Abstract
The aim of this study was to assess the antimicrobial resistance and the prevalence of genes determining the presence of extended-spectrum beta-lactamase (ESBL) enzymes in Escherichia coli isolated from two major rivers of the Podhale region in southern Poland. In total, 196 E. coli isolates were analyzed—98 from each river—Białka and Zakopianka, collected in 8 campaigns, over the period of two years. Antimicrobial resistance was assessed using disk diffusion method and PCR tests were conducted to detect the ESBL genes. In E. coli isolated from Białka, the resistance to amoxicillin/clavulanic acid was detected most frequently (54.08%) and ESBL was detected in 14.29% of strains. In strains isolated from Zakopianka, most frequent resistance was observed toward ticarcillin (51.02%), while ESBL was observed in 16.33% of isolates. In the total pool of isolates, the resistance to amoxicillin/clavulanic acid was most frequent (48.98% of isolates) and ESBL producers comprised 15.30% of E. coli isolates derived from both rivers. Multidrug resistance was observed less frequently in strains derived from Białka (4 isolates resistant to 10 and more antimicrobials) than from Zakopianka, where 10 isolates were resistant to 10 and more antibiotics. Out of the tested ESBL genes blaTEM was detected most frequently (45.4% of isolates), whereas blaCTX-M1 and blaCTX-M3 were recorded in one isolate.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Increasing consumption of antimicrobial agents throughout Europe results in their discharge to surface waters through sewage treatment plants via human urine and feces (Łuczkiewicz et al. 2010). Although their concentration in sewage is significantly lower than the one used in therapy, it most definitely affects microorganisms and results in the selection of resistant strains (Zabłotni and Jaworski 2014). The overuse of antimicrobial agents in medicine, agriculture and animal breeding causes the selection of resistance not only in pathogenic bacteria but also in commensal and environmental strains (van den Bogaard and Stobberingh 2000). Subsequently, such strains may serve as a source of resistance genes that can be transferred to pathogenic strains of the same species and to other bacterial species due to horizontal gene transfer (Zabłotni and Jaworski 2014). All this resulted in widespread prevalence of antimicrobial-resistant bacteria in a broad range of environments, including soil or surface water (Baquero et al. 2008). Therefore, the probability of exposure to the resistant strains of bacteria increases, since an infection might occur through contact or ingestion of contaminated surface water, for instance during recreational activities (Blaak et al. 2014). This may result in both risks related to difficult in treatment infections or exposure to harmless strains of e.g., commensal-resistant bacteria, thus resulting in asymptomatic carriage of such strains (Blaak et al. 2015). Such commensals are often opportunistic pathogens, e.g., Escherichia coli, which can be transferred to immunocompromised people or those more vulnerable to infection, or may cause infection in asymptomatic carriers while they become more susceptible to infection (Blaak et al. 2015). Since surface water (most frequently rivers) often acts as a receiver of effluent from treatment plants and sewage directly discharged from households or can be contaminated by animal feces, it should be considered as one of the possible routes for transfer of fecal species that are either resistant to antibiotics or carry the resistance-conferring genes (Wolny-Koładka and Lenart-Boroń 2016).
One of the most important mechanisms of resistance observed in E. coli is the production of extended-spectrum beta-lactamase enzymes (ESBL). The ESBL-producing strains are of particular concern due to the fact that they are resistant to all penicillins, cephalosporins and aztreonam. Moreover, they can exhibit cross-resistance to trimethoprim/sulfamethoxazole and quinolones (Picozzi et al. 2014). The ESBL-conferring genes are located on large plasmids, which can carry the genes for resistance to numerous other groups of antimicrobials (Rawat and Nair 2010). Therefore, ESBL-producing E. coli isolates not only can be characterized by very broad antibiotic resistance, but also can be a source of rapidly spread resistance genes among strains of different species. All this causes significant problems in effective therapy in the case of possible infection and creates the need for monitoring the occurrence of such strains in the environment coupled with molecular analyses in order to determine the most prevalent ESBL-determining genes in such strains.
Podhale is a cultural region in southern Poland, which is also one of the most popular Polish touristic areas both in winter (ski resorts) and in summer (health resorts, mountain climbing and hiking). This resulted in recent intensive development of tourist infrastructure and thus increased consumption of water resources, large amount of produced waste and the emission of pollutants as well as large amounts of sewage discharged to rivers (Lenart-Boroń et al. 2016).
The aim of this study was to assess the prevalence of antimicrobial-resistant fecal strains of E. coli in waters of two main rivers in the Podhale region and to evaluate the presence of genes responsible for the production of extended-spectrum β-lactamases in waterborne E. coli isolated from those rivers.
The samples of water were collected over two winter seasons (December–March) 2014/2015 and 2015/2016 along two rivers—Białka and Zakopianka in Podhale, southern Poland.
Materials and methods
Study area and sampling
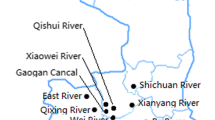
The sampling sites were situated along the course of two main rivers of Podhale—Białka and Zakopianka, which turns into Dunajec. The Białka river along its course flows through the protected areas—its springs are located in the Tatra National Park—while the remaining part is protected under the Natura 2000 programme (the Białka valley—habitat protection area). The river Zakopianka in its lower course turns into Dunajec, which is also protected under the Natura 2000 programme. The major sources of point pollution reaching both rivers are sewage discharge sites—mostly from households but also discharge from the treatment plants—in Bukowina Tatrzańska (Białka river) and Zakopane (Zakopianka). Both rivers flow through popular touristic localities, i.e., Białka through a few large ski resorts in Podhale—Jurgów, Bukowina and Białka Tatrzańska—while Zakopane itself is one of the most frequently visited tourist destinations, both in winter and in summer.
Water samples were collected at eleven sites—six along the course of Białka and five along Zakopianka, selected based on their characteristics (Fig. 1; Table 1). Sampling was conducted in eight campaigns over two winter seasons (the period of the highest tourist traffic in the considered region). Water samples were collected into 1000-ml sterile polypropylene bottles, and temperature and electrolytic conductivity (EC25 °C) were measured onsite with a handheld multimeter (YSI Pro 2030, USA).
Laboratory analyses
Enumeration of E. coli was conducted with the membrane filtration method using TBX agar (incubation at 44 °C, 48 h). Blue-green colonies were preliminarily identified as E. coli, then purified with plate streaking and their species was confirmed using MALDI-TOF mass spectrometry. Ninety-eight bacterial isolates were selected from different sites of each river for further analyses (Table 1).
Antimicrobial resistance of E. coli was tested using the disk diffusion method following the recommendations of the European Committee on Antimicrobial Susceptibility Testing (EUCAST 2016) and in the case of antimicrobial agents not specified by the EUCAST guidelines, i.e., cephalotin—Kronvall et al. (1984), cephazolin—Turnidge (2011), cephamandole—Barry et al. (1983) and tetracycline—Sader et al. (2007). Cartridges of antimicrobial disks were obtained from Oxoid (Great Britain). Bacterial isolates were transferred to sterile saline solutions to prepare 0.5 MacFarland suspension standards, which were then streaked onto Mueller–Hinton II agar (BTL, Poland), and antimicrobial disks were applied. The presence of ESBL was confirmed with the double-disk synergy test (Drieux et al. 2008). After incubation for 18–20 h at 36 ± 1 °C, the diameters of growth inhibition zones around the antimicrobial disks were measured and the results were compared with the breakpoint values recommended by the EUCAST (2016). Quality control was performed using the E. coli strain ATCC 25922.
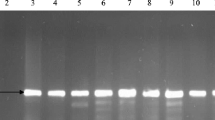
In order to determine the presence of ESBL genes, DNA was extracted from all 196 strains isolated from the studied rivers and the control E. coli strain ATCC 25922 using the Genomic Mini DNA extraction kit (A&A Biotechnology, Poland) in accordance with the manufacturer’s instructions. PCR tests were then conducted using specific primers: blaCTXM3 (Costa et al. 2006), blaCTXM9 (Simarro et al. 2000), blaOXA, blaSHV and blaTEM (Sáenz et al. 2004). Primer sequences and annealing temperatures are given in Table 2. The reactions were performed in a 25 µl volume containing 50 ng of DNA template, 12.5 pM of each primer, 2.5 mM of dNTP, 1 × PCR buffer and 1 U DreamTaq DNA polymerase (Thermo Scientific, US). The following temperature profile was used for the reactions: initial denaturation at 95 °C for 5 min, followed by 35 cycles of 94 °C for 45 s, annealing for 45 s at temperatures corresponding to individual primers, then extension at 72 °C for 1 min with final extension at 72 °C for 10 min and then storage at 4 °C. PCR amplifications were performed in T100™ Thermal Cycler (Bio-Rad, USA). The PCR products were electrophoresed for 60 min in 1 × TBE, 1% agarose gel, stained with Simply Safe (0.5 mg/ml; EurX, Poland), visualized in UV light and documented by the Gel Doc system (Bio-Rad, US).
Statistical analysis
Basic descriptive statistics were calculated, and the significance of differences in the prevalence of E. coli in the examined sites was determined using a one-way ANOVA test conducted in Statistica v. 10 (StatSoft, US). The differences in the prevalence of bacterial resistance between the samples collected from both rivers were estimated using a Chi-square test.
Results and discussion
As listed in Table 1, the mean number of fecal E. coli in the Białka river ranged from 3 CFU/100 ml at the outflow from the Tatra National Park (site Łysa Polana, B1) to more than 2 million CFU/100 ml by the discharge from the sewage treatment plant in Bukowina Tatrzańska (STP, B3). This point source of contamination affects the quality of water downstream, among others at the intake for artificial snowing of slopes of the Kotelnica ski station. Detailed analysis of contamination of the Białka river and the co-occurring mechanisms are discussed by Lenart-Boroń et al. (2016). In the case of the second examined river, Zakopianka, the greatest number of E. coli, was also related to the discharge from the local treatment plant (after STP, Z3), where the mean value was 47,700 CFU/100 ml. Similarly as in the case of Białka, this source of contamination contributed to the increased prevalence of E. coli downstream of the treatment plant and the entire Zakopane locality (i.e., 32,300 CFU/100 ml at the site Poronin bridge, Z4).
Currently in Poland, there are no regulations allowing to classify surface waters based on the microbiological contamination. However, comparison of the mean values of E. coli concentration obtained in this analysis with the Regulation of the Minister of the Environment (2004), which was later repealed, shows that three samples (B3, Z3 and Z4) should be qualified as 5th class of water, i.e., water of bad quality, whose values of biological quality indicators demonstrate that due to significant anthropogenic pressure most of biological populations disappeared. Unsatisfactory quality (class 4th) was detected in the samples B4 (Kotelnica intake) and Z1 (before the hospital), which means that as a result of anthropogenic pressure biological populations in those waters were subjected to major qualitative and quantitative changes. On the other hand, comparing the concentrations of E. coli with the so-called Bathing Water Directive (Directive 2006/7/EC 2006) indicates that the quality of water is poor only in the site B3 (STP), sufficient in Z3 and Z4 (after STP and Poronin bridge) and excellent in the other sites.
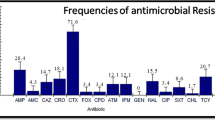
In the course of the conducted study, ninety-eight E. coli strains were isolated from each river, resulting in 196 isolates selected for further studies, i.e., assessment of resistance to antimicrobials important in treating human infections, coupled with the detection of extended-spectrum beta-lactamases (ESBL) and determination of the presence of ESBL-encoding genes. As listed in Table 3, the resistance to most commonly used in Poland groups of antibiotics (Łuczkiewicz et al. 2010), i.e., penicillins: ampicillin, amoxicillin/clavulanic acid and ticarcillin, all three belonging to the class of beta-lactams, was most frequently detected in strains isolated from both rivers. The prevalence of resistance to different antimicrobials was similar in strains from both rivers, with the exception of aminoglycosides, i.e., amikacin, netilmycin and tobramycin. Only in the case of those three antibiotics, the differences in the number of resistant strains were statistically significant (Chi-square values of 5.12, 7.80 and 8.88, respectively). In the case of strains isolated from Białka, aminoglycosides netilmycin and tobramycin were the most efficient, as only 1.02% of E. coli isolates demonstrated the resistance to those antimicrobials (Table 3). Ciprofloxacin (the group of fluoroquinolones) was the most efficient antibiotic toward strains isolated from Zakopianka (5.10% of resistant strains). The percentage of ESBL-producing strains was also similar in both rivers (i.e., 11.22% in Białka and 12.25 in Zakopianka). The multidrug-resistant (MDR) strains comprised 16.3% (n = 16) in Białka and as many as 25.5% (n = 25) in the Zakopianka river (Table 4). Those strains are resistant to at least one antimicrobial agent belonging to three or more classes (http://ecdc.europa.eu/en/activities/diseaseprogrammes/ARHAI/Pages/public_consultation_clinical_microbiology_infection_article.aspx). In the case of extensively drug-resistant strains (XDR), i.e., those resistant to at least one antimicrobial agent from the maximum of two different classes, their share was 25.5 and 30.6% (n = 25 in Białka and n = 30 in Zakopianka, respectively).
The pattern of resistance observed in this study is widely reported by other authors. For instance, retrospective analysis of the prevalence of drug-resistant E. coli over the period of 2004–2007 by Daniluk et al. (2008) on clinical E. coli strains showed that the share of ampicillin-resistant isolates fluctuates around 40%, which is similar to the results obtained in this study. On the other hand, Sacha et al. (2007) in their studies on the mechanisms of resistance to β-lactam antibiotics in cefotaxime-resistant bacterial species observed that antibiotics from the group of aminoglycosides, especially amikacin, were most efficient, as only 4.2% of E. coli strains were resistant to this antimicrobial agent. In the case of studies conducted on environmental samples, the results of this study are generally concurrent with those of other authors in terms of most frequently observed resistance, as the resistance to ampicillin among E. coli strains was also most frequently observed by e.g., Łuczkiewicz et al. (2010, 2011) in surface water samples and samples collected from a sewage treatment plant, or by Reinthaler et al. (2010) in E. coli isolated from the samples of sewage, sludge and receiving waters of treatment plants. Aminoglycosides proved to be the most efficient antimicrobials in research of the previously mentioned authors, with no resistance to amikacin and tobramycin found by Łuczkiewicz et al. in their study of 2010, and no resistance to amikacin and gentamycin found by Łuczkiewicz et al. in 2011.
On the other hand, the prevalence of ESBL-producing strains in surface water and wastewater samples varies in the literature. Reinthaler et al. (2010) reported ESBL-positive E. coli strains in more than 60% of the sludge samples tested, while Łuczkiewicz et al. (2010) reported the presence of ESBL producers only once per a total on 153 isolates of E. coli. Wolny-Koładka and Lenart-Boroń (2016) did not detect the presence of ESBL-producing E. coli strains in water of the Nowohucki Reservoir in Poland.
The percentage of MDR strains observed in the study is disturbingly high, as compared to research conducted by other authors, since Łuczkiewicz et al. (2011) recorded MDR strains at the level of 9% in surface water of Gdańsk and Puck bays and at the same level in a sewage treatment plant in Gdańsk (Łuczkiewicz et al. 2010). The percentage of MDR E. coli strains observed by Blaak et al. (2015) in different types of surface water reached 11 with 26% of isolates resistant to at least one antimicrobial. In this study, the proportion of isolates resistant to at least one antimicrobial agent is significantly larger, i.e., 86.7% in Białka and 82.7% in Zakopianka. Moreover, MDR strains were present in all sampling sites along the Białka river and in all sites along the Zakopianka. What also should be mentioned is that among strains isolated from both rivers there were four (Białka) and ten (Zakopianka) resistant to ten or more antimicrobials. The two resistant to as many as 14 and 15 antibiotics were isolated from the site Z3 (after the STP), but the remaining ones were derived from all other sites along the river. These results indicate that, similarly to the observations of Blaak et al. (2015), given the functions of the analyzed rivers Białka and Zakopianka, there is high danger of transmission of antimicrobial-resistant strains to humans through contact with surface water and with artificial snow produced thereof. Data showing the content of fecal E. coli in the tested waters and the prevalence of antimicrobial-resistant and MDR strains indicate that the distance of sampling sites from sources of contamination affects not only the sanitary quality of water but also the proportion of strains resistant to antimicrobials (Table 5). However, XDR and MDR strains, as well as one strain resistant to even 7 antimicrobial agents, were detected in the sampling site located at the border of the Tatra National Park (B1). Although this was the site with most clean water (the lowest concentration of E. coli), two XDR strains and one MDR were detected. This indicates that virtually entire region of the Tatras is threatened by the drug-resistant E. coli. Such ubiquity could be a result of heavy tourist traffic throughout this area, including high mountains and the National Park.
Among the tested ESBL-encoding genes, blaTEM was the most prevalent one, as it was detected in 89 (45.4%) strains—42.9% of strains from Białka and 47.9% from Zakopianka. This observation is concurrent with Bradford (2001), according to whom TEM is the most commonly detected β-lactamase in Gram-negative bacteria. Also TEM and SHV-type ESBLs are most frequently found in E. coli and K. pneumoniae. Baraniak (2010) demonstrated that the TEM family is most divergent group of ESBL enzymes in Poland, represented by at least 10 variants of β-lactamases, among which eight were identified only in Poland. Evolution of TEM occurs by acquiring subsequent point mutations, which results in the formation of increasingly specialized enzymes that enable adaptation of bacterial strains to various environments, where they get in contact with different β-lactam antimicrobials. Epidemiological analysis of TEM-producing E. coli strains, carried out by Baraniak (2010), revealed a variety of epidemiological phenomena in Polish hospitals. Clonal disease outbreaks were identified in some of the tested facilities and a horizontal spread of plasmids carrying ESBL-conferring genes among strains of different Enterobacteriaceae species was demonstrated.
Out of the remaining genes, only blaCTX-M1 and blaCTX-M3 were recorded in one ESBL-positive isolate (derived from the site Z2—after sewage discharge from the hospital in Zakopane), which also possessed blaTEM gene (Table 5). What should also be noted is that substantially higher percentage of strains carrying ESBL-conferring genes was detected compared to those in which this mechanism was observed in phenotypic assays. Similar observations were described by Wolny-Koładka and Lenart-Boroń (2016), who detected various ESBL-determining genes (among which blaTEM was also most frequent one) in 38% of isolates, among which none exhibited ESBL phenotype. Based on these observations, it can be concluded that it is important to conduct both types of ESBL detection assays (i.e., phenotypic and PCR based), since the double-disk synergy test is important from an epidemiological point of view, allowing to monitor and control potential infection outbreaks (Gniadkowski et al. 2009). What can be seen is that despite the presence of the resistance-determining genes, the disk diffusion test may not indicate the occurrence of any specific resistance mechanism (Idzik et al. 2000). Therefore, it is worth to conduct the PCR-based tests in order to assess the potentially dangerous strains that may spread the ESBL genes among environmental strains of E. coli and other Enterobacteriaceae species.
Conclusion
This study shows that due to multiple sources of contamination, the analyzed major rivers of Podhale are severely contaminated with fecal bacteria. Numerous strains of fecal E. coli were characterized by multidrug resistance as well as the production of ESBL, mostly due to the presence of blaTEM gene. Such high prevalence and spread of multidrug-resistant E. coli along both rivers are disturbing, particularly that they are also present in sites located in the protected sections of the Białka river. Considering the fact that waters of both rivers are used for recreational purposes and for artificial snowing of ski slopes located along both rivers, the threat of transmission of drug-resistant bacteria to humans, as well as the transfer of ESBL-determining genes to strains of other Enterobacteriaceae species is significant. This indicates the need to take actions aimed at limiting the number of water contamination sources, decreasing the pollutant load and reduce the risk of transmission to humans.
Change history
19 September 2017
The text in the original article on page 246 left hand column contains errors.
References
Baquero F, Martínez JL, Cantón R (2008) Antibiotics and antibiotic resistance in water environments. Curr Opin Biotechnol 19:260–265
Baraniak A (2010) Molecular epidemiology and evolution of Enterobacteriaceae strains producing extended spectrum β-lactamases (ESBL) in Poland. PhD dissertation, Faculty of Biology, Warsaw University
Barry AL, Jones RN, Thornsberry C (1983) Evaluation of the cefonicid disk criteria, including disk quality control guidelines. J Clin Microbiol 17(2):232–239
Blaak H, de Kruijf P, Hamidjaja RA, van Hoek AHAM, de Roda Husman AM, Schets FM (2014) Prevalence and characteristics of ESBL-producing E. coli in Dutch recreational waters influenced by wastewater treatment plants. Vet Microbiol 171:448–459
Blaak H, Lynch G, Italiaander R, Hamidjaja RA, Schets FM, de Roda Husman AM (2015) Multidrug-resistant and extended spectrum β-lactamase-producing Escherichia coli in dutch surface water and wastewater. PLoS ONE 10(6):e0127752
Bradford PA (2001) Extended-spectrum β-lactamases in the twenty first century: characterization, epidemiology, and detection of this important resistance threat. Clin Microbiol Rev 14(4):933–951
Costa D, Poeta P, Saenz Y, Vinue L, Rojo-Bezares B, Jouini A, Zarazaga M, Rodrigues J, Torres C (2006) Detection of Escherichia coli harbouring extended-spectrum β-lactamases of the CTX-M, TEM and SHV classes in faecal samples of wild animals in Portugal. J Antimicrob Chemother 58(6):1311–1312
Daniluk T, Ściepuk M, Fiedoruk K, Zaremba ML, Rożkiewicz D, Ołdak E (2008) Prevalence rate and antibiotic susceptibilities of Escherichia coli and Klebsiella pneumoniae strains isolated from hospitalized children in the University Children Hospital during 2004–2007. Nowiny Lekarskie 77(4):265–272 (in Polish)
Directive 2006/7/EC of the European Parliament and of the Council of 15 February 2006 concerning the management of bathing water quality and repealing Directive 76/160/EC
Drieux L, Brossier F, Sougakoff W, Jarlier V (2008) Phenotypic detection of extended-spectrum beta-lactamase production in Enterobacteriaceae: review and bench guide. Clin Microbiol Infect 14(suppl. 1):90–103
European Committee on Antimicrobial Susceptibility Testing (2016) Clinical breakpoints – bacteria (v. 6.0). http://ecdc.europa.eu/en/activities/diseaseprogrammes/ARHAI/Pages/public_consultation_clinical_microbiology_infection_article.aspx
Gniadkowski M, Żabicka D, Hryniewicz W (2009) Recommended selection of the antimicrobial susceptibility tests for bacteria. Determination of Gram-negative rods susceptibility. National Reference Centre for Antimicrobial Susceptibility, Warsaw
Idzik D, Wojtyczka R, Kępa M, Wydmuch Z, Pacha J, Głab S, Wróblewska I, Skowronek-Ciołek O (2000) Evaluation of methicillin-resistance in Staphylococcus aureus by the agar disk diffusion method and PCR. Exp Med Microbiol 52(4):327–332
Kronvall G, Petersson AC, Ljunggren K, Soltesz V (1984) Single-strain regression analysis for quality control of cephalotin-susceptibility testing and determination of interpretive breakpoints. Acta Pathol Microbiol Immunol Scand B 92(1):13–22
Lenart-Boroń A, Wolanin A, Jelonkiewicz Ł, Chmielewska-Błotnicka D, Żelazny M (2016) Spatiotemporal variability in microbiological water quality of the Białka River and its relation to the selected physicochemical parameters of water. Water Air Soil Poll 227:22. doi:10.1007/s11270-015-2725-7
Łuczkiewicz A, Jankowska K, Fudala-Książek S, Olańczuk-Neyman K (2010) Antimicrobial resistance of fecal indicators in municipal wastewater treatment plant. Water Res 44:5089–5097
Łuczkiewicz A, Jankowska K, Kurlenda J, Olańczuk-Neyman K (2011) Identification and antimicrobial susceptibility of fecal coliforms isolated from surface water. Pol J Environ Stud 20(4):941–950
Picozzi SCM, Casellato S, Rossini M, Paola G, Tejada M, Costa E, Carmignani L (2014) Extended-spectrum beta-lactamase-positive Escherichia coli causing complicated upper urinary tract infection: urologist should act in time. Urol Ann 6(2):107–112
Rawat D, Nair D (2010) Extended-spectrum β-lactamases in Gram negative bacteria. J Glob Infect Dis 2(3):263–274
Regulation of the Minister of Environment of 11 February 2004 on the classification for presenting the condition of surface and groundwater, methods of monitoring and interpretation of the results and presenting the condition of these waters. J Laws Repub Pol (2004) No. 32 item 284
Reinthaler FF, Feierl G, Galler H, Haas D, Leitner E, Mascher F, Melkes A, Posch J, Winter I, Zarfel G, Marth E (2010) ESBL-producing E. coli in Austrian sewage sludge. Water Res 44:1981–1985
Sacha P, Jakoniuk P, Wieczorek P, Żórawski M (2007) Mechanisms of resistance to β-lactam antibiotics in Escherichia coli, Klebsiella pneumoniae, Proteus mirabilis and Enterobacter cloacae isolates resistant to cefotaxime. J Med Sci 76(4):314–321 (in Polish)
Sader HS, Ferraro MJ, Reller B, Schreckenberger PC, Swenson JM, Jones RN (2007) Reevaluation of clinical and laboratory standards institute disk diffusion breakpoints for tetracyclines for testing Enterobacteriaceae. J Clin Microbiol 45(5):1640–1643
Sáenz Y, Brinas L, Domínguez E, Ruiz J, Zarazaga M, Vila J, Torres C (2004) Mechanisms of resistance in multiple-antibiotic-resistant Escherichia coli strains of human, animal, and food origins. Antimicrob Agents Chemother 48(10):3996–4001
Simarro E, Navarro F, Ruiz J, Miró E, Górmez J, Mirelis B (2000) Salmonella enterica serovar Virchow with CTX-M-like β-lactamase in Spain. J Clin Microbiol 38(12):4676–4678
Turnidge JD (2011) Cefazolin and Enterobacteriaceae: rationale for revised susceptibility testing breakpoints. Clin Infect Dis 52(7):917–924
van den Bogaard AE, Stobberingh EE (2000) Epidemiology of resistance to antibiotics—links between animals and humans. Int J Antimicrob Agents 14:327–335
Wolny-Koładka K, Lenart-Boroń A (2016) Phenotypic and molecular assessment of drug resistance profile and genetic diversity of waterborne Escherichia coli. Water Air Soil Poll 227:146
Zabłotni A, Jaworski A (2014) Sources of antibiotics in natural environments and their biological role. Adv Hyg Exp Med 68:1040–1049
Acknowledgements
This study was funded by statutory measures of the University of Agriculture in Kraków.
Author information
Authors and Affiliations
Corresponding author
Additional information
Editorial responsibility: A. Galuszka.
An erratum to this article is available at https://doi.org/10.1007/s13762-017-1558-x.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Lenart-Boroń, A. Antimicrobial resistance and prevalence of extended-spectrum beta-lactamase genes in Escherichia coli from major rivers in Podhale, southern Poland. Int. J. Environ. Sci. Technol. 14, 241–250 (2017). https://doi.org/10.1007/s13762-016-1155-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13762-016-1155-4