Abstract
Tuberculosis (TB) and HIV have been closely linked since the emergence of AIDS; TB enhances HIV replication by accelerating the natural evolution of HIV infection which is the leading cause of sickness and death of peoples living with HIV/AIDS. To improve their life the co-infected patients are started to take antiretroviral treatment as patient started to take ART it is common to measure CD4 and other clinical outcomes which is correlated with survival time. However, the separate analysis of such data does not handle the association between the longitudinal measured out come and time-to-event where the joint modeling does to obtain valid and efficient survival time. Joint modeling of longitudinally measured CD4 and time-to death to understand their association. Furthermore, the study identifies factors affecting the mean change in square root CD4 measurement over time and risk factors for the survival time of HIV/TB co-infected patients. The study consists of 254 HIV/TB co-infected patients who were 18 years old or older and who were on antiretroviral treatment follow up from first February 2009 to fist July 2014 in Jimma University Specialized Hospital, West Ethiopia. First, data were analyzed using linear mixed model and survival models separately. After having appropriate separate models using Akaki information criteria, different joint models employed with different random effects longitudinal model and different shared parameters association structure of survival model and compared with deviance information criteria score. The linear mixed model showed functional status, weight, linear time and quadratic time effects have significant effect on the mean change of CD4 measurement over time. The Cox and Weibull survival model showed base line weight, baseline smoking, separated marital status group and base line functional status have significant effect on hazard function of the survival time whereas the joint model showed subject specific base line value; subject specific linear and quadratic slopes of CD4 measurement of were significantly associated with the survival time of co-infected patient at 5% significance levels. The longitudinally measured CD4 count measurement marker process is significantly associated with time to death and subject specific quadratic slope growth of CD4 measurement, base line clinical stage IV and smoking is the high risk factors that lower the survival time of HIV/TB co-infected patients. Since the longitudinally measured CD4 measurement is correlated with survival time joint modeling are used to handle the associations between these two processes to obtain valid and efficient survival time.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
HIV/AIDS and tuberculosis co-infection present special challenges to the expansion and effectiveness of directly observed treatment short-course programs and the Stop TB Strategy. TB accounts for one-quarter of AIDS deaths worldwide and is one of the most common causes of morbidity in people living with HIV and AIDS. Currently, approximately 34 million people are infected with HIV, and at least one-third of them are also infected with TB [1].
Globally the number of TB patients who had been diagnosed with HIV status reached 2.1 million in 2010, equivalent to 34% of notified cases of TB. Of the 8.8 million incident cases globally an estimated 1.1 million (13%) were found to be co-infected with HIV [2]. Overall, the African region accounted for a staggering 82% of all new TB cases co-infected with HIV. Among the TB patients 46% of them are those living with HIV globally and 42% TB patients in the African region were living with HIV in 2010. Among the PLWHA enrolled in HIV care worldwide in 2010 the treatment success and death rates reported for HIV positive TB cases in 2009 were 72 and 20% respectively [2].
HIV infection is now the most common predictor of TB incidence and the other way round, TB is a common infection in sub-Saharan Africa. Thus, these countries continue taking the leading position in HIV/TB morbidity and mortality rate, where the TB epidemic is primarily driven by HIV infection. Ethiopia is one among these countries most heavily affected by HIV and TB co-infection [3,4,5].
Among 76 thousand TB was the causes of deaths in Ethiopia 30% were among HIV positive patients. However, WHO recommends different collaborative activities for HIV/TB co-infections where one is initiation of antiretroviral therapy in order to reduce the risks of death and HIV-related morbidities, or in improvement of quality of people living with HIV [6].
1.1 Joint Modeling Approaches
A common objective in longitudinal studies is to characterize the relationship between a longitudinal response process and time-to-event. Considerable recent interest has focused on so called joint models, where models for the event time distribution and longitudinal data are taken to depend on a common set of latent random effects. In the literature, precise statement of the underlying assumptions typically made for these models has been rare [7].
It is commonly found in the collection of medical longitudinal data that both repeated measures and time-to-event data are collected where both types of data are associated through unobserved random effects. Due to the association exists between these two process the joint models were developed to enable a more accurate method to model both processes simultaneously. Whereas the independent modeling of the longitudinal and time-to- event data can cause biased estimates [8, 9, 19].
Often the longitudinal studies produce two types of outcome, namely a set of longitudinal out come and the time-to-event of interest, such as death, development of a disease or dropout from the study. Two typical examples of this setting are HIV and cancer studies. In case of HIV patients who have been infected are monitored until they develop AIDS or die, and they are regularly measured for the condition of the immune system using markers such as the CD4 lymphocyte count or the estimated viral load. Similarly in cancer studies the event outcome is the death or metastasis and patients also provide longitudinal measurements of antibody levels or of other markers of carcinogenesis, such as the PSA levels for prostate cancer [11].
The joint modeling enables the simultaneous study of a longitudinal marker and a correlated time-to-event. The shared random-effect in the joint modeling define a mixed model for the longitudinal marker and a survival model for the time-to-event including characteristics of the mixed model as covariates received the main interest. Indeed, they extend naturally the survival model with time-dependent covariates and offer a flexible framework to explore the link between a longitudinal biomarker and a risk of event [12].
Joint modeling often assumes a proportional hazards model for the time to event and a linear mixed-effects model for the longitudinal data. Under this framework, different approaches have been proposed in the literature including some likelihood based methods with an assumption on the distribution of the random effects and that of the measurement errors Wulfsohn and Tsiatis [13] and Tsiatis et al. [14] also proposed a two-stage approach in which, based on an approximation to the hazard function for the event times, the usual partial likelihood for the Cox model can be used. In this approach, the observed covariate history is estimated using empirical Bayees methodology, which requires fitting as many mixed effects models as there are event times in the data set.
The approach that this study used to build a joint model is simultaneously modeling the longitudinal CD4 measurements and the time-to-death processes by linking those using shared random effects parameter model. In the proposed model, to characterize the longitudinal CD4 measurements a linear mixed effects model that incorporates patient specific CD4 intercept and slopes is used for the longitudinal sub-model while Cox PH model is used to describe the time-to-death survival data for the survival sub-model. Then, the two sub-models are linked through shared parameters [15], with different forms, since these random effects characterize the subject specific longitudinal process. Because, the standard maximum likelihood method involves integrating out the shared parameters from the log-likelihood function which is difficult when dealing with high dimensional variables [16], a Bayesian estimation procedure and a Markov chain Monte Carlo (MCMC) algorithm is used to fit the joint model. At last, the convergence of the Gibbs sampler is monitored by examining time series plots of the parameters over iteration.
2 Materials and Methods
2.1 Data Source
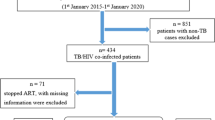
The data for the study was obtained from Jimma University Specialized Hospital from HIV and TB outpatient Clinic, South West of Ethiopia. All HIV/TB co-infected patients who were 18 years old and above taking ART at any time in between first February 2009 to first July 2014 having at least one CD4 count measurement were eligible for the analysis. Therefore, among 856 total co-infected patients during the time period 254 who fulfill the eligibility criteria were considered.
2.2 Variables of the Study
The two outcome variables considered for the study were the survival outcome which was time to death measured from the time of co-infection to death or censored in month. However, time to death were censored for those co-infected patients who were lost the follow, transferred to another Hospital and did not died at first July 2014 (at end of the study). Hence, the longitudinal outcome was CD4 measurement counts per mm\(^{3}\) of blood which was measured within 6 months interval which act as bio marker for co-infected patients.In general the independent covariates considered for the separate longitudinal and survival modeling as well for the joint modeling are listed in the following Table 1 below.
Notice that WHO Clinical Stage which is classified into four; I, II, III and IV; where Stage I indicates asymptomatic disease, Stage II indicates mild disease, Stage III indicates advanced disease and Stage IV indicates severe disease. Hence disease severity increases from Stage I to Stage IV. Functional Status of the patients is also categorical covariate with three categories: Working, Ambulatory and Bedridden. Working patients are those patients who can able to work day to day while ambulatory patients are those patients who can able to work some time but bedridden patients cannot able to work due to the infectious disease.
2.3 Model Specification
2.3.1 Linear Mixed Modeling
Longitudinal outcomes may arise in two common situations; one is when the measurements taken from the same subject at different times and the other is when the measurements taken on related subjects (clusters) in both cases measurements are likely to be correlated. The linear mixed model (LMM) is used to model longitudinal outcomes by accounting with and between subject sources of variations. This is due to the measurement taken from the same subject at different time points or the measurements taken from the same clusters are likely to be correlated.
In this study before the joint modeling to have an appropriate longitudinal sub-model for the longitudinally measured square root of CD4 the LMM were employed to identify the covariates that have significant effects on the mean change of square root of CD4 measurements over time. Therefore, the longitudinal data modeling was begins with exploratory data analysis to determine the mean change of square root transformed CD4 measurement over time. Let \(y_{i1}, y_{i2}\), âĂę, \(y_{in}\) were the square root of CD4 measurement measured at time \(t_{i1},t_{i2}\), âĂę., \(t_{in}\) the linear mixed model of the data which is proposed by Laird and Ware [17] is expressed as:
where \(\mathbf {y}_i\) is the \(n_i\times 1\) vector of observed response values, \({\beta }\) is the \(p\times 1\) vector of fixed-effects parameters, \(\mathbf {X}(t)\) is the \(n_i\times p\) observed design matrix corresponding to the fixed-effects, \(\mathbf {b}_i\) is the \(q\times 1\) vector of random-effects parameters, \(\mathbf {Z}_i\) is the \(n_i\times q\) observed design matrix corresponding to the random-effects, and \({\varepsilon }_i\) is the \(n_j\times 1\) vector of residuals for the response. The corresponding assumptions for model (1) are \(\mathbf {b}_j\sim N_{q}(\mathbf {0}, \mathbf {D})\) and \({\varepsilon }_i\sim N(\mathbf {0}, {\Sigma })\); where \(\mathbf {D}\) and \(\Sigma \) are the variance-covariance matrix for \(\mathbf {b}_i\) and \({\varepsilon }_i\) for outcome variable respectively.
In this model, \({\mu _{i}}(t)=\mathbf {X^{T}}(t){\beta }\) represent is the mean square root of CD4 measurement and \(\mathbf {U}_{1i}(t) = \mathbf {Z}^{T}_{i}(t)\mathbf {b}_{i}\) incorporates the random effects part which is the true individual level CD4 measurement trajectories after they have been adjusted for the overall mean. Here, in mixed effects models, random effects bi is introduced for each subject to incorporate the correlation between the repeated measurements within subject and it facilitates the subject specific inference whereas represents the between sources of variations.
2.3.2 Survival Data Modeling
Survival models are seeks to explain how the risk, or hazard, of an event occurring at a given time is affected by covariates of theoretical interest. Hazard function of survival model is used to explain the probability that an individual will experience an event. This study also considered different Survival models to explain how the risk, or hazard, of death occurring at a given time is affected by covariates of theoretical interest in the study area.
The most commonly used semi-parametric survival model which do not requires the distributional assumption of the survival time is Cox proportional hazard model proposed by Cox [18] which expresses the hazard of an event at time t as:
where \(\mathbf {W}\) is the matrix of baseline covariates which may or may not have the same element with linear mixed effects covariates \(\mathbf {X}(t)\), \({\gamma }\) is the vector of parameters and the term \({\lambda }_{0}(t)\)is the baseline hazard where the effects of covariates are zero. The only assumption of this model is that the hazards ratio \(\psi =\frac{\lambda _{i}(t)}{\lambda _{j}(t)}\) does not change over time (i.e., proportional hazards).
In addition to Cox PH model different parametric survival models are also considered for the study by assuming different parametric distribution for the time to death to have an appropriate survival model for the co-infected patients. However, the hazard model for the parametric model is also same to the Cox PH but the only difference is that the baseline hazard for the parametric survival model is modeled parametrically which have a specified parametric time distribution. In case if the proportional hazard survival model is no longer valid an alternative method modeling survival data is accelerated failure time (AFT) model which is express as:
where \(\zeta _{i}\thicksim F\) and F is parametric error distribution and \({\sigma }\) is scale parameter. The proportional hazard model of the Weibull in our cases were obtained AFT model using the relation between weibull PH and Weibull AFT which is expressed as: \(\hat{\lambda } = exp\left( \frac{-\,\hat{{\gamma }}_{0}}{scale}\right) \), \(\hat{{\alpha }}=\frac{1}{scale}\), \(\hat{\gamma }=\frac{-\,\hat{{\gamma }}_{AFT}}{Scale}\) and where their standard error for the estimated parameters was obtained by using delta methods for the construction confidence interval. Here, \(\hat{\lambda } ,\hat{\alpha }\) and \(\hat{\gamma }\) represents the scale, shape and estimated coefficient of the covariates of Weibull PH model respectively.
2.4 The Joint Modeling Structure
Recently, joint modeling research has expanded very rapidly in Biostatistics and medical research. This is due to model enables the simultaneous study of a longitudinal marker and a correlated time-to-event.
The main aim of this study was also to relate longitudinally measured CD4 biomarker with time to death for HIV/TB co-infected patient to understand the association between the two processes. Therefore, after having appropriate separate models the longitudinal sub-model have the same specification as the separate linear mixed model (1). The survival sub-model includes shared parameter association function to the specified Cox PH model (2). This shared association parameter associates the longitudinally measured CD4 measurement random effects with time-to-death of co-infected patients which can be expressed as follows.
where \(\lambda _{0}(t)\) is the base line hazards rate, \(\mathbf {U}_{2i}(t)\) defines the nature association structure of the shared parameters between the two processes which have a multivariate distribution function. The three association structure values of \(\mathbf {U}_{2i}(t)\) considered for this study were \(\mathbf { U}_{2i}(t) =\alpha ^{T}m_{i}(t)\), \(\mathbf {U}_{2i}(t) = {\alpha }^{T}\mathbf {b}_{i}\) and \(\mathbf {U}_{2i}(t) ={\alpha }^{T}({\beta }_{b}+\mathbf {b}_{i})\). Here, the values of \( m_{i}(t)\) denotes current underlying value of the longitudinally measured CD4 measurement marker processes at the same time point; \({\alpha }\) measures the strength of association vectors between two processes; \(\mathbf {b}_{i}\) is random effect parameters of the longitudinal part and \({\beta }_{b}\) is fixed effect parameters corresponding to the random effects. Finally, DIC score of the fitted association structure of the model was considered to have the appropriate joint model.
2.4.1 Joint Model Estimation Methods
Given the random effects, the longitudinal process is assumed to be independent from the event time. Let \(\varvec{\Theta }_{1}\) and \(\varvec{\Theta }_{2}\) be the vector of parameters defined in linear mixed model and survival model respectively. Assuming the independence between the longitudinal and the survival processes conditionally to the random effects their joint density function is expressed as:
where their joint log likely hood function is expressed as:
where \(\lambda (\mathbf {T}/\mathbf {Y}_{i})\) is the survival hazard function \(S(\mathbf {T}/\mathbf {Y}_{i})=\int _{0}^{t}\lambda (U/\mathbf {Y}_{i})dU\) is the survival function and \(f_{Y}\) and \(f(\eta _{i})\) represents the density function for the longitudinal and shared parameters respectively. However, the computation of the above joint likelihood can be highly intensive. Therefore, Bayesian approach using Markov chain Monte Carlo (MCMC) algorithm was considered for the computation of the model parameters. Under Bayesian approach, model parameters are treated as random variables and assigns probability to each, which is the major difference to the likelihood approach. Therefore, the estimation and inference of the parameters were based on posterior distribution which was obtained based on Bayesian theorem which expressed as:
where \(f({\theta }/\mathbf {y})\) is is the posterior probability distribution of \(\theta \), \(f(y/{\theta })\) is the likelihood function and \(f({\theta })\) is the prior probability distribution of \(\theta \) In our case all the standard prior distribution for all parameters which was aided by Rizopoulos [19] for the JMbayes package was considered.
2.5 Model Selection Techniques
To have an appropriate separate longitudinal and survival model Akaki information criteria (AIC) proposed by Akaike [20] and Bayesian information criteria (BIC) proposed by Schwarz [21] of the model which can be expresses as follows were considered.
where log Lik is the log likelihood function, npar is the number of parameter in the model and N is total number of observation considered to estimate the model. The model with smaller values of AIC and BIC values is considered as an appropriate model.
As explained under joint model selection techniques the joint model estimation was based on the Bayesian approach. Therefore, to have an appropriate joint model the deviance information criterion (DIC) which is proposed by Spiegelhalter et al. [22] which based on the posterior distribution of the deviance statistic was considered. where the values of DIC is expressed as:
where \(f (\mathbf {y}|{\theta })\) is the likelihood function for the observed data vector y given the parameter vector \(\theta \), and h(y) is some standardizing function of the data alone. In this approach, the fit of a model is summarized by the posterior expectation of the deviance, \(\bar{D} = E_{\theta |y}[D]\), while the complexity of a model is captured by the effective number of parameters, pD. Where pD is expressed as:
That is the expected deviance minus the deviance evaluated at the posterior expectations. The DIC is then defined analogously to the AIC as the expected deviance plus the effective number of parameters, i.e.
Since small values of \(\bar{D }\) indicate good fit while small values of pD indicate a parsimonious model, small values of the sum (DIC) indicate preferred model.
3 Results
3.1 Descriptive Results
Among the total co-infected patients during the time period 83 (32.67%) were died whereas 171 (67.33%) were censored co-infected patients. The estimated average age of died co-infected patients were estimated 32.72 years with standard deviation value of 9.44 years while the estimated average age of censored patients were 31.98 years with standard deviation of 8.54 years.
Some of demographic information and some basic base line covariate from the co-infected patients were presented on Table 2 below. Regarding sex of co-infected patients 139 (54.80%) were males and 47 (56.60%) death were also occurred in male group in comparison with female group. More than half 147 (57.90%) of the co-infected patients belongs to orthodox religious group wereas 18 (6.70%) belongs to protestant religious group. Of the total deaths in religious group categories 49 (59.00%) of deaths were occurred in orthodox religious group wereas 4 (4.8%) of deaths occurred in protestant religious group in comparison with Muslim religious group.
When we look at the educational level category of the co-infected patients larger number 109 (42.90%) were attended their primary education while only 17 (6.70%) attended their tertiary education. The Table 2 also shows among the deaths occurred in marital status categories 31 (37.30%) occurred in married group whereas 3 (3.60%) in widowed marital status group in comparison with other marital status groups.
Among the base line clinical stage categories, 8 (3.20 %) were at Stage I, 23 (8.70%) were at Stage II, 124 (48.80%) were at Stage III and the rest 99 (39.30%) were Stage IV. Whereas, among total deaths occurred in these category groups,39 (47.00%) were occurred in both clinical Stage III and IV in comparison with clinical Stage I and II at baseline. Of the functional status categories, 103 (40.50%) were able to work; 126 (49.60%) were at ambulatory and 25 (9.90 %) were at bedridden funct ional status. of the total deaths occurred in this category groups 48(57.80%) were occurred in ambulatory whereas 20 (24.10%) were occurred in working functional status categories.
The longitudinal outcome was the square root of CD4 cells measurement per \(mm^{3}\) of blood which was measured approximately every 6 months. To handle the longitudinal outcome with linear mixed model the square root transformed value were checked for normality and it has the following descriptive summary result on Table 3 at each time of measurement.
As can be observed from Table 3 the square root of CD4 measurement by the censoring status of the patients, the censored patients have an increasing mean square root of CD4 count up to 30 months and started to decreases after 30 months. However, even if the square root of CD4 measurement have an increasing values up 24 months it decreased at month of 30 and become decreases after 30 months for died patients.The variation of the measurement for censored patients there was no this much big measurement variation among the patients between the measurement time up to 30 months and it has decreasing value after 30 months.
3.2 Separate Longitudinal Analysis
Before the joint modeling it is important to explore the appropriate linear mixed model that predicts the mean change of CD4 measurements over time. Therefore, the longitudinal analysis of CD4 measurement started after exploratory analysis of square root transformed CD4 with checking the normality assumption as follows.
3.2.1 Exploring Individual Profile and the Mean Structure Over Time
To understand the association between the CD4 measurement and time individual profile plots were employed. To explore the mean change of square root of CD4 measurement over time loess smoothing techniques over individual profile plots were used since we have unbalanced longitudinal data (Fig. 1).
As indicated on the plots the individual profile plots suggested that there was within and between variations of change in square root CD4 measurements over time. However, the red line which shows the mean structure of CD4 count measurement over time with loess smoothing techniques suggested that the non linear change of the mean square root of CD4 measurement over time. Therefore, it is an important to consider the linear and quadratic time effects in the linear mixed model.
3.2.2 Linear Mixed Model Results
After having appropriate fixed effects of the model using linear model stepwise automatic variables selection with explored time effects as factors that predict the CD4 measurement. After having the appropriate fixed effects the linear mixed model was fitted with different random effects to have an appropriate random effect for the linear mixed model.
As can be observed from the AIC and BIC of the summary of Table 4 model seven linear mixed models were explored. The fitted summary result of Table 4 indicates the quadratic random effects to the model worsen the model since it have large AIC and BIC values than the remaining model, but when we look at the improvement of the model with quadratic random effects with inclusion of linear time effects and random intercepts separately to the model there was an improvement of the model since they have lower AIC and BIC values. Finally we reached on the appropriate linear mixed which have minimum AIC and BIC values than the remaining model with considering random intercept, linear and quadratic time effects for the linear mixed model that appropriately predicts the mean change of CD4 measurements over time.
The fitted result of linear mixed model of Table 5 indicates at base line the CD4 measurement of working functional status group were 2.633 greater and bedridden functional status patients group was 3.174 lower CD4 measurements in comparison with ambulatory functional status group. Furthermore, the model showed with unit increase in weight of the patients increases the mean square root CD4 measurements by 0.063204454 and linear time also have positive effects where as the quadratic time effects have negative effects on the mean change of square roots of CD4 measurements. However, the linear and quadratic mean change of square root of CD4 measurements differs by the functional status of the patients.
3.3 Separate Survival Model Results
After having the appropriate linear mixed model for the CD4 measurement the next step is to determine the appropriate survival model that predicts time to death of the patients for the joint modeling. Therefore, regardless of survival time distribution using stepAIC automatic variable selection smoking status, base line weight, and baseline functional status, type of TB, baseline WHO clinical stage and marital status were extracted to be included in the survival model among the candidate variable considered on Table one.
To have an appropriate survival model all survival models were compared using AIC and BIC of the models. As displayed on Table 6 the null model was the model fitted without covariate whereas the full model was the model with all covariates considered for the model. However, when compare the models using their AIC and BIC values among the parametric models Weibull have smaller AIC and BIC values than the remaining parametric models and we considered this model as appropriate to represent the parametric survival model. However, when we compare the Weibul with the semi-parametric model (Cox PH) model the Cox PH has lowest AIC and BIC value than the parametric survival model. Therefore, the semi-parametric survival model was the preferred model to model time-to-death of co-infected patients in the study area (Tables 7, 8).
3.3.1 The Fitted Cox PH and Weibull PH Models
As can be observed from the fitted survival Cox and Weibull PH models the base line weight, working functional status groups in comparison with ambulatory functional status groups have negative effect on the hazard rate of survival time. However, the hazard rate was higher in bed ridden functional status group, smokers group, separated marital status group in comparison ambulatory functional status group, none smoker group and divorced marital status group respectively. Furthermore, the estimated shape parameter for the Weibull \(=0.9074\) which is less than one shows that the death rate from co-infection was decreasing over time.
3.4 Joint Model Results
After having appropriate separate models that predicts the mean change of CD4 measurements over time and time to death of co-infected patients the next step is to explore an appropriate joint model that associates the longitudinally measured CD4 biomarker with time to death of co-infected patients. As explained under methodology part joint models were fitted under the Bayesian approach using R under JMbayes package and the results are based on single MCMC sampling chains of 75,000 iterations each, following a 35,000 iteration burn-in period.
The results of the joint models were given on the table below and compared with DIC and PD values of the models. As can be observed from the joint models each candidate linear mixed model was associated with survival models with the three candidate association structures. The results shows that among the all explored candidate models sharing the three random effects to survival sub model have minimum PD and DIC values than the remaining models therefore we consider these joint models as an appropriate joint model furthermore when we look at among the three association structure candidate of these groups the model sharing the random intercept, linear and quadratic slopes have minimum PD and DIC values therefore we consider these model as an appropriate final joint model that associates the two process.
The summary of the selected joint model on Table 9 showed the longitudinal sub-model specification was the same to the selected linear mixed model where as the survival sub-model specification incorporates the association parameters that relates the linear mixed model with to survival model using the shared parameters.
As the report of this table or Table 9 the estimated joint model parameter of the posterior estimates of the regression coefficients \(\beta \)and \(\gamma \) together with their 95% credible intervals all of the estimated \(\beta \) values which was significant in the classical separate longitudinal part were here also have significant effects on the CD4 count measurements process of HIV/TB co-infected patients at 5% level of significance. But when we look at the estimated significance of parameters at 5% significance level WHO clinical stage IV, weight and smoking have significant effect on the hazard function of time-to-death of co-infected patient when we look at the classical separate survival model covariate in relation with survival sub-model of the joint modeling the none significance of clinical stages in separate survival analysis have significant effect on hazard function of time to death since clinical stage IV have positive effect significance on the hazard function.
As explained under the methodology the main aim of the joint modeling was to associate the longitudinally measured CD4 measurement to time to death of co-infected patients to understand their association. Therefore, the estimated joint model confirms as the subject specific baseline values and subject specific linear time slope of CD4 were negatively associated whereas the subject specific quadratic slope is positively associated with hazard rate of death of the patients at 5% level of significance (Table 10).
The estimated posterior values of survival sub-model together with the hazard ratio were reported table below. As can be observed from the summary result of table the hazard rate of clinical stage IV patient group was 15.51 times greater in comparison with clinical stage I patient group at base line. Similarly the death rate was higher in patient group of smoker in comparison with none smoker patient group whereas the death rate was lower in windowed marital status patients group in comparison with divorced marital status patient groups.
When we look for the hazard rate association of random effects the unit increment in patient specific baseline square root CD4 lowers hazard rate by 0.9432 and the patient specific slope reduce the hazard rate by 0.16563 for patients with steeper increase in linear longitudinally measured square root of CD4 measurement whereas the patient specific quadratic slope increases the hazard rate by 1.321543 for HIV/TB co-infected patient in the study area at 5% level of significance.
4 Conclusion
In this study, the linear mixed model with subject specific baseline value and subject specific linear and quadratic time slope of square root CD4 measurement was an appropriate fit for the longitudinally measured square root of CD4. However, weight and functional status were factors that affect the mean square root of CD4 measurements whereas the mean change over time in linear and quadratic of square root of CD4 was differ by the functional status of the patients at 5% level of significance.
The Cox PH model was an appropriate mode to fit time to death of patients. The fitted result of Cox and Weibull PH showed weight, functional status, smoking status and marital status of the patients were the factors that affects the hazard rate of time to death of HIV/TB co-infected patients at 5% level of significance in the study area.
The joint model sharing random intercepts and both time slopes of longitudinal model to the survival model was an appropriate for the joint modeling of the data. The estimated association parameters showed subject specific baseline value and the subject specific linear time slope of square root of CD4 measurement was negatively associated whereas the subject specific quadratic time slope of square root was negatively associated with hazard rate of time to death of HIV/TB co-infected patients in the study area at 5% level of significance.
References
USAID fact sheets (2014) The twin epidemics: HIV and TB co-infection. http://www.usaid.gov/news-information/fact-sheets/twin-epidemics-hiv-and-tb-co-infection. Accessed 15 Sep 2014
World Health Organization (2012) Global tuberculosis report. http://apps.who.int/iris/bitstream/10665/75938/1/9789241564502_eng.pdf. Accessed 20 July 2014
World Health Organization (2008) Global tuberculosis control: surveillance, planning, financing. WHO, Geneva
Demissie M, Lindtjørn B, Tegbaru B (2000) Human immunodeficiency virus (HIV) infection in tuberculosis patients in Addis Ababa. Ethiop J Health Dev 14(3):277–282
Yassin M, Takele L, Gebresenbet S, Girma E, Lera M, Lendebo E, Cuevas LE (2004) HIV and tuberculosis co-infection in the southern region of Ethiopia: a prospective epidemiological study. Scand J Infect Dis 36:670–673
WHO TB-HIV (2009) TB/HIV facts. http://www.who.int/tb/challenges/hiv/factsheet_hivtb_2009update.pdf. Accessed 18 June 2014
Tsiatis Anastasios A, Davidian Marie (2012) Joint modeling of longitudinal and time-to-event data: an overview. J Stat Sin 14(2004):809–834
Little RJA, Rubin DB (2002) Statistical analysis with missing data, 2nd edn. Series in probability and statistics. Wiley, New York
Ratclie SJ, Guo W, Ten Have TR (2004) Joint modeling of longitudinal and survival data via a common frailty. Biometrics 60(4):892–899
Tseng Y-K, Hsieh F, Wang J-L (2005) Joint modeling of accelerated failure time and longitudinal data. Department of Statistics, University of California, Davis, CA 95616, USA
Rizopoulos D (2010) Joint modelling of longitudinal and time-to-event data: challenges and future directions. Erasmus Medical Center, Rotterdam
Tapsoba JD, Lee SM, Wang CY (2009) Joint modeling of survival time and longitudinal data with subject-specific change points in the covariates. Graduate Institute of Applied Statistics, Feng Chia University, Taiwan and Division of Public Health, Fred Hutchinson Cancer Research Center, Seattle, USA
Wulfsohn M, Tsiatis A (1997) Joint model for survival and longitudinal data measuredwith error. Biometrics 53:330–339
Tsiatis A, DeGruttola V, Wulfsohn M (1995) Modeling the relationship of survival to longitudinal data measured with error: application to survival and CD4 counts in patients with AIDS. J Am Stat Assoc 90:27–37
Wu L (2010) Mixed effects models for complex data. CRC Press, Boca Raton
Xin H, Gang L, Robert M, Jianxin P (2009) A general joint model for longitudinal measurements and competing risks survival data with heterogeneous random effects, probability and statistics group. J Lifetime Data Anal 17(1):80100
Laird N, Ware J (1982) Random-effects models for longitudinal data. Biometrika 38:963–974
Cox DR (1972) Cox proportional-hazards regression for survival data. J R Stat Soc 34(2):187–220
Rizopoulos D (2014) The R package JMbayes for fitting joint models for longitudinal and time-to-event data using MCMC. Erasmus Medical Center, Rotterdam. arXiv:1404.7625v1
Akaike Hirotugu (1974) A new look at the statistical model identification. IEEE Trans Autom Control J Biom 19(6):716723
Schwarz Gideon E (1978) Estimating the dimension of a model. Ann Stat 6(2):461464. https://doi.org/10.1214/aos/1176344136
Spiegelhalter DJ, Best NG, Carlin BP, Van der Linde A (2002) Bayesian measures of model complexity and fit (with discussion). J R Stat Soc Ser B 64:583–616
Acknowledgements
We would like to thank Jimma University Specialized Hospital administration office, HIV and TB Outpatient Clinics of the Hospital for allowing us the data used for the study. This work was financially supported by the College of Natural Sciences, Jimma University, Jimma, Ethiopia.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Temesgen, A., Gurmesa, A. & Getchew, Y. Joint Modeling of Longitudinal CD4 Count and Time-to-Death of HIV/TB Co-infected Patients: A Case of Jimma University Specialized Hospital. Ann. Data. Sci. 5, 659–678 (2018). https://doi.org/10.1007/s40745-018-0157-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40745-018-0157-0