Abstract
Purpose of Review
Our goal is to show how readily available Pathomics tissue analytics can be used to study tumor immune interactions in cancer. We provide a brief overview of how Pathomics complements traditional histopathologic examination of cancer tissue samples. We highlight a novel Pathomics application, Tumor-TILs, that quantitatively measures and generates maps of tumor infiltrating lymphocytes in breast, pancreatic, and lung cancer by leveraging deep learning computer vision applications to perform automated analyses of whole slide images.
Recent Findings
Tumor-TIL maps have been generated to analyze WSIs from thousands of cases of breast, pancreatic, and lung cancer. We report the availability of these tools in an effort to promote collaborative research and motivate future development of ensemble Pathomics applications to discover novel biomarkers and perform a wide range of correlative clinicopathologic research in cancer immunopathology and beyond.
Summary
Tumor immune interactions in cancer are a fascinating aspect of cancer pathobiology with particular significance due to the emergence of immunotherapy. We present simple yet powerful specialized Pathomics methods that serve as powerful clinical research tools and potential standalone clinical screening tests to predict clinical outcomes and treatment responses for precision medicine applications in immunotherapy.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Human diseases are characterized by correlating host symptoms with clinical examination, radiology, and laboratory testing. The microscopic examination of tissue samples from biopsies and surgical procedures is routinely performed to diagnose diseases based on tissue and cellular pathobiology. This fascinating microscopic world of human biology and disease has been mostly restricted to pathologists and clinical researchers until the introduction of digital pathology. Digital pathology has opened the gates to a significantly larger and more diverse audience through high-resolution whole slide images (WSIs) that support the exploration of complex virtual landscapes of human tissues from every part of the body during normal development and disease. In turn, digital pathology has benefited from extraordinary advances in technology, software, image analysis, machine learning, and computer vision that have led to the development of powerful computational image analysis methods, collectively referred to as Pathomics.
Pathomics has the potential to transform our understanding of human biology and disease by providing numerous opportunities to perform quantitative analysis of tissues and cells. For example, Pathomics with classical image analysis can characterize tissue microarchitecture, microanatomic structures, different types of cells, and other histologic features of disease by analyzing phenotypic traits like color, size, shape, texture, and orientation. In addition, a variety of novel computational methods for digital pathology have recently emerged due to recent advances in artificial intelligence (AI) and computer vision. Currently, Pathomics tissue analytics has reached a level of critical mass that is capable of supporting large interdisciplinary collaborative research efforts to study human pathobiology across clinical medicine, academia, and industry.
This report focuses on Pathomics applications that harvest and analyze data from WSIs to study tumor immune interactions to help advance our understanding of cancer pathobiology. We describe a specialized Pathomics application, Tumor-TILs, to illustrate how complex relationships and patterns in cancer immunopathology are easily explored in hematoxylin and eosin (H&E) WSIs of breast, pancreatic, and lung cancer. We narrowly focused on this specialized form of Pathomics tissue analytics since these Tumor-TILs methods are readily available, open source, and easily implemented to study tumor-infiltrating lymphocytes (TILs) in these types of virtual cancer tissue samples. Moreover, these methods perform automated analyses in a scalable fashion in preparation for when WSIs of cancer tissue samples become much more widely available. We will (1) briefly describe the much larger world of Pathomics within the context of traditional histopathologic properties of tumors in order to present (2) recently developed Tumor-TILs Pathomics applications to characterize TILs in breast, pancreatic, and lung cancer and (3) propose more advanced ensemble Pathomics analytic pipelines to collect and integrate data from multiple scales of magnification to further improve our understanding about cancer immunopathology.
Our primary goal is to demonstrate how automated Pathomics tissue analytics can be realistically utilized to quantitatively study tumor immune interactions in a systematic manner in an effort to emphasize the potential value of Pathomics in precision medicine. Related to this goal, we want to show how Pathomics provides an opportunity to perform analyses in large studies, since there is no sustainable way to manually evaluate thousands to tens of thousands of tissue samples , even if the costs, required time, resources, and access to technical skills and expertise were irrelevant. We present Tumor-TILs analyses to provide an example of how WSIs from large studies can be readily analyzed to provide immediate insight into cancer immunopathology to support the clinical use of Pathomics applications for precision medicine and immunotherapy. Our secondary goal is to provide a vision about integrating clinical informatics, radiomics, pharmacogenomics, and Pathomics tissue analytics with patient outcomes and treatment response data in order to improve the stratification of patient management, treatment selection, and survival.
Pathomics Applications in Cancer Histopathology
Pathology is the study of diseases by examining tissue samples, where microscopic abnormalities in the structures of tissues and cells are associated with diagnostic criteria to render histopathologic diagnoses. The implementation of commercial glass scanners in clinical and research pathology laboratories has led to the increased availability of H&E WSIs and greater access to cancer tissue samples through virtual microscopy, leading to the development of powerful computational methods to support the establishment of precision medicine in the twenty-first century (see recent reviews in refs [1••, 2•, 3•]). Digital pathology gave birth to Pathomics to harvest and quantitatively analyze exquisitely complex biological data with classical image analysis and computer vision applications.
Pathomics tissue analytics applications have matured over the last several years, where computational pathology tools are now readily available to make transformative impacts in biomedical research, clinical trials, and the practice of medicine. Pathomics applications have been implemented to collect, analyze, and quantitatively measure a wide variety of histopathologic features in human diseases like cancer (see refs [1••, 2•, 3•, 4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21] for motivations and methods). A lot of excitement surrounds the development of Pathomics tools for clinical decision support (CDS) systems to help improve our ability to (1) define and delineate human diseases with quantitative metrics to describe salient histopathologic features, (2) predict outcomes by identifying patterns and relationships to further elucidate our understanding about the biological behavior of malignant and non-malignant diseases, and (3) select optimal treatment strategies by using Pathomics to stratify patients and predict treatment response. For example, Pathomics may very useful to study challenging cases of cancer by quantitatively measuring size, shape, texture, and other features to help provide better clinical insight, especially for cases with non-specific and overlapping histologic features that are regularly encountered and easily misinterpreted in daily practice despite considerable expertise and experience.
Substantial advancements in AI through deep learning and computer vision have been particularly important to the development of highly flexible Pathomics applications to study cancer with automated feature extraction. AI applications in Pathomics have overcome various obstacles associated with developing engineered and hand-crafted features based on classical image analysis, which rely on defining various types of parameters associated with color, size, shape, contrast, and texture to characterize complex histopathologic alterations in microanatomic tissue architecture associated with cancer and other human diseases [22••, 23, 24]. Emerging methods for Pathomics tissue analytics in cancer tissues samples with deep learning and computer vision appear to complement traditional histopathologic evaluation in ways that are fundamentally similar to how pathologists utilize pattern recognition and the association of features to characterize diseases. Pathomics tissue analytics is also leveraging concurrent advances in computational processing power, software, and infrastructure in order to deploy automated pipelines to systematically study cancer pathobiology in virtual tissue samples in a practical, robust, and reliable manner for large-scale interdisciplinary clinical research.
Currently available types of Pathomics analyses can be categorized in a manner that parallels multiscale examination, where various regions of interests (ROIs), specific types of objects, and other microscopic properties of tissues are identified, measured, and harvested to identify potential biomarkers and perform downstream correlative analyses. Since digital pathology was originally established to support the virtual examination of histologic tissue samples with software to supplant microscopes, we summarize how Pathomics tissue analytics can be utilized to complement histopathologic evaluation at multiple scales of magnification, as shown in Table 1. Table 1 shows how Pathomics is useful to detect, label, and classify microanatomic regions of tissue, microarchitectural structures, cells, and nuclei. For example, Pathomics data can (1) identify meaningful regions of interest ROIs in WSIs by classifying areas with different patterns of growth and increased mitotic activity [25], (2) measure histologic differentiation by quantifying the distributions of different types of cells in histologically distinct architectural configurations, and/or (3) define cytomorphologic features of cells and nuclei in terms of color, size, geometry, chromatin density, texture, and proliferative activity [12, 13]. Automated Pathomics have also been utilized to systematically study the ecosystems and microenvironments of many different types of cancer through higher-order spatial analyses to characterize human pathobiology (described in recent digital pathology white papers [26, 27] and related refs [1••, 2•, 3•, 28,29,30,31,32,33,34,35,36,37,38,39, 40•, 41]).
Tumor Immune Interactions with Pathomics
Current advances in Pathomics tissue analytics are focused on predicting the biological behavior of human disease and cancer for precision medicine applications. One of the main challenges in Pathomics is to identify the types of data products that can be useful to identify patterns, relationships, or biomarkers to provide clinically actionable information [42]. Current Pathomics applications are incredibly powerful and useful for understanding how and why the dynamic microscopic landscapes of tissues change during the initiation and progression of disease. In order for Pathomics to be clinically useful, quantitative measurements of categorical observations of various phenotypic features of biological phenomena must support the development of computational models that predict changes associated with treatment and outcomes. When Pathomics tissue analytics become more routinely available, advanced understanding of human pathobiology will result from fully integrating Pathomics with clinical data from electronic health records, imaging, and clinical laboratory testing.
Each tissue sample from every patient is unique. Even though clinical history and other countless sources of variability exist in human biology, microscopic evaluation is used to identify morphologic nuances in tissues and cells during the course and progression of various human diseases. Pathomics tissue analytics offers an exciting and useful opportunity to augment traditional histopathologic evaluation on a per tissue sample basis to coincide with the clinical adoption of precision medicine. Even though tumor immune interactions in breast, pancreatic, and lung cancer have been carefully studied and actively explored by pathologists and biologists for many decades, we narrowly focus on the role of Pathomics to capture and quantify tumor immune interactions in H&E WSIs of these kinds of cancer tissue samples as practical examples that can meaningfully impact the role of precision medicine in immunotherapy.
We show straightforward examples of recently developed first-generation specialized deep learning Pathomics applications, referred to as Tumor-TILs analyses, that construct detailed spatial maps of tumor immune interactions in WSIs. Tumor-TIL maps are generated by combining automated detection of cancer regions with automated detection of lymphocytes by using deep learning algorithms in computer vision (see refs [43•, 44, 45] for technical descriptions). Briefly, Tumor-TILs analyses partition WSIs into tiled patches that are analyzed with two distinct deep learning computer vision algorithms, where image patches are designated as either (1) tumor or non-tumor or (2) lymphocytes or no lymphocytes and the remaining patches are considered as background (non-tumor and non-lymphocyte). The algorithmic predictions from each deep learning computer vision model are spatially mapped as probabilities with continuous values from 0 to 1. After evaluating the algorithmic performance of each model individually, the results from each model are combined to generate Tumor-TIL maps that can be used to explore cancer immunopathology.
Tumor-TIL maps represent massive amounts of computational analytics and Pathomics data in a very simple manner in order to help facilitate rapid understanding about cancer immunopathology. The spatial distribution and relative magnitude of lymphocytes are shown within the appropriate histologic context of the tumor microenvironment (TME) by overlaying deep learning algorithmic results on WSIs alongside corresponding H&E WSIs in order to complement traditional microscopic examination. In this manner, Tumor-TILs maps readily complement the routine visual assessment of tissue samples by pathologists with built-in quality assurance and control (QA/QC) for each step of the analyses. This is particularly important if deep learning Pathomics analyses are going to be used to predict treatment response and clinical outcomes for patient care. Since current Pathomics tools can clearly provide interpretable and clinically meaningful data from automated deep learning analyses of WSIs, we want to focus on how these tools can be used to study the functional immunologic status of the TME in WSIs of breast, pancreatic, and lung cancer.
TILs as a Pathomics Biomarker
Tumor-TILs analyses help identify and quantify the role of TILs in cancer. TILs represent an intrinsic prognostic biomarker in H&E tissue samples that can be readily evaluated in a wide variety of solid tumors with the potential to refine diagnostic characterization, patient stratification, and treatment selection. Marrying Pathomics with quantifying an intrinsic H&E biomarker in WSIs during the age of precision medicine and immunotherapy is an opportunity that cannot be missed since Tumor-TIL maps provide a simple and easily interpretable global view tumor immune interactions in the TME. Moreover, Tumor-TILs represents the capacity to perform automated, systematic, and uniform analyses in a scalable manner that can be compared to utilizing immunohistochemistry (IHC) to characterize the expression of biomarkers in tumors. Pathologists and biologists profile immune subtypes of cancer with a variety of methods in anatomic pathology, where inflammatory responses are characterized in terms of cellular components, spatial distribution, and relative magnitude. Pathologists also use terminology like “hot” and “cold” to describe the overall functional immune status of the TME in cancer tissue samples.
Inflammatory responses are typically evaluated with diverse terminology to describe immunohistopathology in tissue samples (e.g., inflamed, reactive, and necrotic), cellular components (e.g., “acute” for neutrophils, “chronic” for lymphoplasmacytic cells, and “histiocytic” and “granulomatous” for macrophages), magnitude (e.g., minimal, mild, moderate, and severe), and distribution (e.g., sparse, focal, and diffuse). Tumor-TILs analyses were developed to augment qualitative and semi-quantitative assessment by addressing the tangible demand to develop a quantitative method to capture and evaluate tumor immune interactions in solid tumors with standardized, cost-effective, and scalable methodology (see relevant literature and reviews from the International Immuno-Oncology Biomarker Working Group for TIL assessment guidelines in breast cancer and other solid tumors [22••, 43•, 44,45,46,47,48,49,50,51]).
Since the prognostic and predictive value of TILs as a biomarker in breast cancer is slowly gaining acceptance as a reportable biomarker that needs to be evaluated during routine histopathologic evaluation, Tumor-TILs may be very practical and useful example of Pathomics to help pathologists score TILs and further evaluate cancer pathobiology (in a manner that is similar to scoring and evaluating the expression of IHC biomarkers in different histologic types of cancer). However, the implementation of TILs as a routine and clinically important H&E biomarker has yet to be adopted, similar to the way that Pathomics have yet to be clinically implemented to quantitatively characterize aforementioned histopathologic features of tumors. Therefore, we present Fig. 1 to provide examples of how Tumor-TILs analyses in breast cancer can be immediately helpful to pathologists and provide useful insight about the functional immune status in breast cancer cases. Figure 1a–d and e–h represent two separate cases of breast cancer, where the panels are assembled in a manner to help ascertain the potential clinical value of TILs as a biomarker. This example also demonstrates the value of automated methods like Tumor-TILs as a practical and useful form of Pathomics to study and evaluate clinically meaningful tumor immune interactions in cancer by characterizing the spatial distribution of TILs in WSIs.
Tumor-TILs analyses in breast cancer. Deep learning Pathomics pipelines perform Tumor-TILs analyses in breast cancer, where panels a–d show analyses for specimen BRCA TCGA-AR-A24U-01Z-00-DX1 and panels e–h for specimen BRCA TCGA-AR-A250-01Z-00-DX1. Tumor-TIL maps depicted as follows: top left panels (a, e) show H&E WSIs at low magnification; top right panels (b, f) show automated tumor detection presented as a spatial probability distribution from 0 to 1 to evaluate algorithmic performance (non-tumor tissue colored blue); bottom left panels (c, g) show automated lymphocyte detection presented as a spatial probability distribution from 0 to 1 (non-lymphocyte tissue colored blue); and bottom left panels (d, h) show Tumor-TILs by combining tumor and lymphocyte detection (tumor colored yellow, lymphocytes colored red, and background non-tumor/non-lymphocyte tissue colored gray). Panels d and h show the spatial distribution of lymphocytes within the histologic context of cancer in order to identify peritumoral and intratumoral TILs and facilitate qualitative and quantitative interpretation of tumor immune interactions (to perform TILs scoring in the overall tumor region and spatially distinct areas). In the first case (d), spatial Tumor-TILs analyses show an abundance of peritumoral and intratumoral tumor immune interactions, whereas the second case (h) shows predominantly peritumoral TILs in breast cancer. The peritumoral distributions of TILs in (h) suggest that the tumor may be exerting immunosuppressive effects on TILs , which can help guide the selection of checkpoint inhibitor immunotherapy to activate the cytotoxicity of these lymphocytes. Tumor-TILs analyses have been performed in large collections of WSIs to evaluate both algorithmic performance and functional immune responses to support precision medicine in cancer
The 4-way panels show spatially distributed tumor immune interactions in WSIs of cancer tissue samples. These examples of Tumor-TIL maps present a global perspective of tumor immune interactions at low magnification. We can also overlay high resolution heat maps on H&E WSIs of cancer tissue sample to evaluate tumor and lymphocyte detection in order to perform quality assurance and control (QA/QC) of algorithmic performance. These examples combine a low magnification view of a H&E WSIs with separate panels showing overlaid probabilities of tumor and lymphocyte detection that are subsequently combined to generate Tumor-TIL maps. Performing Tumor-TILs analyses in large datasets of WSIs creates the ability to view the exquisite variability in the magnitude and spatial distribution of immune responses in the TME to support clinical research about TILs per the recommendations from the International Immuno-Oncology Biomarker Working Group [22••, 46, 47, 52,53,54,55]. Figure 1 also demonstrates how straightforward Pathomics analyses like tumor and lymphocyte detection in WSIs fundamentally complements traditional histopathologic examination to reliably characterize the functional immunologic state of the TME. The simplicity of this approach with incorporated QA/QC allows both pathologists and non-pathologists alike to seriously consider the value and potential of Pathomics tissue analytics to enhance visual examination and further our collective understanding of human pathobiology.
Tumor-TILs analyses augment the ability of pathologists to observe immune and non-immune mediated cancer pathobiology in a way that is not humanly possible across tens of thousands of samples. By predicting which patches of tissue contain tumor, lymphocytes, or both, we are effectively observing the ground cover of TILs in these types of cancer in an effort to accurately describe the percent of a cancer tissue sample that contains peritumoral and intratumoral TILs, lymphoid aggregates, and non-tumor associated lymphocytes. Tumor-TILs analyses have been implemented to measure the variability of these aspects of cancer immunopathology in publicly available datasets like the Cancer Genome Atlas (TCGA), as well as WSI datasets from other institutions in collaborative research projects. Automated Tumor-TILs analyses are also being expanded to catalog tumor immune interactions in several other types of cancer.
The overall goal of automated Tumor-TILs analyses is to introduce how clinical research and hospital laboratories can realistically use Pathomics to analyze H&E WSIs of cancer tissue samples from every patient. Automated Pathomics tissue analytic pipelines like Tumor-TILs show how massive amounts of data can be harvested from WSIs to generate clinically useful data products. If we can reliably correlate Tumor-TILs analyses with clinical outcomes, treatment responses, and survival, clinicians may gain the ability to predict prognosis, stratify patients, and identify candidates for immunotherapy and combinations with classical chemotherapy regimens and other treatment modalities.
Due to the growing acceptance of the role of host immunity in cancer and immunotherapy, systematically mapping tumor immune interactions with readily available Pathomics methods appear very necessary to comprehensively evaluate this aspect of tumor pathobiology. Similarly, the incorporation of TILs into the conventional classification of cancer also appears very likely, so methods like Tumor-TILs that can be readily incorporated will be hard to ignore when digital pathology becomes more widely adopted by clinical laboratories. Tumor-TIL maps are already being used to guide the microscopic examination of tissue samples on glass slides and/or WSIs to examine salient tumor immune interactions, but we need to explore how to use these analyses to develop more advanced types of Pathomics tissue analytics to investigate multidimensional relationships between other Pathomics features and histologic growth patterns, cytonuclear features of cancer cells, cancer staging, disease progression, metastasis, recurrence, treatment response, and survival.
In summary for this section, Tumor-TILs can be readily implemented to support various kinds of research to study cancer initiation, development, dissemination, and treatment response in the neoadjuvant and adjuvant setting by harvesting and cataloging massive amounts of important clinical research data. Tumor-TILs also provides numerous opportunities to explore and study the relationships between cancer immunopathology within the context of histologic growth patterns, cellular differentiation, cytonuclear features, cytogenetics, gene expression, and other types of omics to characterize the molecular profiles of various kinds of cancer. Since Tumor-TILs analyses advocate a solid starting point to promote capturing additional immune-related histopathologic parameters to better understand cancer pathobiology [22••], we present Fig. 2 as a proof of concept to demonstrate how Pathomics appears quite useful for studying tumor immune interactions in a highly complex disease like pancreatic cancer.
Tumor-TILs analyses in pancreatic cancer. Deep learning Pathomics pipelines perform Tumor-TILs analyses in pancreatic cancer ductal adenocarcinoma (PDAC), where panels a–d show analyses for specimen PAAD TCGA-IB-A5SS-01Z-00-DX1 and panels e–h for specimen PAAD TCGA-IB-7890-01Z-00-DX1 (Tumor-TIL maps are depicted in the same manner as Fig. 1). In both cases (panels a, h), Tumor-TILs analyses show predominantly peritumoral tumor immune interactions in pancreatic cancer with subtle differences in the magnitude of the responses. Beyond providing the ability to qualitatively score the relative strength of the immune responses, these maps can guide pathologists to areas of interest in glass slides and WSIs to further review other histopathologic features, such as using the abundance of tertiary lymphoid aggregates in the periphery beyond the invasive boundary of the tumor to guide the search for lymphovascular invasion (LVI). Tumor-TIL maps may also provide clues about tumor heterogeneity in terms of differential tumor immune interactions in the main tumor mass that can be compared to the functional role of TILs with infiltrative clusters of tumor cells. Tumor-TIL maps can also serve as a screening tool to help pathologists consider a particular diagnosis since these algorithms are trained with numerous PDAC samples to perform tumor detection
Practical Translational Research Opportunities With Tumor-TILs
Tumor-TILs have been presented as a proof of concept to demonstrate the power of Pathomics to capture and analyze complex data from pathology images in a uniform manner. We hope that we have shown how these readily available methods can be deployed to analyze hundreds to thousands of WSIs to investigate the role of TILs in cancer immunopathology so far. We also hope to have demonstrated how Pathomics can quantitatively score the number of TILs and characterize their spatial distribution within the appropriate histologic context of cancer, which can be incredibly powerful for cohort discovery and real-time tissue analytics in surgical pathology. Beyond these very important immediate uses, we also want to show how the automated reporting of an overall score of the percent of TILs in a tumor region with a description of their spatial distribution can transform the identification and stratification of patients who might benefit from readily available immunotherapeutic options. To that end, we hope that the selected figures have substantiated the concept of computationally characterizing and scoring TILs as a biomarker with Pathomics, which can be readily utilized as simplistic standalone screening tests to identify hot and cold immune subtypes of cancer and other types of nuances in immunopathologic responses on a case by case basis.
In this respect, Tumor-TIL maps were developed as a visual tool and prototype for a potential Pathomics TILs screening test to help researchers and pathologists quickly evaluate the functional immune status of tumors in order to further guide the evaluation of TILs at higher magnification in relevant ROIs (e.g., TILs with respect to variable morphologic growth patterns and cytonuclear features associated with worse clinical outcomes and poor survival). Therefore, we discuss how Tumor-TILs can be leveraged to drive the development of more powerful ensemble Pathomics by incorporating computational image analysis methodology in this section [12, 13, 56,57,58]. For example, ensemble models can support scientific inquiry by using Tumor-TILs in conjunction with engineered and hand-crafted features to identify particular tumor niches for further laser capture microdissection and molecular testing. Beyond the spatial distributions of TILs, the TILs themselves can be further characterized by size, shape, color, texture, and chromatin density to guide the development of multiplex IHC panels and molecular tests to identify different subsets of immune cells and the expression of immunotherapy drug targets in a non-destructive and cost-effective manner. Other interesting applications include downstream analyses that leverage Tumor-TILs Pathomics data with nearest neighbor distances, clustering, topography, and fractal occupancy analyses to determine the prognostic and predictive significance of the spatial distributions of TILs [22••, 45, 50, 59,60,61,62,63,64,65,66].
As mentioned previously, we can report Tumor-TILs analyses in manner that parallels how the expression of IHC biomarkers in a histologic tissue sections are routinely reported. Similarly, we can envision how Tumor-TILs may be used to suggest further reflex testing during diagnostic workup while being integrated with laboratory data from chemistry, hematology, flow cytometry, human leukocyte antigen (HLA), microbiology, molecular laboratory testing, and clinicopathologic patient data in order to stratify patients and formulate treatment strategies based on computational phenotype. Potential scenarios could involve using first-generation Tumor-TILs to help overcome the problems associated with PD-1/PD-L1 testing assays for immune checkpoint inhibitors [64•, 67, 68] by defining salient tumor immune niches for PD-1/PD-L1 scoring and/or reporting PD-1/PD-L1 IHC scores in conjunction with Pathomics TILs scores integrated with electronic healthcare data to devise an appropriate treatment strategy. Relatively straightforward applications of readily available Pathomics applications like Tumor-TILs can be easily implemented to perform an additional level of time-, cost-, and resource-efficient data collection and analyses in daily practice, clinical trials, and translational clinical research. In the current cycle of development, first-generation Tumor-TILs Pathomics are being used as value added tools in a wide range of clinical research studies to explore and discover various diagnostic, prognostic, and predictive biomarkers.
Up to this point, we presented Tumor-TILs within a narrow focus to help convey realistic avenues for research and clinical adoption with readily available first-generation examples of Pathomics tissue analytics. However, guiding the actual design and selection of immunotherapeutic regimens on a per patient basis with a computational Pathomics marker is a challenging and complex matter when we consider how there are more than 100 different immunophenotypes in human blood [60, 69,70,71,72]. This clinical scenario gets even more complicated by a wide level of variability across individuals during functional physiologic states related to normal homeostasis and clinically detectable adaptations to disease. Thus, it is unsurprising that immunopathology is just as variable in the TME of different types and subtypes of cancer [46, 47, 52, 53, 73,74,75] and a partial explanation of why immune checkpoint treatments have improved overall survival in a subset of patients when compared to standard therapy, which limits our ability to determine the patients who will actually benefit from specific types of immunotherapy.
In this scenario, Tumor-TILs may be useful as first-order analyses that capture the diversity and heterogeneity of immune responses in a scalable and cost-effective manner in WSIs. Tumor-TIL maps can be readily utilized as base layers and incorporated in more advanced Pathomics tissue analytics to help further explore and define the complex relationships between cancer and immune cells in the TME. As stated in the end of the previous section, several types and directions of scientific investigation can be performed by correlating these analyses with a wide spectrum of clinicopathologic, molecular, radiologic, and treatment data to develop more advanced Pathomics applications for precision medicine. As increasingly large and complex datasets from clinical trials become available, Pathomics biomarkers may become as important as specific histopathologic features of cancer and IHC biomarkers.
Ensemble Pathomics for Advanced Tumor-TILs Analyses
There has never been an opportunity to study tumor immune interactions by using automated Pathomics image analysis methods to interpret the role of complex immune responses in cancer without employing other laboratory methods like IHC, flow cytometry, genomic sequencing, and gene expression. In this section, we envision practical ensemble Pathomics methods to further support and potentially accelerate scientific discovery and clinical translation by leveraging Tumor-TILs analyses in WSIs. We envision how ensemble Pathomics tissue analytics can conceivably parallel the impact of IHC and molecular subtyping that fundamentally transformed the classification and subclassification of tumors in anatomic pathology.
We believe that advanced Pathomics applications can be feasibly developed and implemented to further unravel the role of TILs along a trajectory that mirrors how IHC biomarkers that were once rigorously tested for use during diagnostic classification are being actively revisited to explore prognostic value and utility for serving as therapeutic drug targets. Since current versions of Tumor-TILs analyses and mapping provide a global view of tumor immune interactions, we present recently developed prototypes that are geared towards improving the characterization of tumor immune interactions in WSIs of cancer tissue samples. Beyond the examples that we present to study tumor immune interactions and TILs, first-generation Tumor-TILs models are being integrated with classical image analysis to also calculate tumor area, perimeter, greatest dimension, and irregularity of the invasive boundary in order to interpret TILs in various types of histologic contexts. We can use these analyses to determine whether the spatial distribution of lymphocytes is intramural, peritumoral, organized as tertiary lymphoid aggregates, loosely scattered, focal, or scant in WSIs to complement how pathologists evaluate and report the assessment of inflammatory responses and TILs with descriptions of spatial location, strength, and tumor immune interactions in relevant diagnostic ROIs (e.g., central tumor, invasive edge, diffusively infiltrative clusters, and tumor deposits).
Therefore, we present Fig. 3 as a prototype that combines Tumor-TILs analyses with mapping microarchitectural growth patterns of adenocarcinoma in lung cancer to study tumor immune interactions within the context of histologic subtype. Using Fig. 3, we can study the spatial distribution of TILs with respect to the spatial distribution of growth patterns of lung adenocarcinoma to quantitatively measure the variability of tumor immune interactions in a more refined histologic context of tumor landscape. Even though this tool needs further refinement and QA/QC, our intention is to show how automated Tumor-TILs plus the classification of histologic subtype can feasibly support current and future clinical trials in lung cancer with immunotherapy by identifying and quantifying treatment responses with checkpoint inhibitor therapy. Figure 3 also demonstrates a proof of concept of a scalable Pathomics method can be enhanced to potentially discover novel biomarkers with ensemble Pathomics for precision medicine applications, immunotherapy, and beyond.
Tumor-TILs analyses combined with histologic subclassification in lung cancer. Working prototype of a deep learning Pathomics pipeline to perform Tumor-TILs analyses and histologic subclassification in lung adenocarcinoma, where Panels a–d show analyses for specimen LUAD TCGA-69-8255-01Z-00-DX1 and panels e–h for specimen LUAD TCGA-69-A59K-01Z-00-DX1 (Tumor-TIL maps are depicted in the same manner as Fig. 1). In both cases (panels a, h), spatial Tumor-TILs analyses show a robust presence of intratumoral TILs in lung cancer but show differences in the presence of peritumoral lymphocytes due to other histopathologic features. The histologic subclassification of these two specimens is primarily lung adenocarcinoma acinar type, where a shows the presence of necrosis associated with very strong lymphocytic responses in comparison to the absence of necrosis in e. Tumor-TILs analyses in d show small nests of viable tumor adjacent to the area of necrosis as well as lymphoid aggregates that can presumably send reinforcements. This image demonstrates the value of pursuing further investigation into characterizing tumor immune interactions and scoring the exquisite variability of TILs with respect to growth pattern and tumor differentiation
Current deep learning models analyze tiled patches due to limited computational resources. However, access to supercomputing and cloud computing resources has led to new opportunities to leverage the advances in digital pathology from the past two decades. Beyond the opportunities to ensemble and automate existing Pathomics methods to extract, quantitatively characterize, and catalog various properties of tissues in vast collections of WSIs, previous computational limitations are being surpassed to analyze and map the features of every single cell in WSIs at significantly higher resolution. Tumor-TILs and other types of ensembled Pathomics can analyze hundreds of thousands to millions of cells and objects in WSIs without a conceivable limit as technology advances even further. We envision using advanced ensembled Pathomics to compute tumor cellularity based on calculations of the number of segmented nuclei in a tumor region; quantitatively characterize tumor differentiation by computing the percent of cells in specific types of architectural configurations; and generate high resolution maps based on the classification of all of the different types of cells in human tissues with accompanying engineered and hand-crafted features that measure color, size, shape, texture, orientation, and chromatin density.
We also propose incorporating Tumor-TILs into advanced ensemble Pathomics methods to capture as many salient histopathologic features as possible for every WSI at every academic institution, healthcare system, and tumor registry. We imagine being able to utilize Pathomics pipelines to analyze WSIs with corresponding data from tumor registries to study tumor heterogeneity, metastatic cancer, therapeutic efficacy, and drug resistance in order to develop models to help select treatment based on predicted survival [22••, 50, 61, 64•, 76, 77]. However, laying the foundation for the deployment of Pathomics for clinical decision support systems will still require a fair amount of time considering that the Food and Drug Administration (FDA) recently aproved the clinical use of a commercial slide scanner and digital pathology system just a few years ago [30, 31, 36]. Before Pathomics data can be used to provide meaningful clinical information to physicians and patients to help predict patient survival, likelihood of disease recurrence, and progression in the very near future, Pathomics methodology that are still early in the development cycle have to be extensively tested and validated in appropriate clinical settings. As shown in Figs. 1, 2, and 3, current Tumor-TILs Pathomics methods extract and present information that cannot be performed at scale by pathologists using microscopes, IHC, sequencing, and gene expression to characterize tumor immune interactions in every tissue sample.
However, currently accepted methods to evaluate tumor immune interactions in tissue samples focus on the manual assessment of stromal TILs (sTILs), which have been shown to be important prognostic and predictive biomarkers in triple-negative (TNBC) and HER2-positive breast cancer. One of the main reasons that sTILs have become so important is due to consistent reproducibility of scoring by pathologists with the use of standardized guidelines developed by the clinical research community in three large studies [40•, 50, 64•, 65, 78]. Despite the acceptance of sTILs as prognostic and predictive biomarkers, the assessment of sTILs is challenging within the complex landscape of cancer, which results in scoring discordance due to the heterogeneity of the distribution of lymphocytes. Discrepancies also arise from the lack of consensus about the precise boundary of tumors, variable presence of tumor-associated stroma, associations of lymphocytes with other microanatomic structures, presence of other kinds of immune cells [50, 65]. Even though these kinds of variations during the evaluation of sTILs have mitigable effects in estimating risk in early TNBC, multiple areas are scored and averaged during the evaluation of sTILs to improve consistency and minimize the effects of observer variability, scoring discrepancies, and cutoffs that could affect treatment selection [50, 65].
Due to the specific interest in sTILs, we wanted to demonstrate the ability to integrate Tumor-TILs analyses with another open source Pathomics method, Hover-Net [79]. We combined the capabilities of Hover-Net, which was developed to perform simultaneous segmentation and classification of all of the nuclei in colorectal cancer, by labeling breast cancer cells, lymphocytes, and stromal cells (e.g., connective tissues, endothelium, and nerves) in a highly curated and quality controlled set of images with segmented nuclei [80] in order to calculate the populations of these cells in Tumor-TIL maps. Figure 4 shows the concept of an integrated Pathomics tissue analytics workflow that starts with deep learning analyses to perform automated tumor and lymphocyte detection in breast cancer to generate Tumor-TIL maps combined with Hover-Net as proof of concept to further support the fundamental value of mapping and quantifying tumor immune interactions in cancer. Our primary goal is to show how we can enhance the capabilities of existing Pathomics methods and analyses like Tumor-TIL maps to gain even further insight at the cellular level, which opens the door to numerous downstream research opportunities. A secondary reason for including our implementation of Hover-Net was to show the generalizability of Pathomics methods that can be customized in a practical manner to answer a wide spectrum of research questions.
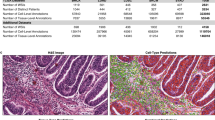
Tumor-TILs analytics combined with nuclear segmentation and classification with Hover-Net in H&E WSIs. Working prototype of an integrated workflow that starts with deep learning analyses to perform automated tumor and lymphocyte detection in breast cancer to generate Tumor-TIL maps, as shown in panels a–d for specimen BRCA TCGA-E2-A9RU-01Z-00-DX1 based on the format in Fig. 1. Implementation of Hover-Net is shown in a representative area in e–g to demonstrate simultaneous segmentation and classification of nuclei. Panel e shows a × 200 representative area of the H&E WSI with panel f showing the segmented and classified nuclei overlaid on the H&E and g showing the masks of the nuclei (lymphocytes denoted in red, stroma and endothelium denoted in blue, and cells with tumor-like features in green). We present this application of Hover-Net as a proof of concept to demonstrate one of many opportunities to integrate Tumor-TILs with other open source Pathomics methods to analyze H&E WSIs at higher resolution. This example of ensemble multiscale Pathomics methods can support scoring the number of TILs in distinct areas (e.g., stromal TILs in central tumor and invasive margin), calculating tumor cellularity in terms of the numbers of cancer cells, IHC quantification, and classical image analysis (color, size, shape, texture, and orientation). Going from the global view from Tumor-TIL maps to cellular level analyses, Hover-Net calculated 2312 nuclei in -g to show that 89% of the cells were classified as lymphocytes. Hover-Net has also been implemented to calculate the number of tumor and stromal cells (not shown since we are focusing on TILs). The green cells in this image are reactive macrophages which have features that can be confused with cancer cells (e.g., increased nuclear size, nucleoli, and irregularities in shape and chromatin texture)
Tumor-TILs provides a global overview of tumor immune interactions, whereas Hover-Net focuses on nuclear segmentation and cell classification at much higher magnification and resolution. This kind of ensemble Pathomics methods supports scoring the number of TILs in distinct areas (e.g., stromal TILs in central tumor and invasive margin), calculating tumor cellularity in terms of the numbers of cancer cells, and classical image analysis to further characterize each classified cell in terms of color, size, shape, texture, and orientation. For example, Hover-Net analyses calculated 2312 nuclei in the representative area shown in Fig. 4e, where 89% of the cells were classified as lymphocytes (Hover-Net has also been implemented to characterize the number of tumor and stromal cells, not shown since we are focusing on TILs).
Even though these methods still require considerable refinement and QA/QC, both Figs. 3 and 4 show how the overall spatial patterns of distribution of TILs can be further interpreted in various histologic contexts at various scales of magnification and resolution with respect to histologic subtype, tumor differentiation, and actual numbers of cancer and immune cells. By utilizing deep learning based nuclear segmentation [80] with classification of different types of nuclei with Hover-Net [79], we can further refine our characterization of the spatial relationships between tumor and immune cells in the TME with a level of precision that has been dreamt about for decades by cancer researchers and pathologists. Ensemble Pathomics can be integrated to systematically capture histopathologic features from multiple scales of magnification en route to developing more comprehensive Pathomics pipelines to catalog quantitative measurements of various phenotypic properties of tissues, cells, and nuclei to support better TILs scoring, IHC quantification, and integrated analyses with other types of data.
Our goals were to introduce Pathomics methods that can be readily implemented to study tumor immune interactions on a large scale, but we have to responsibly address that clinical adoption will require rigorous testing and evaluation. Pathomics algorithms will need to be classified as virtual laboratory tests, which requires evaluating sensitivity, specificity, processing speed (e.g., turnaround time), costs, and limitations. In ways that are no different than any other kind of laboratory test, these kinds of deep learning computer vision Pathomics algorithms must be extensively validated in multiple independent clinical studies before deployment. Currently, the FDA Office of Science and Engineering Laboratories (OSEL) is designing a Medical Device Development Tool (MDDT) to evaluate Pathomics methods to evaluate TILs for clinical use in breast cancer as a necessary mechanism to evaluate both simple and advanced ensemble Pathomics applications due to the challenges associated with so many unknowns within such a complex clinicopathologic scenario (please see refs. [50, 65]). We hope that we have shown how Tumor-TILs Pathomics may serve as a useful tool to view the microscopic world of tumor immune interactions at the cellular and molecular level to support the effort to truly characterize functional immunopathology in every cancer patient for future precision medicine applications.
Conclusions
Our goals are to motivate and inspire future physicians, scientists, engineers, mathematicians, and whomever else to explore the mysteries of the microscopic world of human diseases and cancer with Pathomics tissue analytics. The frontier of cancer research focused on biomarker discovery to help improve treatment and achieve better clinical outcomes for all patients is a highly active area that stands to immediately benefit from Pathomics. We hope the examples that we provided have shown how Pathomics applications can be realistically deployed to identify subtle and complex spatial relationships to study cancer immunopathology and human pathobiology. Moreover, we want to stress that Pathomics tissue analytics can be employed to perform broad types of correlative clinical research that cannot be performed by human beings. There are several readily available opportunities that must be explored with Tumor-TILs and other related methods, starting with using Pathomics tissue analytics for cohort discovery, laboratory QA/QC, and clinical trials before these methods make their way into the clinic. Even though Tumor-TILs provide a global view about the heterogeneity of cancer immunopathology, we believe that these kinds of Pathomics tissue analytics also serve as incredibly useful starting points for precision medicine to develop customized treatment strategies for patients. Despite all of the exciting work and tangible outlooks for the future in terms of countless possibilities for collaborative and correlative research across clinical and scientific disciplines, we also want to stress the need to collectively validate and test Pathomics tissue analytics methodology in multiple collections of WSIs to address key issues that will facilitate the acceptance and clinical adoption of Pathomics in precision medicine.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
•• Bera K, Schalper KA, Rimm DL, Velcheti V, Madabhushi A. Artificial intelligence in digital pathology — new tools for diagnosis and precision oncology. Nat Rev Clin Oncol. 2019;16(11):703–15. https://doi.org/10.1038/s41571-019-0252-y. Digital Pathology and computational image analysis can support Precision Oncology to guide patient stratification selection and select treatment. Morphometric image analysis of tissue samples can provide insight into the relationships between cancer, stromal, and immune cells. This paper evaluates various computational approaches for digital pathology to provide a broad framework across deep learning neural networks, artificial intelligence, and hand-crafted engineered features with an emphasis on biomarker development. Challenges related to the implementation of artificial intelligence in Precision Oncology, such as the need for curated datasets for validation, regulatory approval, and fair reimbursement strategies are also discussed.
• Niazi MKK, Parwani AV, Gurcan MN. Digital pathology and artificial intelligence. The Lancet Oncology. 2019;20(5):e253–e61. https://doi.org/10.1016/s1470-2045(19)30154-8. This paper addresses the integration of digital slides into the workflow of pathology to enable the implementation of advanced algorithms to extend the view of pathologists beyond traditional microscopy of glass slides. Advances in machine learning and the synergy of artificial intelligence and digital pathology are discussed to show for surpassing human limits to integrate data and increase our understanding of human disease.
• Serag A, Ion-Margineanu A, Qureshi H, McMillan R, Saint Martin M-J, Diamond J, et al. Translational AI and deep learning in diagnostic pathology. Frontiers in Medicine. 2019;6(185). https://doi.org/10.3389/fmed.2019.00185. This paper reviews the different approaches of deep learning in emerging applications in pathology and the public grand challenges that have driven innovation. This innovation has led to the exponential growth of technology for tissue and cellular imaging and development of practical applications with the potential to transform diagnostic pathology.
Almeida JS, Iriabho EE, Gorrepati VL, Wilkinson SR, Gruneberg A, Robbins DE, et al. ImageJS: personalized, participated, pervasive, and reproducible image bioinformatics in the web browser. J Pathol Inform. 2012;3:25. https://doi.org/10.4103/2153-3539.98813.
Dong N, Kampffmeyer M, Liang X, Wang Z, Dai W, Xing E. Reinforced auto-zoom net: towards accurate and fast breast cancer segmentation in whole-slide images. Cham: Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support. Springer; 2018. p. 317–25.
Foran DJ, Yang L, Chen W, Hu J, Goodell LA, Reiss M, et al. ImageMiner: a software system for comparative analysis of tissue microarrays using content-based image retrieval, high-performance computing, and grid technology. J Am Med Inform Assoc. 2011;18(4):403–15. https://doi.org/10.1136/amiajnl-2011-000170.
Ghaznavi F, Evans A, Madabhushi A, Feldman M. Digital imaging in pathology: whole-slide imaging and beyond. Annual Review of Pathology: Mechanisms of Disease. 2013;8(1):331–59. https://doi.org/10.1146/annurev-pathol-011811-120902.
Hamilton PW, Bankhead P, Wang Y, Hutchinson R, Kieran D, McArt DG, et al. Digital pathology and image analysis in tissue biomarker research. Methods. 2014;70(1):59–73. https://doi.org/10.1016/j.ymeth.2014.06.015.
Hedvat CV. Digital microscopy: past, present, and future. Archives of pathology & laboratory medicine. 2010;134(11):1666–70. https://doi.org/10.1043/2009-0579-RAR1.1.
Huang H, Tosun AB, Guo J, Chen C, Wang W, Ozolek JA, et al. Cancer diagnosis by nuclear morphometry using spatial information. Pattern Recogn Lett. 2014;42:115–21.
Irshad H, Veillard A, Roux L, Racoceanu D. Methods for nuclei detection, segmentation, and classification in digital histopathology: a review—current status and future potential. IEEE Rev Biomed Eng. 2014;7:97–114.
Kothari S, Phan JH, Osunkoya AO, Wang MD. Biological interpretation of morphological patterns in histopathological whole-slide images. ACM BCB. 2012;2012:218–25. https://doi.org/10.1145/2382936.2382964.
Kothari S, Phan JH, Stokes TH, Wang MD. Pathology imaging informatics for quantitative analysis of whole-slide images. J Am Med Inform Assoc. 2013;20(6):1099–108. https://doi.org/10.1136/amiajnl-2012-001540.
Kumar A, Rao A, Bhavani S, Newberg JY, Murphy RF. Automated analysis of immunohistochemistry images identifies candidate location biomarkers for cancers. Proc Natl Acad Sci. 2014;111(51):18249–54.
Lopez C, Callau C, Bosch R, Korzynska A, Jaen J, Garcia-Rojo M, et al. Development of automated quantification methodologies of immunohistochemical markers to determine patterns of immune response in breast cancer: a retrospective cohort study. BMJ Open. 2014;4(8):e005643. https://doi.org/10.1136/bmjopen-2014-005643.
Madabhushi A, Agner S, Basavanhally A, Doyle S, Lee G. Computer-aided prognosis: predicting patient and disease outcome via quantitative fusion of multi-scale, multi-modal data. Comput Med Imaging Graph. 2011;35(7–8):506–14.
Montalto MC. Pathology RE-imagined: the history of digital radiology and the future of anatomic pathology. Archives of pathology & laboratory medicine. 2008;132(5):764–5. https://doi.org/10.1043/1543-2165(2008)132[764:PRTHOD]2.0.CO;2.
Pantanowitz L. Digital images and the future of digital pathology. Journal of Pathology Informatics. 2010;1(1):15. https://doi.org/10.4103/2153-3539.68332.
Qi X, Xing F, Foran DJ, Yang L. Robust segmentation of overlapping cells in histopathology specimens using parallel seed detection and repulsive level set. IEEE Trans Biomed Eng. 2012;59(3):754–65. https://doi.org/10.1109/TBME.2011.2179298.
Sadimin ET, Foran DJ. Pathology imaging informatics for clinical practice and investigative and translational research. N Am J Med Sci (Boston). 2012;5(2):103–9.
Sertel O, Kong J, Catalyurek UV, Lozanski G, Saltz JH, Gurcan MN. Histopathological image analysis using model-based intermediate representations and color texture: Follicular lymphoma grading. J Signal Process Syst. 2009;55(1–3):169.
•• Galon J, Bruni D. Tumor immunology and tumor evolution: intertwined histories. Immunity. 2020;52(1):55–81. https://doi.org/10.1016/j.immuni.2019.12.018. This article reviews the involvement of immune cells at different stages of tumor progression, which can provide improved prognostic and/or predictive information and therapeutic opportunities. This paper also discusses the inclusion of immune-related parameters to the stratification of tumors that is currently based on histopathology and molecular profiles.
Sergi CM. Digital pathology: the time is now to bridge the gap between medicine and technological singularity. Interactive Multimedia - Multimedia Production and Digital Storytelling. 2019. https://doi.org/10.5772/intechopen.84329.
Shimizu H, Nakayama KI. Artificial intelligence in oncology. Cancer Sci. 2020;111(5):1452–60. https://doi.org/10.1111/cas.14377.
Srivastava A, Kulkarni C, Huang K, Parwani A, Mallick P, Machiraju R. Imitating pathologist based assessment with interpretable and context based neural network modeling of histology images. Biomed Inform Insights. 2018;10:1178222618807481. https://doi.org/10.1177/1178222618807481.
Aeffner F, Zarella MD, Buchbinder N, Bui MM, Goodman MR, Hartman DJ, et al. Introduction to digital image analysis in whole-slide imaging: a white paper from the digital pathology association. Journal of pathology informatics. 2019;10:9. https://doi.org/10.4103/jpi.jpi_82_18.
Zarella MD, Bowman D, Aeffner F, Farahani N, Xthona A, Absar SF, et al. A practical guide to whole slide imaging: a white paper from the digital pathology association. Arch Pathol Lab Med. 2019;143(2):222–34. https://doi.org/10.5858/arpa.2018-0343-RA.
Azuaje F, Kim S-Y, Perez Hernandez D, Dittmar G. Connecting histopathology imaging and proteomics in kidney cancer through machine learning. J Clin Med. 2019;8(10):1535. https://doi.org/10.3390/jcm8101535.
Bukhari SUK, Mehtab U, Hussain SS, Syed A, Armaghan SU, Shah SSH. The assessment of computer vision algorithms for the diagnosis of prostatic adenocarcinoma in surgical specimens. medRxiv. 2020. https://doi.org/10.1101/2020.07.14.20152116.
Campanella G, Hanna MG, Geneslaw L, Miraflor A, Werneck Krauss Silva V, Busam KJ, et al. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat Med. 2019;25(8):1301–9. https://doi.org/10.1038/s41591-019-0508-1.
Chennubhotla C, Clarke LP, Fedorov A, Foran D, Harris G, Helton E, et al. An assessment of imaging informatics for precision medicine in cancer. Yearb Med Inform. 2017;26(1):110–9. https://doi.org/10.15265/IY-2017-041.
Coudray N, Ocampo PS, Sakellaropoulos T, Narula N, Snuderl M, Fenyö D, et al. Classification and mutation prediction from non-small cell lung cancer histopathology images using deep learning. Nat Med. 2018;24(10):1559–67. https://doi.org/10.1038/s41591-018-0177-5.
Ehteshami Bejnordi B, Veta M, Johannes van Diest P, van Ginneken B, Karssemeijer N, Litjens G, et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. Jama. 2017;318(22):2199–210. https://doi.org/10.1001/jama.2017.14585.
Goldenberg SL, Nir G, Salcudean SE. A new era: artificial intelligence and machine learning in prostate cancer. Nat Rev Urol. 2019;16(7):391–403. https://doi.org/10.1038/s41585-019-0193-3.
Gupta R, Kurc T, Sharma A, Almeida JS, Saltz J. The emergence of Pathomics. Current Pathobiology Reports. 2019;7(3):73–84. https://doi.org/10.1007/s40139-019-00200-x.
Litjens G, Sánchez CI, Timofeeva N, Hermsen M, Nagtegaal I, Kovacs I, et al. Deep learning as a tool for increased accuracy and efficiency of histopathological diagnosis. Sci Rep. 2016;6:26286. https://doi.org/10.1038/srep26286.
Nawaz S, Yuan Y. Computational pathology: exploring the spatial dimension of tumor ecology. Cancer Lett. 2016;380(1):296–303. https://doi.org/10.1016/j.canlet.2015.11.018.
Parwani AV. Next generation diagnostic pathology: use of digital pathology and artificial intelligence tools to augment a pathological diagnosis. Diagn Pathol. 2019;14(1):138. https://doi.org/10.1186/s13000-019-0921-2.
Parwani AV, Amin MB. Convergence of digital pathology and artificial intelligence tools in anatomic pathology practice: current landscape and future directions. Adv Anat Pathol. 2020;27(4):221–6. https://doi.org/10.1097/PAP.0000000000000271.
• Pell R, Oien K, Robinson M, Pitman H, Rajpoot N, Rittscher J, et al. The use of digital pathology and image analysis in clinical trials. J Pathol Clin Res. 2019;5(2):81–90. https://doi.org/10.1002/cjp2.127. This publication provides a solid overview about the potential of digital pathology and image analysis to provide greater accuracy, reproducibility, and standardization of criteria for entry and endpoints for clinical trials. Potential applications, current challenges, and limitations are discussed in the context of new opportunities to identify, extract, and quantify histopathologic features in comparison to manual assessment by pathologists in order to address routine adoption of quantitative tissue morphometrics in clinical trials.
Riordan DP, Varma S, West RB, Brown PO. Automated analysis and classification of histological tissue features by multi-dimensional microscopic molecular profiling. PLoS One. 2015;10(7):e0128975. https://doi.org/10.1371/journal.pone.0128975.
Liñares-Blanco J, Munteanu CR, Pazos A, Fernandez-Lozano C. Molecular docking and machine learning analysis of abemaciclib in colon cancer. BMC Molecular and Cell Biology. 2020;21(1):52. https://doi.org/10.1186/s12860-020-00295-w.
• Le H, Gupta R, Hou L, Abousamra S, Fassler D, Torre-Healy L, et al. Utilizing automated breast cancer detection to identify spatial distributions of tumor-infiltrating lymphocytes in invasive breast Cancer. Am J Pathol. 2020;190(7):1491–504. https://doi.org/10.1016/j.ajpath.2020.03.012. This paper describes the quantitative assessment of spatial maps of tumor-infiltrating lymphocytes generated with deep learning Pathomics pipelines in diagnostic whole slide tissue images of breast cancer. The combined maps provide insight about the structural patterns and facilitate improved quantification of tumor-infiltrating lymphocytes for further downstream analyses (like predicting survival). The open-source tools and dataset of 1090 invasive breast cancer images and maps from The Cancer Genome Atlas are publicly available for download.
Le H, Samaras D, Kurc T, Gupta R, Shroyer K, Saltz J, editors. Pancreatic cancer detection in whole slide images using noisy label annotations. Medical Image Computing and Computer Assisted Intervention – MICCAI 2019. Cham: Springer International Publishing; 2019.
Saltz J, Gupta R, Hou L, Kurc T, Singh P, Nguyen V, et al. Spatial organization and molecular correlation of tumor-infiltrating lymphocytes using deep learning on pathology images. Cell Rep. 2018;23(1):181.
Galon J, Pagès F, Marincola FM, Angell HK, Thurin M, Lugli A, et al. Cancer classification using the Immunoscore: a worldwide task force. J Transl Med. 2012;10:205. https://doi.org/10.1186/1479-5876-10-205.
Galon J, Mlecnik B, Bindea G, Angell HK, Berger A, Lagorce C, et al. Towards the introduction of the 'Immunoscore' in the classification of malignant tumours. J Pathol. 2014;232(2):199–209. https://doi.org/10.1002/path.4287.
Hendry S, Salgado R, Gevaert T, Russell PA, John T, Thapa B, et al. Assessing tumor-infiltrating lymphocytes in solid tumors: a practical review for pathologists and proposal for a standardized method from the International Immuno-Oncology Biomarkers Working Group: Part 2: TILs in melanoma, gastrointestinal tract carcinomas, non-small cell lung carcinoma and mesothelioma, endometrial and ovarian carcinomas, squamous cell carcinoma of the head and neck, genitourinary carcinomas, and primary brain tumors. Adv Anat Pathol. 2017;24(6):311–35. https://doi.org/10.1097/PAP.0000000000000161.
Hendry S, Salgado R, Gevaert T, Russell PA, John T, Thapa B, et al. Assessing tumor-infiltrating lymphocytes in solid tumors: a practical review for pathologists and proposal for a standardized method from the international Immunooncology biomarkers working group: part 1: assessing the host immune response, TILs in invasive breast carcinoma and ductal carcinoma in situ, metastatic tumor deposits and areas for further research. Adv Anat Pathol. 2017;24(5):235–51. https://doi.org/10.1097/PAP.0000000000000162.
Amgad M, Stovgaard ES, Balslev E, Thagaard J, Chen W, Dudgeon S, et al. Report on computational assessment of tumor infiltrating lymphocytes from the International Immuno-Oncology Biomarker Working Group. NPJ breast cancer. 2020;6:16. https://doi.org/10.1038/s41523-020-0154-2.
Salgado R, Denkert C, Demaria S, Sirtaine N, Klauschen F, Pruneri G, et al. The evaluation of tumor-infiltrating lymphocytes (TILs) in breast cancer: recommendations by an International TILs Working Group 2014. Ann Oncol. 2015;26(2):259–71. https://doi.org/10.1093/annonc/mdu450.
Fridman WH, Pages F, Sautes-Fridman C, Galon J. The immune contexture in human tumours: impact on clinical outcome. Nat Rev Cancer. 2012;12(4):298–306. https://doi.org/10.1038/nrc3245.
Gajewski TF, Schreiber H, Fu YX. Innate and adaptive immune cells in the tumor microenvironment. Nat Immunol. 2013;14(10):1014–22. https://doi.org/10.1038/ni.2703.
Galon J, Angell HK, Bedognetti D, Marincola FM. The continuum of cancer immunosurveillance: prognostic, predictive, and mechanistic signatures. Immunity. 2013;39(1):11–26. https://doi.org/10.1016/j.immuni.2013.07.008.
Wein L, Savas P, Luen SJ, Virassamy B, Salgado R, Loi S. Clinical validity and utility of tumor-infiltrating lymphocytes in routine clinical practice for breast cancer patients: current and future directions. Front Oncol. 2017;7:156. https://doi.org/10.3389/fonc.2017.00156.
Gurcan MN, Boucheron LE, Can A, Madabhushi A, Rajpoot NM, Yener B. Histopathological image analysis: a review. IEEE Rev Biomed Eng. 2009;2:147–71. https://doi.org/10.1109/RBME.2009.2034865.
Gurcan MN, Pan T, Shimada H, Saltz J. Image analysis for neuroblastoma classification: segmentation of cell nuclei. Conf Proc IEEE Eng Med Biol Soc. 2006;1:4844–7. https://doi.org/10.1109/iembs.2006.260837.
Gurcan MN, Tomaszewski J, Overton JA, Doyle S, Ruttenberg A, Smith B. Developing the quantitative histopathology image ontology (QHIO): a case study using the hot spot detection problem. J Biomed Inform. 2017;66:129–35. https://doi.org/10.1016/j.jbi.2016.12.006.
Kather JN, Suarez-Carmona M, Charoentong P, Weis C-A, Hirsch D, Bankhead P, et al. Topography of cancer-associated immune cells in human solid tumors. Elife. 2018;7:e36967. https://doi.org/10.7554/eLife.36967.
Loi S, Drubay D, Adams S, Pruneri G, Francis PA, Lacroix-Triki M, et al. Tumor-infiltrating lymphocytes and prognosis: a pooled individual patient analysis of early-stage triple-negative breast cancers. J Clin Oncol. 2019;37(7):559–69. https://doi.org/10.1200/jco.18.01010.
Norton K-A, Gong C, Jamalian S, Popel AS. Multiscale agent-based and hybrid modeling of the tumor immune microenvironment. Processes (Basel). 2019;7(1):37. https://doi.org/10.3390/pr7010037.
Wortman JC, He T-F, Solomon S, Zhang RZ, Rosario A, Wang R, et al. Fractal dimension, occupancy and hotspot analyses of B cell spatial distribution predict clinical outcome in breast cancer. bioRxiv. 2019:678607. https://doi.org/10.1101/678607.
Yu CC, Wortman JC, He T-F, Solomon S, Zhang RZ, Rosario A et al. Physics approaches to the spatial distribution of immune cells in tumors. arXiv e-prints 2019. https://ui.adsabs.harvard.edu/abs/2019arXiv191111846Y.
• Gonzalez-Ericsson PI, Stovgaard ES, Sua LF, Reisenbichler E, Kos Z, Carter JM, et al. The path to a better biomarker: application of a risk management framework for the implementation of PD-L1 and TILs as immuno-oncology biomarkers in breast cancer clinical trials and daily practice. J Pathol. 2020;250(5):667–84. https://doi.org/10.1002/path.5406. PD‐1/PD‐L1 targeted immunotherapy has rapidly become the standard of care in oncology for several tumor types, including triple negative breast cancer. However, there are concerns about the performance of immunohistochemical PD‐L1 assays, reproducibility between assays, and interobserver variability. Since high numbers of tumor‐infiltrating lymphocytes are associated with response to PD‐1/PD‐L1 inhibitors in patients with breast cancer, systematic implementation of combined PD‐L1 and TIL analyses as a more comprehensive immuno‐oncological biomarker is proposed for patient selection for PD‐1/PD‐L1 immunotherapy.
Kos Z, Roblin E, Kim RS, Michiels S, Gallas BD, Chen W, et al. Pitfalls in assessing stromal tumor infiltrating lymphocytes (sTILs) in breast cancer. NPJ breast cancer. 2020;6(1):17. https://doi.org/10.1038/s41523-020-0156-0.
Wortman JC, He T-F, Rosario A, Wang R, Schmolze D, Yuan Y, et al. Occupancy and fractal dimension analyses of the spatial distribution of cytotoxic (CD8+) T cells infiltrating the tumor microenvironment in triple negative breast Cancer. Biophys Rev Lett. 2020:1–16. https://doi.org/10.1142/S1793048020500022.
McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM. Reporting recommendations for tumor marker prognostic studies (REMARK). J Natl Cancer Inst. 2005;97(16):1180–4. https://doi.org/10.1093/jnci/dji237.
Sauerbrei W, Taube SE, McShane LM, Cavenagh MM, Altman DG. Reporting recommendations for tumor marker prognostic studies (REMARK): an abridged explanation and elaboration. J Natl Cancer Inst. 2018;110(8):803–11. https://doi.org/10.1093/jnci/djy088.
Lapuente-Santana Ó, Eduati F. Toward systems biomarkers of response to immune checkpoint blockers. Frontiers in oncology. 2020;10:1027. https://doi.org/10.3389/fonc.2020.01027.
Choi J, Maeng HG, Lee SJ, Kim YJ, Kim DW, Lee HN, et al. Diagnostic value of peripheral blood immune profiling in colorectal cancer. Ann Surg Treat Res. 2018;94(6):312–21. https://doi.org/10.4174/astr.2018.94.6.312.
Lepone LM, Donahue RN, Grenga I, Metenou S, Richards J, Heery CR, et al. Analyses of 123 peripheral human immune cell subsets: defining differences with age and between healthy donors and cancer patients not detected in analysis of standard immune cell types. J Circ Biomark. 2016;5:5. https://doi.org/10.5772/62322.
Ogino S, Nowak JA, Hamada T, Phipps AI, Peters U, Milner DA Jr, et al. Integrative analysis of exogenous, endogenous, tumour and immune factors for precision medicine. Gut. 2018;67(6):1168–80. https://doi.org/10.1136/gutjnl-2017-315537.
Calì B, Molon B, Viola A. Tuning cancer fate: the unremitting role of host immunity. Open Biol. 2017;7(4):170006. https://doi.org/10.1098/rsob.170006.
Diaz-Cano SJ. General morphological and biological features of neoplasms: integration of molecular findings. Histopathology. 2008;53(1):1–19. https://doi.org/10.1111/j.1365-2559.2007.02937.x.
Galon J, Costes A, Sanchez-Cabo F, Kirilovsky A, Mlecnik B, Lagorce-Pagès C, et al. Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science. 2006;313(5795):1960–4.
Parra ER, Francisco-Cruz A, Wistuba II. State-of-the-art of profiling immune contexture in the era of multiplexed staining and digital analysis to study paraffin tumor tissues. Cancers (Basel). 2019;11(2). https://doi.org/10.3390/cancers11020247.
Roma-Rodrigues C, Mendes R, Baptista PV, Fernandes AR. Targeting tumor microenvironment for Cancer therapy. Int J Mol Sci. 2019;20(4):840. https://doi.org/10.3390/ijms20040840.
Denkert C, Wienert S, Poterie A, Loibl S, Budczies J, Badve S, et al. Standardized evaluation of tumor-infiltrating lymphocytes in breast cancer: results of the ring studies of the international immuno-oncology biomarker working group. Mod Pathol. 2016;29(10):1155–64. https://doi.org/10.1038/modpathol.2016.109.
Graham S, Vu QD, Raza SEA, Azam A, Tsang YW, Kwak JT, et al. Hover-Net: simultaneous segmentation and classification of nuclei in multi-tissue histology images. Med Image Anal. 2019;58:101563. https://doi.org/10.1016/j.media.2019.101563.
Hou L, Gupta R, Van Arnam JS, Zhang Y, Sivalenka K, Samaras D, et al. Dataset of segmented nuclei in hematoxylin and eosin stained histopathology images of ten cancer types. Scientific Data. 2020;7(1):185. https://doi.org/10.1038/s41597-020-0528-1.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors attest that we have no conflicts of interest.
Ethics Statement for Human and Animal Rights
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Current State of the Science of Disease Modeling
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gupta, R., Le, H., Van Arnam, J. et al. Characterizing Immune Responses in Whole Slide Images of Cancer With Digital Pathology and Pathomics. Curr Pathobiol Rep 8, 133–148 (2020). https://doi.org/10.1007/s40139-020-00217-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40139-020-00217-7