Abstract
Drought has become more frequent in Central Europe causing large losses in cereal yields, especially of spring crops. The development of new varieties with increased tolerance to drought is a key tool for improvement of agricultural productivity. Material for the study consisted of 100 barley recombinant inbred lines (RILs) (LCam) derived from the cross between Syrian and European parents. The RILs and parental genotypes were examined in greenhouse experiments under well-watered and water-deficit conditions. During vegetation the date of heading, yield and yield-related traits were measured. RIL population was genotyped with microsatellite and single nucleotide polymorphism markers. This population, together with two other populations, was the basis for the consensus map construction, which was used for identification of quantitative trait loci (QTLs) affecting the traits. The studied lines showed a large variability in heading date. It was noted that drought-treatment negatively affected the yield and its components, especially when applied at the flag leaf stage. In total, 60 QTLs were detected on all the barley chromosomes. The largest number of QTLs was found on chromosome 2H. The main QTL associated with heading, located on chromosome 2H (Q.HD.LC-2H), was identified at SNP marker 5880–2547, in the vicinity of Ppd-H1 gene. SNP 5880–2547 was also the closest marker to QTLs associated with plant architecture, spike morphology and grain yield. The present study showed that the earliness allele from the Syrian parent, as introduced into the genome of an European variety could result in an improvement of barley yield performance under drought conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Barley (Hordeum vulgare L.) is not only one of the most important crops from an economic point of view (FAOSTAT 2014), but it is also an excellent species for genome mapping and map-based analyses (Costa et al. 2001; Mansour et al. 2014). Its diploid nature, low chromosome number and a high degree of self-fertility mean that barley is a common subject for genetic studies examining drought resistance of crops (Tondelli et al. 2006; Talamè et al. 2007).
Several genetic maps based upon different genetic marker techniques have been published (Wenzl et al. 2006; Zhou et al. 2015). Among various types of DNA markers, microsatellites (SSR) and single nucleotide polymorphism (SNP) have been widely used for genome analyses (Cockram et al. 2010; Cuesta-Marcos et al. 2010; Honsdorf et al. 2014; Varshney et al. 2007). The first high-density gene map based on SNP markers contained 2.943 SNP loci in 975 marker bins and covered a genetic distance of 1099 cM (Close et al. 2009). The availability of high-throughput SNP genotyping has facilitated the genetic studies of agronomically important traits (Close et al. 2004; Wang et al. 2010b). Recently, research containing a detailed overview of the functional portions of the barley genome has been published (International Barley Genome Sequencing Consortium 2012). This highly resoluted genetic map together with the sequence data has a tremendous potential for candidate gene discovery using conservation of the grass genome synteny (Mayer et al. 2009).
Abiotic stresses reduce average yields for most crops (Boyer 1982; Bray 1997). Among the stresses, water deficit is the most devastating on a global scale (Zhao and Runnings 2009). Aspinall et al. (1964) and Samarah (2005) have reported stage-specific drought responses in the crops. Appropriate irrigation conditions during the stem elongation phase are indispensable for the formation of fertile florets at anthesis as the final number of grains is determined during this period (Miralles and Slafer 1995). Water deprivation in this critical developmental stage affects numerous aspects of plant metabolism leading to impairment of many biochemical pathways (Moran et al. 1994; Loggini et al. 1999; Farooq et al. 2009).
The simplest solution to survive in dry environments is an escape from drought (Passioura 1996; Richards 1996). The short life cycle of crop plants might be considered as an important trait related to water deficit adaptation (Araus et al. 2002). The majority of barley cultivars vary significantly in their response to water scarcity (Zare 2012). Sources of drought tolerance can be found in landraces from geographical regions with challenging climates close to their domestication origin (Ellis et al. 2000; Górny 2001; Nevo and Chen 2010).
Heading date in barley depends on vernalisation requirements (Takahashi and Yasuda 1956; Sasani et al. 2012), photoperiodic response (Roberts et al. 1988; Laurie et al. 1995) and earliness per se genes (Gallagher et al. 1991; Sameri et al. 2006). Quantitative trait loci (QTLs) associated with heading date have been mapped on all barley chromosomes (e.g. Hayes et al. 1993; Laurie et al. 1995; Tinker et al. 1996; Bezant et al. 1997; Qi et al. 1998; Pillen et al. 2003) and many allele-specific markers for some candidate genes controlling these processes are known (Turner et al. 2005; Faure et al. 2007; Szűcs et al. 2007).
The major photoperiod response locus has been identified by RFLP analysis on the short arm of chromosome 2H. Dominant alleles at Ppd-H1 accelerate flowering under long day conditions, whereas no effect has been detected under short day conditions. Laurie et al. (1995) and Turner et al. (2005) have shown that the late-flowering allele is recessive. Two main single nucleotide polymorphisms have been detected which differentiate alleles involved in the plant sensitivity to day length. Non-synonymous (G – Ppd-H1/ A – ppd-h1) SNP within the CCT domain has been suggested as an explanation for recessive form of the allelic variation (Turner et al. 2005). Another study revealed that a polymorphism in the photoperiodic response in barley varieties might be associated with the SNP48 situated in the exon 6 of the Ppd-H1 coding region) (Jones et al. 2008). A second major photoperiodic response locus (Ppd-H2) has been mapped to the chromosome 1H (Laurie et al. 1995). The Ppd-H2 affects the flowering time under short day conditions. A candidate gene (HvFT3) for this locus has been proposed by Faure et al. (2007).
The vernalisation and photoperiodic pathways correspond to each other to promote flowering in crops (Distelfeld et al. 2009). A study conducted using barley spring crosses revealed loci for flowering time in the regions connected with vernalisation response (Bezant et al. 1997). Three genes, Vrn-H1, Vrn-H2 and Vrn-H3, located on the chromosomes 5H, 4H and 7H, respectively, have been proposed as the major vernalisation response genes (Cockram et al. 2007).
The aim of the present study was to detect QTLs determining yield and yield-forming traits in a recombinant inbred line (RIL) population developed from a hybrid between European and Syrian genotypes (adapted to dry environments) under optimal and water stress conditions, with special attention being paid to earliness.
Materials and methods
Plant material
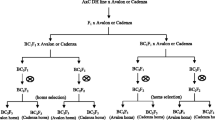
Material for the study covered RIL population of spring barley (Hordeum vulagre L.) derived from the cross Lubuski × Cam/B1/CI08887//CI05761. The parent Cam/B1/CI08887//CI05761 (hereafter referred as CamB) is the Syrian breeding line supplied to Dr. A. Górny by Drs S. Grando and S. Ceccarelli from ICARDA in Aleppo and Lubuski is an old Polish cultivar derived from a Heines-Haisa/Skrzeszowicki hybrid. The examined population was developed by means of the single seed descent (SSD) technique (up to F8) (Goulden 1939) associated with in vitro culture of immature embryos (Surma et al. 2013). Out of 150 developed RILs 100 were randomly chosen for the present experiments.
Greenhouse experiments
The greenhouse experiments with the Lubuski × CamB population were conducted (was grown in three replicates) in two growing seasons (2012, 2013) during April–August. In both years, three water regimes were applied: (1) C – optimal water supply for the whole vegetation period, (2) DI – drought stress beginning at the three-leaf stage (13 in the BBCH scale) and maintained for 10 days, (3) DII – drought stress beginning at the flag leaf stage (37 in the BBCH scale) and maintained for 14 days, which created six environments denoted as: C 2012, C 2013, DI 2012, DI 2013, DII 2012 and DII 2013. Ten plants were grown in pots containing 9 kg of soil. Air moisture and temperature were monitored by special device (LOG32 - Temperature-humidity logger with integrated USB-interface and automatic PDF-creation). Control of the soil moisture was provided by a hand-held device (FOM/mts) designed for field measurements of the soil moisture and temperature (Malicki et al. 1996). The weighing method was used as an additional control of the irrigation system. The soil moisture was kept at 2.2 and 3.2 pF in optimal and drought conditions, respectively (ESM 1). Three groups of traits were observed: associated with morphology of the main and lateral spikes (grain weight per main spike - GWSm, number of grains per main spike - NGSm, number of spikelets per main spike - NSSm, length of main spike - LSm, grain weight per lateral spike - GWSl, number of grains per lateral spike - NGSl, number of spikelets per lateral spike- NSSl, length of lateral spike - LSl), with plant architecture (length of main stem - LSt, number of productive tillers per plant - NPT), and with grain yield (1000-grain weight - TGW, Grain weight per plant - GWP). Duration of the vegetative growth period was expressed as the number of days from sowing to heading (heading date - HD). The measured traits are listed in ESM 2.
Genotyping
In the present studies consensus map constructed by Mikołajczak et al. (2016) was used for QTL analysis. Briefly: A set of 78 barley SSR markers developed by Varshney et al. (2007) was used in the experiment. SNP genotyping was carried out at the Southern California Consortium using the Illumina GoldenGate array 1 (Illumina Inc., San Diego, CA) that analyses 1.536 genome-wide single nucleotide polymorphisms; details of this array (BOPA - barley oligo pool assays) are described by Close et al. (2009).
JoinMap 3.0 software (Van Ooijen and Voorrips 2001) was used for the map construction. Once the individual genetic map was obtained, the consensus map was constructed. The complete dataset consisted of 819 markers mapped in the Maresi × CamB (MCam), Lubuski × CamB (LCam) and Georgie × Harmal (GH) populations. Details on the development of the map construction are given in Mikołajczak et al. (2016).
Statistical analysis
Observations for RILs were processed by analysis of variance in a mixed model with fixed effects for year, drought and year × drought interaction, and with random effects for line and interaction of line with year and/or drought treatment. The residual maximum likelihood (REML) algorithm was used to estimate variance components for random effects and the F-statistic was computed to assess the significance of the fixed effects. Ordinary, mean values computed for RILs in all specific (years × drought) - combinations were used for construction of principal component biplots. Pearson correlation coefficients between all the analysed traits were calculated. QTL analysis was performed for the consensus linkage map (Mikołajczak et al. 2016) with the mixed-model approach described by Malosetti et al. (2013), including optimal genetic correlation structure selection and the significance threshold estimation. The interval mapping was conducted with a step size of 2 cM by selecting the QTL candidate and then using them iteratively as cofactors until the list of QTL was not changed. The threshold for the − log10(P-value) statistic was computed by the method of Li and Ji (2005) to ensure the genome-wide error rate was less than 0.01. The windows for not selecting two close QTLs and for exclusion of cofactors were set at 10 and 30 cM, respectively. Selection of the set of QTL effects in the final model was performed at P < 0.05; the P-values for the Wald test were computed as the mean from the values obtained by adding and dropping the QTL main and interaction effects in the model. All the above computations were performed in Genstat 16 (VSN Int. 2013).
QTL annotation
All SNP sequences taken from Close et al. (2009) (Supplementary material file BOPA1 SNP 1471-2164-10-582-S19.xls) were mapped using NCBI Blast for Windows to barley genomic space in Ensembl Plants ver. 2.28 (reference repeat masked sequence Hordeum_vulgare.082214v1.28.dna_rm.toplevel.fa, maximum EValue = 1e-060, minimum 95 % identity of the SNP sequence). The SNP mapping positions were used to obtain a projection of two LOD QTL support intervals (see Xu 2010) onto the genomic sequence; all genes located in projected intervals were listed and annotated using Gene Ontology (GO) terms. For QTL interpretation, we applied a method similar to the one implemented by Cantalapiedra et al. (2015).
Early and late heading subgroups of plants
According to SNP 5880–2547 segregation (Mansour et al. 2014; Muñoz-Amatriaín et al. 2011), RILs were divided into two subgroups—early heading (group A – allele from CamB) and late heading plants (group B – allele from Lubuski).
Results
Phenotypic evaluation
The average heading dates and the mean values of morphological traits for parental cultivars and RILs in the six environments are presented in ESM 3a and ESM 3b as supplementary material, respectively. Parental genotypes are classified as early (CamB) and late (Lubuski) according to the large differences between their heading dates in all experiments. The heading (HD) of the Syrian genotype grown under well-watered conditions was about 15–19 days earlier than of the European one. The HD for parents increased both in DI conditions (by about 3 - 6 days) and in DII conditions (by about 1 - 6 days) as compared to the well-watered conditions. The data analysis across two years showed highly significant differences among lines for HD. RILs with longer vegetation periods than the late-heading parent were noticed among the studied population in all environments. For all studied traits, Lubuski showed significantly higher values under well-watered conditions compared to CamB (with the exception of LSt in 2012, 2013 and NPT in 2013). The Syrian cultivar showed higher values for TGW than the European parent in DI over the two years. On the other hand, Lubuski showed higher GWP in all environments. The comparison of the parental genotypes for traits connected with the plant architecture showed that CamB formed more productive tillers under water-stress conditions applied at the three-leaf stage.
In RILs, NPT increased both under drought I and drought II conditions. For all observed traits (with the exceptions: HD and NPT), greater decrease were noticed under drought II.
Lines of the LCam population were significantly differentiated in terms of all analysed traits (Table 1). In all case, the variance components for all types of interactions were smaller than that for lines. For HD, variance components were significant for all types of interaction (i.e. for line × year, line × water regimes, line × year × water regimes). On the other hand, no interaction component was significant for NGSm and LSl.
As shown in biplots (Fig. 1), RIL plants grown in drought II were affected more than in drought I, as they are further away from control plants superior—in both years—by spike morphology traits and LSt.
Correlations of all traits with HD were significant (P < 0.001) in at least one environment, with no correlation significant in DII 2012 (Table 2). The highest correlation coefficient was found for NSSl in drought I in 2012 (r = 0.762), whereas the correlation between HD and NGSl (DII 2013) was the weakest (r = 0.236). Significant negative correlations between HD and NPT, revealed also in 2013 biplot, were observed across three environments (DI 2012, DI 2013, C 2013) which indicates that early heading lines developed more productive tillers, especially in DI conditions. No significant association was found between days to heading and 1000-grain weight, except for the control conditions in 2012 (negative correlation). Positive and significant correlations were recorded between HD and spike traits: GWSm, NGSm, NSSm, LSm, GWSl, NGSl, NSSl in both years in DI and C conditions. This indicates that late heading lines developed longer spikes with more spikelets and—as a consequence—more grains. Moreover, a positive correlation was found between GWP and heading stage, which indicates that early heading lines were characterised by lower yield.
QTL analyses
A total of 60 QTLs were detected on all chromosomes (Table 3). The largest number of QTLs were found on chromosome 2H (23 QTLs). Only three QTLs were detected on chromosome 1H. The largest number of QTLs were found for NGSm and LSm (nine QTLs). The lowest number of QTLs were found for GWP (one QTL). The QTL × E interaction was found for 68 % of QTLs detected. All QTLs for HD and NSSm showed QTL × E interaction (ESM 4).
QTLs for earliness and yield-forming traits
Four QTLs for heading date (HD) were found on chromosomes 2H (Q.HD.LC-2H), 3H (Q.HD.LC-3H.1), 5H (Q.HD.LC-5H.3) and 7H (Q.HD.LC-7H.2). In the vicinity of the Q.HD.LC-2H, 12 QTLs for plant architecture, spike morphology and grain yield were detected (Q.GWSl.LC-2H-1, Q.GWSm.LC-2H-1, Q.GWP.LC-2H, Q.HD.LC-2H, Q.LSl.LC-2H, Q.LSm.LC-2H-1, Q.NGSl.LC-2H-1, Q.NGSm.LC-2H-1, Q.NPT.LC-2H-1, Q.NSSl.LC-2H, Q.NSSm.LC-2H-1 and Q.LSt.LC-2H) for the region on the short arm of chromosome 2H, whereas one QTL (Q.HD.LC-3H.1) was found for the region detected on chromosome 3H. On chromosome 5H, close to the Q.HD.LC-5H.3, were found QLSm.LC-5H.3, QLSl.LC-5H.3 and QGWSm.LC-5H.3 and also three QTLs were identified in the vicinity of the Q.HD.LC-7H.2: Q.LSm.LC-7H.2, Q.LSl.LC-7H.2 and Q.TGW.LC-7H.2.
For HD the major was QTL on chromosome 2H, which showed the most significant effect and explained a large proportion of the phenotypic variation. This QTL was mapped at the marker 5880–2547 at the position of 10.74 cM (Fig. 2). In almost all environments the alleles from the Syrian parent reduced days to heading; DII 2012 was an exception to this rule, and in this environment the percentage of explained variation was a low (9.55 %).
According to SNP 5880–2547 segregation RILs were divided into two subgroups – early heading (group A – allele from CamB) and late heading plants (group B – allele from Lubuski). The different developmental pattern for these subgroups was noticed in the stress conditions. An extreme delay in heading was observed for early heading lines in DII conditions (Fig. 3).
In the region of Q.HD.LC-2H, marked by SNP 5880–2547 (10.14 cM) and 7747–1056 (14.78 cM), QTLs for all yield-related traits, except TGW, were found (Fig. 2). In all cases they appeared to be the most significant QTLs for a particular trait, with the LogP statistic ranging from 8.00 for GWP to 24.73 for LSm. All of them showed a significant interaction with environment, but the sign of the allelic effects was consistent over environments. For yield-forming traits, except NPT, alleles contributed by Lubuski increased the traits.
Q.HD.LC-3H.1 on chromosome 3H at SNP 10353–119 showed a minor positive effect contributed by a Syrian parent allele. That QTL explained only 2.64–7.42 % of phenotypic variance, and its additive effect was significant only in DI 2012 and C 2012. No QTL associated with other traits was found in this region (Table 3).
The Q.HD.LC-5H.3 located on the linkage group 5H.3 at SNP 314–559 with CamB allele causing later heading was significant only in 2012. In the region of this QTL, marked by SNP 314-559 and ConsensusGBS0138-2, QTLs for LSm, GWSm and LSl were also found, with alleles from Lubuski increasing the trait values, and the variance explained from 0.5 to 34.4 % (Table 3).
The Q.HD.LC-7H.2 was detected on chromosome 7H at SNP 1213–1959. It explained from 2.19 to 24.11 % of the phenotypic variation, with the Lubuski allele increasing the number of days from sowing to heading. Effect of that QTL was significant only in 2012 in DII and C conditions. At the same position QTL for TGW (Q.TGW.LC-7H.2) was localised and significant also only in 2012. That QTL explained 4.23-10.12 % of the phenotypic variance and allele contributed by Lubuski reduced the TGW. Near Q.HD.LC-7H.2 the QTLs for LSm and LSl were also found. These QTLs (Q.LSm.LC-7H.2 and Q.LSl.LC-7H.2, both with stabile effects) explained 4.20–13.99 % and 5.62–12.17 % of the phenotypic variation, respectively. Both QTLs were characterised by Lubuski alleles increasing the traits (Table 3).
Functional annotation of QTLs
For a biological interpretation of the QTL regions identified on the basis of linkage analysis, we refer to the annotation of SNP using the Ensembl Plants barley gene space according with the approach used in Mikołajczak et al. (2016). The analysis revealed two of the main GO biological processes (defense response and protein ubiquitination) overrepresented in the annotation genes for traits: grain weight per main spike, grain weight per plant and length of main spike (Table 4). The largest number of genes (15) annotated with the previously mentioned terms was noticed for “protein ubiquitination”. Functional annotation analysis also showed six other biological processes overrepresented in the annotation of genes occurring in the QTL regions (defense response, lipid transport, metabolic process, oxidation-reduction process, protein phosphorylation, protein ubiquitination, response to oxidative stress, transmembrane transport (Table 5).
Discussion
The present study examined the mapping population derived from the cross Lubuski × Cam/B1/CI08887//CI05761 and special attention was given to earliness. The Syrian genotype, when compared to the European cultivar, was generally characterised by earlier heading and lower yield. Drought stress conditions caused the reduction of the studied traits (with the exceptions: HD and NPT). Grain yield was the most decreased under drought stress applied at the flag leaf stage. This was due to reduced spikelets and grain numbers per spike. These results are in agreement with the results reported by Zinselmeier et al. (1999) and Samarah et al. (2009) who demonstrated the impact of drought during the flowering period on grain yield.
The effect of water scarcity on yield varies depending on the plant development stage. This is why we noted different mean values for traits observed in drought I and drought II. It is noteworthy that in DI and DII treatments plants were observed to have more tillers than in the well-watered conditions. It may be explained by the emergence of new tillers during re-watering period. A similar phenomenon has also been noticed in other works, e.g. by Aspinall et al. (1964) and Loss and Siddique (1994), but in most studies a significant decrease of the number of productive tillers under drought conditions has been observed (Samarah 2005; Shirazi et al. 2010; Tsenov et al. 2015).
In our study, stress conditions caused a delay in heading. These findings are in agreement with other studies (Winkel et al. 1997; Wopereis et al. 1996; Farooq et al. 2011) but, on the other hand, our results are also in contrast to the results showed by Desclaux and Romet (1996), Slafer et al. (2005) and Richards (2006), where the drought conditions caused an acceleration of plant growth and development. In the present studies the highest delay in the appearance of the heading was triggered by drought stress conditions II, especially for early heading lines. This phenomenon can be associated with the survival strategies of this group of plants, which were, in general, at an advanced stage of development at the time of stress application. The results of the study confirmed the assumption that the drought escape can be an effective strategy only when a plant has completed its life cycle before the environment conditions become unfavourable.
QTLs for earliness
Earliness affects the plant adaptation to the environmental changes and it is a trait affected by numerous QTLs (Yano et al. 2000; Sameri et al. 2011). In our study, four QTLs for earliness were detected on chromosomes 2H, 3H, 5H and 7H. The localisation of these QTLs on barley chromosomes is consistent with previously identified QTLs (Hayes et al. 1993; Laurie et al. 1995; Thomas et al. 1995; Bezant et al. 1996; Tinker et al. 1996; Qi et al. 1998; Pillen et al. 2003). Some of them were found in genomic regions that have been reported to harbour genes involved in flowering time regulation. In our study, the main effects were shown by QTL detected on chromosome 2H at SNP 5880–2547 (11_21015). On the short arm of that chromosome the major photoperiod response locus (Ppd-H1) which causes early flowering under day length has been mapped (Laurie et al. 1995). The 2HS region association with the earliness was also observed in numerous other studies. Ren et al. (2010) identified three QTLs determining the heading date on chromosomes 2H (and also on 3H and 7H), which is in agreement with our results. QTL analysis of the Steptoe/ Morex population conducted by Mansour et al. (2014) revealed the QTL also located at SNP 11_21015. All these findings support the notion that the region on the short arm of chromosome 2H is tightly associated with heading date. SNP 11_21015 has been mapped close to markers 12_30871 and 12_30872 (Muñoz-Amatriaín et al. 2011) which are SNPs in Ppd-H1. Borrás-Gelonch et al. (2012), following a series of experiments involving environments with artificially extended photoperiod, reported the two QTLs affected earliness on chromosome 2H. The regions on 2H were also the main determinants of heading date in autumn-sown experiments conducted using mapping populations grown under Mediterranean conditions (Moralejo et al. 2004; Cuesta-Marcos et al. 2008). Comadran et al. (2011) reported five QTLs for heading date (located on 1H, 2H, 3H and 5H). Their research revealed that the highest effect was shown by two QTLs detected in the centromeric region of 2H, where another major gene affecting heading date (eam6) had previously been reported (Cuesta-Marcos et al. 2009).
Although our study was based on the analysis of a population derived from spring barley parents, QTL analysis revealed some associations with chromosome regions harbouring genes related to vernalisation requirements. Vernalisation response has been shown to be strongly influenced by photoperiod (Roberts et al. 1988; Wang et al. 2010b). Epistatic interaction among major loci of vernalisation response, photoperiod reaction and earliness per se may be responsible for the fact that a large number of genomic regions have been identified as determinants of heading date (Karsai et al. 2001). In our study Q.HD.LC-3H.1 was detected on chromosome 3H at SNP 10353–119 and in the vicinity (0.24 cM) of microsatellite Bmag603. Wang et al. (2010b) revealed that a flowering time candidate gene (HvFT2) had been located 3 cM from this SSR marker. Another locus connected with heading date was found in our study on chromosome 5H at SNP 314–559 positioned at 59.03 cM (Q.HD.LC-5H.3). This QTL was located in a similar position as QTL for heading date reported by Marquez-Cedillo et al. (2001) and Thomas et al. (1995) and the vernalisation response gene (Vrn-H1) found by Laurie et al. (1995). According to Muñoz-Amatriaín et al. (2011), Vrn-H1 contains SNP 12_30883 and is mapped on the long arm of chromosome 5H between SNPs 11_21247 (7639-122) and 11_11080 (ABC03900-1-2-406), the latter being located in the consensus map used in our studies in the distance of 0.6 cM from SNP 314-559. Our data also showed that in the vicinity of SNP 314–559 another SNP 7523–440 (11_21241) was located, which was linked to the locus Vrn-H1 in the study conducted by Malosetti et al. (2011).
QTLs for agronomic traits
Several yield-related QTLs have been mapped to the short arm of chromosome 2H, including plant height (Karsai et al. 1997), kernel weight (Han and Ullrich 1993), number of seeds per spike (Kjaer et al. 1991) and tiller number (Eshghi et al. 2011). In the present study, QTLs with large effects for yield, plant height, number of productive tillers, length of spike, spikelet number and number and weight of grain were found near SNP 5880–2547 on chromosome 2H. Results obtained in numerous studies have shown that loci associated with the length of spikes are placed on all the barley chromosomes (Hori et al. 2003; Sameri et al. 2006; Baghizadeh et al. 2007; Wang et al. 2010a, b). The localisation of the QTL for earliness on chromosome 2H coincided with QTLs for spike morphology. The QTL for the length of the main spike (LSm) was discovered in genomic regions associated with earliness, except the one which was found on chromosome 3H. Interestingly, the QTL for LSm was found in our study both on chromosome 5H (Q.LSm.LC-5H.3) and on chromosome 7H (Q.LSm.LC-7H.2), where were identified regions related to heading stage. In the present study, the QTL for number of grains per main spike was mapped at the marker 5880–2547 on chromosome 2H. The QTLs affecting the number of grains per spike on chromosome 2H have been reported by Mohammadi and Baum (2008) and Mehravaran et al. (2014), and in our investigation, SNP 5880–2547 was also the nearest marker for QTLs related to grain weight per main and lateral spike and grain weight per plant. These results are in agreement with the findings of Peighambari et al. (2005) who found the QTL for grain yield on chromosome 2H. In other studies QTLs for grain yield were identified on almost all the barley chromosomes (Cuesta-Marcos et al. 2009; Mansour et al. 2014; Mehravaran et al. 2014). Stem length is an important morphological character directly linked with the productive potential of barley plants. In the present study, we did not detect any QTL for the length of stem close to SNP 5880–2547 associated with earliness. However, the QTL analysis revealed Q.LSt.LC-2H at SNP 7747-1056, 2.2 cM shifted from from SNP 5880–2547. In the region of QTL for HD detected on 3H no QTL for stem length was found. It should be noted that on 3H sdw1/denso locus causing reduction of plant height was localized and several studies revealed that this locus may also be associated with flowering time (Barua et al. 1993; Laurie et al. 1994; Bezant et al. 1996; Kuczyńska et al. 2013, 2014).
In our experiment, two QTLs associated with the numbers of spikelets per spikes (Q.NSSm.LC-2H and Q.NSSl.LC-2H) were found on chromosome 2H at the SNP 5880–2547. These results are in agreement with the QTL localisation previously reported by Li et al. (2005) and Baghizadeh et al. (2007). Additionally, these authors revealed QTLs for these traits also on chromosomes 1H, 5H and 7H.
In the current study, we have identified locus associated with the number of productive tillers on 2H (Q.NPT.LC-2H) at the same position as the main QTL for heading date — position 10.7 cM). Tiller number is a key component of barley grain yield (Sakamoto and Matsuoka 2004). Fertile tillers contribute significantly to grain yield improvement, but those tillers without fertile spikes decrease the harvest index (Mäkelä and Muurinen 2011). In our study we noticed an increase in the number of productive tillers triggered by drought conditions which could be explained by the secondary tiller development process, commonly observed in the field conditions (Aspinall et al. 1964).
QTLs related to drought
A recent study revealed that QTLs related to drought stress tolerances were placed on chromosome 2H and 5H (Fan et al. 2015). The pivotal importance of the genomic regions for drought tolerance was also reported by Mehravaran et al. (2014) on chromosomes 2H, 5H and 7H. The authors suggested that these regions may be used as an important target for improving drought tolerance of barley.
The association of heading date and drought tolerance has been reported by Xu et al. (2005), Araus et al. (2002), Kigel et al. (2011), Schmalenbach et al. (2014). Similar results were obtained in the present studies. We identified QTLs related to heading date on chromosomes 2H, 5H and 7H, where QTLs for drought tolerance have been reported in other studies. QTLs connected with yield structure were found near QTLs identified for earliness, which is also in agreement with other studies (Wang et al. 2010a; Honsdorf et al. 2014; Mansour et al. 2014; Mehravaran et al. 2014).
In the present study, early heading barley plants did not realise the drought escape strategy, and other mechanisms also associated with water scarcity tolerance appeared to be ineffective. On the other hand, we observed an increase in productive tillers forming after drought during re-watering, especially in the Syrian parent. As early heading lines tend to have low quality yield, the enhancement of productive tillers seems to be a promising strategy.
Functional annotations
The overrepresentation of genes annotated as “defense response” for traits: grain weight per main spike and grain weight per plant did not allow for an unambiguous interpretation. This GO term was descript by QuickGO (http://www.ebi.ac.uk/QuickGO) as “reactions, triggered in response to the presence of a foreign body or the occurrence of an injury, which result in restriction of damage to the organism attacked or prevention/recovery from the infection caused by the attack”, which can be assigned to a every type of plant reaction associated with biotic or abiotic stresses. Noteworthy, the second type of overrepresented GO term was related to protein ubiquitination as “the process in which one or more ubiquitin groups are added to a protein”. Ubiquitin is well established as the major modifier of signalling in eukaryotes. The main characteristic of ubiquitination is the conjugation of ubiquitin onto lysine residues of acceptor proteins (Stone and Callis 2007). In most cases, the targeted protein is degraded by the 26S proteasome, the major proteolysis machinery in eukaryotic cells. The ubiquitin–proteasome system is responsible for removing most abnormal peptides and short-lived cellular regulators. This allows cells to respond rapidly to intracellular signals and changing environmental conditions. These types of biological processes are crucial to sustain cellular functions under drought. In Arabidopsis thaliana more than 1400 genes encode components of the ubiquitin/26S proteasome (Ub/26S) pathway (Smalle and Vierstra 2004). Approximately 90 % of these genes encode subunits of the E3 ubiquitin ligases, which confer substrate specificity to the pathway. This mechanism can be observed in the gibberellin-dependent signalling pathway that regulates the flowering process (Cheng et al. 2004). Gibberellins (GAs), one kind of endogenous growth regulator, play an essential role not only in reproductive development of plants but also in stem and spike growth regulation (Kumar et al. 2003; Tyagi and Singh 2006; Janowska and Andrzejak 2010). Moreover, treatment of GA causes stem elongation, expansion and proliferation and cell wall thickening increased cell division and cell elongation (Taiz and Zeiger 1998). Similar processes may be observed in spike growth and development. Our plant material was differential in terms of spike length both in well-watered and drought conditions, which may suggest that the effect of GA can be a major factor related to spike growth irrespective of irrigation conditions.
The annotation of QTL regions by genes occurring in the projected support intervals showed the six other biological processes, one of which may play a key role in the drought stress. “Response to oxidative stress” was annotated for five genes occurring in the regions of identified QTLs. Prolonged drought stress results in oxidative damage due to the over production of reactive oxygen species (ROS) (Smirnoff 1993). ROS seem to have a dual effect under drought stress conditions that depend on their overall cellular amount. If kept at relatively low levels they are likely to function as components of a stress-signalling pathway, triggering stress defense/acclimation responses. However, when reaching a certain level of phytotoxicity, ROS become damaging, initiating unwell-watered led oxidative cascades that harm cellular membranes and other cellular components resulting in oxidative stress and eventually cell death (Dat et al. 2000) and—as a consequence—the wilting process noticed in water scarcity conditions.
Conclusions
Four QTLs for HD were detected on chromosomes 2H, 3H, 5H and 7H. Hence, the major was QTL located on the short arm of 2H chromosome at SNP marker 5880–2547, in the vicinity of Ppd-H1 gene. In the region of SNP 5880–2547 QTLs associated with plant architecture, spike morphology and grain yield were localised. The present study showed that the earliness allele from the Syrian parent, as introduced into the genome of a European variety could result in an improvement of barley yield performance under drought conditions. Screening barley cultivars for growth duration under terminal drought stress is needed to evaluate drought escape in barley grown under such conditions. In order to use these QTLs for improvement of agronomic traits, further complementary studies in different environments and genetic contexts should be performed.
References
Araus JL, Slafer GA, Reynolds MP, Royo C (2002) Plant breeding and drought in C3 cereals: what should we breed for? Ann Bot 89:925–940
Aspinall D, Nicholls PB, May LH (1964) The effects of soil moisture stress on the growth of barley. I. Vegetative development and grain yield. Aust J Agric Res 15:729–745
Baghizadeh A, Taleei AR, Naghavi MR (2007) QTL analysis for some agronomic traits in barley (Hordeum vulgare L.). Int J Agric Biol 9:372–374
Barua UM, Chalmers KJ, Thomas WTB, Hackett CA, Lea V, Jack P, Forster BP, Waugh R, Powell W (1993) Molecular mapping of genes determining height, time to heading, and growth habit in barley (Hordeum vulgare). Genome 36:1080–1087
Bezant J, Laurie D, Pratchett N, Chojecki J, Kearsey M (1996) Marker regression mapping of QTL controlling flowering time and plant height in a spring barley (Hordeum vulgare L.) cross. Heredity 77:64–73
Bezant JH, Laurie D, Pratchett N, Chojecki J, Kearsey M (1997) Mapping QTL controlling yield and yield components in a spring barley (Hordeum vulgare L.) cross using marker regression. Mol Breed 3:29–38
Borrás-Gelonch G, Denti M, Thomas WTB, Romagosa I (2012) Genetic control of pre-heading phases in the Steptoe × Morex barley population under different conditions of photoperiod and temperature. Euphytica 183:303–321
Boyer JS (1982) Plant productivity and environment. Science 218(4571):443–448
Bray EA (1997) Plant responses to water deficit. Trends Plant Sci 2:48–54
Cantalapiedra CP, Boudiar R, Casas MA, Igartua E, Contreras-Moreira B (2015) BARLEYMAP: physical and genetic mapping of nucleotide sequences and annotation of surrounding loci in barley. Mol Breed 35:13
Cheng H, Qin LQ, Lee SC, Fu XD, Richards DE, Cao DN, Luo D, Harberd NP, Peng JR (2004) Gibberellin regulates Arabidopsis floral development via suppression of DELLA protein function. Development 131:1055–1064
Close TJ, Wanamaker SI, Caldo RA, Turner SM, Ashlock DA, Dickerson JA, Wing RA, Muehlbauer GJ, Kleinhofs A, Wise RP (2004) A new resource for cereal genomics: 22K barley GeneChip comes of age. Plant Physiol 134:960–968
Close TJ, Bhat PR, Lonardi S, Wu Y, Rostoks N, Ramsay L, Druka A, Stein N, Svensson JT, Wanamaker S, Bozdag S, Roose ML, Moscou MJ, Chao S, Varshney R, Szucs P, Sato K, Hayes PM, Matthews DE, Kleinhofs A, Muehlbauer GJ, DeYoung J, Marshall DF, Madishetty K, Fenton RD, Condamine P, Graner A, Waugh R (2009) Development and implementation of high-throughput SNP genotyping in barley. BMC Genomics 10:582
Cockram J, Chiapparino E, Taylor SA, Stamati K, Donini P, Laurie DA, O’Sullivan DM (2007) Haplotype analysis of vernalization loci in European barley germplasm reveals novel VRN-H1 alleles and a predominant winter VRN-H1/VRN-H2 multi-locus haplotype. Theor Appl Genet 115:993–1001
Cockram J, White J, Zuluaga DL, Smith D, Comadran J, Macaulay M, Luo Z, Kearsey MJ, Werner P, Harrap D, Tapsell C, Liu H, Hedley PE, Stein N, Schulte D, Steuernagel B, Marshall DF, Thomas WT, Ramsay L, Mackay I, Balding DJ, AGOUEB Consortium, Waugh R, O’Sullivan DM (2010) Genome-wide association mapping to candidate polymorphism resolution in the unsequenced barley genome. Proc Natl Acad Sci U S A 107(50):21611–21616
Comadran J, Russell JR, Booth A, Pswarayi A, Ceccarelli S, Grando S, Stanca AM, Pecchioni N, Akar T, Al-Yassin A, Benbelkacem A, Ouabbou H, Bort J, van Eeuwijk FA, Thomas WTB, Romagosa I (2011) Mixed model association scans of multi-environmental trial data reveal major loci controlling yield and yield related traits in Hordeum vulgare in Mediterranean environments. Theor Appl Genet 122:1363–1373
Costa JM, Corey A, Hayes PM, Jobet C, Kleinhofs A, Kopisch-Obusch A, Kramer SF, Kudrna D, Li M, Riera-Lizarazu O, Sato K, Szucs P, Toojinda T, Vales MI, Wolfe RI (2001) Molecular mapping of the Oregon Wolfe Barleys: a phenotypically polymorphic doubled-haploid population. Theor Appl Genet 103:415–424
Cuesta-Marcos A, Igartua E, Ciudad F, Codesal P, Russell J, Molina-Cano J, Moralejo M, Szűcs P, Gracia M, Lasa J, Casas A (2008) Heading date QTL in a spring × winter barley cross evaluated in Mediterranean environments. Mol Breed 21:455–471
Cuesta-Marcos A, Casas AM, Hayes PM, Gracia MP, Lasa JM, Ciudad F, Codesal P, Molina-Cano JL, Igartua E (2009) Yield QTL affected by heading date in Mediterranean grown barley. Plant Breed 128:46–53
Cuesta-Marcos A, Szűcs P, Close TJ, Filichkin T, Muehlbauer GJ, Smith KP, Hayes PM (2010) Genome-wide SNPs and re-sequencing of growth habit and inflorescence genes in barley: implications for association mapping in germplasm arrays varying in size and structure. BMC Genomics 11:707
Dat J, Vandenabeele S, Vranová E, Van Montagu M, Inzé D, Van Breusegem F (2000) Dual action of the active oxygen species during plant stress responses. Cell Mol Life Sci 57:779–795
Desclaux D, Romet P (1996) Impact of drought stress on the phenology of two soybean (Glycin max L.Merr) cultivars. Field Crops Res 46:61–70
Distelfeld A, Li C, Dubcovsky J (2009) Regulation of flowering in temperate cereals. Curr Opin Plant Biol 12:178–184
Ellis RP, Forster BP, Robinson D, Handley LL, Gordon DC, Russell JR, Powell W (2000) Wild barley: a source of genes for crop improvement in the 21st century? J Exp Bot 51:9–17
Eshghi R, Ojaghi J, Baraty M, Rahimi M, Salayeva S (2011) QTL mapping for yield and its components in barley (Hordeum vulgare L.). The 7th National Biotechnology Congress of I.R, Tehran
Fan Y, Shabala S, Ma Y, Xu R, Zhou M (2015) Using QTL mapping to investigate the relationships between abiotic stress tolerance (drought and salinity) and agronomic and physiological traits. BMC Genomics 16:43
FAOSTAT (2014) http://faostat3.fao.org/home
Farooq M, Wahid A, Kobayashi N, Fujita D, Basra SMA (2009) Plant drought stress: effects, mechanisms and management. Agron Sustain Dev 29:185–212
Farooq M, Bramley H, Palta JA, Siddique KHM (2011) Heat stress in wheat during reproductive and grain filling phases. Crit Rev Plant Sci 30:491–507
Faure S, Higgins J, Turner A, Laurie DA (2007) The FLOWERING LOCUS T-like gene family in barley (Hordeum vulgare). Genetics 176:599–609
Gallagher LW, Soliman KM, Vivar H (1991) Interactions among loci conferring photoperiod insensitivity for heading time in spring barley. Crop Sci 31:256–261
Górny AG (2001) Variation in utilization efficiency and tolerance to reduced water and nitrogen supply among wild and cultivated barley. Euphytica 117:59–66
Goulden CH (1941) Problems in plant selection. Proceeding of the Seventh Genetics Congress. Cambridge University Press, Cambridge, pp 132–133
Han F, Ullrich SE (1993) Mapping of quantitative trait loci associated with malting quality in barley. Barley Genet Newsl 23:84–97
Hayes PM, Liu BH, Knapp SJ, Chen F, Jones B, Blake T, Franckowiak J, Rasmusson D, Hearnden PR, Eckermann PJ, McMichael GL, Hayden MJ, Eglinton JK (1993) Quantitative trait locus effects and environmental interaction in a sample of North American barley germ plasm. Theor Appl Genet 87(3):392–401
Honsdorf N, March TJ, Berger B, Tester M, Pillen K (2014) High-throughput phenotyping to detect drought tolerance QTL in wild barley introgression lines. PLoS One 9(5):e97047. doi:10.1371/journal.pone.0097047
Hori K, Kobayashi T, Shimizu A, Sato K, Takeda K, Kawasaki S (2003) Efficient construction of high-density linkage map and its application to QTL analysis in barley. Theor Appl Genet 107:806–813
VSN International (2013) GenStat for Windows 16th Edition VSN International. Hemel Hempstead
International Barley Genome Sequencing Consortium (2012) A physical, genetic and functional sequence assembly of the barley genome. Nature 491:711–716
Janowska B, Andrzejak R (2010) Effect of gibberellic acid spraying and soaking of rhizomes on the growth and flowering of calla lily (Zantedeschia Spreng.). Acta Agrobot 63(2):155–160
Jones H, Leigh FJ, Mackay I, Bower MA, Smith LMJ, Charles MP, Jones G, Martin K, Jones MK, Brown TA, Powell W (2008) Population-based resequencing reveals that the flowering time adaptation of cultivated barley originated east of the Fertile Crescent. Mol Biol Evol 25:2211–2219
Karsai I, Meszaros K, Hayes PM, Bedo Z (1997) Effects of loci on chromosomes 2(2H) and 7(5H) on developmental patterns in barley (Hordeum vulgare L.) under different photoperiod regimes. Theor Appl Genet 94:612–618
Karsai I, Meszaros K, Lang L, Hayes PM, Bedo Z (2001) Multivariate analysis of traits determining adaptation in cultivated barley. Plant Breed 120:217–222
Kigel J, Konsens I, Rosen N, Rotem G, Kon A, Fragman-Sapir O (2011) Relationships between flowering time and rainfall gradients across Mediterranean-desert transects. Isr J Ecol Evol 57:91–109
Kjaer B, Haahr V, Jensen J (1991) Associations between 23 quantitative traits and 10 genetic markers in a barley cross. Plant Breed 106:261–274
Kuczyńska A, Surma M, Adamski T, Mikołajczak K, Krystkowiak K, Ogrodowicz P (2013) Effects of the semi-dwarfing sdw1/denso gene in barley. J Appl Genet 54:381–390
Kuczyńska A, Mikołajczak K, Ćwiek H (2014) Pleiotropic effects of the sdw1 locus in barley populations representing different rounds of recombination. Electron J Biotechnol 17(5):217–223
Kumar P, Raghava SPS, Misra RL, Singh KP (2003) Effect of GA3 on growth and yield of China aster. J Orna Hortic 6(2):110–112
Laurie DA, Pratchett N, Bezant JH, Snape JW (1994) Genetic analysis of a photoperiod response gene on the short arm of chromosome 2 (2H) of Hordeum vulgare. Heredity 72:619–627
Laurie DA, Pratchett N, Bezant JH, Snape JW (1995) RFLP mapping of five major genes and eight quantitive trait loci controlling flowering time in a winter × spring barley (Hordeum vulgare L.). Genome 38:575–585
Li J, Ji L (2005) Adjusting multiple testing in multilocus analyses using the eigenvalues of a correlation matrix. Heredity 95:221–227
Li JZ, Huang XO, Heinrichs F, Ganal MW (2005) Analysis of QTLs for yield, yield components, and malting quality in a BC3-DH population of spring barley. Theor Appl Genet 110:356–363
Loggini B, Scartazza A, Brugnoli E, Navari-Izzo F (1999) Antioxidative defense system, pigment composition, and photosynthetic efficiency in two wheat cultivars subjected to drought. Plant Physiol 119:1091–1099
Loss SP, Siddique KHM (1994) Morphological and physiological traits associated with wheat yield increases in Mediterranean environments. Adv Agron 52:229–276
Mäkelä P, Muurinen S (2011) Uniculm and conventional tillering barley accessions under northern growing conditions. J Agric Sci 150(03):335–344
Malicki MA, Plagge R, Roth CH (1996) Improving the calibration of dielectric TDR soil moisture determination taking into account the solid soil. Eur J Soil Sci 47:357–366
Malosetti M, Van Eeuwijk FA, Boer MP, Casas AM, Elia M (2011) Gene and QTL detection in a three-way barley cross under selection by a mixed model with kinship information using SNPs. Theor Appl Genet 122:1605–1616
Malosetti M, Ribaut J-M, van Eeuwijk FA (2013) The statistical analysis of multi-environment data: modeling genotype-by-environment interaction and its genetic basis. Front Physiol 4:44
Mansour E, Casas AM, Gracia AP, Molina-Cano JL, Moralejo M, Cattivelli L, Thomas WTB, Igartua E (2014) Quantitative trait loci for agronomic traits in an elite barley population for Mediterranean conditions. Mol Breed 33:249–265
Marquez-Cedillo LA, Hayes PM, Kleinhofs A, Legge WG, Rossnagel BG, Sato K, Ullrich SE, Wesenberg DM (2001) QTL analysis of agronomic traits in barley based on the doubled haploid progeny of two elite North American varieties representing different germplasm groups. Theor Appl Genet 103:625–637
Mayer KF, Taudien S, Martis M, Šimková H, Suchánková P, Gundlach H, Wicker T, Petzold A, Felder M, Steuernagel B, Scholz U, Graner A, Matthias Platzer M, Doležel J, Stein N (2009) Gene content and virtual gene order of barley chromosome 1H. Plant Physiol 151:496–505
Mehravaran L, Fakheri B, Sharifi-Rad J (2014) Localization of quantitative trait loci (QTLs) controlling drought tolerance in Barley. Int J Biosci 5(7):248–259
Mikołajczak K, Ogrodowicz P, Gudyś K, Krystkowiak K, Sawikowska A, Frohmberg W, Górny A, Kędziora A, Jankowiak J, Józefczyk D, Karg G, Andrusiak J, Krajewski P, Szarejko I, Surma M, Adamski T, Guzy–Wróbelska J, Kuczyńska A (2016) Quantitative trait loci for yield and yield–related traits in spring barley populations derived from crosses between European and Syrian cultivars. PLoS ONE 11(5):e0155938
Miralles DJ, Slafer GA (1995) Yield, biomass and yield components in dwarf, semi-dwarf and tall isogenic lines of spring wheat under recommended and late sowing dates. Plant Breed 114:392–396
Mohammadi M, Baum M (2008) QTL analysis for morphological traits in doubled haploid population of barley. Sci Technol Agric Nat Res 45(A):111–120
Moralejo M, Swanston JS, Munoz P, Prada D, Elia M, Russell JR, Ramsay L, Cistue L, Codesal P, Casas AM, Romagosa I, Powell W, Molina-Cano JL (2004) Use of new EST markers to elucidate the genetic differences in grain protein content between European and North American two-rowed malting barleys. Theor Appl Genet 110:116–125
Moran JF, Becana M, Iturbe-Ormaetxe I, Frechilla S, Klucas RV, Aparicio-Tejo P (1994) Drought induces oxidative stress in pea plants. Planta 194:346–352
Muñoz-Amatriaín M, Moscou MJ, Bhat PR, Svensson JT, Bartoš J, Suchánková P, Šimková H, Endo TR, Fenton RD, Lonardi S, Castillo AM, Chao S, Cistué L, Cuesta-Marcos A, Forrest KL, Hayden MJ, Hayes PM, Horsley RD, Makoto K, Moody D, Sato K, Vallés MP, Wulff BBH, Muehlbauer GJ, Doležel J, Close TJ (2011) An improved consensus linkage map of barley based on flow-sorted chromosomes and single nucleotide polymorphism markers. Plant Genome 4:238–249
Nevo E, Chen G (2010) Drought and salt tolerances in wild relatives for wheat and barley improvement. Plant Cell Environ 33:670–685
Passioura JB (1996) Drought and drought tolerance. Plant Growth Regul 20:79–83
Peighambari SA, Yazdi Samadi B, Nabipour N, Charmet G, Sarrafi A (2005) QTL analysis for agronomic traits in a barley doubled haploid population grown in Iran. Plant Sci 169:1008–1013
Pillen K, Zacharias A, Léon J (2003) Advanced backcross QTL analysis in barley (Hordeum vulgare L.). Theor Appl Genet 107:340–352
Qi X, Niks RE, Stam P, Lindhout P (1998) Identification of QTLs for partial resistance to leaf rust (Puccinia hordei) in barley. Theor Appl Genet 96:1205–1215
Ren XF, Sun DF, Guan WW, Sun GL (2010) Inheritance and identification of molecular markers associated with a novel dwarfing gene in barley. BMC Genet 11:89
Richards RA (1996) Defining selection criteria to improve yield under drought. Plant Growth Regul 20:157–166
Richards RA (2006) Physiological traits used in the breeding of new cultivars for water-scarce environments. Agric Water Manag 80:197–211
Roberts EH, Summerfield RJ, Cooper JP, Ellis RH (1988) Environmental control of flowering in barley (Hordeum vulgare L.). I. Photoperiod limits to long day responses, photoperiod-insensitive phases and effects of low-temperature and short-day vernalization. Ann Bet 62:127–144
Sakamoto T, Matsuoka M (2004) Generating high-yielding varieties by genetic manipulation of plant architecture. Curr Opin Biotechnol 15(2):144–147
Samarah NH (2005) Effects of drought stress on growth and yield of barley. Agron Sustain Dev 25:145–149
Samarah NH, Alqudah AM, Amayreh JA, McAndrews GM (2009) The effect of late-terminal drought stress on yield components of four barley cultivars. J Agron Crop Sci 195(6):427–441
Sameri M, Takeda K, Komatsuda T (2006) Quantitative trait loci controlling agronomic traits in recombinant inbred lines from a cross of oriental- and occidental-type barley cultivars. Breed Sci 56:243–252
Sameri M, Pourkheirandish M, Chen G, Tonooka T, Komatsuda T (2011) Detection of photoperiod responsive and non-responsive flowering time QTL in barley. Breed Sci 61:183–188
Sasani S, Hemming MN, Oliver SN, Greenup A, Reza Tavakkol-Afshari R, Sayed MA, Schumann H, Pillen K, Naz AA, Leon J (2012) AB-QTL analysis reveals new alleles associated to proline accumulation and leaf wilting under drought stress conditions in barley (Hordeum vulgare L.). BMC Genet 13:61
Schmalenbach I, Zhang L, Reymond M, Jiménez-Gómez JM (2014) The relationship between flowering time and growth responses to drought in the Arabidopsis Landsberg erecta x Antwerp-1 population. Front Plant Sci 5:609
Shirazi M, Gyamfi J, Ram T, Bachiri H, Rasyid B, Rehman A, Khan M, Mujtaba S, Ali M, Shreen A (2010) Selection of some suitable drought tolerant wheat genotypes using carbon isotopes discrimination (cid) technique. Pak J Bot 42:3639–3644
Slafer GA, Araus JL, Royo C, Garcia Del Moral LF (2005) Promising eco-physiological traits for genetic improvement of cereal yields in Mediterranean environments. Ann Appl Biol 146:61–70
Smalle J, Vierstra RD (2004) The ubiquitin 26S proteasome proteolytic pathway. Annu Rev Plant Biol 55:555–590
Smirnoff N (1993) The role of active oxygen in the response of plants to water deficit and desiccation. New Phytol 125:27–58
Stone SL, Callis J (2007) Ubiquitin ligases mediate growth and development by promoting protein death. Curr Opin Plant Biol 10:624–632
Surma M, Adamski T, Święcicki W, Barzyk P, Kaczmarek Z, Kuczyńska A, Krystkowiak K, Mikołajczak K, Ogrodowcz P (2013) Preliminary results of in vitro culture of pea and lupin embryos for the reduction of generation cycles in single seed descent technique. Acta Soc Bot Pol 82:231–236
Szűcs P, Skinner JS, Karsai I, Cuesta-Marcos A, Haggard KG, Corey AE, Chen TH, Hayes PM (2007) Validation of the VRN-H2/VRN-H1 epistatic model in barley reveals that intron length variation in VRN-H1 may account for a continuum of vernalization sensitivity. Mol Genet Genome 277:249–261
Taiz L, Zeiger E (1998) Plant physiology, 2nd edn. Sinauer, Sunderland, p 792
Takahashi R, Yasuda S (1956) Genetic studies of spring and winter habit of growth in barley. Bericht des Ohara Instituts für Landwirtschaftliche Biologie, Okayama 10:245–308
Talamè V, Ozturk NZ, Bohnert HJ, Tuberosa R (2007) Barley transcript profiles under dehydration shock and drought stress treatments: a comparative analysis. J Exp Bot 58:229–240
Thomas WTB, Powell W, Waugh R, Chalmers KJ, Barua UM, Jack P, Lea V, Forster BP, Swanston JS, Ellis RP, Hanson PR, Lance RCM (1995) Detection of quantitative trait loci for agronomic, yield, grain and disease characters in spring barley (Hordeum vulgare L.). Theor Appl Genet 91:1037–1047
Tinker NA, Mather DE, Rossnagel BG, Kasha KJ, Kleinhofs A, Hayes PM, Falk DE, Ferguson T, Shugar LP, Legge WG, David HL (1996) Regions of the genome that affect agronomic performance in two-row barley. Crop Sci 36:1–0531062
Tondelli A, Francia E, Barabaschi D, Aprile A, Skinner JS, Stockinger EJ, Stanca AM, Pecchioni N (2006) Mapping regulatory genes as candidates for cold and drought stress tolerance in barley. Theor Appl Genet 112:445–454
Tsenov N, Atanasova D, Stoeva I, Tsenova E (2015) Effects of drought on grain productivity and quality in winter bread wheat. Bulg J Agric Sci 21(3):592–598
Turner A, Beales J, Faure S, Dunford RP, Laurie DA (2005) The pseudo-response regulator Ppd-H1 provides adaptation to photoperiod in barley. Science 310:1031–1034
Tyagi AK, Singh CN (2006) Effect of GA3 and IBA on flowering and bulb production in tuberose (Polianthes tuberosa Linn.) cultivar Pearl Double. J Orna Hortic 9(2):152
Van Ooijen JW, Voorrips RE (2001) JoinMap® 3.0, software for the calculation of genetic linkage maps PRI. Wageningen
Varshney RK, Marcel TC, Ramsay L, Russell J, Röder MS, Stein N, Waugh R, Langridge P, Niks RE, Graner A (2007) A high density barley microsattelite consensus map with 775 SSR loci. Theor Appl Genet 114:1091–1110
Wang CL, Chen YH, Ku LX, Wang TG, Sun ZH et al (2010a) Mapping QTL associated with photoperiod sensitivity and assessing the importance of QTL × environment interaction for flowering time in maize. PLoS One 5:e14068. doi:10.1371/journal.pone.0014068
Wang G, Schmalenbach I, von Korff M, Léon J, Kilian B, Rode J, Pillen K (2010b) Association of barley photoperiod and vernalization genes with QTLs for flowering time and agronomic traits in a BC2DH population and a set of wild barley introgression lines. Theor Appl Genet 120:1559–1574
Wenzl P, Li H, Carling J, Zhou M, Raman H, Paul E, Hearnden P, Maier C, Xia L, Caig V, Ovesna J, Cakir M, Poulsen D, Wang J, Raman R, Smith KP, Muehlbauer GJ, Chalmers KJ, Kleinhofs A, Huttner E, Kilian A (2006) A high density consensus map of barley linking DArT markers to SSR, RFLP and STS loci and agricultural traits. BMC Genomics 7:206
Winkel T, Renno JF, Payne WA (1997) Effect of the timing of water deficit on growth, phenology and yield of pearl millet (Pennisetum glaucum (L.) R. Br.) grown in Sahelian conditions. J Exp Bot 48:1001–1009
Wopereis MCS, Kropff MJ, Maligaya AR, Tuong TP (1996) Drought-stress responses of two lowland rice cultivars to soil water status. Field Crop Res 46:21–39
Xu Y (2010) Molecular plant breeding. CAB International, Wallingford
Xu JL, Lafitte HR, Gao YM, Fu BY, Torres R, Li ZK (2005) QTLs for drought escape and tolerance identified in a set of random introgression lines of rice. Theor Appl Genet 111(8):1642–1650
Yano M, Katayose Y, Ashikari M, Yamanouchi U, Monna L, Fuse T, Baba T, Yamamoto K, Umehara Y, Nagamura Y, Sasaki T (2000) Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell 12:2473–2484
Zare M (2012) Evaluation of drought tolerance indices for the selection of Iranian barley (Hordeum vulgare L.) cultivars. Afr J Biotechnol 11:15975–15981
Zhao MS, Running SW (2009) Drought-induced reduction in global terrestrial net primary production from 2000 through 2009. Science 329:940
Zhou G, Zhang Q, Zhang X-q, Tan C, Li C (2015) Construction of high-density genetic map in barley through restriction-site associated DNA sequencing. PLoS ONE 10(7):e0133161
Zinselmeier C, Jeong BR, Boyer JB (1999) Starch and the control of kernel number in maize at low water potentials. Plant Physiol 121:25–36
Acknowledgments
This work was supported by the European Regional Development Fund through the Innovative Economy Programme 2007–2013, project WND-POIG.01.03.01–00–101/08 POLAPGEN-BD “Biotechnological tools for breeding cereals with increased resistance to drought”.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by: Andrzej Górny
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Ogrodowicz, P., Adamski, T., Mikołajczak, K. et al. QTLs for earliness and yield-forming traits in the Lubuski × CamB barley RIL population under various water regimes. J Appl Genetics 58, 49–65 (2017). https://doi.org/10.1007/s13353-016-0363-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-016-0363-4