Abstract
Introduction
Bacterial meningitis in infants is an infrequent but life-threatening condition. Empiric therapy should begin as soon as meningitis is thought likely. Consequently, the causative microorganisms may not always be detected using culturing techniques, as cerebrospinal fluid (CSF) cultures are influenced by antibiotics. Nucleic acid amplification tests, such as polymerase chain reaction (PCR) (multiplex panels), may overcome this limitation but require a priori knowledge of the likely pathogen present within the sample. With this in mind, we investigated to what extent a culture-free, broad-range 16S rRNA gene next-generation sequencing (NGS) platform (MYcrobiota) could add to the microbiological diagnosis of meningitis.
Methods
Retrospective cohort study at level III neonatal intensive care unit. Included were all infants with suspected meningitis admitted between 10 November 2017 and 31 December 2020. A comparison was made of the bacterial pathogen detection rate between MYcrobiota and conventional bacterial culture.
Results
In a 3-year period, 37 CSF samples (diagnostic and follow-up) from 35 infants with proven or possible meningitis were available for MYcrobiota testing. MYcrobiota detected the presence of bacterial pathogens in 11 samples (30%), in contrast with the conventional CSF culture, which detected bacteria in 2 of 36 samples (5.6%).
Conclusion
Addition of 16S rRNA sequencing to conventional culturing greatly improved the identification of the aetiology of bacterial meningitis compared to culturing of CSF samples alone.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Why carry out this study? |
Administration of antibiotics prior to lumbar puncture may lead to negative cerebrospinal fluid (CSF) culture results |
PCR-based techniques can detect small amounts of pathogen DNA independently from the growth of the microorganism causing the disease |
The recently developed 16S rRNA gene PCR/NGS platform generates a highly accurate and comprehensive overview of the microbial composition of clinical samples or, alternatively, confirms the absence of bacterial DNA in culture-negative clinical samples |
What was learned from the study? |
The 16S rRNA gene PCR/NGS platform confirms more cases of bacterial meningitis compared to conventional culture methods of CSF |
Culture-free techniques require a more prominent place in regular diagnostics of neonatal meningitis |
Introduction
Bacterial meningitis in infants is a rare but seriously life-threatening condition with high mortality but with sometimes subtle clinical signs at start, which should be treated promptly. Neonatal sepsis is frequently accompanied by meningitis and a lumbar puncture (LP) is often performed in neonates with a suspicion of neonatal sepsis since the cerebrospinal fluid (CSF) culture can show growth when the blood culture remains negative [1, 2]. The presence of increased leukocyte counts or culturing microorganisms from CSF is indicative for the presence of bacterial meningitis. However, cultures of CSF might be negative because of the low amount of CSF obtained at the LP and pretreatment with antibiotics. Although exact numbers are lacking, several studies show a sensitivity of CSF culture < 10% [2, 3]. Diagnosing bacterial meningitis has important implications for the duration, choice, and dosing of antibiotics and for the follow-up and prognosis of the infant.
As alternative to conventional culture, polymerase chain reaction (PCR)-based detection methods have been developed and implemented for routine diagnostics to detect and identify multiple pathogens in a single test with high sensitivity and specificity [4, 5]. Although these kind of syndromic testing panels are comprehensive, no syndromic testing panel is able to cover all possible causative agents of bacterial meningitis. Especially in the case of nosocomial sepsis/meningitis, the spectrum of potential causative bacteria is too large to be covered by target-specific PCR or syndromic testing. To overcome this limitation, the presence of bacterial DNA can be detected and identified using a broad-range PCR strategy that targets the prokaryotic 16S rRNA gene followed by Sanger sequencing, or more recently next-generation sequencing (NGS).
The MYcrobiota platform is a non-commercial consolidated tool that includes a validated, quantitative micelle 16S rRNA gene PCR/NGS methodology [6, 7] and a dedicated bioinformatics pipeline that was specifically designed for use in clinical diagnostic laboratories [8]. This platform enables the detection, quantification, and characterization of bacterial operational taxonomic units (OTUs), a group of very similar 16S rRNA gene sequences (> 97% similarity), present within clinical samples and corresponding negative extraction controls to correct results for inevitable bacterial DNA contamination derived from laboratory reagents and/or the laboratory environment. The MYcrobiota platform possesses a much lower limit of detection compared to conventional 16S rRNA gene sequencing methods and allows detection of bacteria at very low amounts and can confirm the absence of bacterial OTUs in culture-negative samples [6, 8].
In this study, we explored the utility of MYcrobiota for use in the diagnosis of bacterial meningitis by processing a total of 37 CSF samples obtained from infants with proven or possible meningitis and then comparing the results to conventional culture results.
Methods
Setting
This retrospective study was conducted at the Leiden University Medical Center (LUMC). The level III neonatal intensive care unit (NICU) admits approximately 600 infants per year. Parents of infants had the option to opt out of evaluation studies. This study was conducted in accordance with the Helsinki Declaration. This study was approved by the medical ethics review committee of the LUMC (B20.002). Informed consent was waived for this study.
Subjects and Data Collection
All infants admitted at the NICU in whom CSF was collected for diagnosing bacterial meningitis between November 10, 2017, until December 31, 2020, were identified. This period was chosen because of the introduction of a new laboratory information system, which enabled the selection of relevant infants. Relevant patient data were retrieved from the electronic patient data management system. Microbiological data were retrieved from the laboratory information system. Clinical data were collected before the MYcrobiota results were available.
Definitions
Pleocytosis > 15 leukocytes/μl in infants 0–28 days old, > 9 leukocytes/μl in infants 29–60 days old, > 7 leukocytes/μl in infants > 60 days old, not corrected for the presence of erythrocytes [9].
CSF contamination with peripheral blood > 500 erythrocytes/μl of CSF [9].
Positive blood culture Blood culture, collected between 2 days before and 2 days after collection of CSF, showing growth of microorganisms that can cause bacterial meningitis.
Positive CSF culture CSF culture, showing growth of microorganisms that can cause bacterial meningitis.
Potential causative bacterial microorganisms for meningitis Streptococcus agalactiae, Streptococcus pneumoniae, Neisseria meningitidis, Haemophilus influenzae, Enterobacteriales, Staphylococcus aureus, Listeria monocytogenes, Pseudomonas aeruginosa [10].
Proven meningitis CSF culture positive.
Possible meningitis CSF culture negative; pleocytosis present or CSF culture negative; pleocytosis absent but blood culture positive.
Meningitis unlikely CSF culture negative; pleocytosis absent and blood culture negative or not available.
Standard Procedures
If an infant was primarily suspected of bacterial meningitis, a blood culture and CSF were obtained. If no LP had been performed, but a blood culture showed bacterial growth, CSF was collected to rule out meningitis. When an infant was transferred from another hospital, the LP was repeated if indicated by the treating physician. CSF was analysed for cell count, glucose, and protein levels, and a bacterial culture was performed. The decision to treat an infant for bacterial meningitis was made by the treating physician based on a combination of factors such as clinical illness and laboratory and culture results of blood and CSF.
Laboratory Techniques
Routine Diagnostics
The CSF samples received were aliquoted for bacterial culture and molecular diagnostics, and remaining CSF was stored. The amount used for conventional culture was one droplet for each culture plate and enrichment broth. CSF was not centrifuged for bacterial culture because of the small amounts collected. A gram stain was made, and CSF was inoculated at 35 °C for 5 days on trypcase soy agar plates with 5% sheep blood (TSS, bioMérieux, Marcy-l’Etoile, France) under aerobic conditions, on chocolate agar PolyViteX plates (bioMérieux) under 5% CO2 conditions, on TSS under anaerobic conditions, in thioglycolate with resazurine broth (bioMérieux), and in tryptone soya broth with XV factor (MP products, Groningen, The Netherlands). Molecular viral diagnostics (herpes simplex virus 1 and 2, varicella zoster virus, enterovirus, and parechovirus) were performed if requested by the treating physician using real-time PCRs.
MYcrobiota Platform
Additional testing was performed only on remnant CSF samples from patients with proven or possible meningitis that were stored at − 80 °C. Nucleic acids were extracted from 200 μl of CSF samples (replenished with Tris–EDTA buffer if less was available) and eluted in a 100-μl volume using the MagNA Pure 96 DNA and Viral NA small volume Kit (2.0) with the Pathogen Universal 200 protocol on a MagNA Pure 96 instrument (Roche, Mannheim, Germany). MYcrobiota analysis (non-commercial) was then performed as previously described by Boers et al. [8]. Additional BLAST-ing of representative OTU sequences was performed against the NCBI 16S rRNA gene database to classify OTUs at the lowest taxonomic level as possible.
Data Analysis
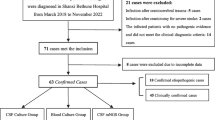
Infants were divided into subgroups, based on the likelihood of meningitis: proven, possible, or meningitis unlikely (see definitions and Fig. 1). Infants with viral or fungal meningitis were excluded. Samples on which conventional meningitis diagnostics had been performed were analysed (initial samples). In cases of proven meningitis, follow-up samples were analysed as well if available. Comparative analysis was performed using R, version 3.6.3 [11]. The analysis was descriptive. Data are presented as median with the boundaries of the interquartile range (IQR), if appropriate. Data were analysed anonymously and in a time frame in which results were not considered relevant anymore to the treatment of the infant; therefore, MYcrobiota testing results were not reported back to the parents.
Results
Patient Demographics
From November 10, 2017, until December 31, 2020, CSF was collected for culture from 162 infants. In 55 infants (34%) viral diagnostics had been performed; none tested positive. One infant was excluded because of Candida albicans meningitis. Five infants (3.1%) had culture-proven meningitis, 38 infants (24%) had culture-negative possible meningitis and in 118 infants (73%) meningitis was unlikely (Fig. 1). In 32 of 43 infants (74%) with proven (n = 2) and possible meningitis (n = 30), CSF was available for MYcrobiota analysis. An additional five follow-up samples from infants with culture-proven meningitis were available.
Of the 32 infants whose initial CSF was available, median age at collection of CSF was 7 days (IQR 3–22) and median time from NICU admission to CSF collection was 39 h (IQR 12–155). None of the infants had a CSF drain at the time of CSF collection. In the 48 h preceding CSF collection, in 31 infants (97%) antibiotics had been administered. Blood cultures from 28 infants (88%) showed growth of a pathogenic microorganism (Table 1). Both infants with proven meningitis and 11 infants with possible meningitis had been treated for bacterial meningitis (41%).
Comparing MYcrobiota Results to Conventional Bacterial Culture in Diagnostic Samples
The MYcrobiota platform detected the presence of bacterial OTUs in 7 of the 32 (22%) diagnostic CSF samples (Fig. 2-1 and Table 1). Four samples tested positive for streptococci OTUs, in concordance with the blood culture results from these infants, from which S. agalactiae was cultured. Two samples tested positive for Enterobacteriaceae spp. OTUs, which was in concordance with the result of the CSF culture growing Citrobacter koseri in one infant and growth of K. pneumoniae in the blood culture from the other infant. One sample tested positive for a Yersiniaceae spp. OTU; however, blood and CSF cultures were negative.
In the two infants with a CSF culture-proven meningitis and CSF available, MYcrobiota could confirm both diagnoses. In total, 13 infants had been treated for bacterial meningitis, in whom MYcrobiota was able to confirm the presence of bacteria in six samples (46%) (Fig. 2-2).
Nineteen infants had not been treated for bacterial meningitis and in 18 infants (95%) the MYcrobiota platform confirmed the absence of bacterial OTUs. However, in one sample low amounts of Yersiniaceae spp. were detected (as described above). The infant was not treated for bacterial meningitis, and antibiotic treatment was stopped after 1 week because of negative cultures and favourable clinical course.
Comparing MYcrobiota Results to Conventional Bacterial Culture in Follow-Up Samples
Five follow-up samples collected from four infants with proven meningitis were available for comparison (Table 2, Fig. 2-3). All were collected when the infants received targeted antibiotic therapy. Four follow-up samples resulted in negative culture results; no culture was performed on the remaining sample. MYcrobiota tested positive for the expected bacterial OTUs in four out of five follow-up samples. The MYcrobiota negative follow-up sample had to be diluted by a factor of 20 before it could be processed because of a low amount of CSF available, which increases the limit of detection.
Discussion
In this study, we investigated 37 CSF samples from 35 infants with suspected bacterial meningitis using MYcrobiota and compared the results to conventional bacterial culture. Our results show the added value of testing CSF samples from infants with suspected meningitis using the MYcrobiota platform as this method results in a higher positivity rate compared to conventional culture (30% vs. 5.6%), and it allows for the detection of microorganisms in CSF samples despite the use of antibiotics prior to sample collection as was also clearly demonstrated in our follow-up cohort. Also, it ruled out meningitis in all children with possible meningitis in accordance with the clinical judgement.
Compared to CSF culture, MYcrobiota confirmed the CSF culture results of the two proven bacterial meningitis and identified potential bacteriological aetiologies of meningitis in five additional cases with CSF culture-negative results. In all but one case, the detected bacterial OTUs by MYcrobiota were in concordance with the bacteria detected with conventional blood and/or CSF cultures. Interestingly, the discrepant sample (repeatedly) tested positive for Yersiniaceae spp. OTU using MYcrobiota, whereas the blood and CSF cultures remained negative. Further determination using nanopore sequencing showed that the sample contained Serratia liquefaciens, which is a very rare causative agent of meningitis [12]. This infant was not treated for bacterial meningitis and the significance of this finding remains unclear as the infant recovered clinically well. Contamination during the process between collecting the sample and DNA isolation cannot be ruled out. False-positive results of MYcrobiota from contamination are uncommon because of the extraction of the results of a negative control sample, which is illustrated by the fact that all sample results of this study were negative or contained a single bacterium species. Furthermore, a high positive predictive value of the MYcrobiota platform has been shown previously [7, 13]. Nevertheless, clinical correlation of positive results is mandatory before cessation or initiation of antibiotic treatment based on test results.
The improved clinical sensitivity of MYcrobiota compared to CSF culture is also confirmed by the higher detection rate (80% vs. 0%) in follow-up samples from culture-proven meningitis despite several days of antibiotic therapy. Again, all detected bacterial OTUs by MYcrobiota were in concordance with the bacteria detected with diagnostic CSF culture results, but none of these bacteria were detected in these follow-up samples using culture due to the administrated antibiotic therapy and despite the need for sample dilution in one sample. Although the clinical sensitivity of MYcrobiota seems high, it remains unclear whether the seven infants treated for meningitis but with negative MYcrobiota results did not have bacterial meningitis or that the MYcrobiota platform gave false-negative results. Larger studies are needed to investigate this but are likely similarly hampered by the small sample volumes that can be taken and stored after culture in infants. In the remaining 18 infants who were not treated, MYcrobiota confirmed the absence of bacterial OTUs, suggesting clinical decision making is safe but may be supported and facilitated by diagnostic tools such as MYcrobiota.
Other culture-free techniques for diagnosing meningitis include targeted PCR assays, multiplex PCR assays, broad range 16S rRNA gene PCR/Sanger sequencing assays, and NGS, with varying sensitivity and specificity [4]. Conventional 16S rRNA gene PCR/Sanger sequencing assays have the same advantage as MYcrobiota of being a broad range strategy that removes the need for an a priori knowledge of the potential bacterial pathogen before a test is performed. However, in contrast to MYcrobiota, these methods rely on relatively high concentrations of bacteria in samples and are therefore unlikely to have adequate sensitivity for exclusion of meningitis [14, 15]. Another alternative for diagnosing meningitis is the use of commercial meningitis multiplex PCR panels, which also have the advantage of having short processing times [5]. For example, the BioFire FilmArray Meningitis/Encephalitis Panel is extensive and includes viral pathogens as well; however, results are limited to the included pathogens. Comparing the pathogens of our study with the pathogens of this panel, K. pneumoniae and C. koseri would have been missed by this commercial multiplex PCR panel.
MYcrobiota has some inherent technical limitations. Like all short-read 16S rRNA gene NGS methods, MYcrobiota currently lacks accurate prokaryotic identification at the species level for most OTUs due to the lack of the discriminative power of the partial 16S rRNA gene used. Implementation of third generation sequencing techniques, such as nanopore sequencing, can overcome this limitation by generating (near) full-length 16S rRNA gene sequences. In addition, nanopore sequencing can collect and analyze sequence data in real time, which can significantly shorten the time to result compared to the current configuration of the MYcrobiota platform based on NGS. Future versions of MYcrobiota should demonstrate whether 16S rRNA gene nanopore sequencing can performed timely, accurately, and cost-effectively in a clinical microbiological diagnostic setting.
Some limitations apply to our study design. First, the sensitivity of the MYcrobiota platform is hampered by the amount of sample available; in this study only remnant samples were used, making dilution a necessity for some samples. In addition, due to the low numbers of infants, and only using samples with sufficient volume in storage, our cohort may have some form of bias, which impeded reliable statistics; therefore, we chose not to perform statistical tests and only used descriptive terms. Furthermore, in samples which were contaminated with peripheral blood, it is unknown whether the microorganisms found were present in CSF or present in blood. Lastly, our definition of pleocytosis is conservative; other studies might use other limits [16]. This may have led to inclusion of children without meningitis in our cohort of possible meningitis and hence to a larger proportion of MYcrobiota-negative cases of possible meningitis.
Conclusions
In conclusion, obtaining accurate quantitative 16S rRNA gene NGS-data using the MYcrobiota platform enables the identification of bacteria present within CSF samples that were not always identified using conventional bacterial culture methods or, alternatively, confirms the absence of 16S rRNA gene copies in culture-negative samples. Therefore, MYcrobiota is a promising tool to aid in the diagnosis of bacterial meningitis, but further studies are needed to evaluate its performance on treatment decisions, cost-effectiveness, and, ultimately, patient outcomes.
References
Visser VE, Hall RT. Lumbar puncture in the evaluation of suspected neonatal sepsis. J Pediatr. 1980;96(6):1063–7.
Garges HP, Moody MA, Cotten CM, Smith PB, Tiffany KF, Lenfestey R, Li JS, Fowler VG Jr, Benjamin DK Jr. Neonatal meningitis: what is the correlation among cerebrospinal fluid cultures, blood cultures, and cerebrospinal fluid parameters? Pediatrics. 2006;117(4):1094–100.
Wang Y, Guo G, Wang H, Yang X, Shao F, Yang C, Gao W, Shao Z, Zhang J, Luo J, Yang Y, Kong F, Zhu B. Comparative study of bacteriological culture and real-time fluorescence quantitative PCR (RT-PCR) and multiplex PCR-based reverse line blot (mPCR/RLB) hybridization assay in the diagnosis of bacterial neonatal meningitis. BMC Pediatr. 2014;14:224.
Pammi M, Flores A, Versalovic J, Leeflang MM. Molecular assays for the diagnosis of sepsis in neonates. Cochrane Database Syst Rev. 2017;2:CD011926.
Leber AL, Everhart K, Balada-Llasat JM, Cullison J, Daly J, Holt S, Lephart P, Salimnia H, Schreckenberger PC, DesJarlais S, Reed SL, Chapin KC, LeBlanc L, Johnson JK, Soliven NL, Carroll KC, Miller JA, Dien Bard J, Mestas J, Bankowski M, Enomoto T, Hemmert AC, Bourzac KM. Multicenter evaluation of BioFire FilmArray meningitis/encephalitis panel for detection of bacteria, viruses, and yeast in cerebrospinal fluid specimens. J Clin Microbiol. 2016;54(9):2251–61.
Boers SA, Hays JP, Jansen R. Micelle PCR reduces chimera formation in 16S rRNA profiling of complex microbial DNA mixtures. Sci Rep. 2015;5:14181.
Boers SA, Hays JP, Jansen R. Novel micelle PCR-based method for accurate, sensitive and quantitative microbiota profiling. Sci Rep. 2017;7:45536.
Boers SA, Hiltemann SD, Stubbs AP, Jansen R, Hays JP. Development and evaluation of a culture-free microbiota profiling platform (MYcrobiota) for clinical diagnostics. Eur J Clin Microbiol Infect Dis. 2018;37(6):1081–9.
Thomson J, Sucharew H, Cruz AT, Nigrovic LE, Freedman SB, Garro AC, Balamuth F, Mistry RD, Arms JL, Ishimine PT, Kulik DM, Neuman MI, Shah SS, H.S.V.S.G. Pediatric Emergency Medicine Collaborative Research Committee. Cerebrospinal fluid reference values for young infants undergoing lumbar puncture. Pediatrics. 2018;141(3).
Ku LC, Boggess KA, Cohen-Wolkowiez M. Bacterial meningitis in infants. Clin Perinatol. 2015;42(1):29–45 (vii–viii).
R Core Team. R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. 2019. https://www.R-project.org/.
Mahlen SD. Serratia infections: from military experiments to current practice. Clin Microbiol Rev. 2011;24(4):755–91.
Boers SA, Reijnen L, Herpers BL, Hays JP, Jansen R. Detection of bacterial DNA in septic arthritis samples using the MYcrobiota platform. J Clin Rheumatol. 2019;25(8):351–3.
Liu CL, Ai HW, Wang WP, Chen L, Hu HB, Ye T, Zhu XH, Wang F, Liao YL, Wang Y, Ou G, Xu L, Sun M, Jian C, Chen ZJ, Li L, Zhang B, Tian L, Wang B, Yan S, Sun ZY. Comparison of 16S rRNA gene PCR and blood culture for diagnosis of neonatal sepsis. Arch Pediatr. 2014;21(2):162–9.
Abelian A, Mund T, Curran MD, Savill SA, Mitra N, Charan C, Ogilvy-Stuart AL, Pelham HRB, Dear PH. Towards accurate exclusion of neonatal bacterial meningitis: a feasibility study of a novel 16S rDNA PCR assay. BMC Infect Dis. 2020;20(1):441.
Chapter 6 bacterial meningitis. In: Sharland M, editor. Manual of childhood infections: the blue book. Oxford University Press. 2016.
Acknowledgements
The authors would like to thank Kawtar Benchamach and Margriet Kraakman, analysts at the Department of Medical Microbiology at the LUMC, for their technical and laboratorial assistance. They did not receive funding for their assistance. In addition, we are grateful to all children and their parents for being able to use the data for these analyses.
Funding
The authors have not declared a specific grant for this research from any funding agency in the public, commercial, or not-for-profit sectors. The rapid service fee for publication in this journal was waived.
Author Contributions
Alieke van der Hoeven conceptualized and designed the study, collected data, carried out the initial analyses, drafted the initial manuscript, and critically reviewed and revised the manuscript. Martha van der Beek conceptualized and designed the study, drafted the initial manuscript, and critically reviewed and revised the manuscript. Vincent Bekker conceptualized and designed the study, and critically reviewed and revised the manuscript. Erin Meijers and Marco Ivens carried out technical and laboratorial analyses and critically reviewed and revised the manuscript. Els Wessels and Aloysius Kroes critically reviewed and revised the manuscript for important intellectual content. Stefan Boers conceptualized and designed the study and critically reviewed and revised the manuscript for important intellectual content.
Disclosures
The Alieke van der Hoeven, Martha T. van der Beek, Vincent Bekker, Erin Meijers, Marco J.R. Ivens, Els Wessels, Aloysius C.M. Kroes and Stefan A. Boers have no relevant financial or non-financial interests to disclose. Alieke van der Hoeven changed affiliation after completion of the manuscript and is now employed by Atalmedial.
Compliance with Ethics Guidelines
This study was conducted in accordance with the Helsinki Declaration. This study was approved by the medical ethics review committee of the LUMC (B20.002). Informed consent was waived for this study.
Data Availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
van der Hoeven, A., van der Beek, M.T., Bekker, V. et al. Improved Diagnostics in Bacterial Neonatal Meningitis Using a Next-Generation Sequencing Platform. Infect Dis Ther 12, 1921–1933 (2023). https://doi.org/10.1007/s40121-023-00844-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40121-023-00844-8