suppressor cells in pancreatic and breast cancer murine models. Int J Mol Sci. 2021. https://doi.org/10.3390/ijms22115587.
Yin Y, et al. Aurora kinase A regulates liver regeneration through macrophages polarization and Wnt/β-catenin signalling. Liver Intern: Official Journal of the International Association For the Study of the Liver. 2022;42(2):468–78.
Taube JM, et al. The Society for Immunotherapy of Cancer statement on best practices for multiplex immunohistochemistry (IHC) and immunofluorescence (IF) staining and validation. J Immunother Cancer. 2020. https://doi.org/10.1136/jitc-2019-000155corr1.
Lin Y, Xu J, Lan H. Tumor-associated macrophages in tumor metastasis: biological roles and clinical therapeutic applications. J Hematol Oncol. 2019;12(1):76.
Fares J, et al. Molecular principles of metastasis: a hallmark of cancer revisited. Signal Transduct Target Ther. 2020;5(1):28.
Bakir B, et al. EMT, MET, plasticity, and tumor metastasis. Trends Cell Biol. 2020;30(10):764–76.
Mohan V, Das A, Sagi I. Emerging roles of ECM remodeling processes in cancer. Semin Cancer Biol. 2020;62:192–200.
Grzelak CA, et al. Elimination of fluorescent protein immunogenicity permits modeling of metastasis in immune-competent settings. Cancer Cell. 2022;40(1):1–2.
Sun L, et al. Activating a collaborative innate-adaptive immune response to control metastasis. Cancer Cell. 2021. https://doi.org/10.1016/j.ccell.2021.08.005.
Simoni Y, et al. Bystander CD8 T cells are abundant and phenotypically distinct in human tumour infiltrates. Nature. 2018;557(7706):575–9.
Mazzaschi G, et al. Low PD-1 expression in cytotoxic CD8 tumor-infiltrating lymphocytes confers an immune-privileged tissue microenvironment in NSCLC with a prognostic and predictive value. Clin Cancer Res: an Official J Am Assoc Cancer Res. 2018;24(2):407–19.
Champiat S, et al. Hyperprogressive disease is a new pattern of progression in cancer patients treated by anti-PD-1/PD-L1. Clin Cancer Res: an Official Journal of the American Association For Cancer Research. 2017;23(8):1920–8.
Agrawal B, Gupta N, Konowalchuk JD. MUC1 mucin: a putative regulatory (Checkpoint) molecule of T cells. Front Immunol. 2018;9:2391.
Saleh R, Elkord E. Acquired resistance to cancer immunotherapy: role of tumor-mediated immunosuppression. Semin Cancer Biol. 2020;65:13–27.
Weisshaar N, et al. Rgs16 promotes antitumor CD8 T cell exhaustion. Sci Immunol. 2022;7(71):1873.
Acknowledgements
We sincerely thank all colleagues in the Department of Liver Surgery, Renji Hospital, School of Medicine for their help in this study.
Funding
This study was supported by the Project of the Shanghai Municipal Health Commission (20204Y0012), Seed Fund of Renji Hospital (RJZZ18-010), Shenkang three-year action plan (SHDC2020CR2003A, SHDC2020CR5012), Innovative Research Team of High-Level Local Universities in Shanghai (SSMU-ZDCX20180802), National Natural Science Foundation of China (81972205), and the Project of Shanghai key clinical specialties (shslczdzk05801).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by [Yanze Yin], [Changjie Yang] and [Jiafeng Xu]. The first draft of the manuscript was written by [Yanze Yin] and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Informed consent was obtained from all individual participants included in the study.This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Ethics Committee of Shanghai Renji hospital, Shanghai Jiao Tong University (No:KY2020-190).
Consent for publication
The authors affirm that human research participants provided informed consent for publication of the images in all Figure(s).
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Fig S1.
MUC1 expression level in liver cancer. (A, B) MUC1 expression level analysis of HCC (A) patients from LIHC of TCGA and GTEx, and (B) patients from Shanghai Renji Hospital (n=80). Scale bars, 50 μm. Fig S2. MUC1 associates with tumor immune microenvironment. (A) Correlation analysis of MUC1 and immune cells from TCGA using ssGSEA. Fig S3. Correlation of MUC1 and immune cells in lung metastasis. (A) Correlation analyses between MUC1 and metastasis-associated immune cells.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yin, Y., Yang, C., Xu, J. et al. MUC1 promotes lung metastases of liver cancer by impairing anti-tumor immunity. Discov Onc 14, 18 (2023). https://doi.org/10.1007/s12672-023-00627-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-023-00627-0