Abstract
Background
Atherosclerotic plaque is considered the hallmark of atherosclerotic lesions in coronary atherosclerosis (CAS), the primary pathogenesis in coronary artery disease (CAD), which develops and progresses through a complex interplay between immune cells, vascular cells, and endothelial shear stresses. Early diagnosis of CAS is critical for avoiding plaque rupture and sudden death. Therefore, identifying new CAD biomarkers linked to vessel wall functions, such as RNA molecules with their distinct signature, is a promising development for these patients. With this rationale, the present study investigated the expression level of the vascular-related RNA transcripts (lncRNA ANRIL, miRNA-126-5p, CDK4, CDK6, TGF-β, E-cadherin, and TNF-α) implicated in the cellular vascular function, proliferation, and inflammatory processes.
Methods
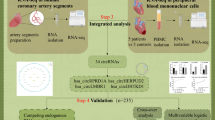
A case-control study design with a total of 180 subjects classified participants into two groups; CAD and control groups. The relative expression levels of the seven transcripts under study—selected using online bioinformatics tools and current literature—were assessed in the plasma of all study participants using RT-qPCR. Their predictive significance testing, scoring of disease prioritization, enrichment analysis, and the miRNA‐mRNA regulatory network was investigated.
Results
The relative expression levels of all seven of the circulating vascular-related transcripts under study were statistically significant between CAD patients and controls. Receiver operating characteristic (ROC) analysis results indicated the statistical significance of all the transcripts under study with CDK4 showing the highest area under the curve (AUC) equivalent to 0.91, followed by E-cadherin (0.90), miRNA-126-5p (0.83), ANRIL (0.82), TNF-α (0.63), TGF-β (0.62), and CDK6 (0.59), in descending order. A strong association was detected between most of the transcripts studied in CAD patients with a significant Spearman’s correlation coefficient with a two-tailed significance of p < 0.001. Network analysis revealed a strong relationship between the five circulating vasculature transcripts studied and their target miRNAs and miR-126-5p, but not for ANRIL.
Conclusion
The seven circulating vascular-related RNA transcripts under study could serve as potential CAD biomarkers, reflecting the cellular vascular function, proliferation, and inflammatory processes in CAD patients. Therefore, blood transcriptome analysis opens new frontiers for the non-invasive diagnosis of CAD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
The seven circulating vascular-related RNA transcripts under study (lncRNA ANRIL, miRNA-126-5p, CDK4, CDK6, TGF-β, E-cadherin, and TNF-α) could serve as potential coronary artery disease (CAD) biomarkers. |
The studied transcripts are a promising non-invasive panel that can be used for reflecting the cellular vascular function, proliferation, and inflammatory processes in CAD patients. |
Blood transcriptome analysis via investigating different types of RNA transcripts in CAD patients opens new frontiers for the non-invasive diagnosis of CAD. |
1 Introduction
Coronary artery disease (CAD), a subtype of cardiovascular disease, is considered one of the leading causes of death worldwide [1]. Several risk factors are associated with CAD, including smoking, obesity, high blood pressure, high blood levels of lipids, family history, and genetic predisposition [2]. Previous epidemiological studies indicated that CAD susceptibility is 40–60% of genetic origin [3]. In Egypt, cardiac genetic and genomic changes are currently considered a priority with the rise of the Egyptian Collaborative Cardiac Genomics (ECCO-GEN) project supported by the Aswan Heart Center and the Magdi Yacoub Global Heart Foundation [4].
Atherosclerosis is the primary pathogenesis of coronary vasculature [5]. Coronary atherosclerosis (CAS) is characterized by various clinical manifestations, such as acute coronary syndrome, stable angina, and heart failure [6]. CAS is a prolonged, inflammatory, multifaceted condition where atherosclerotic plaque is considered the hallmark of atherosclerotic lesions [7]. CAS develops and progresses through a complex interplay between immune cells, vascular cells, and endothelial shear stresses [8].
Immune cells produce both pro- and anti-inflammatory cytokines and thus are involved in the initiation of the atherogenic process and affect plaque formation as well as stability [9]. Vascular smooth muscle cells (VSMCs) and vascular endothelial cells (EC) are major constituents of the arterial wall. They are essential in maintaining blood flow via vascular wall remodeling [5]. VSMC proliferation and phenotype switch are prominent contributors to atherosclerotic plaque formation [10], where plaque content of VSMCs is inversely proportional to plaque vulnerability [10]. Furthermore, endothelial cell activation and dysfunction are significant features during atherogenesis, where endothelial dysfunction plays a central role in the formation, progression, and complications of atherosclerotic plaque [11].
Early diagnosis of CAS is critical for avoiding plaque rupture that may result in acute myocardial infarction (MI) or sudden cardiac death [12]. Therefore, identifying coronary artery biomarkers linked to vessel wall functions is a hot spot in CAS research. Earlier studies indicated that RNAs have a distinct signature in the blood of CAD patients [13,14,15]. These RNA transcripts are probably linked to genes involved in inflammation and cell cycle control [16,17,18].
Long ncRNA (lncRNA), a class of non-coding RNA (ncRNA) molecules with lengths of >200 nucleotides, are thought to serve an essential role in the development of human diseases; however, only a few have been well characterized with regards to their functional roles [19]. ANRIL, also known as cyclin-dependent kinase inhibitor 2B antisense RNA 1 (CDKN2B-AS1), is implicated in various malignant and nonmalignant diseases [20] and is known previously to have a role in cardiovascular diseases (CVDs) by regulating inflammation responses and cell cycle [11]. Regarding CAD, ANRIL is expressed in vascular endothelial cells and smooth muscles and has a vital role in plaque formation [19,20,21,22].
Another subtype of ncRNAs is the microRNAs (miRNAs), which have a role in regulating gene expression on post-transcription or post-translation levels [23] and were also found to be altered in CAD patients and implicated in atherosclerosis development [24]. MiRNA-126-5p was among the early discovered miRNAs expressed specifically in the vasculature and have critical functions in vascular development [25]. Recent studies have unveiled the potentially important function of miRNA-126-5p in vascular diseases, including atherosclerosis, coronary artery disease, stroke, and diabetic vasculopathy. The action of miRNA-126-5p reflects its function in angiogenesis and inflammation [24,25,26].
Several publications illustrated the interplay and cross-regulation between different RNA classes, especially in chronic diseases [27,28,29,30]. Following on from these, the present study investigates the expression level of some of the vascular-related RNA transcripts (lncRNA ANRIL, miRNA-126-5p, CDK4, CDK6, TGF-β, E-cadherin, and TNF-α) implicated in the cellular vascular function, proliferation, and inflammatory processes in the plasma of CAD patients to elucidate their role as potential CAD biomarkers.
2 Study Participants and Methodological Approach
2.1 Study Participants
A case-control study design with 180 participants was classified into two groups. The CAD group included 90 patients presenting with either complaints or signs of CAD revealed by examination, ECG, and echocardiography and recruited from the Suez Canal University Hospital (SCUH) cardiology outpatient clinic from June 2020 to December 2021. Patients were enrolled from both genders, and their ages ranged from 18 to 85 years. Patient history was taken for each patient, including demographic data, CAD family history, past history, and lifestyle habits. We excluded patients with other chronic CVD or congenital heart disease. The risk factors were detected by clinical diagnosis and/or treatment for hyperlipidemia, diabetes mellitus, and hypertension. All patients under study underwent a clinical examination, including vital signs and general and local examinations. For calculating body mass index (BMI) (kg/m2), anthropometrical height and body weight were estimated. Two measurements were performed for blood pressure and resting heart rate using an automatic sphygmomanometer, and average values were recorded. left ventricular (LV) size, LV systolic function (ejection fraction), and diastolic dysfunction was evaluated by echocardiography with two independent operators. The left ventricular ejection fraction (LVEF) and wall motion score index (WMSI) were estimated for all CAD patients. CAD patients under study were selected based on at least one of the following findings: positive stress imaging, whether by stress echocardiography or exercise stress ECG or single-photon emission computerized tomography (SPECT), wall motion alterations in resting echocardiography, prior history of admission to a coronary care unit with CAD or previous coronary stenting or prior coronary angiography diagnosing obstructive CAD.
The control group included 90 healthy volunteers from the SCUH blood bank of both genders, aged over 18 years old, with no current treatments or complaints from any CVD or chronic disease. All control subjects aged over 45 years old were subjected to echocardiography to assess their cardiac condition and to ensure they were free from any CAD signs.
2.2 Selection of Circulating Vasculature-Related Transcripts Using Bioinformatics Tools
The transcripts under study were selected using online bioinformatics tools such as HMDD (http://www.cuilab.cn/) [31], miR2Disease (http://www.mir2disease.org/) [32], LncRNAdisease (http://www.rnanut.net/lncrnadisease/) [33], and LncBase v3 (https://diana.e-ce.uth.gr/lncbasev3) [34]. In addition, current literature results for the most common ncRNAs and their target genes involved in vascular heart disease were considered [35,36,37].
2.3 Plasma Samples Collection
A vacutainer tube containing ethylene diamine tetraacetic acid (EDTA) anticoagulant was used to collect 3 mL of blood from all study participants. Plasma was obtained via centrifugation for 15 minutes at 3500 rpm, and qiazol was added to plasma with a ratio of 5:1 and stored at − 80 °C until transcript analysis. To ensure there was no sample hemolysis, the blood sample was slowly withdrawn and slowly shacked (7–8 times) inside the EDTA vacutainer. Fresh samples were centrifuged within 1 hour after withdrawal. In case of any visible hemolysis, the sample was assessed using spectrophotometer and samples with a significant degree of hemolysis were discarded.
2.4 Total RNA Including miRNA Extraction and Quality Analysis
Qiagen miRNeasy Mini kit (cat no 217004, QIAGEN, Hilden, Germany) was used for total RNA extraction using the manufacturer protocol. NanoDrop 2000 1C spectrophotometer (NanoDrop Tech., Inc. Wilmington, DE, USA) was used to determine RNA concentration and purity.
2.5 Circulating Vasculature-Related Transcripts Expression Analysis Using Quantitative Real-Time PCR Assay
The expression levels of the seven selected vasculature circulating transcripts were assessed in the plasma of all study participants. RT-qPCR was done via two steps:
-
(a)
Reverse transcription (RT), where miScript II RT Kit (Qiagen, Catalog no. 218161) was used to generate complementary DNA (cDNA) from total RNA, in which mRNAs, miRNAs, and lncRNAs were polyadenylated by poly(A) polymerase and converted into cDNA. RT was carried out at 37 °C for 1 hour, followed by inactivation of the reaction by briefly incubating at 95 °C using a Veriti™ 96-Well Thermal Cycler (Applied Biosystems, USA)
-
(b)
qPCR, where the cDNA premix was used as a template for relative quantification of the seven transcripts under study (ANRIL, miR-126-5p, CDK4, CDK6, E-cadherin, TNF-α, and TGF-β) using miScript SYBR Green PCR Kit (Qiagen, cat. no 218076) with a universal reverse primer and a specific forward primer sequence for miRNAs, while other genes had their specific forward and reverse primers as presented in Table 1. For ANRIL, Taqman ncRNA assay CDKN2B-AS1 (Thermofisher, USA) was used following the manufacturer's instructions. Minimum information for publication of quantitative RT-PCR experiments (MIQE) guidelines were followed throughout the experiment. The reactions were run in triplicate with ‘No-RT’, and ‘No-template’ controls present in each run. The qPCR cycling conditions started at 95 °C for 5 minutes, followed by 40 cycles of denaturation, annealing, and elongation at 95 °C (15 s), 55 °C (1 min), and 72 °C (1 min), respectively. Endogenous control β-actin was used to normalize ANRIL and mRNAs, while SNORD68 was used to normalize miRNA-126-5p to enable data analysis using the ΔΔCT method of relative quantification.
2.6 Circulating Vasculature-Related Transcripts Predictive Significance Testing
The discriminative power of the vasculature circulating transcripts was analyzed individually and in groups using receiver operating characteristic (ROC) curves to evaluate the diagnostic power of the transcripts under study as biomarkers in CAD. Statistical significance was considered for a p value of < 0.05.
2.7 Scoring of Disease Prioritization for Circulating Vasculature-Related Transcripts Under Study in Relation to CAD
For disease prioritization, the gene set option in GeneAnalytics software [38] was used to perform the disease relationship analysis using the information available in MalaCards database, the human disease database. The following parameters determined the disease-matching score: the number of transcripts studied that matched CAD normalized by the total number of transcripts associated explicitly with CAD, the quality and type of the transcript–CAD relations, and each gene in CAD has a score derived from its relationship to the disease, as annotated in MalaCards sources. We also chose the anatomical disease categorization option and selected CVDs; then, we presented the score of the top 12 CVDs associated with the transcripts under study. As for gene prioritization concerning CAD, VarElect software [39] was used for the prioritization of the genes for the transcripts under study based on their contribution to CAD. The VarElect scoring system depends on giving each gene a relevance score depending (in part) on its current weight with CAD based on previous research. The weight with CAD was determined by the frequency with which the gene appears in association with CAD (term frequency) compared with all genes under study (inverse document frequency).
2.8 Enrichment Analysis of Circulating Vasculature-Related Transcripts in CAD
GeneAnalytics software was used to conduct the functional enrichment analysis to annotate the seven vasculature circulating transcripts into different pathways and gene ontology (GO) terms (biological process, cellular component, and molecular function). The results were prioritized by relevance to the analyzed transcripts under study. Considering pathway enrichment analysis, GeneAnalytics software was also used to analyze the seven transcripts under study—pathway relationships using PathCards available information from the biological pathways database. PathCards applies a gene-content algorithm enabling the unification of thousands of pathways from multiple pathway sources available in GeneCards and clusters them into SuperPaths. Considering GO analysis, novel and supporting information were retrieved regarding the functional roles of the genes under study in biological processes such as development, differentiation, diseases, and pathways, as well as in molecular functions such as binding or catalysis. The presented score in each item is a transformation of the resulting p value, corrected for multiple comparisons using the false discovery rate (FDR) method, with higher scores indicating a better match.
2.9 Vasculature-Related miRNA–mRNA Regulatory Network Construction
The targets of the vasculature circulating transcripts in CAD were predicted using miRTargetLink 2.0 (Version 2.0, https://ccb-compute.cs.uni-saarland.de/) [40]. Statistically homogenous and significant differentially expressed transcript targets were kept, and different filters were used to retrieve strong validated targets only.
2.10 Statistical Analysis
SPSS software version 23.0 and Excel 2016 were used for data analysis. G*Power 3.1.9.2 (http://www.gpower.hhu.de/A) [41] was used for sample size calculation. Gene expression fold change of the transcripts was estimated using the LIVAC method (= 2−ΔΔCq) [42]. Quantitative data were expressed as median and quartiles or means ± standard deviation, while qualitative data were expressed as numbers and percentages. Student-t, Two-sided Chi-square, ANOVA, Kruskal-Wallis, and Mann-Whitney U tests were used as appropriate. Correlation analysis by Spearman's rank test was carried out. Different statistical methods were used to decrease the confounding effect as follows:
-
1.
Stratification for producing different groups within which some of the confounders do not vary. Then, the exposure–outcome association was evaluated within each stratum of the confounder. So, within each stratum, the confounder could not confound because it does not vary across the exposure–outcome.
-
2.
ROC curve of the circulating transcripts and other study variables.
-
3.
Spearman’s correlation analysis for the correlation between CAD and various variables in the study.
3 Results
3.1 Main Features and Risk Factors Among the Study Population
The main feature data from all study participants are presented in Table 2. The average age of participants was 49.4 ± 6.1 and 53.9 ± 9.7 years among controls and CAD groups, respectively. The CAD group showed significantly higher ages (p < 0.001) than the control group. On the contrary, there was no significant difference among different ages. The male gender was more represented among the study participants, with 54 (60.0%) and 65 (72.2%) males in the control and the CAD group, respectively, with a non-statistical difference (p > 0.05) among the two groups. Regarding CAD risk factors, smokers were significantly (p = 0.011) higher in the CAD group with a total of 51 (56.7%) smokers compared with the control group, which included 33 (36.7%) smokers. Family history was found in 57 (63.3%) patients compared with 35 (38.9%) participants in the control group, with a significant difference between the two groups. Concerning dyslipidemia, CAD patients had a significantly higher incidence, representing 80.0% of the CAD population compared with 7.8% in the control group. The average (mean ± SD) BMI of the CAD group was 30.4 ± 6.2, with no statistical difference (p = 0.158) compared with the control group (29.3 ± 2.7). At the same time, there was a statistically significant difference among different obesity grades, as shown in Table 2.
3.2 Comorbidities and Cardiac Findings in CAD Patients
Table 3 shows the comorbidities and cardiac findings among the CAD patients in the study. Clinical examination revealed an average ± SD for body surface area (BSA) of 1.83 ± 0.17, systolic blood pressure of 126.78 ± 16.46, and diastolic blood pressure of 80.89 ± 14.8. At the same time, echocardiography revealed an average ± SD for LVEF (46.02 ± 12.99) and WMSI (1.5 ± 0.42) with statistically significant results. Concerning comorbidities, there was no statistical significance between CAD and either diabetes, hypertension, or ischemic heart disease among our study patients. Regarding LVEF grades, the chi-squared test revealed a statistically significant difference among the different grades. The diastolic function grades are represented in grades normal 50 (26.3%), mild 30 (15.8%), moderate 4 (2.1%), and severe 6 (3.2%), with a highly significant difference between them.
3.3 Relative Expression Analysis of Circulating Vasculature-Related Transcripts
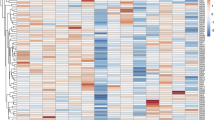
The relative expression patterns of ANRIL, miR-126-5p, CDK4, CDK6, E-cadherin, TNF-α, and TGF-β were assessed using qRT‐PCR and are shown in Fig. 1 as a heatmap. The transcripts’ relative expression is presented as median (Q1 and Q3) in Fig. 2. The scores were as follows: ANRIL 3.6 (1.9–5.9), miR-126 4.2 (1.7–8.2), CDK4 7.9 (4.33–10.9), CDK6 1.8 (− 0.6 to 3.1), E-cadherin 7.7 (4.1–11.4), TNF-α 2.1 (0.1–4.4), and TGF-β − 0.1 (− 3.1 to 2.2) (Fig. 2).
3.4 Circulating Vasculature-Related Transcripts Predictive Significance by ROC Analysis
ROC, including AUC and confidence interval levels, are presented in Table 4. The results indicated the statistical significance of all the circulating transcripts under study. CDK4 showed the highest AUC equivalent to 0.91, followed by E-cadherin (0.90), miR-126-5p (0.83), ANRIL (0.82), TNF-α (0.63), TGF-β (0.62), and CDK6 (0.59) in descending order.
The combined predictive ROC curve analysis for the transcripts under study is presented in Fig. 3, showing high predictive significant results for each group. Groups were combined based on the different roles of these transcripts on vasculature pathogenesis. In group A, CDK4 and CDK6 are implicated in vascular smooth muscle proliferation, while E-cadherin is responsible for the proliferation and integrity of endothelial cells. In group B, ANRIL is implicated in vascular smooth muscle proliferation, while miR-126-5p is responsible for the proliferation and integrity of endothelial cells. In group C, TNF-α and TGF-β are implicated in the vascular inflammatory process with opposite atherogenic and athero-protective actions.
Combined predictive significance by ROC curve analysis for different circulating vasculature transcripts under study. A Combined ROC curve analysis for E-cadherin, CDK4, and CDK6. B Combined ROC curve analysis for ANRIL and miR-126-5p. C Combined ROC curve analysis for TNF-α and TGF-β. ROC receiver operating characteristic
3.5 Correlation Analysis Among the Circulating Vasculature-Related Transcripts’ Expression Levels and CAD Patients’ Clinical Features
The seven circulating vasculature-related transcripts showed various distributions among all CAD patients, as shown in Table 5. Although none of the transcripts under study is a potential target gene for miR-126-5p or ANRIL, a strong association was detected between most of the transcripts in CAD patients with a significant Spearman’s correlation coefficient with a two-tailed significance of p < 0.001. The most significant correlation was between ANRIL and CDK4 (r = 0.74), ANRIL and CDK6 (r = 0.70), ANRIL and E-cadherin (r = 0.68), miR-126-5p and CDK4 (r = 0.55), E-cadherin and CDK4 (r = 0.64), and TNF-α and CDK6 (r = 0.54). This could suggest new predicted targets for both miR-126-5p and ANRIL that need further validation in future studies.
Considering the relation between the seven circulating vasculature-related transcripts under study and the clinical findings shown in Table 6, all the studied genes were statistically significant with CAD. ANRIL, CDK4, CDK6, and E-cadherin showed a positive statistically significant relationship with age. ANRIL, CDK4, CDK6, E-cadherin, and TNF-α showed a positive, statistically significant relationship with gender. All the genes studied showed positive, statistically significant results with smoking except TGF-β. MiR-126-5p, CDK4, and E-cadherin showed positive, statistically significant results with family history. Dyslipidemia correlated significantly with all genes under study except TGF-β. Finally, ANRIL and CDK4 showed a significant correlation concerning hypertensive patients.
As for correlations of the clinicopathological features among CAD patients shown in Table 7, age correlated significantly with gender, smoking, family history, diabetes, hypertension, and ischemic heart disease. Gender correlated significantly with age, smoking, family history, hypertension, and BSA. Family history correlated significantly with age, gender, ischemic heart disease, and WMSI. Dyslipidemia correlated significantly with BMI, obesity, obesity grade, and LVEF. BMI, obesity, and obesity grade correlated significantly with each other, dyslipidemia, DM, ischemic heart disease, BSA, and WMSI. Both LVEF and WMSI correlated significantly with each other and with ischemic heart disease.
3.6 Disease and Prioritization Scores for the Circulating Vasculature-Related Transcripts Studied in Relation to CAD
Anatomical disease categorization revealed a total of 1215 diseases related to the seven circulating transcripts under study, with 120 diseases related to the cardiovascular system. The top scores are shown in Fig. 4A and are mostly related to CAD pathogenesis.
Disease and prioritization scores for the transcripts under study in relation to CAD. A Disease score for the transcripts among cardiovascular diseases. Scores were calculated using GeneAnalytics tool (https://geneanalytics.genecards.org). B Prioritization score of the gene variants of the seven circulating vasculature transcripts in relation to CAD. C Average disease-causing likelihood score for the seven circulating vasculature transcripts in relation to CAD. The color scale presented from red to blue where blue reflects lower scores, and red indicates higher ones. Scores were calculated using VarElect tool (https://ve.genecards.org/). CAD coronary artery disease, CVD cardiovascular disease, Familial AA familial aortic aneurysm, Lipoprotein QTL lipoprotein quantitative trait locus
Concerning the strength of the connection between each transcript under study and CAD, the results presented in Fig. 4B show the prioritization score of the gene variants among the seven circulating transcripts. This list is based on the association between CAD-shared pathways and these biomarkers, paralog relations, interaction networks, domain sharing, and joint publications. Furthermore, analyzing the average likelihood for these seven biomarkers to cause disease (shown in Fig. 4C) revealed that CDK4 and CDK6 had the highest scores, equivalent to 84.53% and 78.44%, respectively. E-cadherin scored 70.62%, followed by TNF-α with a score of 70.41%, and finally TGF-β with 42.49%. The score for ANRIL and miR-126-5p was not detected by the tool used. This score reflects the principle that a variant in the original genes of those transcripts with high mutation intolerance is more likely to cause CAD.
3.7 Enrichment Analysis of the Circulating Vasculature-Related Transcripts in CAD
All the functions and pathways targeted by the seven circulating vasculature-related transcripts under study in CAD were identified via enrichment analysis based on annotated gene targets in GO.
Our analysis found the biological processes previously known to be involved in CAD distinctly enriched. The top 10 enriched pathways were associated with response to organic substance regulation of cell population proliferation, positive regulation of fibroblast proliferation, positive regulation of mononuclear cell migration, regulation of gene expression, negative regulation of production of miRNAs involved in gene silencing by miRNA, negative regulation of cell–cell adhesion, positive regulation of protein kinase B signaling, negative regulation of myoblast differentiation, and positive regulation of ERK1 and ERK2 cascade as represented in Fig. 5A according to their matching scores.
Function and pathway enrichment analysis for the seven circulating vasculature-related transcripts under study according to matching score. A Top 10 biological processes. B Top 10 molecular functions. C Top 10 cellular components. D Top 10 pathways. The color scale presented from red to blue where blue reflects higher scores, and red indicates lower ones. All scores were calculated using GeneAnalytics tool (https://geneanalytics.genecards.org)
Considering molecular function enrichment analysis shown in Fig. 5B, the top ten functions were RNA polymerase II CTD heptapeptide repeat kinase activity, cyclin-dependent protein serine/threonine kinase activity, cyclin binding, FBXO family protein binding, type III transforming growth factor (TGF)-β receptor binding, cytokine activity, type II TGF-β receptor binding, type I TGF-β receptor binding, γ-catenin binding, and GTPase activating protein binding according to their matching scores.
Moreover, we revealed that the top ten cellular components for the seven circulating vasculature-related genes under study were cyclin-dependent protein kinase holoenzyme complex, mediator complex, cyclin D2-CDK4 complex, Flotillin complex, apical junction complex, phagocytic cup, catenin complex, cell surface, cytoplasmic side of the plasma membrane, and extracellular space according to their matching scores shown in Fig. 5C.
Finally, enrichment of specific pathways for the seven circulating vasculature-related transcripts under study revealed pathways relevant to mitotic G1-G1/S phases, developmental lineage pathway of dendritic cells, cyclins and cell cycle regulation, Sertoli-Sertoli cell junction dynamics, cell cycle role of SCF complex in cell cycle regulation, EV release from cardiac cells and their functional effects, T-cell antigen receptor (TCR) signaling pathway, RXR and RAR heterodimerization with other nuclear receptors, and ERK signaling according to their matching scores (Fig. 5D).
3.8 Regulatory Network Construction for the Vasculature-Related miRNA–mRNA Targets
Network analysis revealed a strong relationship between five of the circulating vasculature-related transcripts under study and their target miRNAs and miR-126-5p, but not for ANRIL. The miRNA-target gene network comprised the six circulating vasculature-related transcripts under study where it initially revealed 407 targets (Supplementary file 1, see electronic supplementary material) and then filtered to select only strongly validated targets that revealed a final set of 82 targets (11 mRNA and 71 miRNAs) using miRTargetLink 2.0 (Fig. 6). Also, we constructed a miRNA-target gene network comprising the four top biomarkers according to ROC curve analysis, which were CDK4, E-cadherin, miR126-5p, and ANRIL in descending order (Fig. 6). The network revealed 128 targets that were then filtered to select only strongly validated targets, with a final set of 50 targets (11 mRNA and 39 miRNA).
Vasculature-related miRNAs-target gene network analysis for the circulating vasculature-related transcripts under study in CAD. The targets used were only the strongly validated miRNAs in the literature using ‘miRTargetLink 2.0 (https://ccb-compute.cs.uni-saarland.de/mirtargetlink2)’. A The seven circulating vasculature transcripts’ miRNA-target gene network. B The top four circulating vasculature transcripts according to ROC curve analysis miRNA-target gene network. CAD coronary artery disease, ROC receiver operating characteristic
4 Discussion
Blood transcriptome analysis via investigating different types of RNA transcripts in CAD patients opens new frontiers for the non-invasive diagnosis of CAD. These RNA transcripts are seen as potential biomarkers for early detection of CAD in high-risk individuals or for improving the diagnosis of CAD in areas where invasive or advanced imaging methods are unavailable. They can also potentially serve as therapeutic targets and/or markers to monitor the suitability and efficacy of new or existing therapies [43].
Previously, lncRNA-ANRIL, miR-126-5p, CDK6, TNF-α, and TGF-β expression in CAD patients were studied either separately or in a maximum combination of two [44,45,46,47,48,49]. In the present study, the preceding transcripts, in addition to CDK4 and E-cadherin, are examined, aiming to elucidate their role as potential biomarkers for CAD.
The study results showed upregulation of ANRIL, miR-126-5p, CDK4, CDK6, E-cadherin, and TNF-α expression levels, while TGF-β expression was downregulated in the plasma of CAD patients. Upregulated expression of ANRIL, miR-126-5p, CDK6, TNF-α, and downregulation of TGF-β in CAD is in line with previous studies [44,45,46,47,48]. Nevertheless, downregulation of ANRIL in CAD patients was also documented in other studies [49,50,51]. This discrepancy could be attributed to different Chr9p21 locus genotypes as the Chr9p21 rs10757278/rs2383206-G risk allele associated with increased ANRIL expression is prevalent in Egyptian CAD patients [52, 53]. In fact, and to our knowledge, this study addressed for the first time the upregulated expression level of CDK4 and E-cadherin in CAD patients. All the transcripts showed a significant correlation with CAD. CDK4, E-cadherin, miR-126-5p, and ANRIL showed the highest predictive values for CAD, while the combined predictive values of TNF-α–TGF-β and CDK4–CDK6–E-cadherin showed 100% discrimination between CAD and controls.
Previous research stated that either the ncRNAs or the gene product of the transcripts under study participate in the pathogenesis of atherosclerosis, plaque formation, and stability [47, 54,55,56,57,58,59], mainly by inducing vascular endothelial cells (VEC) injury, and promotion of VSMC proliferation, migration, as well as the transformation from the contractile to the synthetic phenotype [60].
Increasing ANRIL expression in VECs participates in inflammation and accelerates endothelial injury via the TNF-α-NF-κB-ANRIL/YY1-IL6/8 signaling pathway [54]. ANRIL regulated the expression of VEGF, an important angiogenic factor reported to induce the migration and proliferation of endothelial cells, enhanced vascular permeability, and accelerated endothelial injury [61].
ANRIL facilitates the adhesion and proliferation of mononuclear cells to injured endothelial cells, thus promoting its infiltration into the endothelial layer and its differentiation to macrophages, indorsing extracellular matrix (ECM) degradation, and ultimately leading to superficial erosion or rupture of atheromatic plaques [52].
Moreover, ANRIL promotes VSMC proliferation, migration, and apoptosis via regulating CDK4 and CDK6 kinase activity. It also inhibits CDK4/6 inhibitors p16INK4a and p15INK4b from the CDKN2A/CDKN2b locus by recruiting PRC1/PRC2/EZH2 protein complex, and activation of DNA methyltransferase1 (DNMT1), resulting in hypermethylation and inactivation of CDKN2A/CDKN2b genes [62,63,64].
CDK4 and CDK6 are master regulators of the cell cycle. Their activity is restricted to the G1-S phase, controlled by the regulatory subunits D-type cyclins and CDK inhibitor p16INK4a [65, 66].
Inhibition of p16ink4a and p15INK4b leads to the assembly of the functioning D1-CDK4/6 complexes, which phosphorylates and inactivates the cell cycle suppressor proteins Rb (Retinoblastoma protein) family (p105Rb, p107, p130Rb2) [65]. The phosphorylated RB releases the transcription factor E2F1 from the Rb/E2F complexes, activating the transcription of genes whose products are required for DNA synthesis and S-phase progression, thereby allowing VSMC proliferation [67, 68].
MiR-126-5p, a flow-responsive miRNA, has a variable role in coronary atherogenesis depending on the stage of the disease. Upregulation of this miRNA may be athero-protective in the early phases of plaque development by promoting endothelial cell proliferation via targeting the negative regulator of EC proliferation, Dlk1 (delta-like 1 homolog) [69]. The high miR-126-5p expression could serve as a promoter of plaque destabilization later [45]. MiR-126-5p secreted by the EC has a paracrine effect on VSMC via modulating its gene expression and function toward atherogenic phenotype [70].
E-cadherin, a cell surface adhesion molecule, is implicated in mechanical adhesion, regulation of cell division, migration, differentiation, and apoptosis of VSMCs [71]. Increased expression of E-cadherin is coincident with VSMC proliferation. E-cadherin regulates cell proliferation by adjusting the expression of the cell cycle activator cyclin D1, mediated by the E-cadherin/catenin/TCF signaling pathway [57, 72].
TNF-α, a pro-atherosclerotic cytokine produced by several cells such as macrophages, T-lymphocytes, and dendritic cells, is known to be activated in atherogenesis. TNF-α boosts endothelial dysfunction, promoting VSMC proliferation and switching to a synthetic phenotype, which increases the production of ECM, and nucleic acid-enriched exosomes [73, 74].
Conversely, TGF-β inhibits migration and proliferation of VSMC by targeting signal transducer and activator of transcription 3 (STAT3) and NF-κB pathways [75].
TGF-β stimulates endothelial migration and proliferation at low concentrations but inhibits both at higher concentrations [76] by regulating the expression of cell cycle inhibitors p16INK4A and p15INK4B. These proteins are expressed at low levels in proliferating cells and are rapidly induced by TGF-β [77].
In the present study, upregulation and positive correlation between ANRIL and TNF-α agree with previously described research findings [49].
However, the expression pattern and correlations between the studied transcripts could be attributed to dyslipidemia and smoking, as nearly all RNAs, except for CDK6 and TGF-β, show significant correlations with both, creating a link between these risk factors and CAD via atherosclerosis. Smoking and chronic dyslipidemia led to enhanced oxidative stress, resulting in increased ox-LDL [70, 78]. Ox-LDL significantly promotes ANRIL expression in human aortic smooth muscle cells [44]. A high level of LDL cholesterol significantly increased miR-126-5p in patients with risk factors for CAD [79].
Ox-LDL stimulated monocyte/macrophage release of TNF-α through the NF-κB signaling pathway [80]. TNF-α induces the expression of ANRIL, CDK4, and CDK6 through the TNF-α/NF-kb signaling pathway [81, 82].
In addition, ox-LDL downregulates TGF-β expression by enhancing TGF‐β-producing-Treg (Regulatory T) cell apoptosis through the Fas/FasL pathway [83].
The TGF-β/SMADs pathway regulates the expression of E-cadherin through induction of the transcriptional repressors SNAIL, ZEB, and TWIST. These repressors bind to the promoter region of the E-cadherin gene and suppress its transcription [84].
Hence, it could be concluded that ox-LDL indirectly enhances the expression of E-cadherin by downregulating TGF-β.
Regarding other risk factors, ANRIL, CDK4, CDK6, and E-cadherin showed a positive relationship with age. ANRIL correlation with age is in line with the previous study [85]. Furthermore, ANRIL is positively correlated with CDK4, CDK6, and E-cadherin in agreement with a previous study [86], explaining the correlation of CDK4, CDK6, and E-cadherin with age.
On the other hand, a positive correlation between ANRIL and TGF-β could be related to a positive correlation between TNF-α and TGF-β, which could be attributed to the balanced expression of pro-and anti-inflammatory cytokines, characteristic of the chronic, low-degree inflammation in an atherosclerotic arterial wall [87].
According to Cesari et al., circulating levels of inflammatory markers could predict future cardiovascular events, both in the general population and in adults with a known CVD. The present study combined predictive ROC curve analysis of TNF-α and TGF-β, showing 100% prediction of CAD [88]. This result is supported by the analysis of connection strength between each transcript and CAD. TNF-α and TGF-β show the highest percentage of gene variants implicated in CAD, probably due to their roles in mediating the inflammatory processes related to atherosclerosis and CAD pathogenesis [87].
Anatomical disease categorization analysis showed that the transcripts studied are related to CAD, mainly myocardial infarction, aortic aneurysm, and essential hypertension, whereas CDK4, CDK6, and E-cadherin had the highest scores related to the likelihood of causing disease [89, 90], thus explaining the high combined predictive value of these transcripts for CAD.
Interestingly, the examined transcripts showed high categorization to lipoprotein quantitative trait locus. Further studies are needed to elucidate the association between them.
Function and pathway enrichment analysis of the circulating vasculature transcripts in the present study were associated with response to an organic substance, regulation of different cell population proliferation, regulation of gene expression, and mononuclear cell migration, consistent with many previous results [19, 20, 91,92,93,94,95].
Considering the analysis of cellular components and molecular function of the studied transcripts, it indicated their confinement to components and molecules mediating and executing cell proliferation, namely CDK4/6 cyclin-dependent protein kinase holoenzyme complex mediating RNA polymerase II CTD heptapeptide repeat kinase activity, mediator complex which interacts with and transmits signals from transcription factors to RNA polymerase II, cyclin binding and formation of cyclin D2-CDK4 complex arbitrating serine/threonine kinase activity, which phosphorylates and inactivates RBp, which is also supported by previous results [68, 96,97,98].
Finally, enrichment of specific pathways for the seven circulating vasculature-related genes revealed pathways relevant to mitotic G1-G1/S phases, cyclins and cell cycle regulation, and dendritic cell developmental lineage pathway TNF-α and TGF-β that were also previously reported [65, 99].
In the present study, we constructed a vasculature-related miRNA‐mRNA regulatory network which identified a strong relationship between miR-126-5p, CDK4, and E-cadherin, with 50 targets (11 mRNA and 39 miRNAs). Further research is needed to validate the role of the identified transcript targets as CAD biomarkers.
The primary limitation of the current study is that it was a mono-center study; further studies with a larger sample size from multicentric sources are advised to confirm the study results on a broader scale, however this study is considered a preliminary proof-of-concept study and further larger studies will be done. The scarcity of research studies investigating CDK4 and E-cadherin as biomarkers of CAD established an opportunity to highlight this literature gap and to emphasize the need for further research in this area. A further limitation is that not all the control subjects were assessed using echocardiography. Only subjects aged over 45 years were assessed to ensure they are free from any CAD signs. Furthermore, as the CAD patients in this study were under treatment, there is a possibility that these drugs may have interfered with the expression of the evaluated transcripts. Finally, it is vital that functional in vitro and in vivo studies are performed to validate the study results and emphasize these vasculature-related transcripts as CAD biomarkers.
References
Murray C, Lozano R, Davis A, Liang X, Zhou M, Vollset S, Ozgoren AA, Abdalla S, Abd-Allah F, Aziz M, Abera SF. Global, regional, and national age-sex specific all-cause and cause-specific mortality for 240 causes of death, 1990–2013: a systematic analysis for the Global Burden of Disease Study 2013. Lancet. 2015;385(9963):117–71.
McPherson R, Tybjaerg-Hansen A. Genetics of coronary artery disease. Circ Res. 2016;118(4):564–78.
Vilahur G, Badimon JJ, Bugiardini R, Badimon L. Perspectives: the burden of cardiovascular risk factors and coronary heart disease in Europe and worldwide. European Heart Journal Supplements. 2014;16(suppl_A):A7–11.
El-Attar EA, Helmy Elkaffas RM, Aglan SA, Naga IS, Nabil A, Abdallah HY. Genomics in Egypt: current status and future aspects. Front Genet. 2022;13: 797465. https://doi.org/10.3389/fgene.2022.797465.
Bentzon JF, Otsuka F, Virmani R, Falk E. Mechanisms of plaque formation and rupture. Circ Res. 2014;114(12):1852–66.
Soehnlein O, Libby P. Targeting inflammation in atherosclerosis—from experimental insights to the clinic. Nat Rev Drug Discov. 2021;20(8):589–610.
Falk E. Pathogenesis of atherosclerosis. J Am Coll Cardiol. 2006;47:C7–12. https://doi.org/10.1016/j.jacc.2005.09.068.
Beck-Joseph J, Lehoux S. molecular interactions between vascular smooth muscle cells and macrophages in atherosclerosis. Front Cardiovasc Med. 2021;8.
Ilhan F, Kalkanli ST. Atherosclerosis and the role of immune cells. WJCC. 2015;3(4):345.
Grootaert MO, Bennett MR. Vascular smooth muscle cells in atherosclerosis: time for a re-assessment. Cardiovasc Res. 2021;117(11):2326–39.
Lalonde S, Codina-Fauteux VA, de Bellefon SM, Leblanc F, Beaudoin M, Simon MM, Dali R, Kwan T, Lo KS, Pastinen T, Lettre G. Integrative analysis of vascular endothelial cell genomic features identifies AIDA as a coronary artery disease candidate gene. Genome Biol. 2019;20(1):1–3.
Sakamoto A, Cornelissen A, Sato Y, Mori M, Kawakami R, Kawai K, Ghosh SK, Xu W, Abebe BG, Dikongue A, Kolodgie FD. Vulnerable plaque in patients with acute coronary syndrome: identification, importance, and management. US Cardiol Rev. 2022.
Elashoff MR, Wingrove JA, Beineke P, Daniels SE, Tingley WG, Rosenberg S, Voros S, Kraus WE, Ginsburg GS, Schwartz RS, Ellis SG. Development of a blood-based gene expression algorithm for assessment of obstructive coronary artery disease in non-diabetic patients. BMC Med Genom. 2011;4(1):1–4.
Huan T, Zhang B, Wang Z, Joehanes R, Zhu J, Johnson AD, Ying S, Munson PJ, Raghavachari N, Wang R, Liu P. A systems biology framework identifies molecular underpinnings of coronary heart disease. Arterioscler Thromb Vasc Biol. 2013;33(6):1427–34.
Mohler ER III, Sarov-Blat L, Shi Y, Hamamdzic D, Zalewski A, MacPhee C, Llano R, Pelchovitz D, Mainigi SK, Osman H, Hallman T. Site-specific atherogenic gene expression correlates with subsequent variable lesion development in coronary and peripheral vasculature. Arterioscler Thromb Vasc Biol. 2008;28(5):850–5.
Sluimer JC, Kisters N, Cleutjens KB, Volger OL, Horrevoets AJ, van den Akker LH, Bijnens AP, Daemen MJ. Dead or alive: gene expression profiles of advanced atherosclerotic plaques from autopsy and surgery. Physiol Genom. 2007;30(3):335–41.
Aziz H, Zaas A, Ginsburg GS. Peripheral blood gene expression profiling for cardiovascular disease assessment. Genom Med. 2007;1(3):105–12.
Morris KV, Mattick JS. The rise of regulatory RNA. Nat Rev Genet. 2014;15(6):423–37.
Song C, Qi Y, Zhang J, Guo C, Yuan C. CDKN2B-AS1: an indispensable long non-coding RNA in multiple diseases. Curr Pharm Des. 2020;26(41):5335–46.
Razeghian-Jahromi I, Karimi Akhormeh A, Zibaeenezhad MJ. The role of ANRIL in atherosclerosis. Dis Markers. 2022;9:2022.
Holdt LM, Beutner F, Scholz M, Gielen S, Gäbel G, Bergert H, Schuler G, Thiery J, Teupser D. ANRIL expression is associated with atherosclerosis risk at chromosome 9p21. Arterioscler Thromb Vasc Biol. 2010;30(3):620–7.
Yari M, Bitarafan S, Broumand MA, Fazeli Z, Rahimi M, Ghaderian SM, Mirfakhraie R, Omrani MD. Association between long noncoding RNA ANRIL expression variants and susceptibility to coronary artery disease. Int J Mol Cell Med. 2018;7(1):1.
O’Brien J, Hayder H, Zayed Y, Peng C. Overview of microRNA biogenesis, mechanisms of actions, and circulation. Front Endocrinol. 2018;3(9):402.
Latronico MV, Condorelli G. MicroRNAs and cardiac pathology. Nat Rev Cardiol. 2009;6(6):418–29.
Yu B, Jiang Y, Wang X, Wang S. An integrated hypothesis for miR-126 in vascular disease. Med Res Arch. 2020;8(5).
Abdallah HY, Hassan R, Fareed A. et al. Identification of a circulating microRNAs biomarker panel for non-invasive diagnosis of coronary artery disease: case–control study. BMC Cardiovasc Disord. 2022; 22:286. https://doi.org/10.1186/s12872-022-02711-9
Russo F, Fiscon G, Conte F, Rizzo M, Paci P, Pellegrini M. Interplay between long noncoding RNAs and MicroRNAs in cancer. In: Computational cell biology. New York: Humana Press; 2018. p. 75–92.
Morceau F, Chateauvieux S, Gaigneaux A, Dicato M, Diederich M. Long and short non-coding RNAs as regulators of hematopoietic differentiation. Int J Mol Sci. 2013;14(7):14744–70.
Ratti M, Lampis A, Ghidini M, Salati M, Mirchev MB, Valeri N, Hahne JC. MicroRNAs (miRNAs) and long non-coding RNAs (lncRNAs) as new tools for cancer therapy: first steps from bench to bedside. Targ Oncol. 2020;15(3):261–78.
Yamamura S, Imai-Sumida M, Tanaka Y, Dahiya R. Interaction and cross-talk between non-coding RNAs. Cell Mol Life Sci. 2018;75(3):467–84.
Huang Z, Shi J, Gao Y, Cui C, Zhang S, Li J, Zhou Y, Cui Q. HMDD v3. 0: a database for experimentally supported human microRNA–disease associations. Nucleic Acids Res. 2019;47(D1):D1013–7.
Jiang Q, Wang Y, Hao Y, Juan L, Teng M, Zhang X, Li M, Wang G, Liu Y. miR2Disease: a manually curated database for microRNA deregulation in human disease. Nucleic Acids Res. 2009;37(suppl_1):D98–104.
Bao Z, Yang Z, Huang Z, Zhou Y, Cui Q, Dong D. LncRNADisease 2.0: an updated database of long non-coding RNA-associated diseases. Nucleic Acids Res. 2019;47(D1):D1034–7.
Karagkouni D, Paraskevopoulou MD, Tastsoglou S, Skoufos G, Karavangeli A, Pierros V, Zacharopoulou E, Hatzigeorgiou AG. DIANA-LncBase v3: indexing experimentally supported miRNA targets on non-coding transcripts. Nucleic Acids Res. 2020;48(D1):D101–10.
Ghafouri-Fard S, Gholipour M, Taheri M. The emerging role of long non-coding RNAs and circular RNAs in coronary artery disease. Front Cardiovasc Med. 2021;23(8): 632393.
Zhang Y, Zhang L, Wang Y, Ding H, Xue S, Qi H, Li P. MicroRNAs or long noncoding RNAs in diagnosis and prognosis of coronary artery disease. Aging Dis. 2019;10(2):353.
Cardona-Monzonís A, García-Giménez JL, Mena-Mollá S, Pareja-Galeano H, Guía-Galipienso FD, Lippi G, Pallardó FV, Sanchis-Gomar F. Non-coding RNAs and coronary artery disease. Non-coding RNAs Cardiovasc Dis. 2020:273–85.
Ben-Ari Fuchs S, Lieder I, Stelzer G, Mazor Y, Buzhor E, Kaplan S, Bogoch Y, Plaschkes I, Shitrit A, Rappaport N, Kohn A. GeneAnalytics: an integrative gene set analysis tool for next generation sequencing, RNAseq and microarray data. Omics. 2016;20(3):139–51.
Stelzer G, Plaschkes I, Oz-Levi D, Alkelai A, Olender T, Zimmerman S, Twik M, Belinky F, Fishilevich S, Nudel R, Guan-Golan Y. VarElect: the phenotype-based variation prioritizer of the GeneCards Suite. BMC Genom. 2016;17(2):195–206.
Kern F, Aparicio-Puerta E, Li Y, Fehlmann T, Kehl T, Wagner V, Ray K, Ludwig N, Lenhof HP, Meese E, Keller A. miRTargetLink 2.0—interactive miRNA target gene and target pathway networks. Nucleic Acids Res. 2021;49(W1):W409–16.
Faul F, Erdfelder E, Lang AG, Buchner A. G* Power 3: A flexible statistical power analysis program for the social, behavioral, and biomedical sciences. Behav Res Methods. 2007;39(2):175–91.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25(4):402–8. https://doi.org/10.1006/meth.2001.1262.
McCaffrey TA, Toma I, Yang Z, Katz R, Reiner J, Mazhari R, Shah P, Tackett M, Jones D, Jepson T, Falk Z. RNA sequencing of blood in coronary artery disease: involvement of regulatory T cell imbalance. BMC Med Genom. 2021;14(1):1–20.
Zhang C, Ge S, Gong W, Xu J, Guo Z, Liu Z, Gao X, Wei X, Ge S. LncRNA ANRIL acts as a modular scaffold of WDR5 and HDAC3 complexes and promotes alteration of the vascular smooth muscle cell phenotype. Cell Death Dis. 2020;11(6):1–3.
Knoka E, Trusinskis K, Mazule M, Briede I, Crawford W, Jegere S, Kumsars I, Narbute I, Sondore D, Lejnieks A, Erglis A. Circulating plasma microRNA-126, microRNA-145, and microRNA-155 and their association with atherosclerotic plaque characteristics. J Clin Transl Res. 2020;5(2):60.
Witten A, Martens L, Schäfer AC, Troidl C, Pankuweit S, Vlacil AK, Oberoi R, Schieffer B, Grote K, Stoll M, Markus B. Monocyte subpopulation profiling indicates CDK6-derived cell differentiation and identifies subpopulation-specific miRNA expression sets in acute and stable coronary artery disease. Sci Rep. 2022;12(1):1–3.
Subirana I, Fitó M, Diaz O, Vila J, Francés A, Delpon E, Sanchis J, Elosua R, Muñoz-Aguayo D, Dégano IR, Marrugat J. Prediction of coronary disease incidence by biomarkers of inflammation, oxidation, and metabolism. Sci Rep. 2018;8(1):1–7.
Dabek J, Mazurek U, Gasior Z, Wilczok T, Kulach A, Kucia-Kuzma S. Transcriptional activity of genes encoding Transforming Growth Factor β and its receptors in peripheral blood mononuclear cells from patients with acute coronary syndromes. Int J Cardiol. 2006;111(2):275–9.
Hu Y, Hu J. Diagnostic value of circulating lncRNA ANRIL and its correlation with coronary artery disease parameters. Braz J Med Biol Res. 2019;12:52.
Liu ZF, Hu WW, Li R, Gao Y, Yan LL, Su N. Expression of lncRNA-ANRIL in patients with coronary heart disease before and after treatment and its short-term prognosis predictive value. Eur Rev Med Pharmacol Sci. 2020;24(1):376–84.
Li J, Chen J, Zhang F, Li J, An S, Cheng M, Li J. LncRNA CDKN2B-AS1 hinders the proliferation and facilitates apoptosis of ox-LDL-induced vascular smooth muscle cells via the ceRNA network of CDKN2B-AS1/miR-126-5p/PTPN7. Int J Cardiol. 2021;1(340):79–87.
Holdt LM, Hoffmann S, Sass K, Langenberger D, Scholz M, Krohn K, Finstermeier K, Stahringer A, Wilfert W, Beutner F, Gielen S. Alu elements in ANRIL non-coding RNA at chromosome 9p21 modulate atherogenic cell functions through trans-regulation of gene networks. PLoS Genet. 2013;9(7): e1003588.
Shendy HA, Hassanein SI, Gad MZ. “Desert” gene (Chr9p21) variants as novel markers for coronary artery disease. Anatol J Cardiol. 2017;18(2):84.
Chen L, Qu H, Guo M, et al. ANRIL and atherosclerosis. J Clin Pharm Ther. 2020;45:240–8.
Dzau VJ, Braun-Dullaeus RC, Sedding DG. Vascular proliferation and atherosclerosis: new perspectives and therapeutic strategies. Nat Med. 2002;8(11):1249–56.
Boon RA, Dimmeler S. Commentary on cutting edge science. Arterioscler Thromb Vasc Biol. 2014;34:e15–6.
Bedel A, Nègre-Salvayre A, Heeneman S, Grazide MH, Thiers JC, Salvayre R, Maupas-Schwalm F. E-cadherin/β-catenin/T-cell factor pathway is involved in smooth muscle cell proliferation elicited by oxidized low-density lipoprotein. Circ Res. 2008;103(7):694–701.
Toma I, McCaffrey TA. Transforming growth factor-β and atherosclerosis: interwoven atherogenic and atheroprotective aspects. Cell Tissue Res. 2012;347(1):155–75.
Wang M, Li C, Cai T, Zhang A, Cao J, Xin H. Circ_CHFR promotes PDGF-BB-induced proliferation, invasion and migration in VSMCs via miR-149-5p/NRP2 axis. J Cardiovasc Pharmacol. 2021.
Xu T, Jia J, Xu N, Ye C, Zheng F, Yuan Y, Zhu GQ, Zhan YY. Apelin receptor upregulation in spontaneously hypertensive rat contributes to the enhanced vascular smooth muscle cell proliferation by activating autophagy. Ann Transl Med. 2021;9(8).
Zhang B, Wang D, Ji TF, Shi L, Yu JL. Overexpression of lncRNA ANRIL up-regulates VEGF expression and promotes angiogenesis of diabetes mellitus combined with cerebral infarction by activating NF-κB signaling pathway in a rat model. Oncotarget. 2017;8(10):17347.
Almontashiri NA. The 9p21. 3 risk locus for coronary artery disease: a 10-year search for its mechanism. J Taibah Univ Med Sci. 2017;12(3):199–204.
Zhuang J, Peng W, Li H, Wang W, Wei Y, Li W, Xu Y. Methylation of p15INK4b and expression of ANRIL on chromosome 9p21 are associated with coronary artery disease.
Witkiewicz AK, Knudsen KE, Dicker AP, Knudsen ES. The meaning of p16ink4a expression in tumors: functional significance, clinical associations and future developments. Cell Cycle. 2011;10(15):2497–503.
Baker SJ, Reddy EP. CDK4: a key player in the cell cycle, development, and cancer. Genes Cancer. 2012;3(11–12):658–69.
Wang M, Fu Z, Wu J, Zhang J, Jiang L, Khazan B, Telljohann R, Zhao M, Krug AW, Pikilidou M, Monticone RE. MFG-E8 activates proliferation of vascular smooth muscle cells via integrin signaling. Aging Cell. 2012;11(3):500–8.
Narasimha AM, Kaulich M, Shapiro GS, Choi YJ, Sicinski P, Dowdy SF. Cyclin D activates the Rb tumor suppressor by mono-phosphorylation. Elife. 2014;3.
Harbour JW, Luo RX, Dei Santi A, Postigo AA, Dean DC. Cdk phosphorylation triggers sequential intramolecular interactions that progressively block Rb functions as cells move through G1. Cell. 1999;98(6):859–69.
Schober A, Nazari-Jahantigh M, Wei Y, Bidzhekov K, Gremse F, Grommes J, Megens RT, Heyll K, Noels H, Hristov M, Wang S. MicroRNA-126-5p promotes endothelial proliferation and limits atherosclerosis by suppressing Dlk1. Nat Med. 2014;20(4):368–76.
Zhou MS, Chadipiralla K, Mendez AJ, Jaimes EA, Silverstein RL, Webster K, Raij L. Nicotine potentiates proatherogenic effects of oxLDL by stimulating and upregulating macrophage CD36 signaling. Am J Physiol Heart Circul Physiol. 2013;305(4):H563–74.
Stepniak E, Radice GL, Vasioukhin V. Adhesive and signaling functions of cadherins and catenins in vertebrate development. Cold Spring Harb Perspect Biol. 2009;1(5): a002949.
Frismantiene A, Philippova M, Erne P, Resink TJ. Cadherins in vascular smooth muscle cell (patho) biology: Quid nos scimus? Cell Signal. 2018;1(45):23–42.
Kong P, Cui ZY, Huang XF, Zhang DD, Guo RJ, Han M. Inflammation and atherosclerosis: signaling pathways and therapeutic intervention. Signal Transduct Target Ther. 2022;7(1):1–24.
Ait-Oufella H, Taleb S, Mallat Z, Tedgui A. Recent advances on the role of cytokines in atherosclerosis. Arterioscler Thromb Vasc Biol. 2011;31(5):969–79.
Gao P, Wu W, Ye J, Lu YW, Adam AP, Singer HA, Long X. Transforming growth factor β1 suppresses proinflammatory gene program independent of its regulation on vascular smooth muscle differentiation and autophagy. Cell Signal. 2018;1(50):160–70.
Bobik A. Transforming growth factor-βs and vascular disorders. Arterioscler Thromb Vasc Biol. 2006;26(8):1712–20.
Blain S. Switching cyclin D-Cdk4 kinase activity on and off. Cell Cycle. 2008;7(7):892–8.
Hasheminasabgorji E, Jha JC. Dyslipidemia, diabetes and atherosclerosis: role of inflammation and ROS-redox-sensitive factors. Biomedicines. 2021;9(11):1602.
Sun X, Zhang M, Sanagawa A, Mori C, Ito S, Iwaki S, Satoh H, Fujii S. Circulating microRNA-126 in patients with coronary artery disease: correlation with LDL cholesterol. Thromb J. 2012;10(1):1–5.
Hodes GE, Kana V, Menard C, Merad M, Russo SJ. Neuroimmune mechanisms of depression. Nat Neurosci. 2015;18(10):1386–93.
Zhou X, Han X, Wittfeldt A, Sun J, Liu C, Wang X, Gan LM, Cao H, Liang Z. Long non-coding RNA ANRIL regulates inflammatory responses as a novel component of NF-κB pathway. RNA Biol. 2016;13(1):98–108.
Wu X, Wu MY, Jiang M, Zhi Q, Bian X, Xu MD, Gong FR, Hou J, Tao M, Shou LM, Duan W. TNF-α sensitizes chemotherapy and radiotherapy against breast cancer cells. Cancer Cell Int. 2017;17(1):1–2.
Kuan R, Agrawal DK, Thankam FG. Treg cells in atherosclerosis. Mol Biol Rep. 2021;48(5):4897–910.
Lamouille S, Xu J, Derynck R. Molecular mechanisms of epithelial–mesenchymal transition. Nat Rev Mol Cell Biol. 2014;15(3):178–96.
Vausort M, Wagner DR, Devaux Y. Long noncoding RNAs in patients with acute myocardial infarction. Circ Res. 2014;115(7):668–77.
Zhang Y, Alexander PB, Wang XF. TGF-β family signaling in the control of cell proliferation and survival. Cold Spring Harb Perspect Biol. 2017;9(4): a022145.
Bäck M, Yurdagul A, Tabas I, et al. Inflammation and its resolution in atherosclerosis: mediators and therapeutic opportunities. Nat Rev Cardio. 2019;16(7):389.
Cesari M, Penninx BW, Newman AB, Kritchevsky SB, Nicklas BJ, Sutton-Tyrrell K, Rubin SM, Ding J, Simonsick EM, Harris TB, Pahor M. Inflammatory markers and onset of cardiovascular events: results from the Health ABC study. Circulation. 2003;108(19):2317–22.
Yasuno K, Bilguvar K, Bijlenga P, Low SK, Krischek B, Auburger G, Simon M, Krex D, Arlier Z, Nayak N, Ruigrok YM. Genome-wide association study of intracranial aneurysm identifies three new risk loci. Nat Genet. 2010;42(5):420–5.
Ye S, Willeit J, Kronenberg F, Xu Q, Kiechl S. Association of genetic variation on chromosome 9p21 with susceptibility and progression of atherosclerosis: a population-based, prospective study. J Am Coll Cardiol. 2008;52(5):378–84.
McCartney A, Migliaccio I, Bonechi M, Biagioni C, Romagnoli D, De Luca F, Galardi F, Risi E, De Santo I, Benelli M, Malorni L. Mechanisms of resistance to CDK4/6 inhibitors: potential implications and biomarkers for clinical practice. Front Oncol. 2019;23(9):666.
Roda G, Jharap B, Neeraj N, Colombel JF. Loss of response to anti-TNFs: definition, epidemiology, and management. Clin Transl Gastroenterol. 2016;7(1): e135.
Huynh LK, Hipolito CJ, Ten Dijke P. A perspective on the development of TGF-β inhibitors for cancer treatment. Biomolecules. 2019;9(11):743.
Li X, Liu C, Zhao X, Wang R, Gu N, Shen H, Li X, Wang L, Li C. Effects of CDK6 regulated by miR-298 on proliferation and apoptosis of thyroid cancer cells. Oncol Lett. 2020;19(4):2909–15.
Georgopoulos NT, Kirkwood LA, Walker DC, Southgate J. Differential regulation of growth-promoting signalling pathways by E-cadherin. PLoS ONE. 2010;5(10): e13621.
Kõivomägi M, Swaffer MP, Turner JJ, Marinov G, Skotheim JM. G1 cyclin–Cdk promotes cell cycle entry through localized phosphorylation of RNA polymerase II. Science. 2021;374(6565):347–51.
Allen BL, Taatjes DJ. The Mediator complex: a central integrator of transcription. Nat Rev Mol Cell Biol. 2015;16(3):155–66.
Hu X, Lou T, Yuan C, Wang Y, Tu X, Wang Y, Zhang T. Effects of lncRNA ANRIL-knockdown on the proliferation, apoptosis and cell cycle of gastric cancer cells. Oncol Lett. 2021;22(2):1–2.
Merad M, Sathe P, Helft J, Miller J, Mortha A. The dendritic cell lineage: ontogeny and function of dendritic cells and their subsets in the steady state and the inflamed setting. Annual review of immunology. 2013;31.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB). No other funding was received for conducting this study.
Conflicts of interest statement
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Ethics approval and consent to participate
Ethics Committee approval no. 4504 from the Suez Canal University, Faculty of Medicine, Ismailia, Egypt was received, and the study was conducted according to the Declaration of Helsinki’s guidelines. Informed consent was obtained from all individual participants included in the study.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Code availability
Not applicable.
Consent for publication
Not applicable.
Authors' Contributions
AHY, TEDLA and AM designed the study, Fareed A collected the clinical data, AAKK collected the patients’ samples, AHY, TEDLA, AM, AS, and AAKK carried out the experiments, AHY, TEDLA, AM, and FA analyzed and interpreted the patient data. All authors discussed the results, contributed to the final manuscript and approved it.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Abdallah, H.Y., Fareed, A., Abdelmaogood, A.K.K. et al. Introducing Circulating Vasculature-Related Transcripts as Biomarkers in Coronary Artery Disease. Mol Diagn Ther 27, 243–259 (2023). https://doi.org/10.1007/s40291-022-00622-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40291-022-00622-1