Abstract
Background
Pancreatic ductal adenocarcinoma (PDAC) is characterized by a highly metastatic potential and a heterogeneous tumor microenvironment. It exhibits limited sensitivity to conventional therapies, necessitating a deeper understanding of its pathogenesis. The role of the intratumoral microbiome in regulating cancer development in PDAC has been the subject of debate. Previous investigations into intra-tumor microbiomes have yielded uncertain results due to sample size limitations and insufficient decontamination procedures. Further research is imperative to elucidate the intricate relationship between intra-tumor microbiomes, the immune landscape of PDAC, and overall prognosis.
Results
Our findings revealed that the intratumor microbiota in PDAC tissue exhibited lower diversity and distinct communities compared to non-tumor tissues. The top microorganisms distinguishing between patients with long or short survival were used to construct the risk signature. We found that Stenotrophomonas is implicated in short survival of PDAC patients, while Neorickettia and Mediterraneibacter are correlated with long survival. This microbiome-based PDAC subtyping, grounded in prognosis-related signatures, exhibited significant correlations with distinct clinical prognoses and immune microenvironments. Microorganisms associated with negative prognoses were linked to pro-tumor immune activation, while those associated with positive prognoses were linked to anti-tumor immune response activation and a more favorable prognosis.
Conclusions
Our PDAC subtyping approach, based on a microbiome-derived prognostic risk signature, unveiled compelling associations between the PDAC microbiota and disparities in both clinical prognosis and the tumor microenvironment. These findings suggest that microbiota may serve as a promising biomarker for predicting the prognosis of PDAC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Despite a notable decline in cancer mortality in recent years, attributed to reduced smoking rates and advancements in early screening and treatment, cancer remains a prominent global cause of death [1]. Pancreatic cancer, characterized by its high mortality rate, presents a significant challenge. The 5-year survival rate for all stages stands at a mere 9%, dropping to a mere 3% for patients with metastatic disease. Unfortunately, pancreatic cancer tends to be diagnosed at an advanced stage, often rendering available therapies ineffective. Consequently, efforts in prevention, early diagnosis, and therapeutic interventions have yet to yield satisfactory outcomes, resulting in a poor prognosis [2]. Moreover, pancreatic cancer demonstrates notable resistance to most conventional treatment options, encompassing chemotherapy, radiotherapy, and targeted therapies. This resistance can be attributed to a myriad of genetic and epigenetic alterations, as well as the intricate nature of the tumor microenvironment in pancreatic cancer [3,4,5]. Thus, there is an urgent call to develop comprehensive strategies for the diagnosis and treatment of pancreatic cancer. This includes the identification of high-risk groups for screening, early detection via biomarkers, and the formulation of personalized treatment approaches. Furthermore, the exploration of the tumor microenvironment as a potential target becomes paramount in overcoming resistance to current therapies [6].
The tumor microenvironment encompasses intra-tumor microbes, which necessitates a combined analysis with microbiome studies. The field of microbiomes involves the scrutiny of microbial species, their abundance, functionalities, and their interconnectedness within microbial communities and their hosts [7, 8]. It is widely recognized that microbial communities in tumors encompass not only bacteria [9, 10], but also fungi [11] and viruses [12], and they play a substantial role in influencing the development, progression, metastasis, and treatment response of various cancer types [13]. Recent investigations have underscored the significant involvement of intra-tumor microbes in the development of pancreatic cancer and their impact on the efficacy of therapeutic interventions [2, 14, 15]. Nevertheless, the reliability and consistency of research findings in the field of microbiomes are often challenged by various confounding factors, including sample geography, sequencing technology, and disease state. It is, therefore, imperative to conduct more comprehensive and in-depth studies to elucidate the relationship between microorganisms and pancreatic cancer and to develop novel strategies and directions for future pancreatic cancer treatment.
Recent studies have brought to light the crucial role of intra-tumor microbes in the development and treatment of pancreatic cancer. The progress in assessing microbial communities within paraffin-embedded tissue has paved the way for retrospective investigations [16]. Various methods for extracting reliable microbial data from high-throughput sequencing of tumor samples are currently being proposed and refined [10, 17]. While there have been several attempts to explore the intra-tumor microbiome in pancreatic cancer [2, 14, 15], concerns about inadequate decontamination and limitations in sample size have raised questions about their reliability. Furthermore, a deeper understanding of the intricate relationship between intra-tumor microbiomes, the immune landscape, and the prognostic implications of pancreatic cancer is imperative.
In our study, we employed a calculation method to extract tumor microbial information and identify crucial microbes, drawing on established approaches from previous studies. This method is applicable in a wide range of scenarios involving the extraction of microbial information from high-throughput sequencing resources. Our study specifically concentrated on tumor microbes and their impact on the progression of pancreatic cancer, as well as their interactions within the tumor microenvironment. This research represents a foundational stride towards refining future approaches for the prevention and treatment of pancreatic cancer.
2 Methods
2.1 Data access
Pancreatic ductal adenocarcinoma data were downloaded from The Cancer Genome Atlas (TCGA), in which 144 samples were included. We downloaded bam documents of original sequencing data of pancreatic ductal adenocarcinoma, containing 142 tumor samples and 2 normal samples. We also downloaded clinical information of 142 patients.
2.2 Data acquisition and overview
We downloaded all the pancreatic ductal adenocarcinoma RNA-seq data from TCGA database using the officially recommended gdc-client software (version 1.2.0) [18] in a bam format. The pancreatic ductal adenocarcinoma RNA-seq data contains 144 samples, with a total data size of about 1 TB. Standardization by STAR software (version 2.5.3a) [19] were conducted at the same time. The count document includes standardized Fragments Per Kilobase of transcript per Million mapped reads (FPKM). Patient information is listed in Table 1.
2.3 Extraction of pancreatic ductal adenocarcinoma microbe
We transduced the bam documents of pancreatic ductal adenocarcinoma downloaded from TCGA to fastq documents through samtools software (version 1.3.1) [20]. Then we used Kraken2 to acquire report document by comparing to the reference gene index PlusPF. This index contains archaebacteria, bacteria, virus, plasmid, human, protozoon, and fungi, which possesses the highest species richness among all databases. One fastq document generates a report document, containing information of all species within the fastq document, and number of sequencing fragments. Bayesian algorithm function RDPclassifier was used to perform taxonomic comparative analysis on OTU of PDAC patients and microbial community composition of each sample was counted.
2.4 Microbial contamination removement
We performed a rigorous filtering and decontamination steps. Microbial reads with low expression (i.e., those that did not meet the "expressed in at least 5 individuals) or microbial reads (i.e., microbial reads that did not meet the criteria of "expressed in at least 5 individuals and the expression of a microorganism is greater than 50"), Human and Archaea reads were filtered out. The further decontamination step used the R package decontam (version 1.12.0) [21] to remove microorganisms with non-linear variation in expression, deleting a total of 85.3% of the original reads were removed.
2.5 Standardization and differential abundance analysis on microbial profile
We combined utilization of observational level variance modeling (VOOM) normalization and surrogate variable analysis (SNM) to preliminarily standardize microbial expression profile [10]. VOOM converts the discrete classification count to Mega log counts per sample (log-cpm) which approximately follows the normal distribution and eliminates heteroscedasticity of data [22]. SNM is a supervised normalization method which can be used to eliminate significant batch effect [23]. After standardization, Mann–Whitney test and PERMANOVA test were used to detect differences between groups.
2.6 Microbial diversity analysis
The microbial reads count extracted by software kraken2 were read at the "species" level and each microorganism was identified as an OTU. The OTU table of KIRC patients was analyzed by using R package vegan (version 2.5-6) (https://cran.r-project.org/package=vegan), i.e., the sequences of all samples were randomly selected to a uniform amount of data, to ensure the homogeneity of the sequenced sequences of the samples. The α-diversity indices were also calculated using vegan, and the Wilcoxon test was used to show the differences between groups. In this study, vegan was used to calculate Bray curtis distance and Jaccard index distance using weighted algorithm and unweighted algorithm, respectively, and PCoA principal coordinate analysis was performed to demonstrate the differences in biological β-diversity between subgroups of PDAC patient samples, and the differences between groups were tested by Adonis test. LEfSe analysis was performed using the online tool at http://www.bic.ac.cn/BIC.
2.7 Differential gene expression analysis
We used DESeq2 (version 1.32.0) [24], using patient gene counts as the input matrix, and the data were normalized to compare gene expression in two different groups of patients. Pearson and Spearman coefficients were used to correlate microbial and gene expression the results of the correlation analysis were tested for significance. The results of correlation analysis were tested for significance.
2.8 Survival analysis
In this study, one-way Cox and multi-way Cox analyses were performed on the sample using R package survival (version 3.3-1). The risk values were obtained based on the scoring of significant variables, and the survival prediction was performed using the risk values and the R package survminer (version 0.4.7) (https://cran.r-project.org/web/packages/survminer/index.html) was used to plot Kaplan–Meier (KM) curves.
2.9 Correlation analysis
The Pearson and Spearman coefficients were used for the correlation between microbial and gene expression. The results of correlation analysis were tested for significance.
2.10 Gene function enrichment analysis
We mainly used R package clusterProfiler (version 4.0.5) [25] for GO enrichment analysis, KEGG enrichment analysis and GSEA enrichment analysis.
2.11 Immuno-infiltration analysis
In this study, immuno-infiltration analysis was performed using R package xcell (version 1.1.0) [26] and R package CIBERSORT (version 0.1.0) [27].
2.12 Weighted correlation network analysis
R package WGCNA (version 1.66) [28] was used to create weighted gene co-expression network analysis and function flashClust was used for analysis of interaction relationship between different co-expression modules.
3 Results
3.1 Overall profile of tumor microbiome
We assembled a dataset consisting of 144 samples of pancreatic ductal adenocarcinoma sourced from The Cancer Genome Atlas (TCGA). We utilized Kraken2 software (version 2.0.8) [29] to align the sequencing results with the reference gene index. The primary objective of our study was to scrutinize the variations in tumor microbes among patients exhibiting different overall survival (OS) rates. For our classification, individuals with an OS of less than 1 year, who eventually succumbed to the disease, were categorized as having a short OS. Conversely, those with an OS extending beyond 3 years were classified as having a long OS. This classification led to the identification of 33 patients with a short OS, and 11 patients with a long OS.
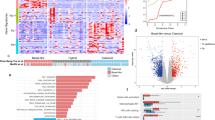
The microbes were transformed into operational taxonomic units (OTUs) and their compositions were compared between patients with short- and long-term OS. Figure 1A and B display the abundance in stacked bar charts of tumor microbes in pancreatic ductal adenocarcinoma patients at both Phylum and Genus levels. These visuals suggest no significant difference in the overall composition of tumor microbes between patients with short OS and long OS. The most abundant phyla were Firmicutes, Proteobacteria, and Actinobacteria, while at the Genus level, Bacillus, Priestia, and Pasteurella showed the highest richness. In order to pinpoint potential differences in the distribution of bacteria and fungi between patients with short and long OS, we generated a superimposed map of species classifications for both mycomonera and fungi (Fig. 1C and D). The analysis indicated that Bacillus remains the most abundant bacterium, with Lactobacillus taking the place of Priestia as the second most abundant. As for fungi, Sporisorium, Fusarium, and Toxoplasma emerged as the top three richest microbes.
Overall profile of tumoral microbiome comparison. A Stacked bar charts of total abundance of the Phylum level microbes in pancreatic ductal adenocarcinoma patients. B Stacked bar charts of total abundance of the Genus level microbes in pancreatic ductal adenocarcinoma patients. Long: samples with long survival. Short: samples with short survival. Vertical coordinates represent relative abundance of microbes, in which each color correlates to a microbial category. Only the 10 microbes with the highest relative abundance are shown. The rest are categorized as “Others”. C Stacked charts of relative abundance of tumor bacteria in long and short survival patients with pancreatic ductal adenocarcinoma. D Stacked charts of relative abundance of fungi in long and short survival patients with pancreatic ductal adenocarcinoma. Horizontal axis shows samples and vertical axis represents relative abundance. Long: samples with long survival. Short: samples with short survival. E Boxplots of \(\alpha\) diversity of tumor bacteria in long and short survival patients with pancreatic ductal adenocarcinoma. F Boxplots of \(\alpha\) diversity of fungi in long and short survival patients with pancreatic ductal adenocarcinoma. Long: long survival. Short: short survival. \(\alpha\) diversity is shown by Shannon index and Simpson index. Mann–Whitney test was used for significance test. G PCoA charts of \(\beta\) diversity of tumor bacteria in long and short survival patients with pancreatic ductal adenocarcinoma. H PCoA charts of \(\beta\) diversity of fungi in long and short survival patients with pancreatic ductal adenocarcinoma. Long: long survival. Short: short survival. Weighted Bray–Curtis and non-weighted Jaccard index distances were used to evaluate \(\beta\) diversity. PERMANOVA test was used for significance test. I Branching plot of differential microbial taxonomy in long and short survival samples. Microbes with significant difference are labelled. J LDA score bar chart calculated by LEFSe according to the microbial relative abundance between long and short survival samples. Microbes with an LDA score (log10) > 2 and p value < 0.05 are shown. Long: long survival samples. Short: short survival samples. K Clustering heatmap of microbial expression in patients with long and short survival. Different colors reflect expression levels between microbes in two groups. A deep color represents a higher level of expression. PCoA, principal coordinate analysis; LDA, Linear discriminant Analysis; LEfSe, linear discriminant analysis effect size
To further explore the microbial diversity in both short and long OS patients, we conducted α diversity analysis on bacteria and fungi employing the Shannon index and Simpson index. As shown in Fig. 1E and F, the α diversity of tumor bacteria in long OS patients appears slightly higher than that in short OS patients, though no statistically significant difference was observed. Similarly, the diversity of fungi in patients showed no significant difference. We then proceeded with β diversity analysis on bacteria and fungi, utilizing the weighted Bray–Curtis and non-weighted Jaccard index distances. As demonstrated in Fig. 1G and H, no significant difference in the diversity of bacteria and fungi was noted between patients with short OS and long OS. Furthermore, the linear discriminant analysis effect size (LEfSe) analysis uncovered a notable distinction in the richness of seven specific microbes between short OS and long OS samples, including Actinomycetes, Thermomonosporaceae, Nitriliruptoria, Lachnospiraceae, Piscirickettsiaceae, and Thiotrichales (Fig. 1I and J). Notably, among them, Actinomycetes exhibited significant differences at all three levels: family, order, and genus.
The Kruskal–Wallis test (K-W test) was employed to discern differential microbes, and multi-check calibration was conducted to ensure accuracy. In this analysis, we scrutinized both biological and non-biological variables. Regarding the biological variable (OS), we pinpointed microbes that exhibited significant changes (P < 0.05). As for the non-biological variable, we focused on the TCGA sample center and identified microbes that did not display significant differences (P > 0.1) among various sample centers. The overlap between these two sets of microbes yielded our alternative differential microbes. Figure 1K presents a microbial expression clustering heatmap for short and long-OS patients. This heatmap illustrates a noteworthy clustering pattern among the total 44 patients based on OS, underscoring discernible disparities in microbe expression profiles between the two groups. Subsequently, from the aforementioned alternative differential microbes, we conducted a manual literature search to identify 11 microbes that potentially exhibit variations in humans for further analysis.
3.2 Microbiomes predict patient survival
We conducted a single-factor Cox’s proportional hazards regression model (Cox) analysis on the 11 previously selected differential microbes, of which 10 exhibited significant effects on survival (P < 0.05) (Table 2). These 10 significant microbes are: Actinomadura, Pseudoxanthomonas, Neorickettsia, Mediterraneibacter, Cedecea, Hafnia, Azotobacter, Stenotrophomonas, Porphyromonas, and Desulfotalea. Figure 2A depicts box plots comparing the expression levels of these microbes between patients with short and long OS. The expression levels of these microbes indeed show notable differences between the two groups, as outlined in Table 3. Upon further examination, the first three microbes (Actinomadura, Pseudoxanthomonas, and Neorickettsia) were found to be higher in patients with short OS, while the remaining seven microbes (Mediterraneibacter, Cedecea, Hafnia, Azotobacter, Stenotrophomonas, Porphyromonas, and Desulfotalea) were more prevalent in patients with long OS. Consequently, we identified these 10 microbes as differential microbes for further analysis. Subsequently, we performed a multi-factor Cox regression analysis on these differential microbes, which revealed that only Stenotrophomonas had a significant impact on survival (P < 0.05).
Differential microbial analysis between patients with different overall survival. A Boxplots of differential microbial expression levels of long and short survival patients. Long: long survival. Short: short survival. Vertical axis represents standardized microbial expression level. Microbes present in the figure are Actinomadura, Stenotrophomonas, Pseudoxanthomonas, Neorickettsia, Porphyromonas, Mediterraneibacter, Cedecea, Hafnia and Desulfotalea, respectively. B Kaplan–Meier survival curve of 98 patients based on predicted risk score
Stenotrophomonas is a partially anaerobic, fermentative, sporeless, gram-negative bacillus in the order Xanthomonadales, family Xanthomonadaceae [30]. It ranks among the most common gram-negative bacilli encountered in clinical laboratories, second only to Pseudomonas aeruginosa. The prototypical species in this genus, Stenotrophomonas maltophilia, is acknowledged as an opportunistic human pathogen, widely distributed in both natural environments and healthcare settings. Previous research has demonstrated a higher prevalence of Stenotrophomonas maltophilia in tumor tissue compared to adjacent normal tissues in patients [31]. Experiments have confirmed that Stenotrophomonas maltophilia activates the TLR-4 mediated NF-κB pathway, resulting in the induction of senescence-associated secretory phenotypes in hepatic stellate cells (HSC). This leads to the subsequent formation of the NLRP3 inflammasome complex and the secretion of multiple inflammatory factors in the liver [31]. This process fosters the progression of hepatic cancer, resulting in a less favorable prognosis for affected patients. Given these findings, we conducted further research in pancreatic cancer to investigate whether Stenotrophomonas exerts a similar effect. We considered five species, including: Stenotrophomonas maltophilia, Stenotrophomonas sp. SAU14A_NAIMI4_5, Stenotrophomonas sp. LM091, Stenotrophomonas sp. 364, and Stenotrophomonas sp. ESTM1D_MKCIP4_1. Through single-factor Cox regression analysis on these five species, followed by subsequent multi-factor Cox regression analysis on those showing significant differences, our results revealed that Stenotrophomonas sp. SAU14A_NAIMI4_5, Stenotrophomonas sp. LM091, and Stenotrophomonas sp. 364 significantly impact patient survival in pancreatic cancer (Table 4).
We employed the Predict function from the R package survival (version 3.3–1) (https://CRAN.R-project.org/package=survival) to construct a risk score model specifically for the three significant microbes in Stenotrophomonas, as identified by the multi-factor Cox regression analysis. A total of 44 patients were utilized for establishing the score model, while the remaining 98 patients were reserved for model validation. The risk score model allocates higher scores to patients deemed at a higher risk, indicative of a shorter survival time. The Kaplan–Meier survival curve (K–M curve) in Fig. 2B shows how the model stratifies the 98 patients into high and low-risk groups based on their predicted risk scores. This risk scoring approach demonstrates a significant differentiation in terms of survival outcomes, underscoring its potential as a valuable prognostic tool in pancreatic cancer.
In order to account for potential influences of patients' conditions and treatment regimens on survival outcomes, we conducted a single-factor Cox regression analysis on the three core Stenotrophomonas microbes, in conjunction with key clinical data available in the TCGA dataset. This clinical information encompassed age, gender, cancer staging, and the treatments being administered (chemotherapy or radiotherapy). Factors found to be significant in this analysis were subsequently subjected to a multi-factor Cox regression analysis. The results unveiled that both Stenotrophomonas sp. LM091 and chemotherapy independently exerted statistically significant impacts on survival outcomes. Given the limited effectiveness of radiotherapy in treating pancreatic cancer, chemotherapy is frequently applied in clinical settings to prolong patient survival. This finding underscores the substantial influence of Stenotrophomonas on the survival rates of individuals with pancreatic ductal adenocarcinoma (Table 4).
3.3 Key microbiome is related with immune-related genes
We obtained the expression profiles of pancreatic ductal adenocarcinoma from TCGA, categorizing a cohort of 44 patients into two groups: 33 patients with a survival duration of less than 1 year, and 11 patients with a survival duration exceeding 3 years. The RNA sequencing (RNA-seq) counts of pancreatic ductal adenocarcinoma were standardized and employed as the input matrix for differential gene analysis between patients with short and long OS using the R package DESeq2 (version 1.32.0) [24]. Through this analysis, a total of 143 differentially expressed genes (DEGs) were identified, applying a threshold of log2 fold change (log2FC) > 1 and adjusted P-value (P-adj) < 0.05. Among these DEGs, 71 were found to be up-regulated genes, and 72 were down-regulated genes. The volcano plot of DEGs is presented in Fig. 3A, highlighting genes associated with inflammation, cell cycle regulation, innate immunity, and adaptive immunity. For example, IL31RA is correlated with inflammation, S100A2 is involved in cell cycle regulation, BPIFB4 is associated with innate immunity, and IGKV3D-20 correlates with adaptive immunity. In Fig. 3B, the DEGs were labeled, and a heatmap displayed the expression levels of these genes. Notably, patients with short and long OS exhibited inverse expression patterns in these immunity-related genes. Genes associated with cell cycle and inflammation displayed higher expression in patients with short OS, while in long OS patients, genes related to innate and adaptive immunity showed higher expression. These observations suggest that the overall survival of PDAC is likely related to the level of immune response.
Relationship between differential microbes and immuno-genes. A Volcano plot of differential genes in long and short survival patients. Horizontal axis represents log2 Fold Change, and vertical axis shows − log10(padj). With each dot representing a gene, red ones mean up-regulated genes, blue ones stand for down-regulated genes and grey ones represent genes that failed to meet threshold conditions. Threshold conditions are log2 Fold Change > 1 and padj < 0.05. B Heatmap of differential genes in long and short survival patients. Red represents up-regulated genes and blue represents down-regulated genes. Long: long survival samples. Short: short survival samples. This figure shows 13 immuno-genes, in which 5 are up-regulated and 8 are down-regulated. Functions of immune-genes are listed at the right. C Correlation heatmap of differential immune-genes and microbes between long and short survival patients. Significant correlation coefficients (p < 0.05) are labelled in the figure. D Correlation heatmap of differential microbes and immune-cells in patients. Vertical axis shows differential microbes and horizontal axis shows immuno-cells. Red represents up-regulated genes and blue represents down-regulated genes, while a deeper color means a higher expression level. The lower three rows represent microbes related to poor prognosis, while the rest represents microbes related to good prognosis. Significant correlation coefficients (p < 0.05) are labelled in the figure. E Correlation heatmap of core microbes and immune-cells in patients. Vertical axis shows core microbes and horizontal axis shows immune-cells. Red represents up-regulated genes and blue represents down-regulated genes, while a deeper color means a higher expression level. Significant correlation coefficients (p < 0.05) are labelled in the figure
To investigate the correlation between differential tumor microbes and immune genes, we conducted Pearson correlation analysis between 13 immune genes and the 10 differential microbes that exhibited significant differences between patients with short and long OS (Fig. 3C). Intriguingly, microbes associated with short OS displayed positive correlations with most genes responsible for inflammation, anti-tumor responses, and innate immunity. Additionally, they exhibited negative correlations with genes related to adaptive immunity. Conversely, microbes linked to long OS demonstrated opposite correlations. Particularly noteworthy is the correlation coefficient of 0.59 between MASP1 and actinomycetes, which is notably higher than the correlations observed between other genes and microbes. MASP1 encodes a serine protease, a crucial component of the complement-activated lectin pathway that plays a pivotal role in both innate and adaptive immune responses [32]. These findings suggest a potential association between actinomycetes and both innate and adaptive immunity in PDAC.
To examine the relationship between microbes and tumor-infiltrating immune cells, we utilized the R package xCell (version 1.1.0) [26] to calculate the relative contents of tumor-infiltrating immune cells in each sample, based on the gene expression profiles of PDAC patients. We then conducted Pearson correlation analysis on 23 representative immune cells and the differential microbes. The correlation heatmap is depicted in Fig. 3D. Notably, microbes associated with poor prognosis, such as Desulfotalea, Porphyromonas, and Stenotrophomonas, exhibit a more negative correlation with the relative contents of immune cells, indicating potential immunosuppressive effects. On the other hand, other microbes linked to a favorable prognosis display more positive correlations with immune cell relative contents, suggesting potential immune activation.
In our examination of the three core Stenotrophomonas microbes most pertinent to survival among the differential microbes, we performed Pearson correlation analysis to probe into the consistency of their relationship with immune cells (Fig. 3E). The outcomes disclosed negative correlations between these Stenotrophomonas microbes associated with poor prognosis and immune cells. This observation suggests a potential causal link to immunosuppression.
3.4 Risk scores stratify patients into different risk groups with distinct immune characteristics
We downloaded the gene expression profiles of PDAC from TCGA. Utilizing the previously established risk score model, we classified patients into either high-risk or low-risk groups based on their respective risk scores. There were 21 patients in the high-risk group and 23 in the low-risk group. Standardized RNA-seq counts were employed as the input matrix, and we conducted differential expression analysis between the two groups using DESeq2. A total of 145 genes were identified using a threshold of log2FC > 1 and P-adj < 0.05. Among these genes, 39 were up-regulated and 106 were down-regulated. The volcano plot of the DEGs is depicted in Fig. 4A. Among the DEGs, several were found to be associated with critical biological processes. For instance, HES2 is recognized as one of the core genes in the NOTCH1-downstream pathway, known for its involvement in cell proliferation. NTS may have a role in the regulation of lipid metabolism, potentially exerting endocrine or paracrine effects. REG3G is implicated in microbe resistance, while IGHM is related to adaptive immunity, indicating its potential involvement in the immune response. In Fig. 4B, the volcano plot features labeled genes, and the heatmap displays the expression levels of these genes. Notably, we observed contrasting expression patterns in immune-related genes between high-risk and low-risk patients. Specifically, in the high-risk group, up-regulated genes are predominantly associated with cell proliferation, while in the low-risk group, up-regulated genes are primarily linked to adaptive immunity and microbe resistance.
Survival risk score distinguished high and low-risk patients with different immune-related characteristics. A Volcano plot of differential genes of patients with high and low risk scores. Horizontal axis shows log2 Fold Change and vertical axis shows -log10 (padj). Comparison is made from high risk scores to low risk scores. With each dot representing a gene, red ones mean up-regulated genes, blue ones stand for down-regulated genes and grey ones represent genes that failed to meet threshold conditions. Threshold conditions are log2 Fold Change > 1 and padj < 0.05. B Heatmap of differential immune-genes of high and low risk patients. Low: low risk samples. Red represents up-regulated genes and blue represents down-regulated genes, while a deeper color means a higher expression level. High: high risk samples. This figure shows 26 genes, in which 2 are up-regulated and 23 down-regulated. Functions of immune-genes are listed at the right. C KEGG pathway bar chart of down-regulated genes in high and low risk patients. D GO pathway enrichment of up-regulated genes in high and low risk patients. E GO pathway enrichment of down-regulated genes in high and low risk patients. The bar plot shows some of the highest scoring pathways in biological process, cellular component and molecular function. F Correlation heatmap of differential genes and microbes of high and low risk patients. Vertical axis represents genes and horizontal axis represents differential microbes. Red represents up-regulated genes and blue represents down-regulated genes, while a deeper color means a higher expression level. The left three columns are microbes related to poor prognosis, while the rest represents microbes related to good prognosis. Significant correlation coefficients (p < 0.05) are labelled in the figure
The 145 DEGs were categorized into up-regulated and down-regulated genes, and their biological functions were evaluated using gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis (Fig. 4C). We employed the R package ClusterProfiler (version 4.0.5) [25] and applied a significance level of P < 0.05. Up-regulated genes were found to be enriched in biological processes such as keratinization, keratinocyte differentiation, skin development, and epidermal cell differentiation (Fig. 4D). On the other hand, down-regulated genes showed enrichment in biological processes including humoral immune response, immunoglobulin production, complement activation classical pathway, and B cell receptor signaling pathway (Fig. 4E). Regarding KEGG analysis, up-regulated genes did not yield enriched pathways, likely due to an insufficient number of up-regulated genes. In contrast, down-regulated genes were primarily enriched in pathways related to pancreatic secretion, protein digestion and absorption, fat digestion and absorption, and neuroactive ligand-receptor interaction (Fig. 4C).
We conducted correlation analysis on the differential genes and microbes in patients to explore the relationship between microbes and biological processes. Given that both gene and microbial expression profiles are continuous, we employed the Pearson correlation coefficient to assess the correlation. The heatmap in Fig. 4F illustrates the association between immune genes and the 10 differential microbes. Microbes linked to poor prognoses, such as Desulfotalea, Porphyromonas, and Stenotrophomonas, exhibited a positive correlation with genes related to cell proliferation (e.g., HES2 and IL1R2) and displayed a negative correlation with genes related to adaptive immunity and bacteria resistance (e.g., IGKV3D-20 and REG3G). Notably, Stenotrophomonas demonstrated a highly negative correlation with adaptive immunity genes. Based on the opposite correlation patterns observed in microbes related to better and worse prognosis, we speculate that poor prognosis-related microbes may promote cell proliferation and inhibit the immune response, thereby facilitating cancer progression and shortening patient survival. Conversely, microbes associated with a positive prognosis were linked to immune response activation and the suppression of cell growth, which hinders cancer progression and prolongs patient survival.
3.5 Weighted correlation network analysis
In the Weighted Correlation Network Analysis (WGCNA) of the transcriptome sequencing results of pancreatic ductal adenocarcinoma, we identified 16 modules, each designated by a distinct color (Fig. 5A). To explore the association between these 16 modules and the differential microbes, core microbes, survival, and the risk score established above, we conducted Pearson correlation analysis (Fig. 5B). Module MEgrey60 exhibited a significant negative correlation with the good prognosis-related microbe Mediterraneibacter. Conversely, Module MEblack demonstrated a notable positive correlation with the good prognosis-related microbe Neorickettsia, and it displayed a negative correlation with all poor prognosis-related microbes.
Characteristics of microbes in high and low-risk patients. A, B Correlation heatmap of WGCNA gene modules and microbes and risk scores. Vertical axis shows differential microbes, core microbes, survival, risk scores and levels. Horizontal axis shows 16 modules obtained by WGCNA. Red represents up-regulated genes and blue represents down-regulated genes, while a deeper color means a higher expression level. Significant correlation coefficients (p < 0.05) are labelled in the figure. C GO analysis bar charts of genes correlated to module MEgrey60. D GO analysis bar charts of genes correlated to module MEblack. The figure shows some of the highest scoring pathways in biological process, cellular component and molecular function. E and F GSEA enrichment analysis of up-regulated (above) and down-regulated (below) genes in high and low risk patients. Five pathways are shown in each graph. Screening parameters: |NES|> 1, P value < 0.05, FDR < 0.25
We performed Gene Ontology (GO) analysis on the genes corresponding to modules MEgrey60 and MEblack, respectively (Fig. 5C and D). Module MEgrey60 comprises 528 genes, and these genes were predominantly enriched in biological processes related to cell division (Fig. 5C). This suggests that MEgrey60 is likely associated with the inhibition of Mediterraneibacter, leading to cancer cell division and a poor prognosis. On the other hand, Module MEblack encompasses 7841 genes, primarily enriched in biological processes related to immunity (Fig. 5D). This implies that this module may be linked to the promotion of Neorickettsia growth, which could inhibit the growth of core microbes and activate the immune system. These effects could collectively contribute to an improved prognosis for patients.
To further elucidate pathways associated with expression patterns of MEgrey60 and MEblack, we conducted Gene Set Enrichment Analysis (GSEA) (Fig. 5E and F). The analysis revealed that MEblack genes are enriched in pathways related to epidermal cell differentiation, epidermis development, interleukin-1 receptor binding, and structural constituent of chromatin (Fig. 5E). On the other hand, MEgrey60 genes are enriched in pathways related to antigen binding, B cell receptor signaling pathway, immunoglobulin complex, phagocytosis, and T cell receptor complex (Fig. 5F). Based on these findings, we hypothesize that microbes within tumor niches in high-risk patients are likely related to cell proliferation and differentiation. The high expression levels of genes responsible for cell proliferation could potentially promote cancer development and contribute to a worsened prognosis in these patients. Conversely, immune genes are highly expressed in low-risk patients. This indicates that tumor microbes in low-risk patients might be related to immune activation, which could inhibit cancer development and extend patient survival. These observations align with the GO analysis.
4 Discussion
In our investigation, we undertook a thorough analysis of RNA-seq data and clinical records obtained from patients afflicted with pancreatic ductal adenocarcinoma, leveraging the extensive resources available from TCGA. We extracted microbiome data from transcriptome records and delved deeper into the intricate interplay between the microbiome, gene expression, and immune cells. This was achieved through a combination of correlation analysis and multi-omics assessments.
Following the extraction of microbial expression profiles from PDAC RNA-seq data, a meticulous process of decontamination and standardization was carried out. Subsequently, patients were stratified into distinct groups based on their respective survival outcomes. Our comprehensive analysis shed light on the broader landscape of tumor microbiology within pancreatic ductal adenocarcinoma. While the assessment of α and β diversity in tumor bacteria and fungi showed marginal disparities, LEFSe analysis pinpointed a notable distinction in the presence of Actinomyces between the two groups. In our pursuit of identifying clinically relevant microbes, we employed a combination of rigorous statistical tests and thorough literature research. The culmination of this effort was the recognition of Stenotrophomonas as a key player with a substantial impact on patient survival. A deeper dive into the Stenotrophomonas genus, coupled with additional Cox regression analyses, led to the identification of three core microbes whose influence on survival was particularly significant. Building on the foundation of these core microbes, we constructed a risk score model designed to prognosticate patient survival. We investigated the relationship between core microbes and clinical variables, finding that one of the core microbes, along with chemotherapy, significantly influenced survival. This finding underscores the independent role of Stenotrophomonas-associated core microbes in shaping patient outcomes, distinct from the impact of chemotherapy alone.
Following the differential expression analysis of PDAC patient gene profiles based on survival duration, we pinpointed 143 genes with notable disparities. Correlation assessments between immune genes, differential microbes, and survival revealed a distinctive pattern. Microbes linked to shorter survival displayed positive correlations with pro-inflammatory, anti-tumor, and innate immunity-related genes, but were negatively correlated with adaptive immunity genes. Conversely, microbes associated with extended survival exhibited an opposing trend. We subsequently stratified patients into high-risk and low-risk groups using the risk score model, identifying 145 differentially expressed genes. Enrichment analyses spotlighted up-regulated genes in pathways linked to cell growth, while down-regulated genes were associated with immune system activation. This implies that high-risk patients might exhibit heightened expression of genes promoting cell proliferation, potentially exacerbating cancer progression. In contrast, low-risk patients demonstrated elevated expression of immune-related genes, suggesting an activated immune response that inhibits cancer advancement. Further examination of the correlation between differential microbes and immune cells unveiled that microbes correlated with poor prognosis had a negative association with immune cell content, implying potential immunosuppressive effects. Conversely, microbes linked to a favorable prognosis exhibited a positive correlation with immune activation. The subsequent Weighted Correlation Network Analysis (WGCNA) identified 16 modules, with MEgrey60 being linked to the inhibition of beneficial microbes and cancer cell proliferation, potentially leading to poorer prognosis. In contrast, Module MEblack was associated with the promotion of beneficial microbe development and inhibition of core microbes, thereby activating the immune system and potentially improving prognosis.
Notably, two of the 10 distinctive microorganisms identified in our study align with those observed in independent data collected by Riquelme et al. [16] from patients afflicted with ductal carcinoma of the pancreas, specifically Pseudoxanthomonas spp. and Actinobacteria spp. This intriguing finding raises the possibility that these two organisms may be indigenous to the pancreas and exert a discernible influence on tumor development. To validate this hypothesis, we plan to conduct in vitro cultures of these bacteria and co-culture them with pancreatic ductal carcinoma cells. This will enable us to ascertain whether the beneficial bacteria extend the survival duration of pancreatic ductal carcinoma cells, while potentially detrimental bacteria may lead to a reduction in cell survival. Should the experimental outcomes substantiate our hypothesis, we will embark on an in-depth exploration of the functions and underlying mechanisms of these distinct microorganisms, employing a mouse model of pancreatic ductal carcinoma.
The microbiome composition can vary significantly depending on the anatomical location of the tumor within the pancreas. Studies have indicated differences in microbial diversity and abundance between tumors located in the head versus the tail of the pancreas. Geller et al. found that the microbiome in pancreatic cancer is distinct from that in healthy pancreatic tissue and that specific bacterial populations are associated with different regions of the pancreas [33]. The presence of bacteria in PDAC tissues has been linked to tumor characteristics and patient outcomes. Pushalkar et al. [34] demonstrated that bacterial colonization in PDAC tissues is associated with an immunosuppressive tumor microenvironment, which can influence tumor growth and patient prognosis. This study highlighted that bacterial load and diversity might differ between tumors in the head and tail of the pancreas, potentially due to variations in the local microenvironment and immune response. The study also indicated that oral antibiotic administration can slow oncogenic progression and may be useful as a chemopreventive measure for high-risk patients or individuals with an increased genetic risk for PDAC development. Differences in the microbiome based on tumor location can have clinical implications for the diagnosis, treatment, and prognosis of PDAC. Riquelme et al. [16] explored how the intratumoral microbiome affects patient survival and response to treatment, suggesting that microbial composition could be used as a biomarker for tailoring therapeutic approaches.
Neoadjuvant therapies, administered before the main treatment (usually surgery) to shrink tumors, play a critical role in the management of PDAC. Recent research has begun to uncover how these therapies may influence the tumor microbiome, potentially affecting treatment outcomes and patient prognosis. Pushalkar et al. [34] indicated that bacterial colonization in PDAC tissues could contribute to an immunosuppressive environment. Thus, modifying the microbiome through neoadjuvant therapies might help reduce immune suppression, enhancing the effectiveness of subsequent surgical and adjuvant treatments. Changes in microbiome composition following neoadjuvant therapy could serve as potential biomarkers for treatment response. Research suggests that certain microbial signatures are associated with different outcomes in PDAC patients. Riquelme et al. [16] found that a more diverse tumor microbiome was associated with improved survival rates, suggesting that monitoring microbiome changes during neoadjuvant therapy could provide insights into patient prognosis and guide personalized treatment strategies.
Research on microbes in pancreatic ductal adenocarcinoma is currently in its nascent stage. Previous studies have been constrained by small sample sizes and a lack of decontamination procedures. In contrast, our investigation offers a comprehensive characterization of intra-tumor microbial profiles. We have identified specific microbial strains that exert a significant impact on patient survival. Furthermore, we have established a robust risk score model for prognostic assessment. Our study sheds light on potential microbial pathways that influence survival outcomes. These findings not only advance our comprehension of tumor-microbe interactions but also provide valuable prognostic insights. Additionally, they pave the way for further research into the intricate interplay between tumor microbes and the immune system in pancreatic ductal adenocarcinoma.
5 Conclusions
To conclude, our study represents a pioneering effort in elucidating the role of the intratumoral microbiome in pancreatic ductal adenocarcinoma (PDAC), an area that has been subject to ongoing debate. We overcame limitations of previous studies, such as sample size constraints and inadequate decontamination procedures, revealing that PDAC tissues harbor a less diverse and distinct microbiota compared to non-tumor tissues. Leveraging microbiota associated with patient survival, we constructed a risk signature for PDAC subtyping. This microbiome-based subtyping demonstrated significant correlations with clinical prognosis and immune environments. Microbes linked to poorer prognosis were associated with pro-tumor immune activation, while those associated with better prognosis were linked to anti-tumor immune response and improved outcomes. Our study highlights the intricate relationship between the PDAC microbiota, clinical prognosis, and tumor microenvironment, suggesting that microbiota could serve as a promising biomarker for predicting PDAC prognosis.
6 Limitations
Several limitations need to be acknowledged. Firstly, this research focused solely on samples of pancreatic ductal adenocarcinoma. Secondly, the presence of contaminations was deduced from the sequencing data, reducing their reliability. Lastly, the analysis was conducted only in silico. Further in vitro experiments are necessary to confirm the interactions between the tumor and the microbiota.
Data availability
The data that support the findings of this study are available from TCGA but restrictions apply to the availability of these data, which were used under license for the current study, and so are not publicly available. Data are however available from the authors upon reasonable request and with permission of TCGA.
References
Williams PA, Zaidi SK, Sengupta R. AACR cancer progress report 2022: decoding cancer complexity, integrating science, and transforming patient outcomes. Clin Cancer Res. 2022;28(19):4178–9.
Chakladar J, Kuo SZ, Castaneda G, Li WT, Gnanasekar A, Yu MA, Chang EY, Wang XQ, Ongkeko WM. The pancreatic microbiome is associated with carcinogenesis and worse prognosis in males and smokers. Cancers (Basel). 2020;12(9):2672.
Long J, Zhang Y, Yu X, Yang J, LeBrun DG, Chen C, Yao Q, Li M. Overcoming drug resistance in pancreatic cancer. Expert Opin Ther Targets. 2011;15(7):817–28.
Wang Z, Li Y, Ahmad A, Banerjee S, Azmi AS, Kong D, Sarkar FH. Pancreatic cancer: understanding and overcoming chemoresistance. Nat Rev Gastroenterol Hepatol. 2011;8(1):27–33.
Gnanamony M, Gondi CS. Chemoresistance in pancreatic cancer: emerging concepts. Oncol Lett. 2017;13(4):2507–13.
Neoptolemos JP, Kleeff J, Michl P, Costello E, Greenhalf W, Palmer DH. Therapeutic developments in pancreatic cancer: current and future perspectives. Nat Rev Gastroenterol Hepatol. 2018;15(6):333–48.
Knippel RJ, Drewes JL, Sears CL. The cancer microbiome: recent highlights and knowledge gaps. Cancer Discov. 2021;11(10):2378–95.
Shreiner AB, Kao JY, Young VB. The gut microbiome in health and in disease. Curr Opin Gastroenterol. 2015;31(1):69–75.
Nejman D, Livyatan I, Fuks G, Gavert N, Zwang Y, Geller LT, Rotter-Maskowitz A, Weiser R, Mallel G, Gigi E, Meltser A, Douglas GM, Kamer I, Gopalakrishnan V, Dadosh T, Levin-Zaidman S, Avnet S, Atlan T, Cooper ZA, Arora R, Cogdill AP, Khan MAW, Ologun G, Bussi Y, Weinberger A, Lotan-Pompan M, Golani O, Perry G, Rokah M, Bahar-Shany K, Rozeman EA, Blank CU, Ronai A, Shaoul R, Amit A, Dorfman T, Kremer R, Cohen ZR, Harnof S, Siegal T, Yehuda-Shnaidman E, Gal-Yam EN, Shapira H, Baldini N, Langille MGI, Ben-Nun A, Kaufman B, Nissan A, Golan T, Dadiani M, Levanon K, Bar J, Yust-Katz S, Barshack I, Peeper DS, Raz DJ, Segal E, Wargo JA, Sandbank J, Shental N, Straussman R. The human tumor microbiome is composed of tumor type-specific intracellular bacteria. Science. 2020;368(6494):973–80.
Poore GD, Kopylova E, Zhu Q, Carpenter C, Fraraccio S, Wandro S, Kosciolek T, Janssen S, Metcalf J, Song SJ, Kanbar J, Miller-Montgomery S, Heaton R, McKay R, Patel SP, Swafford AD, Knight R. Microbiome analyses of blood and tissues suggest cancer diagnostic approach. Nature. 2020;579(7800):567–74.
Aykut B, Pushalkar S, Chen R, Li Q, Abengozar R, Kim JI, Shadaloey SA, Wu D, Preiss P, Verma N, Guo Y, Saxena A, Vardhan M, Diskin B, Wang W, Leinwand J, Kurz E, Kochen Rossi JA, Hundeyin M, Zambrinis C, Li X, Saxena D, Miller G. The fungal mycobiome promotes pancreatic oncogenesis via activation of MBL. Nature. 2019;574(7777):264–7.
Tang KW, Alaei-Mahabadi B, Samuelsson T, Lindh M, Larsson E. The landscape of viral expression and host gene fusion and adaptation in human cancer. Nat Commun. 2013;4:2513.
Zitvogel L, Ma Y, Raoult D, Kroemer G, Gajewski TF. The microbiome in cancer immunotherapy: diagnostic tools and therapeutic strategies. Science. 2018;359(6382):1366–70.
Guo W, Zhang Y, Guo S, Mei Z, Liao H, Dong H, Wu K, Ye H, Zhang Y, Zhu Y, Lang J, Hu L, Jin G, Kong X. Tumor microbiome contributes to an aggressive phenotype in the basal-like subtype of pancreatic cancer. Commun Biol. 2021;4(1):1019.
Nalluri H, Jensen E, Staley C. Role of biliary stent and neoadjuvant chemotherapy in the pancreatic tumor microbiome. BMC Microbiol. 2021;21(1):280.
Riquelme E, Zhang Y, Zhang L, Montiel M, Zoltan M, Dong W, Quesada P, Sahin I, Chandra V, San Lucas A, Scheet P, Xu H, Hanash SM, Feng L, Burks JK, Do KA, Peterson CB, Nejman D, Tzeng CD, Kim MP, Sears CL, Ajami N, Petrosino J, Wood LD, Maitra A, Straussman R, Katz M, White JR, Jenq R, Wargo J, McAllister F. Tumor microbiome diversity and composition influence pancreatic cancer outcomes. Cell. 2019;178(4):795–806.
Dohlman AB, Arguijo Mendoza D, Ding S, Gao M, Dressman H, Iliev ID, Lipkin SM, Shen X. The cancer microbiome atlas: a pan-cancer comparative analysis to distinguish tissue-resident microbiota from contaminants. Cell Host Microbe. 2021;29(2):281–98.
Grossman RL, Heath AP, Ferretti V, Varmus HE, Lowy DR, Kibbe WA, Staudt LM. Toward a shared vision for cancer genomic data. N Engl J Med. 2016;375(12):1109–12.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29(1):15–21.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R. The sequence alignment/map format and SAMtools. Bioinformatics. 2009;25(16):2078–9.
Davis NM, Proctor DM, Holmes SP, Relman DA, Callahan BJ. Simple statistical identification and removal of contaminant sequences in marker-gene and metagenomics data. Microbiome. 2018;6(1):226.
Law CW, Chen Y, Shi W, Smyth GK. voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 2014;15(2):R29.
Mecham BH, Nelson PS, Storey JD. Supervised normalization of microarrays. Bioinformatics. 2010;26(10):1308–15.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15(12):550.
Wu T, Hu E, Xu S, Chen M, Guo P, Dai Z, Feng T, Zhou L, Tang W, Zhan L, Fu X, Liu S, Bo X, Yu G. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation (Camb). 2021;2(3):100141.
Aran D, Hu Z, Butte AJ. xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 2017;18(1):220.
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, Hoang CD, Diehn M, Alizadeh AA. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015;12(5):453–7.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 2008;9:559.
Wood DE, Lu J, Langmead B. Improved metagenomic analysis with Kraken 2. Genome Biol. 2019;20(1):257.
Ryan RP, Monchy S, Cardinale M, Taghavi S, Crossman L, Avison MB, Berg G, van der Lelie D, Dow JM. The versatility and adaptation of bacteria from the genus Stenotrophomonas. Nat Rev Microbiol. 2009;7(7):514–25.
Liu B, Zhou Z, Jin Y, Lu J, Feng D, Peng R, Sun H, Mu X, Li C, Chen Y. Hepatic stellate cell activation and senescence induced by intrahepatic microbiota disturbances drive progression of liver cirrhosis toward hepatocellular carcinoma. J Immunother Cancer. 2022;10(1):514–25.
Matsushita M, Endo Y, Fujita T. MASP1 (MBL-associated serine protease 1). Immunobiology. 1998;199(2):340–7.
Geller LT, Barzily-Rokni M, Danino T, Jonas OH, Shental N, Nejman D, Gavert N, Zwang Y, Cooper ZA, Shee K, Thaiss CA, Reuben A, Livny J, Avraham R, Frederick DT, Ligorio M, Chatman K, Johnston SE, Mosher CM, Brandis A, Fuks G, Gurbatri C, Gopalakrishnan V, Kim M, Hurd MW, Katz M, Fleming J, Maitra A, Smith DA, Skalak M, Bu J, Michaud M, Trauger SA, Barshack I, Golan T, Sandbank J, Flaherty KT, Mandinova A, Garrett WS, Thayer SP, Ferrone CR, Huttenhower C, Bhatia SN, Gevers D, Wargo JA, Golub TR, Straussman R. Potential role of intratumor bacteria in mediating tumor resistance to the chemotherapeutic drug gemcitabine. Science. 2017;357(6356):1156–60.
Pushalkar S, Hundeyin M, Daley D, Zambirinis CP, Kurz E, Mishra A, Mohan N, Aykut B, Usyk M, Torres LE, Werba G, Zhang K, Guo Y, Li Q, Akkad N, Lall S, Wadowski B, Gutierrez J, Kochen Rossi JA, Herzog JW, Diskin B, Torres-Hernandez A, Leinwand J, Wang W, Taunk PS, Savadkar S, Janal M, Saxena A, Li X, Cohen D, Sartor RB, Saxena D, Miller G. The pancreatic cancer microbiome promotes oncogenesis by induction of innate and adaptive immune suppression. Cancer Discov. 2018;8(4):403–16.
Acknowledgements
We would like to express our sincere gratitude to all individuals and organizations who contributed to this research.
Funding
This study is funded by China Postdoctoral Science Foundation (No. 2023T160328, No.2023M731765) and Beijing Municipal Natural Science Foundation (No. Z19J00085).
Author information
Authors and Affiliations
Contributions
J.L., H.X., and X.Lan contributed to the initial idea and conceptualization of the review article. J.L., H.X., and X.Liu were responsible for writing the initial draft of the review article. J.L., H.X., X.Liu, Y. Y., C.N., J.Q., X.K. and L.S. prepared the figures. J.L., H.X., and X.Liu prepared the tables. X.Lan reviewed and revised the manuscript. All authors listed have made a substantial contribution to the work. All authors have read and approved the article.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Leng, J., Xu, H., Liu, X. et al. Intratumoral microbiota of pancreatic ductal adenocarcinoma impact patient prognosis by influencing tumor microenvironment. Discov Onc 15, 443 (2024). https://doi.org/10.1007/s12672-024-01320-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-024-01320-6