Abstract
Root-associated microbial communities have strong influences on the health and development of plants. Through the secretion of root exudates, the soil microbiome is impacted by plants, thereby steering plant-soil reactions. Considering the importance of root exudates in the establishment of symbiotic associations in the rhizosphere, it is quite clear that understanding the interaction between plant roots and the soil microbiome may prove beneficial. Here, we review the interaction between plant root exudates and microbial communities in the soil. The influence of these bioactive molecules on the structure and function of soil microbes is also considered. We additionally, deliberate on how plants determine the soil microbiome and how they extract nutrients from endophytes for the augmentation of their growth and development. A good perspective of the communication between plant roots and the soil microbiome could lead to increased crop production, thereby limiting the need for synthetic fertilizers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The rhizosphere is an intricate ecosystem inhabited by several organisms such as bacteria, fungi, nematodes, and arthropods. Augmented bacterial abundance and activity are features of the rhizosphere relative to bulk soil; it however, reportedly has lower diversity. In a complex manner, plant roots interact with microbial communities in the soil through chemical communication in the rhizosphere. Interactions with the complex microbial community known as the root microbiome have the prospective of inducing plant growth and development. Microorganisms are major contributors to nutrient cycles and facilitate the acquisition of nutrients by plants (Amoo and Babalola 2017; Lareen et al. 2016; Mhlongo et al. 2018).
The root microbiome has been regarded as the ‘secondary genome’ because host plants get compounds and qualities that are consequential from microbes there. For the formation of symbiotic associations with plants, beneficial molecules that can alter some plant traits are produced by rhizobacteria (Lareen et al. 2016; Mhlongo et al. 2018). Numerous organisms possess the ability to impact their local ecosystem thus altering individual functioning. Plants can influence the soil microbiota through the exudation of bioactive molecules into the rhizosphere. Root exudates characterize carbon and nitrogen substrates for the growth of microbes, and they consist of primary and secondary metabolites. Through the alteration of microbial communities, root exudate metabolites can steer plant-soil reactions. Root exudates influence microorganisms present in the rhizosphere by operating as repellants, signaling molecules, stimulants, or attractants (Olanrewaju et al. 2019).
The increasing population of humans has placed more demand on food production, and this has resulted in the overuse of synthetic fertilizers by some farmers. Fertilization has been reported to impact microbiome-mediated processes in plants such as modification of hormones and alteration of resource accessibility, which causes vicissitudes in competition among microbiota. Intensifying fertilizer application can alter the availability of nutrients crucial to plant growth (Amoo et al. 2021; Enebe and Babalola 2021). Understanding how plant roots and the soil microbiome communicate is important because various studies have highlighted the influences root-associated microbial communities have on the health and development of plants. Here, the interactions between exudates produced by plant roots and soil microbes are discussed. We further discuss how hosts determine the microbiome and how plants extract nutrients from endosymbiotic microbes, thereby enhancing their growth and development. We conclude by pinpointing areas of prospective research necessary for the growth and development of plants.
1.1 Plant-root exudates and the soil microbiome
Plant-root exudation is a complex process involving factors that regulate carbon transport via the Munch's pressure flow hypothesis (Canarini et al. 2019). The hypothesis suggests that the movement of sap, which encompasses essential nutrients such as carbohydrates, is propelled by osmotic pressure gradients existing between source and sink tissues (Babst et al. 2022). Carbohydrates are loaded into the phloem at source regions (photosynthetic leaves), creating higher osmotic pressure, and flow toward sink regions (non-photosynthetic areas) where they are unloaded for growth and storage (Hunt et al. 2023).
Root exudates establish an intricate and interconnected association between plants and the microbiome functioning as substrates and signalling molecules for microorganisms. These exudates establish a distinctive environment in the rhizosphere and can attract and instigate pathogenic and symbiotic associations in the rhizosphere (Hu et al. 2018). Previous studies have revealed that modifications in the composition and concentration of root exudates in maize, pea, wheat and sugar beet at different developmental stages have caused changes in the structure of fungal and bacteria (Chaparro et al. 2014; Garcia and Kao-Kniffin 2018).
Vesicle transport, diffusion, and ion channels are some mechanisms by which exudates are released by plant roots (Olanrewaju et al. 2019). Root exudates move outside the root tips through the plasma membrane either by active or passive transport to reach the cell walls (Olanrewaju and Babalola 2022). Low-molecular-weight compounds (Table 1) can pass through the plasma membrane using specific flow carriers. The specific flow carriers for amino acids are Usually Multiple Acids Move In and out Transporters (UMAMIT) (Besnard et al. 2016) or the CAT transporters (cationic amino acid) (Jungnickel et al. 2018). The SWEET transporters (Julius et al. 2017) carry sugars while organic acids use aluminum-activated malate transporter (ALMT) (Palmer et al. 2016). These carriers do not need to be attached to ATP to move through the plasma membrane, they use passive mediated transport. High-molecular-weight compounds (Table 1) use ATP dependent carriers such as ABC transporters and MATE (citrate); thereby passing through the plasma membrane by active transport. Once these compounds exit the plasma membrane, they move between cell walls of the cortex to the soil environment (Canarini et al. 2019).
1.2 Chemical signals influencing microbial community structure and function in the soil microbiome
Employing chemical communication, plants can actively structure their environment. Plants exude organic substances into the soil thereby stimulating the growth of microbial communities in the soil. These microbes utilize this carbon pool as their principal supply. Organic compounds that characterize the majority of the molecules exuded into the soil by plants are emitted through a process known as rhizodeposition (Guerrieri et al. 2019). Root exudates possess biological and chemical influences on their immediate environment playing roles in nutrient acquisition and interaction with soil organisms. Chemical signaling between plants and other organisms inhabiting the soil is the biological effect of root exudates. Through recruiting defensive organisms upon infection by insects/pathogens or attracting beneficial microorganisms for the enhancement of nutrient uptake, plants can modify their rhizobiome (Pascale et al. 2020).
There are favorable chemicals released by the root exudates to microbes in the rhizosphere are called phytohormones, these chemicals have a key role in regulating the physiology of microorganisms in the plant rhizosphere for plant development (Bedini et al. 2018). Some of these classical phytohormones include flavonoids, strigolactones, and salicylic acid.
1.2.1 Flavonoids
Flavonoids are a diverse group of hormones in plants that are involved in growth, development, and reproduction. The absence of these hormones does not result in the immediate death of plants but rather long-term impairment of survivability. Flavonoid solvency and versatility in the soil differs. While glycosylation improves their solvency in water, flavonoid glycosides are rapidly deglycosylated by microorganisms leaving the more hydrophobic aglycone (Slámová et al. 2018). Soil microorganisms use these plant glycosides as a source of energy and control the degrees of the bioactive aglycone (Biernat et al. 2018). This prompts the expansion in microbial community structure and function and increased plant development and advancement (Jacoby et al. 2017). Infected plants such as legumes, soybeans, and alfalfa use flavonoids as the first defensive mechanism at the site of infection (Noor 2023). Flavonoids can add to plant growth and development by fixing the plant structures and tissues by balancing auxin (IAA) action, which can prompt the differentiation of tissues, advancement of callus and tylose development, and closing of the vascular system to prevent pathogen disease. It has also been discovered that the over-application of fertilizers influences the production of flavonoids. Deng et al. (2019) demonstrated that the over-application of nitrogen fertilizers in Cyclocarya paliurus decreases the production of flavonoids and results in the decrease of the plant growth and development.
1.2.2 Strigolactones
Strigolactones signal molecules are released by plants when there are nitrogen and phosphate deficiencies. They respond to the reaction of plants to environmental stress. Carvalhais et al. (2019) demonstrated that strigolactones shape microbial communities and functions in the rhizosphere. They reported that plants producing strigolactones have a major influence on fungal diversity compared to bacterial diversity relative to plants that do not produce strigolactones. They attract arbuscular mycorrhizal fungi for symbiosis with roots in approximately 80% of vascular plants to increase the uptake of the mineral nutrients like phosphate and they, in turn, receive carbohydrates leading to plant growth and development (De Cuyper and Goormachtig 2017). For instance, strigolactones are found in mosses, liverworts, and in the alga Chara coralline, where they advance rhizoid development. Strigolactones also attract rhizobia for nodulation where bacteria fix nitrogen in exchange for carbohydrates. According to Nelson (2021), strigolactones have disadvantages because they can activate seed germination in parasitic weeds. These seeds attach to the plant roots and then obtain nutrients from it.
1.2.3 Salicylic acid
Salicylic acid is believed to be a plant defence strategy against plant pathogenic fungi across different plant genotypes. However, it has been reported to also influence the establishment of bacteria from specific families in the rhizosphere (Veach et al. 2019). According to the study conducted by (Benjamin et al. 2022), it was shown that tomatoes (Solanum lycopersicum) emit salicylic acid, which serves a dual purpose of pathogen defence and facilitation of symbiotic associations with beneficial microorganisms such as mycorrhizal fungi. This suggests that salicylic acid contributes to microbial structure and function. Salicylic acid incites plant defence chemical in Populus in response to an infectious pathogen. Salicylic acid also induces stimulants that influence litter decomposition (Veach et al. 2019) and fungal endophyte colonization in other Populus species, promoting plant development (Omomowo and Babalola 2019).
1.3 Determination of microbiome by host
Root exudates show great variability among various plant species, growth stages, and even cultivars, inside various root developmental stages of individual plants and considering different biotic and abiotic stress factors (Adeleke et al. 2022b; Vieira et al. 2020). As a result of these differences, rhizobacteria with corresponding substrate uptake inclinations are chosen (Vieira et al. 2019; Zhalnina et al. 2018). For instance, C3 (three-carbon compound via the Calvin cycle) and C4 plants (four-carbon compound via the Calvin cycle) show dissimilarities in the kinds of exudates discharged into the rhizosphere (Chen et al. 2016). Predominant sugars in the two kinds of plants contrast, with the secretion of maltose, mannose, and ribose by C3 plants and inositol, erythritol, and ribitol in C4 plants (Olanrewaju et al. 2019). C4 plants (e.g. maize, sugarcane, sorghum) ooze higher quantities of organic carbon and amino acids due to high photosynthetic efficiency and absence of photorespiration while C3 plants (e.g. wheat, rye, oats, rice, cotton, sunflower) produce less organic carbon during photosynthesis due to photorespiration (Olanrewaju et al. 2019). This suggests that the microbial functions and community structure in C4 plants are much more abundant than those in C3 plants.
The potential link between C4 plants, nitrogen fixation, and increased root exudation presents a compelling perspective on nutrient acquisition strategies. C4 plants are known for their efficient water usage and carbon fixation, but a recent investigation reveals that some may also fix nitrogen (Schlüter et al. 2023). C4 plants increase root exudation to attract nitrogen-fixing bacteria, which is consistent with rhizosphere interactions. In nitrogen-deficient conditions, this hypothesized method might help C4 plants improve their nutrient uptake and perhaps depend more on nitrogen fixation than C3 plants. This dynamic interaction shows how plants modify their relationships with microbes to improve nutrient availability, affecting plant ecology and agricultural practices (Aluko et al. 2023).
Genotypic varieties in plants may likewise change rhizosphere microbiome communities, however to an exceptionally less degree (Liu et al. 2019). It was exhibited that even plants that fall under the same carbon fixation metabolic pathway can create distinctive root exudates from one another. For instance, (Vaughan et al. 2013) reported that the diterpene rhizathalene A was constitutively delivered and discharged from the roots of A. thaliana (C3 plants) while Xuan et al. (2016) showed that diterpene momilactone A is delivered and emitted from the foundations of O. sativa (C3 plants) seedlings into the rhizobiome, the molecules released by these plants are antimicrobial agents. Endogenous developments control varieties in root exudate arrangements (Olanrewaju et al. 2019).
According to previous studies, it has been proven that different stages in plant development have different impact on microbiome (Compant et al. 2019). The microbial diversity and function at seeding stages are much more abundant than in other plant stages such as vegetative, plant bolting and flowering since the plant roots discharge more root exudates at early plant stages than in developed plants (Olanrewaju et al. 2019). Microbial communities in young plants grow rapidly and can use simple substrates produced by these young plants. As these young plants mature, the root exudates they release shape the microbial structure and function by attracting beneficial microbes that degrade more complex substrates and reject infection-causing microorganisms. This results in slower growth rates and the dominance of specific microorganisms (Alawiye and Babalola 2019).
The host genotype also determines root exudates that selects for specific microorganisms that could be beneficial for plant growth and development and inhibition of pathogenic microorganisms. Through the secretion of a wide variety of compounds, roots may influence the soil microbial community both emphatically and adversely (David 2023). According to Wu and Yu (2019), the effect of root exudates of Eucalyptus urophylla and Acacia mearnsii on soil microbes under simulated warming climate conditions directly affected the biomass of microorganisms. It also indirectly affected the soil microbial community structure by altering the composition and content of plant root exudates.
1.4 Effects of rhizosphere microbiome on plant growth and development
Various parts of plants harbor microorganisms that are structurally and functionally diverse e.g. the phyllosphere, endosphere, and rhizosphere microbiomes. Several taxa of microbes, such as bacteria, fungi, archaea, and viruses, occupy the rhizosphere of plants and this boosts the chances of interactions influencing nutrient dynamics affecting plant growth. The microbial community found in the rhizosphere play key roles in the growth and reproduction of plants (Garcia and Kao-Kniffin 2018; Lu et al. 2017). Through the production of phytohormones, such as auxins, cytokinins, gibberellins, and abscisic acid, the rhizosphere microbiome increases plant growth, protects against pathogens, and may help tolerate abiotic stresses like drought (Fadiji et al. 2023).
Mutualistic symbionts and saprophytic microorganisms living at the root-soil interfaces are involved in nutrient cycling and the control of abiotic stress. Arbuscular mycorrhiza (AM) is an example of mutualistic symbionts and are also regarded as natural biofertilizers as they provide the host with water, nutrients, and pathogen protection, in exchange for photosynthetic products (Fasusi et al. 2023). AM establish beneficial interactions with most plant species by extending the root system to enhance nutrient uptake properties while saprophytic microorganisms are perceived for their ability to push nitrogen (N) fixation as well as phosphorus (P) rally; these two key procedures support plant development (Liu et al. 2018). According to a study conducted by Míguez-Montero et al. (2019), C. multiflorus, C. scoparius, C. balansae, and C. striatus leguminous species in water and phosphorus-deficient environment showed similar patterns of extending their root system when mineral nutrition was not sufficient in the soil to meet their physiological requirements. The rhizosphere microbiome also enhances stress resilience producing auxins, cytokinins, and gibberellins, increasing hormone levels in the host plant to intensify growth and development.
Microorganisms control diseases in plants through the inhibition of pathogenic microbial invasion by releasing antibiotics, lytic enzymes, or pathogen inhibiting volatile compounds (Compant et al. 2019). Different individuals from the rhizosphere microbiome can alienate soil-borne pathogens previously and during essential disease and optional spread on and in root tissues (Mhlongo et al. 2018). The rhizosphere microorganisms avert plant pathogens by antibiosis (Topolovec-Pintarić 2019) and opposition interference with quorum sensing affecting virulence and induced systemic resistance (Fleitas Martínez et al. 2019).
1.5 Mechanism of nutrient extraction from endosymbiotic microbes by plants
Endosymbiotic microbes refer to microorganisms that form a symbiotic association with a host organism, whereby they inhabit the host's cells or tissues and develop a mutually advantageous relationship (Sharma et al. 2023). Microbial endosymbionts offer their host plants a range of benefits comprising growth promotion, augmented disease protection, and enhanced tolerance against biotic and abiotic stresses. These endophytes promote plant growth by boosting nutrient acquisition. Certain plants are involved in ‘nitrogen-transfer symbioses’ which is a process in which plants partner with prokaryotes that fix nitrogen in connection with roots subsequently conveying nitrogen to plants e.g. actinorhizal and rhizobial symbioses (Martínez-Hidalgo and Hirsch 2017; White et al. 2018). In the context of phosphorus, arbuscular mycorrhizal fungi (AMF) enable plants get phosphorus. This symbiotic interaction allows the fungus to extend hyphae into the soil and release enzymes that break down insoluble phosphorus molecules, making it more available to plants (Agbodjato et al. 2022). In a similar vein, plants use the strategy of releasing siderophores into the rhizosphere to facilitate the uptake of iron. These siderophores act as chelating agents, effectively binding to iron ions and therefore increasing their solubility and accessibility for plant absorption (Roskova et al. 2022). This mechanism enhances the capacity of plant roots to effectively absorb iron, which is essential for several physiological functions. Collectively, these tactics highlight the ability of plants to interact with microorganisms as cooperative partners in mitigating nutritional deficiencies and enhancing overall development and well-being (Singh et al. 2022).
In nutrient-limited soils, the discharge of exudates by plants intensifies. This increase in exudation possibly enhances the activities of microorganisms around plant roots and boosts the ‘microbial mining’ of nutrients. The cultivation of microbial communities’ upsurges through the excretion of more exudates by plants under nutrient-limited conditions. Plants can be deemed to be ‘farming’ microbes by the emission of exudates (Jacoby et al. 2017).The rhizophagy cycle is an emerging ecological phenomenon where plants uptake bacteria or fungi as endophytes from the soil, subsequently utilizing reactive oxygen species (ROS) oxidation to break down these microorganisms (Fig. 1) (Verma et al. 2022). Rhizophagy involves the use of root exudates to enable bacteria enter root cells and survive and reproduce in plant tissues (Chiaranunt and White 2023). Plants produce ROS as a reaction to the presence of endophytes. ROS are highly reactive molecules that activate oxidative processes, leading to the degradation of endophytes and the subsequent release of nutrients, including nitrogen and phosphorus (Adeleke et al. 2022a). The assimilation of these nutrients supports the plant's development. Some bacteria, such as Klebsiella implicated in the rhizophagy cycle, have been reported to fix atmospheric nitrogen, while others lack this nitrogen-fixing ability. These bacteria that are unable to fix nitrogen, e.g., some species of Bacillus, only need to have proficiency in nutrient obtention from the rhizosphere by foraging from fungi, plants, or bacteria (Singh et al. 2018; White et al. 2018). Nutrient solubilization in the rhizosphere is made possible through the action of some symbiotic microorganisms and these solubilized nutrients can be taken up into the roots by root hairs. Bacteria linked to roots are usually capable of motion and can leave the plant root to obtain nutrients from the soil. They are also able to return to the plants to get supplementary carbon and nutrients from root exudates. Bacteria seem to be perfect transporters of micronutrients that are needed for the growth and development of plants. All nutrients obtained or produced by bacteria can be oxidatively mined from them through the process of rhizophagy (Verma and White 2019).
1.6 Perspectives in agriculture: synthetic fertilizers and plant root-soil microbe communications
Various interacting factors can influence the structure of microbial communities in the soil. Farm management practices including fertilization, crop rotation, and tillage modify soil physicochemical properties thereby impacting microbial diversity and community composition (Nwachukwu et al. 2023). The microbial community composition of conventionally managed systems varies from that of organic systems (Amoo and Babalola 2019). Concerning management, the influence of plants on microbial communities extends taxonomy, the network structure is also affected. Through alterations in pH instigated by synthetic fertilizers, soil acidification which implicitly enhances the abundance of acid-tolerant taxa may occur thereby influencing the structure of microbial communities. This could also trigger the alteration of the relative abundance of certain taxa (Li et al. 2017; Schmidt et al. 2019).
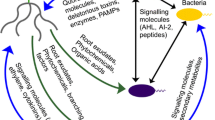
Towards meeting the food demand of an ever-increasing population, there is an over-application of synthetic fertilizers in most cases. When synthetic fertilizers are administered to soils, they get absorbed by plant roots thereby increasing crop yield (Chandini et al. 2019). It has been reported that the quorum-sensing system between the cross-domain and cross-kingdom microorganisms responds to environmental nutrition status. Over-fertilization results in the decrease of microbial communications thereby affecting the coordination of the microorganisms between cross-domain and cross-kingdom. The motility, virulence, and biofilm formation also get affected resulting in a decrease in the microbial community (Fig. 2) (Paluch et al. 2020).
The rhizosphere of the natural system vs that of artificial systems. The rhizosphere of the natural system: Plant roots release exudates which act as chemoattractant or repellent signalling to microorganisms in the rhizosphere. The cross-domain and cross-kingdom quorum-sensing system remains normal, resulting in the growth and production of healthy plants and increased crop yields. The rhizosphere of the artificial system: Plant roots release exudates that act as chemoattractant and repellents to microorganisms in the rhizosphere. The cross-domain and cross-kingdom quorum-sensing system decreases which result in poor growth and development thereby leading to reduced crop yields
1.7 Conclusions and prospects
The intricate interactions between plant roots and their associated complex microbial communities can stimulate plant growth and development. Through the secretion of bioactive molecules into the rhizosphere by plants, the soil microbiome may be influenced resulting in the steering of plant-soil reactions by these root exudate metabolites. Due to the ever-increasing human population, more demand has been placed on food production and this has led to synthetic fertilizers being overused by some farmers. Escalating fertilizer application can alter the accessibility of nutrients crucial for plant growth. Plants can structure their environments using chemical communication through root exudates. These exudates are involved in the nourishment and development of plants and the signaling of molecules that facilitate communication in the rhizobiome. The effects of root-associated microbial communities on plant health and development have been emphasized by several studies and it is of utmost importance to understand the communication between plant roots and the soil microbiome. Such information could lead to an improvement of cultivating techniques that help in the maintenance of advantageous microbial communities and reduction of pathogenic microorganisms in the rhizosphere consequently resulting in enhanced plant growth and development.
References
Adeleke BS, Akinola SA, Adedayo AA, Glick BR, Babalola OO (2022a) Synergistic relationship of endophyte-nanomaterials to alleviate abiotic stress in plants. Front Environ Sci 10:2398
Adeleke BS, Fadiji AE, Ayilara MS, Igiehon ON, Nwachukwu BC, Babalola OO (2022b) Strategies to enhance the use of endophytes as bioinoculants in agriculture. Horticulturae 8(6):498
Agbodjato NA, Assogba SA, Babalola OO, Koda AD, Aguégué RM, Sina H, Dagbénonbakin GD, Adjanohoun A, Baba-Moussa L (2022) Formulation of biostimulants based on arbuscular mycorrhizal fungi for maize growth and yield. Front Agron 4:894489
Alawiye TT, Babalola OO (2019) Bacterial diversity and community structure in typical plant rhizosphere. Diversity 11(10):179
Aluko OO, Liu Z, Sun X (2023) The interplay of carbon and nitrogen distribution: Prospects for improved crop yields. Mod Agric 1:57–75
Amoo AE, Babalola OO (2017) Ammonia-oxidizing microorganisms: key players in the promotion of plant growth. J Soil Sci Plant Nutr 17(4):935–947
Amoo AE, Babalola OO (2019) Impact of land use on bacterial diversity and community structure in temperate pine and indigenous forest soils. Diversity 11(11):217
Amoo AE, Enagbonma BJ, Ayangbenro AS, Babalola OO (2021) Biofertilizer: an eco-friendly approach for sustainable crop production. Food Secur Saf 1:647–669
Babst BA, Braun DM, Karve AA, Frank Baker R, Tran TM, Kenny DJ, Rohlhill J, Knoblauch J, Knoblauch M, Lohaus G (2022) Sugar loading is not required for phloem sap flow in maize plants. Nat Plants 8(2):171–180
Bedini A, Mercy L, Schneider C, Franken P, Lucic-Mercy E (2018) Unravelling the initial plant hormone signalling, metabolic mechanisms and plant defense triggering the endomycorrhizal symbiosis behavior. Front Plant Sci 9:1800
Benjamin G, Pandharikar G, Frendo P (2022) Salicylic acid in plant symbioses: beyond plant pathogen interactions. Biology 11(6):861
Besnard J, Pratelli R, Zhao C, Sonawala U, Collakova E, Pilot G, Okumoto S (2016) UMAMIT14 is an amino acid exporter involved in phloem unloading in Arabidopsis roots. J Exp Bot 67(22):6385–6397
Biernat KA, Li B, Redinbo MR (2018) Microbial unmasking of plant glycosides. Mbio 9(1):e02433-e2417
Canarini A, Kaiser C, Merchant A, Richter A, Wanek W (2019) Root exudation of primary metabolites: mechanisms and their roles in plant responses to environmental stimuli. Front Plant Sci 10:157
Carvalhais LC, Rincon-Florez VA, Brewer PB, Beveridge CA, Dennis PG, Schenk PM (2019) The ability of plants to produce strigolactones affects rhizosphere community composition of fungi but not bacteria. Rhizosphere 9:18–26
Chandini, KR, Kumar R, Prakash O (2019) the impact of chemical fertilizers on our environment and ecosystem. Chief Ed 35:69–86
Chaparro JM, Badri DV, Vivanco JM (2014) Rhizosphere microbiome assemblage is affected by plant development. ISME J 8(4):790–803
Chen J, Wang Q, Li M, Fan L, Li W (2016) Does the different photosynthetic pathway of plants affect soil respiration in a subtropical wetland? Ecol Evol 6:8010–8017
Chiaranunt P, White JF (2023) Plant beneficial bacteria and their potential applications in vertical farming systems. Plants 12(2):400
Compant S, Samad A, Faist H, Sessitsch A (2019) A review on the plant microbiome: ecology, functions and emerging trends in microbial application. J Adv Res 19:29–37
David OG (2023) Metagenomic analysis of rhizospheric soil microbial communities associated with Striga infested maize field. North-West University (South Africa)
De Cuyper C, Goormachtig S (2017) Strigolactones in the rhizosphere: friend or foe? Mol Plant Microbe Interact 30(9):683–690
Deng B, Li Y, Xu D, Ye Q, Liu G (2019) Nitrogen availability alters flavonoid accumulation in Cyclocarya paliurus via the effects on the internal carbon/nitrogen balance. Sci Rep 9(1):2370
Enebe MC, Babalola OO (2021) Soil fertilization affects the abundance and distribution of carbon and nitrogen cycling genes in the maize rhizosphere. AMB Express 11(1):24
Fadiji AE, Yadav AN, Santoyo G, Babalola OO (2023) Understanding the plant-microbe interactions in environments exposed to abiotic stresses: An overview. Microbiol Res 271:127368
Fasusi OA, Babalola OO, Adejumo TO (2023) Harnessing of plant growth-promoting rhizobacteria and arbuscular mycorrhizal fungi in agroecosystem sustainability. CABI Agric Biosci 4(1):26
FleitasMartínez O, Rigueiras PO, da Silva Pires Á, Porto WF, Silva ON, de la Fuente-Nunez C, Franco OL (2019) Interference with quorum-sensing signal biosynthesis as a promising therapeutic strategy against multidrug-resistant pathogens. Front Cell Infect Microbiol 8:444–444
Garcia J, Kao-Kniffin J (2018) Microbial group dynamics in plant rhizospheres and their implications on nutrient cycling. Front Microbiol 9:1516
Guerrieri A, Dong L, Bouwmeester HJ (2019) Role and exploitation of underground chemical signaling in plants. Pest Manag Sci 75(9):2455–2463
Hu L, Robert CA, Cadot S, Zhang X, Ye M, Li B, Manzo D, Chervet N, Steinger T, Van Der Heijden MG (2018) Root exudate metabolites drive plant-soil feedbacks on growth and defense by shaping the rhizosphere microbiota. Nat Commun 9(1):2738
Hunt H, Brueggen N, Galle A, Vanderauwera S, Frohberg C, Fernie AR, Sonnewald U, Sweetlove LJ (2023) Analysis of companion cell and phloem metabolism using a transcriptome-guided model of Arabidopsis metabolism. Plant Physiol 192(2):1359–1377
Jacoby R, Peukert M, Succurro A, Koprivova A, Kopriva S (2017) The role of soil microorganisms in plant mineral nutrition-current knowledge and future directions. Front Plant Sci 8:1617–1617
Julius BT, Leach KA, Tran TM, Mertz RA, Braun DM (2017) Sugar transporters in plants: new insights and discoveries. Plant Cell Physiol 58(9):1442–1460
Jungnickel KEJ, Parker JL, Newstead S (2018) Structural basis for amino acid transport by the CAT family of SLC7 transporters. Nat Commun 9(1):550–550
Lareen A, Burton F, Schäfer P (2016) Plant root-microbe communication in shaping root microbiomes. Plant Mol Biol 90(6):575–587
Li F, Chen L, Zhang J, Yin J, Huang S (2017) Bacterial community structure after long-term organic and inorganic fertilization reveals important associations between soil nutrients and specific taxa involved in nutrient transformations. Front Microbiol 8:187–187
Liu A, Contador CA, Fan K, Lam H-M (2018) Interaction and regulation of carbon, nitrogen, and phosphorus metabolisms in root nodules of legumes. Front Plant Sci 9:1860
Liu F, Hewezi T, Lebeis SL, Pantalone V, Grewal PS, Staton ME (2019) Soil indigenous microbiome and plant genotypes cooperatively modify soybean rhizosphere microbiome assembly. BMC Microbiol 19(1):1–19
Lu H, Sun J, Zhu L (2017) The role of artificial root exudate components in facilitating the degradation of pyrene in soil. Sci Rep 7(1):7130
Martínez-Hidalgo P, Hirsch AM (2017) The nodule microbiome: N2-fixing rhizobia do not live alone. Phytobiomes 1(2):70–82
Mhlongo MI, Piater LA, Madala NE, Labuschagne N, Dubery IA (2018) The chemistry of plant-microbe interactions in the rhizosphere and the potential for metabolomics to reveal signaling related to defense priming and induced systemic resistance. Front Plant Sci 9:112
Míguez-Montero M, Valentine A, Pérez-Fernández M (2019) Regulatory effect of phosphorus and nitrogen on nodulation and plant performance of leguminous shrubs. AoB Plants 12(1):plz047
Nelson DC (2021) The mechanism of host-induced germination in root parasitic plants. Plant Physiol 185(4):1353–1373
Noor R (2023) Microbial pathogenesis and the evasion strategies from the legume plant protective immunity. Legum Sci 5(1):e160
Nwachukwu BC, Ayangbenro AS, Babalola OO (2023) Structural diversity of bacterial communities in two divergent sunflower rhizosphere soils. Ann Microbiol 73(1):1–18
Olanrewaju OS, Babalola OO (2022) The rhizosphere microbial complex in plant health: a review of interaction dynamics. J Integr Agric 21(8):2168–2182
Olanrewaju OS, Ayangbenro AS, Glick BR, Babalola OO (2019) Plant health: feedback effect of root exudates-rhizobiome interactions. Appl Microbiol Biotechnol 103(3):1155–1166
Omomowo OI, Babalola OO (2019) Bacterial and fungal endophytes: tiny giants with immense beneficial potential for plant growth and sustainable agricultural productivity. Microorganisms 7(11):481
Palmer AJ, Baker A, Muench SP (2016) The varied functions of aluminium-activated malate transporters-much more than aluminium resistance. Biochem Soc Trans 44(3):856–862
Paluch E, Rewak-Soroczyńska J, Jędrusik I, Mazurkiewicz E, Jermakow K (2020) Prevention of biofilm formation by quorum quenching. Appl Microbiol Biotechnol 104(5):1871–1881
Pascale A, Proietti S, Pantelides IS, Stringlis IA (2020) Modulation of the root microbiome by plant molecules: the basis for targeted disease suppression and plant growth promotion. Front Plant Sci 10:1741–1741
Roskova Z, Skarohlid R, McGachy L (2022) Siderophores: an alternative bioremediation strategy? Sci Total Environ 819:153144
Schlüter U, Bouvier JW, Guerreiro R, Malisic M, Kontny C, Westhoff P, Stich B, Weber AP (2023) Brassicaceae display variation in efficiency of photorespiratory carbon-recapturing mechanisms. J Exp Bot 10:1093
Schmidt JE, Kent AD, Brisson VL, Gaudin ACM (2019) Agricultural management and plant selection interactively affect rhizosphere microbial community structure and nitrogen cycling. Microbiome 7(1):146
Sharma P, Sangwan S, Kaur H, Patra A, Anamika, Mehta S (2023) Diversity and evolution of nitrogen fixing bacteria. In: Sustainable agriculture reviews 60: microbial processes in agriculture, pp 95–120
Singh S, Gupta P, Bajaj BK (2018) Characterization of a robust serine protease from Bacillus subtilis K-1. J Basic Microbiol 58(1):88–98
Singh P, Chauhan PK, Upadhyay SK, Singh RK, Dwivedi P, Wang J, Jain D, Jiang M (2022) Mechanistic insights and potential use of siderophores producing microbes in rhizosphere for mitigation of stress in plants grown in degraded land. Front Microbiol 13:898979
Slámová K, Kapešová J, Valentová K (2018) “Sweet flavonoids”: glycosidase-catalyzed modifications. Int J Mol Sci 19(7):2126
Topolovec-Pintarić S (2019) Trichoderma: invisible partner for visible impact on agriculture. Trichoderma-the most widely used fungicide. IntechOpen 1:15
Vaughan MM, Wang Q, Webster FX, Kiemle D, Hong YJ, Tantillo DJ, Coates RM, Wray AT, Askew W, O’Donnell C (2013) Formation of the unusual semivolatile diterpene rhizathalene by the Arabidopsis class I terpene synthase TPS08 in the root stele is involved in defense against belowground herbivory. Plant Cell 25(3):1108–1125
Veach AM, Morris R, Yip DZ, Yang ZK, Engle NL, Cregger MA, Tschaplinski TJ, Schadt CW (2019) Rhizosphere microbiomes diverge among populus trichocarpa plant-host genotypes and chemotypes, but it depends on soil origin. Microbiome 7(1):76
Verma SK, White J (2019) Seed endophytes. Springer, Cham
Verma SK, Chen Q, White JF (2022) Evaluation of colonization and mutualistic endophytic symbiosis of Escherichia coli with tomato and Bermuda grass seedlings. PeerJ 10:e13879
Vieira S, Sikorski J, Dietz S, Herz K, Schrumpf M, Bruelheide H, Scheel D, Friedrich M, Overmann J (2019) Drivers of the composition of active rhizosphere bacterial communities in temperate grasslands. ISME J 14
Vieira S, Sikorski J, Dietz S, Herz K, Schrumpf M, Bruelheide H, Scheel D, Friedrich MW, Overmann J (2020) Drivers of the composition of active rhizosphere bacterial communities in temperate grasslands. ISME J 14(2):463–475
White JF, Kingsley KL, Verma SK, Kowalski KP (2018) Rhizophagy cycle: an oxidative process in plants for nutrient extraction from symbiotic microbes. Microorganisms 6(3):95
Wu J, Yu S (2019) Effect of root exudates of Eucalyptus urophylla and Acacia mearnsii on soil microbes under simulated warming climate conditions. BMC Microbiol 19(1):224
Xuan TD, Minh TN, Khanh TD (2016) Allelopathic momilactones A and B are implied in rice drought and salinity tolerance, not weed resistance. Agron Sustain Dev 36(3):52
Zhalnina K, Louie KB, Hao Z, Mansoori N, da Rocha UN, Shi S, Cho H, Karaoz U, Loqué D, Bowen BP, Firestone MK, Northen TR, Brodie EL (2018) Dynamic root exudate chemistry and microbial substrate preferences drive patterns in rhizosphere microbial community assembly. Nat Microbiol 3(4):470–480
Acknowledgements
OOB would like to thank the National Research Foundation of South Africa for a grant (Grant Ref: UID123634; UID132595 OOB) that has supported work in our laboratory. RRM would like to thank the North-West University for the postgraduate bursary that was granted during her MSc program.
Funding
Open access funding provided by North-West University. This work is supported by the National Research Foundation of South Africa (Grant Refs: UID123634; UID132595) awarded to OOB.
Author information
Authors and Affiliations
Contributions
All the mentioned authors contributed substantially and intellectually to the work. OOB designed the research, revised the work critically for important intellectual content, performed quality assurance, provided funding acquisition, project administration, and resources. RRM was involved in formal analysis, investigation, and writing of the original draft of the manuscript. AEA was involved in reviewing and thoroughly editing of original draft, validation, and formal analysis.
Corresponding author
Ethics declarations
Ethics statement
None required.
Conflict of interest
None declared.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Molefe, R.R., Amoo, A.E. & Babalola, O.O. Communication between plant roots and the soil microbiome; involvement in plant growth and development. Symbiosis 90, 231–239 (2023). https://doi.org/10.1007/s13199-023-00941-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13199-023-00941-9