Abstract
Oral squamous cell carcinoma (OSCC) is the sixth most common malignancy worldwide. Abnormal epigenetic modifications, including DNA methylation, are hallmarks of cancer and implicated in the development of various tumors. DNA methylation is catalyzed by the DNA methyltransferase and ten-eleven translocation dioxygenase families, with DNMT3A and TET2 being the most widely studied members, respectively. The correlation of methylation β values and clinical features was conducted in patients with OSCC in The Cancer Genome Atlas database. DNA methylation and protein expression levels of DNMT3A and TET2 in tissues were analyzed with methylation-specific polymerase chain reaction (MSP) and western blotting. To evaluate the effects of DNMT3A and TET2 on the biological characteristics of OSCC, cell proliferation was assessed with 5-ethynyl-2'-deoxyuridine, and cell migration capacity was quantified with wound healing and transwell assays. A survival analysis was performed with the Kaplan–Meier approach. The correlation between different methylation β values and clinical features was revealed. MSP revealed varying methylation degrees of DNMT3A and TET2 in OSCC tissues. Furthermore, western blotting showed that the protein expression levels were significantly different in cancer and surrounding healthy tissue samples. In vitro experiments demonstrated that DNMT3A knockdown and TET2 overexpression could inhibit the proliferation and migration of OSCC. Survival analysis revealed that patients with high DNMT3A methylation levels showed higher survival rates.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Cancer is a global health burden because of the lack of prevention and treatment options. Oral squamous cell carcinoma (OSCC) is a common malignancy, with high rates of metastasis, recurrence, and resistance to conventional chemotherapy. Every year, nearly 400,000 new cases and 200,000 deaths [1] are attributed to OSCC, showing a poor prognosis for most affected patients.
Abnormal epigenetic modification is a hallmark of cancer. Epigenetic modification factors, particularly DNA methylation, affect tumor occurrence, development, invasion, metastasis, and treatment resistance [2]. By changing the functional state of the regulatory regions without altering the DNA sequence, DNA methylation can significantly affect the gene expression. For example, hypermethylation of the gene promoter regions may lead to transcriptional silencing of certain tumor suppressors, while loss of methylation in the promoter regions may lead to decreased genomic stability and potentially activate viral sequences in the genome [3]. DNA methylation patterns are established through a coordinated mechanism catalyzed by various enzymes, including the DNA methyltransferase (DNMT) and ten-eleven translocation (TET) dioxygenase families, with DNMT3A and TET2 being the most extensively studied members, respectively. DNMT3A is an enzyme responsible for the de novo methylation of cytosine residues and a modification frequently associated with gene silencing. TET2 is a demethylase enzyme that initiates a series of cytosine demethylation reactions, which perform antagonistic enzymatic activity.
DNA methylation is a crucial regulatory mechanism in the development and maintenance of acute myeloid leukemia, and DNMT3A mutations are implicated in various hematological tumors. Furthermore, DNMT3A is associated with chemical carcinogenicity and can lead to double-stranded DNA damage [4]. It is also involved in the amino acid metabolism. Methionine dependence is a hallmark of cancer. DNMT3A is a key gene in the methionine metabolism and predicts a poor cancer prognosis [5]. In addition, it is closely related to biological processes [6,7,8,9] and signaling pathways [10,11,12].
The demethylase TET2 is a regulator of normal hematopoiesis, particularly bone marrow cell production [13,14,15]. In addition to frequent mutations in various hematological tumors, TET2 is also associated with immune and inflammatory responses. TET2 plays a vital role in various inflammatory conditions by regulating the immune response [16] and expressions of signaling networks and effectors during the initiation and regression of inflammation [17,18,19]. Simultaneously, TET2 mutations occur in human solid tumors, and a reduced TET2 protein expression is prevalent in several types of cancers [20,21,22,23,24,25].
Although the pathogenic roles of DNMT3A and TET2 in various tumors have been well-documented, their role in OSCC has rarely been studied. Some studies have shown their expressions in OSCC tissues [26,27,28,29,30]; however, their effects on the proliferation or migration of OSCC remain unclear. The aim of the present study was to determine whether or not DNMT3A and TET2 could serve as latent biomarkers for the diagnosis and prognosis of OSCC.
2 Material and methods
2.1 Patients and methylation analysis
Clinical information and methylation chip data of 399 patients with OSCC was obtained from The Cancer Genome Atlas (TCGA) database (https://portal.gdc.cancer.gov/). The methylation β values of DNMT3A and TET2 genes under different clinical characteristics were extracted. The Kolmogorov–Smirnov test were used to assess the mean methylation differences of all probes in each sample under different clinical settings and the methylation differences of each probe in different regions.
2.2 Cell line and culture
OSCC cell line Cal27 was obtained from Procell and maintained in Dulbecco’s Modified Eagle Medium (DMEM; Gibco) supplemented with 10% fetal bovine serum (Gibco) and 1% penicillin_streptomycin at 37 °C under 5% CO2. The cells were subcultured using 0.25% trypsin at 80–90% confluency, and the medium was changed every 2 days.
2.3 Tumor and surrounding healthy tissues
29 primary tumor and surrounding healthy tissue samples were obtained from the First Affiliated Hospital of Harbin Medical University. Qualified pathologists certified all tissues and determined the OSCC diagnosis. Informed consent was obtained from all patients. The study protocol was approved by the institutional ethics committee of the hospital. The study procedures were performed in accordance with the tenets of the Declaration of Helsinki. After collection, all the tissues were immediately preserved in liquid nitrogen for future use.
2.4 Primer design
Primer3 was used to design primers for reverse-transcription polymerase chain reaction (RT-PCR). For the methylation-specific polymerase chain reaction (MSP), we used MethPrimer to design primer sequences.
2.5 DNA extraction, bisulfite conversion, and MSP
The QIAamp DNA Mini Kit (Qiagen) was utilized to obtain genomic DNA from 29 fresh primary tumor tissues. The EZ DNA Methylation-Gold Kit (Zymo Research) was used for bisulfite conversion to detect the methylation status. In MSP, every 20 μL of the reaction mixture contained 2 μL of DNA, and amplification was performed using ABI 7500 (Thermo Fisher Scientific). PCR involved an initial denaturation step at 95 °C for 10 min, followed by 40 cycles at 95 °C for 15 s and 60 °C for 60 s.
2.6 Western blotting
Six pairs of fresh primary tumor and surrounding healthy tissue samples stored in liquid nitrogen were used to extract total protein. Each pair of tissues was cut into 0.03 g pieces, washed with phosphate-buffered saline, and lysed with the radioimmunoprecipitation assay (RIPA) lysis buffer. Finally, the TissueMaster Handheld Homogenizer (Beyotime) was utilized to grind the tissues thoroughly. Proteins in the tissues were separated using 8% sodium dodecyl-sulfate polyacrylamide gel electrophoresis and transferred onto a 0.22 μm nitrocellulose membrane (Pall Corporation). The membrane was incubated overnight with the following primary antibodies (Abcam) after blocking in a 5% skim milk solution for 2 h: DNMT3A (rabbit polyclonal, 1:1000) and TET2 (rabbit polyclonal, 1:1000). β-actin antibody (1:1000) was utilized as the control.
2.7 Transfection and expression validation
Small interfering DNMT3A (si-DNMT3A) was purchased from Hanheng Biotechnology, and the human TET2 overexpression plasmid was purchased from Genechem. Transfection was conducted using the Lipofectamine 2000 (Invitrogen) or jetPRIME (PolyPlus transfection) transfection reagent. For RNA extraction, the AxyPrep Multisource Total RNA Miniprep Kit (Axygen) was used. For protein extraction, the RIPA lysis buffer (Beyotime) was used. Moreover, RT-PCR and western blotting were performed to confirm the knockdown and overexpression. β-actin antibody (1:1000) and GAPDH were utilized as the control.
2.8 Cell proliferation assay
To evaluate cell viability, the Cal27 cell lines were seeded in a 96-well plate and cultured for 24 h. Cell proliferation was assessed with the BeyoClick™ EdU Cell Proliferation Kit (Beyotime), which labels cells in the S phase of the cell cycle. The number of EdU-positive cells was observed under an inverted microscope (Nikon) to determine the extent of cell proliferation.
2.9 Wound healing assay
The Cal27 cell lines were seeded in a six-well plate until the formation of monolayer cells. A 200 μL tip was then utilized to produce a scratch, and the detached cells were washed away. Serum-free DMEM was added to the wells for cell culture. After incubating for 24 and 48 h, the scratches were photographed using an upright microscope (Leica).
2.10 Transwell assay
To assess the migration capacity of Cal27, we used 6 mm transwell chambers with 8-μm pores (Corning). A serum-free medium was added to the chambers, while a regular medium was added to a 24-well plate in which the chambers had been placed. After 48 h, the cells on the lower side of the membrane were fixed with 4% paraformaldehyde and stained with crystal violet. The stained cells were then photographed using an upright microscope.
2.11 Statistical analysis
Statistical analyses were performed using GraphPad Prism 9 software. Fisher’s exact test, Student’s t-test, Kaplan–Meier (KM) approach and one- and two-way analyses of variance were used, as appropriate. Statistical significance was set at a p-value < 0.05. Each experiment was conducted at least thrice.
3 Results
3.1 Methylation β values analysis results of DNMT3A and TET2 genes under different clinical characteristics
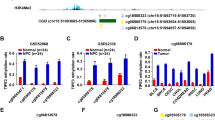
We obtained methylation chip data and clinical information of 399 OSCC samples from the TCGA database. The correlation between the mean β value of all probes and age, gender, TNM stage, and clinical stage was analyzed. Methylation β values of DNMT3A and TET2 were significantly correlated with gender, tumor size, extent of invasion, and clinical stage (P < 0.05; Figs. 1 and 2). We further grouped different CpG regions to examine the methylation β values of probes in different clinical groups. DNMT3A covers 25 probes in the CpG island region, seven probes in the downstream island region, 19 probes in the upper island region, and 15 probes in the Open_Sea region. TET2 covers five probes in the CpG island region, three probes in the downstream island region, three probes in the upper island region, and four probes in the Open_Sea region. Six DNMT3A probes (cg00912598, cg13344237, cg15998962, cg23569120, cg26995204, cg27369452) and one TET2 probe (cg14330655) in the CpG island region had a strong correlation with the clinical stage (P < 0.05; Figs. 3 and 4).
3.2 Correlation of the methylation degrees of DNMT3A and TET2 with the clinicopathological characteristics
The methylation degrees of DNMT3A and TET2 were statistically analyzed in correlation with clinical and pathological features (gender, age, lesion site, tumor size, clinical stage, pathological stage, and lymphatic metastasis). The methylation degrees of DNMT3A and TET2 differed significantly by pathological stage (Table 1, P < 0.05), but were not correlated with other clinicopathological parameters (P > 0.05). The methylation levels of the two genes changed with the degree of tumor differentiation; therefore, different protein expression levels were displayed at different stages.
3.3 Differences in DNMT3A and TET2 protein expressions in OSCC tissues
To assess the DNMT3A and TET2 protein expressions in OSCC, we performed western blotting of six pairs of primary tumor and surrounding healthy tissue samples. The DNMT3A expression was significantly higher in OSCC samples than in surrounding healthy tissue samples (P < 0.05, Fig. 5a), while the TET2 expression was the opposite (P < 0.05, Fig. 5b). Based on these results, we purchased si-DNMT3A and TET2 overexpression plasmids for subsequent cell function experiments.
DNMT3A and TET2 protein expressions in six pairs of OSCC samples and transfection effects of DNMT3A and TET2 a, b: DNMT3A and TET2 protein expression levels in tumor and surrounding healthy tissue samples c, d: Statistical analysis results of protein expressions e, h: DNMT3A and TET2 protein expressions f, i: Statistical analysis results of protein expressions g, j: DNMT3A and TET2 messenger RNA expressions
3.4 Expression validations of DNMT3A and TET2
To determine the transfection effects of si-DNMT3A and TET2 overexpression plasmids, we performed western blotting (P < 0.05, Fig. 5e, h) and RT-PCR (P < 0.05, Fig. 5g, j) to confirm the knockdown and overexpression, respectively. Both knockdown and overexpression showed good results.
3.5 Effects of DNMT3A and TET2 on OSCC cell proliferation
In the EdU cell proliferation experiment, the TET2 overexpression group showed a significantly lower number of proliferating cells compared to the control group (P < 0.05; Fig. 6c). However, unlike the TET2 overexpression experimental group, the DNMT3A knockdown experimental group showed no significant differences (P < 0.05; Fig. 6a). The number of proliferating cells was displayed by the amount of green fluorescence. The results suggested that TET2 played a greater role in the proliferation of OSCC cells compared to DNMT3A.
Proliferation and migration of OSCC cells a: Cells in the proliferation phase in the DNMT3A experimental group c: Cells in the proliferation phase in the TET2 experimental group b, d: Statistical analysis results of proliferation e: Cal27 cell migration in the DNMT3A experimental group g: Cal27 cell migration in the TET2 experimental group f, h: Statistical analysis results of migration i: Cal27 cell migration in the DNMT3A experimental group j: Cal27 cell migration in the TET2 experimental group k, l: Statistical analysis results of migration (scale bar = 100 μm)
3.6 Effects of DNMT3A and TET2 on OSCC cell migration
We conducted wound healing and transwell assays to assess the migratory ability of OSCC cells. The wound healing assay showed wound healing at 24 and 48 h. The DNMT3A-knockdown (P < 0.05, Fig. 6e) and TET2 overexpression experimental groups (P < 0.05, Fig. 6g) showed significantly low wound healing rates. Similarly, in the transwell assay, compared to the control group, both experimental groups showed a reduced number of OSCC cells passing through the transwell to varying degrees (P < 0.05; Fig. 6i, j). These results suggested that DNMT3A and TET2 played vital roles in cell migration inhibition. TET2 displayed a weaker ability to deform OSCC cells, making it more difficult for cells to pass through pores. Compared to the DNMT3A-knockdown experimental group, the TET2 overexpression experimental group showed a stronger ability to inhibit the migration of OSCC cells.
3.7 Association between DNMT3A and TET2 methylation levels and OSCC survival
According to the KM method, cases of high DNMT3A methylation levels showed higher survival rates (P < 0.05; Fig. 7a). However, TET2 methylation levels were not significantly associated with OSCC survival (P > 0.05; Fig. 7b). This result may be owing to the low number of cases and short follow-up time.
4 Discussion
OSCC is the sixth most common malignancy worldwide, and its occurrence and development result from the gradual accumulation of various factors, including epigenetic factors. DNA methylation modifications affect genetic mutations and occur before tumor formation [31]. In general, the epigenetic state of cells is precisely regulated to maintain an appropriate state of differentiation. However, this carefully balanced genomic programming is disturbed in cancer, resulting in uncontrolled cell proliferation, impaired differentiation, and resistance to apoptosis [32]. Therefore, the DNA methylation status of target genes is a potential therapeutic target in cancer treatment and biomarker in early detection and prognostication of cancer. Consequently, it should be explored for improving cancer treatment outcomes.
In this study, the methylation degree revealed that as the cancer became less differentiated, DNMT3A promoter methylation decreased gradually, indicating that its expression was no longer inhibited. The level of global methylation of cancer increased, thereby promoting the malignant transformation of tumors. DNMT3A is involved in the oncogenic process of various tumors by regulating the level of promoter methylation. Yu et al. revealed that miR-26a-5p targeted DNMT3A to reduce the degree of global methylation in non-small cell lung cancer and restore the SFRP1 expression, thereby regulating cell viability and the stem-like phenotype by regulating the Wnt/β-catenin pathway [33]. Lu et al. found DNMT3A promoted the Warburg effect and tumor malignant biological behavior by inhibiting the miR-603 expression. Therefore, it is a potential therapeutic target for ovarian cancer [34]. Pang et al. demonstrated that MYC-recruited DNMT3A induced promoter methylation, resulting in miR-200b silencing, thereby promoting epithelial mesenchymal transformation and mammary globular formation in triple-negative breast cancer cells [35].
In the present study, TET2 functioned as a tumor suppressor gene (TSG) in OSCC. The methylation degree revealed that TET2 was predominantly unmethylated in tissues with a lower degree of malignancy, indicating that the TET2 expression was not inhibited. Further, as the degree of differentiation decreased, the degree of promoter methylation increased, and the expression was inhibited. Promoter methylation of TSGs can be an early occurrence in the development of tumors. This epigenetic alteration may confer advantages to cell growth, in turn promoting malignant transformation. TSGs that undergo abnormal promoter hypermethylation can encode various protein products, including DNA repair factors, cell cycle inhibitors, and cell adhesion receptors [36]. Hitchins et al. reported that promoter hypermethylation of MLH1 and epigenetic silencing were associated with various cancers characterized by defects in mismatch repair [37]. Casalino et al. showed that CDKN2A encoded the cell cycle inhibitor p16INK4A, and loss of the p16INK4A expression because of hypermethylation of its promoter was an early event in breast and lung cancers [38]. Markou et al. selected five TSGs, involved in cancer cell differentiation, proliferation, apoptosis, adhesion, and metastasis for methylation evaluation, which in combination with circulating tumor cells and matched plasma-free DNA provided effective prognostic information for patients with early lung cancer [39]. These examples confirm that abnormal methylation contributes to the development of cancer by affecting genes involved in key cellular processes and suggest that the methylation degree of the promoter of TSGs change continuously during tumor development, which is an effective mean for the early detection, disease surveillance, and evaluation of the therapeutic efficacy.
The present study showed a higher protein expression of DNMT3A in tumor samples than in surrounding healthy tissue samples. The TET2 expression in cancer tissues was significantly reduced, and its recovery could effectively inhibit the malignant biological behavior of OSCC. Altered expression levels of DNMTs, particularly elevated DNMT3A levels, are common in multiple cancer samples and cell lines [40]. TET proteins regulate the equilibrium between DNA methylation and demethylation by regulating the dynamic transformation among cytosine, 5-mC, and 5-hmC [41]. However, missense and truncated TET mutations have been reported in nearly all types of solid tumors [42], and their reduced protein expression and 5-hmC levels are markers of many cancer types, including colorectal cancer [43], glioblastoma [24], cervical cancer [44], and pancreatic cancer [45]. The aforementioned studies are consistent with our findings, showing that modulating the DNMT3A and TET2 expression levels could effectively change the biological behavior of OSCC.
This was the first study to show that DNMT3A and TET2 methylation levels change with the degree of cancer differentiation and that DNMT3A knockdown or TET2 overexpression could effectively inhibit the proliferation and migration of OSCC. Although the oncogenic mechanisms of DNMT3A and TET2 should be further explored, they have a wide range of applications as potential therapeutic targets for cancer.
Data availability
Data will be made available on request.
Code availability
Not applicable.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–12.
Dawson MA, Kouzarides T. Cancer epigenetics: from mechanism to therapy. Cell. 2012;150:12–22.
Kulis M, Esteller M. DNA methylation and cancer. Adv Genet. 2010;70:27–35.
Qiu M, Zhang N, Yao S, Zhou H, Chen X, Jia Y, et al. DNMT3A-mediated high expression of circ_0057504 promotes benzo[a]pyrene-induced DNA damage via the NONO-SFPQ complex in human bronchial epithelial cells. Environ Int. 2022. https://doi.org/10.1016/j.envint.2022.107627.
Bondarev N, Ivanenko K, Khabusheva E, Lebedev T, Manukhov I, Prassolov V. MGL S3 chimeric enzyme drives apoptotic death of EGFR-dependent cancer cells through ERK downregulation. Int J Mol Sci. 2022;23:12807.
Li Y, Gan Y, Liu J, Li J, Zhou Z, Tian R, et al. Downregulation of MEIS1 mediated by ELFN1-AS1/EZH2/DNMT3a axis promotes tumorigenesis and oxaliplatin resistance in colorectal cancer. Signal Transduct Target Ther. 2022;7:87.
Liang Y, Cen J, Huang Y, Fang Y, Wang Y, Shu G, et al. CircNTNG1 inhibits renal cell carcinoma progression via HOXA5-mediated epigenetic silencing of Slug. Mol Cancer. 2022;21:224.
He J, Dong C, Zhang H, Jiang Y, Liu T, Man X. The oncogenic role of TFAP2A in bladder urothelial carcinoma via a novel long noncoding RNA TPRG1-AS1/DNMT3A/CRTAC1 axis. Cell Signal. 2023;102:110527.
Lu W, Lu T, Wei X. Downregulation of DNMT3a expression increases miR-182-induced apoptosis of ovarian cancer through caspase-3 and caspase-9-mediated apoptosis and DNA damage response. Oncol Rep. 2016;36:3597–603.
Li S, Hu J, Li G, Mai H, Gao Y, Liang B, et al. Epigenetic regulation of LINC01270 in breast cancer progression by mediating LAMA2 promoter methylation and MAPK signaling pathway. Cell Biol Toxicol. 2022. https://doi.org/10.1007/s10565-022-09763-9.
Zhou Y, Yang Z, Zhang H, Li H, Zhang M, Wang H, et al. DNMT3A facilitates colorectal cancer progression via regulating DAB2IP mediated MEK/ERK activation. Biochim Biophys Acta Mol Basis Dis. 2022;1868:166353.
Zhao Z, Song J, Tang B, Fang S, Zhang D, Zheng L, et al. CircSOD2 induced epigenetic alteration drives hepatocellular carcinoma progression through activating JAK2/STAT3 signaling pathway. J Exp Clin Cancer Res. 2020;39:259.
Quivoron C, Couronné L, Della Valle V, Lopez CK, Plo I, Wagner-Ballon O, et al. TET2 inactivation results in pleiotropic hematopoietic abnormalities in mouse and is a recurrent event during human lymphomagenesis. Cancer Cell. 2011;20:25–33.
Álvarez-Errico D, Vento-Tormo R, Sieweke M, Ballestar E. Epigenetic control of myeloid cell differentiation, identity and function. Nat Rev Immunol. 2015;15:7–1.
Delhommeau F, Dupont S, Della Valle V, James C, Trannoy S, Massé A, et al. Mutation in TET2 in myeloid cancers. N Engl J Med. 2009;360:2289–92.
Ma S, Wan X, Deng Z, Shi L, Hao C, Zhou Z, et al. Epigenetic regulator CXXC5 recruits DNA demethylase Tet2 to regulate TLR7/9-elicited IFN response in pDCs. J Exp Med. 2017;214:1471–81.
Zhang Q, Zhao K, Shen Q, Han Y, Gu Y, Li X, et al. Tet2 is required to resolve inflammation by recruiting Hdac2 to specifically repress IL-6. Nature. 2015;525:389–93.
Cull AH, Snetsinger B, Buckstein R, Wells RA, Rauh MJ. Tet2 restrains inflammatory gene expression in macrophages. Exp Hematol. 2017;55:56–7.
Fuster JJ, MacLauchlan S, Zuriaga MA, Polackal MN, Ostriker AC, Chakraborty R, et al. Clonal hematopoiesis associated with TET2 deficiency accelerates atherosclerosis development in mice. Science. 2017;355:842–8.
Deng W, Wang J, Zhang J, Cai J, Bai Z, Zhang Z. TET2 regulates LncRNA-ANRIL expression and inhibits the growth of human gastric cancer cells. IUBMB Life. 2016;68:355–63.
Nickerson ML, Das S, Im KM, Turan S, Berndt SI, Li H, et al. TET2 binds the androgen receptor and loss is associated with prostate cancer. Oncogene. 2017;36:2172–82.
Alrehaili AA, Gharib AF, Alghamdi SA, Alhazmi A, Al-Shehri SS, Hagag HM, et al. Evaluation of TET family gene expression and 5-hydroxymethylcytosine as potential epigenetic markers in non-small cell lung cancer. In Vivo. 2023;37:445–54.
Lyu R, Zhu X, Shen Y, Xiong L, Liu L, Liu H, et al. Tumour suppressor TET2 safeguards enhancers from aberrant DNA methylation and epigenetic reprogramming in ERα-positive breast cancer cells. Epigenetics. 2022;17:1180–1.
Lopez-Bertoni H, Johnson A, Rui Y, Lal B, Sall S, Malloy M, et al. Sox2 induces glioblastoma cell stemness and tumor propagation by repressing TET2 and deregulating 5hmC and 5mC DNA modifications. Signal Transduct Target Ther. 2022;7:37.
Gong F, Guo Y, Niu Y, Jin J, Zhang X, Shi X, et al. Epigenetic silencing of TET2 and TET3 induces an EMT-like process in melanoma. Oncotarget. 2017;8:315–23.
Adhikari BR, Uehara O, Matsuoka H, Takai R, Harada F, Utsunomiya M, et al. Immunohistochemical evaluation of Klotho and DNA methyltransferase 3a in oral squamous cell carcinomas. Med Mol Morphol. 2017;50:155–61.
Yakushiji T, Uzawa K, Shibahara T, Noma H, Tanzawa H. Over-expression of DNA methyltransferases and CDKN2A gene methylation status in squamous cell carcinoma of the oral cavity. Int J Oncol. 2003;22:1201–11.
Daniel FI, Rivero ER, Modolo F, Lopes TG, Salum FG. Immunohistochemical expression of DNA methyltransferases 1, 3a and 3b in oral leukoplakias and squamous cell carcinomas. Arch Oral Biol. 2010;55:1024–31.
Supic G, Kozomara R, Zeljic K, Jovic N, Magic Z. Prognostic value of the DNMTs mRNA expression and genetic polymorphisms on the clinical outcome in oral cancer patients. Clin Oral Invest. 2017;21:173–81.
Jäwert F, Hasséus B, Kjeller G, Magnusson B, Sand L, Larsson L. Loss of 5-hydroxymethylcytosine and TET2 in oral squamous cell carcinoma. Anticancer Res. 2013;33:4325–34.
Wang X, Zhang Y, Sun L, Wang S, Nie J, Zhao W, et al. Evaluation of the clinical application of multiple tumor marker protein chip in the diagnostic of lung cancer. J Clin Lab Anal. 2018;32:e22565.
Werner RJ, Kelly AD, Issa JJ. Epigenetics and precision oncology. Cancer J. 2017;23:262–72.
Yu J, Ge Z, Chen S, Li S, Zhang X, Hu J, et al. miR-26a-5p suppresses Wnt/β-catenin signaling pathway by inhibiting DNMT3A-mediated SFRP1 methylation and inhibits cancer stem cell-like properties of NSCLC. Dis Markers. 2022;2022:7926483.
Lu J, Zhen S, Tuo X, Chang S, Yang X, Zhou Y, et al. Downregulation of DNMT3A attenuates the warburg effect, proliferation, and invasion via promoting the inhibition of miR-603 on HK2 in ovarian cancer. Technol Cancer Res Treat. 2022. https://doi.org/10.1177/15330338221110668.
Pang Y, Liu J, Li X, Xiao G, Wang H, Yang G, et al. MYC and DNMT3A-mediated DNA methylation represses microRNA-200b in triple negative breast cancer. J Cell Mol Med. 2018;22:6262–6.
Esteller M. CpG island hypermethylation and tumor suppressor genes: a booming present, a brighter future. Oncogene. 2002;21:5427–35.
Hitchins MP, Ward RL. Constitutional (germline) MLH1 epimutation as an aetiological mechanism for hereditary non-polyposis colorectal cancer. J Med Genet. 2009;46:793–8.
Casalino L, Verde P. Multifaceted roles of DNA methylation in neoplastic transformation, from tumor suppressors to EMT and metastasis. Genes. 2020;11:922.
Markou Α, Londra D, Tserpeli V, Kollias Ι, Tsaroucha E, Vamvakaris I, et al. DNA methylation analysis of tumor suppressor genes in liquid biopsy components of early stage NSCLC: a promising tool for early detection. Clin Epigenet. 2022;14:61.
Subramaniam D, Thombre R, Dhar A, Anant S. DNA methyltransferases: a novel target for prevention and therapy. Front Oncol. 2014;4:80.
Lian H, Li WB, Jin WL. The emerging insights into catalytic or non-catalytic roles of TET proteins in tumors and neural development. Oncotarget. 2016;7:64512–6.
Rasmussen KD, Helin K. Role of TET enzymes in DNA methylation, development, and cancer. Genes Dev. 2016;30:733–7.
El-Harakeh M, Saliba J, SharafAldeen K, Haidar M, El Hajjar L, Awad MK, et al. Expression of the methylcytosine dioxygenase ten-eleven translocation-2 and connexin 43 in inflammatory bowel disease and colorectal cancer. World J Gastroenterol. 2022;28:5845–55.
Gao J, Liu R, Feng D, Huang W, Huo M, Zhang J, et al. Snail/PRMT5/NuRD complex contributes to DNA hypermethylation in cervical cancer by TET1 inhibition. Cell Death Differ. 2021;28:2818–22.
Fujikura K, Alruwaii ZI, Haffner MC, Trujillo MA, Roberts NJ, Hong SM, et al. Downregulation of 5-hydroxymethylcytosine is an early event in pancreatic tumorigenesis. J Pathol. 2021;254:279–82.
Acknowledgements
None.
Funding
None.
Author information
Authors and Affiliations
Contributions
XL made contributions to the conception and design of the work, and provided the acquisition, analysis of mostly data; ZL provided the acquisition, analysis of partly data; QG drafted the work; YP revised the work critically for important intellectual content; YY helped the first author to acquire, analyze data; TH approved the version to be published and provided technical support; WW agree to be accountable for all aspects of the work in ensuring that questions related to the accuracy and provided fund support.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, X., Li, Z., Gao, Q. et al. Correlation of DNA methylation of DNMT3A and TET2 with oral squamous cell carcinoma. Discov Onc 15, 15 (2024). https://doi.org/10.1007/s12672-024-00866-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-024-00866-9