Abstract
Fusarium is a large genus that includes animal and plant pathogenic species as well as saprophytes. Moreover, specimens from this genus are used in biocontrol as well as for industrial applications. We integrated various disciplines on representatives across the genus. Chromosome numbers (CN) were determined using Germ Tube Burst Method (GTBM), and different sequencing platforms were applied to generate high quality assemblies. In several species, remarkable genome plasticity is observed, including variable CNs and the presence of supernumerary chromosomes that differ markedly from the core chromosomes. It appears that several fusion events between core chromosomes happened during speciation. In F. poae, these supernumerary chromosomes (~8 Mb) exhibit marked differences from the core chromosomes: in the core genome only 2.1% consists of transposable elements (TEs) while TEs make up 25% of the supernumerary chromosomes. The TEs in the core genome show clear signs of repeat-induced point mutation (RIP), while no RIP was found in the supernumerary genome. In addition, no paralogous genes are present on the core, but many are found in the supernumerary genome. Exchange of genetic material occurs between the core and supernumerary genomes. Intact TEs from the supernumerary genome integrate into the core chromosomes, where they are subsequently subjected to RIP. In addition, large blocks of sequence (>200 kb) from the supernumerary genome have recently been translocated to the core genome. The reverse also appears to have occurred: genes from the core seem to have undergone duplication followed by translocation of one of the resulting paralogs to the supernumerary genome, where some paralogs may have undergone further duplications. This exchange of genes between the core and supernumerary genomes bestows significant opportunities for adaptation and evolution on the organism. This is reminiscent to the compartmentalization of genetic material in F. graminearum, where non-conserved regions are found at various places on the four core chromosomes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Fusarium comprises a huge set of species (Geiser et al. 2013) that possess a wealth of biological properties. While some species are used for the preparation of industrially applicable enzymes, others cause serious diseases in many agronomically important crops. In addition, some Fusarium strains are also applied as biocontrol agents (Edel-Hermann et al. 2009). Finally, strains of F. venenatum are used for the production of alternatives for meat (Wiebe 2004).
The classification of Fusarium was traditionally based on morphological characteristics, also known as the Morphological Species Concept (MSC). The Biological Species Concept (BSC) was introduced to accommodate the finding that strains with identical phenotype were only capable of crossing with a subset of morphological identical isolates. This has led to the subdivision of several morphological species into multiple biological species as evidenced by the Fusarium fujikuroi Species Complex, FFSC that is composed of at least 11 mating populations (e.g. Martin et al. 2011). A major drawback of this species concept is the fact that many Fusarium species have no known sexual cycle. This was addressed by the introduction of the Phylogenetic Species Concept (PSC), whereby the DNA sequence of one or more loci is used to discern individuals. The power of this PSC was already demonstrated in 1998 by O’Donnell and co-workers, who recognized 45 species in the FFSC, based on the loci for ITS, β-tubulin and mtSSU (O’Donnell et al. 2008). Since then the number of loci has gradually increased to a multilocus sequence analyses (MLSA) spanning 13 loci, encompassing 16.3 kb (O’Donnell et al. 2008). This expansion of MLSA coincided with the emergence of next-generation sequencing (NGS) technologies. NGS has greatly enhanced speed and sequence quality at significantly reduced costs, allowing the extraction of any given locus for both Fusarium systematics and comparative research. Currently hundreds of fungal genomes have been sequenced and are available in the public domain. Typically two technologies are available that either generate large numbers of small sequence reads with high accuracy (e.g. Illumina) or long read technologies, like single molecule real time (SMRT), that lead to long reads often exceeding 10 kb. However, SMRT has the drawback that it yields sequence data with a high error rate, 10–20%. Ideally, the assembly of raw data from either technology would generate a small number of contigs, but both platforms have their complementary pros and cons (Table 1).
The advantages of a completely assembled genome are multifaceted. Factors influencing life style, adaptability to (changing) environments, and evolution often not only rely on the coding capacity of the genome, but rather on genomic features beyond genes. These include non-coding regions as well as repetitive elements. Proper assessment of the role of specific genetic elements such as telomeres, centromeres and repetitive elements will benefit from the availability of a fully assembled genome. To date, the best assembled genomes of Fusarium species are F. graminearum (King et al., 2015); F. fujikuroi (Wiemann et al., 2013) and F. poae (Vanheule et al., 2016). In addition, genome compartmentalization and structural rearrangements within and between chromosomes can only be studied accurately, when a genome is assembled to chromosome-sized contigs. To this end, the chromosome number of isolates and/or species is a prerequisite. Chromosomes in fungi are too small to be studied using cytological karyotyping. This problem was tackled during the previous decade, when cytogenetics was used to examine filamentous fungi (Tsuchiya and Taga 2010). When this technology was applied to F. graminearum, it revealed that this fungus has only four chromosomes (Gale et al. 2005). Similar results were obtained with several other members of the Fusarium sambucinum species complex, as exemplified by F. culmorum (Fig. 1).
Cytology on the chromosomes of F. culmorum isolate IPO39 Figure. The Germ Tube Burst Method was applied on germinating conidia and the emerging hyphae were lysed by osmotic shock. In metaphase, chromosomes stained with DAPI appear as brightly colored molecules. The Nucleolar Organizing region (rDNA) appears as a thread-like extension of one of the chromosomes (white arrow). This end of the chromosome is stained less dense as a consequence of this protrusion
The advantages of a fully assembled genome were recently reviewed by Thomma and co-workers (Thomma et al. 2016). These authors presented seven reasons that underpin the importance of generating complete genome assemblies. Centromeres as well as telomeres play a vital role in mitosis and meiosis, because they are required for proper segregation of chromosomes during cell division and protection against progressive shorting of chromosomes following DNA replication, respectively. Centromeres are often difficult to assemble due to their very high AT content, while telomeres consist of tandem repeats of the hexanucleotide TTAGGG that reside on the ends of chromosomes. Many features that regulate the interaction between fungi and their environment have been shown to reside in the vicinity of either centromere of telomeres. For example, in the rice blast fungus Magnaporthe oryzae the gene encoding the AVR-PITA protein is located near one of the telomeres. This protein is recognized by resistant rice cultivars and the fungus seems to overcome this resistance by loss of the gene (Chuma et al. 2011). In addition, in F. graminearum, genes involved in the secretome of the fungus (e.g. genes encoding extracellular proteins) are preferentially located in the neighborhood of chromosome ends (Cuomo et al. 2007).
The crosstalk between microbes and their host is often established through effector molecules that can be typically characterized as small cysteine–rich secreted proteins with very limited sequence similarity. Effector genes appear to reside in fast-evolving genomic regions, like telomeres where their presence/absence can be accelerated. Similarly, specialized metabolites have important roles in the interactions between host and fungus and/or other microorganisms in the environment. These complex molecules require multiple enzymes involved in sequential chemical conversions leading to the final product. Likewise, many genes are required for synthesis of these biologically relevant compounds. To ensure concerted expression these genes are often clustered into biosynthetic gene clusters, BGCs (van der Lee and Medema 2016; Hoogendoorn et al. 2016). For instance, the cluster involved in the mycotoxin fumonisin encompasses 15 clustered genes in a 40-kb region (~Proctor et al. 2003; Waalwijk et al., 2004). In a fragmented genome with maybe hundreds of contigs, it is likely that one or more of these BGCs will be missed and proper annotation of the specialized metabolism of fungi will clearly benefit from fully assembled genomes.
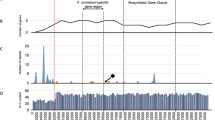
Comparative genomics of Fusarium and other fungal genomes have revealed that genomes can be divided into at least two components, core and supernumerary/accessory genomes, that differ in multiple characteristics: i) evolutionary speed; ii) expression level and iii) gene repertoire. Genes responsible for primary metabolism reside in the core compartment, where evolution occurs at slow speed. This class of genes is (highly) conserved among species. In contrast, many of the BGCs in F. graminearum are found in non-conserved (NC) regions across the four chromosomes (Zhao et al. 2014). In these compartments, the expression of genes is significantly lower than in the remainder of the genome. Aligning of the chromosomes of F. graminearum with those of F. verticillioides, substantiated the role of these NC-regions. As shown in Fig. 2, chromosome 1 of F. graminearum shows synteny with chromosomes 1, 8 and 5 of F. verticillioides with the exception of the NC regions. The NC-regions coincide with the telomeres of the three F. verticillioides chromosomes. Moreover these NC-regions have a higher density of genes encoding secretory proteins and BGCs. In contrast, genes presumed to be involved in primary metabolism, like transcription factors (Fig. 2, line a) or genes encoding ribosomal proteins (Fig. 2, line b) are evenly distributed along all four chromosomes.
(Top) Synteny between and chromosomes 1, 8 and 5 of F. verticillioides and chromosome 1 of F. graminearum. Note that the syntenic regions are intertwined with NC regions that are not conserved between the two species. In F. verticillioides these regions are located at the ends of the chromosomes e.g. (sub) telomeric regions. (Bottom) Positions of transcription factor genes (a); ribosomal protein genes (b); genes putative encoding secreted proteins (c) and BGCs (d) on F. graminearum chromosome 1 are given. Note that TFs and ribosomal genes are mapped along the entire chromosome. Secretome related genes and BGCs preferentially map at or near NC regions. Source: adapted from Zhao et al., BMC Genomics 2014
Adaptation of a population to changing ecological environmental conditions can require changes in gene function and/or expression. On the other hand, mutations in genes can be beneficial or detrimental on the organism. Hence diversification is often preceded by duplication of parts of the genome. However, high-level expression of mutated genes can cause an energetic burden on the organism. If the expression of paralogous genes can be reduced, such burdens will be diminished. Therefore it was hypothesized that these NC regions function as a cradle for evolution, contributing to the ability of fungi to adapt to changing conditions. Modification of histones in nucleosomes by either methylation or acetylation strongly influences the expression of genes in BGCs and methylation profiles across chromosomes provide independent support for the presence of BGCs at their anticipated positions. Histone H3 methylation is associated with gene silencing, and in F. graminearum methylated histones were predominantly found in regions containing BGCs (Connolly et al. 2013).
Intra-and inter-chromosomal rearrangements are powerful mechanisms by which regions of DNA involved in interaction with the host can be brought together, e.g. recombinations that generate novel combinations of genes and BGCs with novel functionalities. Similar mechanisms allow to enrich for effector genes. In the smut fungus Sporisorium scitamineum, evolution of effector genes is driven by tandem gene duplication (Dutheil et al. 2016). These authors also showed that transposable elements (TEs) play an important role in the evolution of clustered genes.
Repetitive elements in particular TEs are powerful elements that can separate or bring together different portions of the genome. However, de novo sequencing using short reads will not allow to read across the TE (Table 1) thereby resulting in (many) contig ending in (parts) of TEs. As no full-size TEs will be obtained, all TEs from the same class will collapse and the estimated genome size will be underestimated. In fact, in F. poae we obtained a genome size of 39 Mb using Illumina Hiseq (Vanheule et al. 2016). Contrarily, when the long read technology was applied on the same isolate, the length of the genome expanded to 46 Mb. Comparison between assemblies from both sequencing platforms also showed major differences e.g. in numbers of contigs (176 for SMRT and 1253 for HiSeq) and N50 (>8 Mb for SMRT vs. 700 Kb for HiSeq). The distribution of TEs across the genome showed a substantial disequilibrium: while TEs covered 2.1% of the four chromosomes in F. poae, in the supernumerary genome they occupied > 25% of the extra 8 Mb (Vanheule et al. 2016). Interestingly, TEs from the same family could be found in both the core genome as well as in the supernumerary genome. However, the copies that are located in the core genome appeared to be subject to repeat-induced point mutations (RIP). This process is unique to fungi and specifically acts as defense system against repetitive sequences by silencing repeated copies. RIP is presumed to occur during the sexual cycle and a hallmark for RIP is the dominance of CpA → TpA mutations (Galagan and Selker 2004). The activity of the RIP process in F. poae showed a strong bias between TEs in the core genome and in the accessory genome (Vanheule et al., 2016). All copies in the core genome are mutated by RIP with an extreme preference for CpA → TpA mutations. In contrast, TEs in the supernumerary genome are not RIPped (Fig. 3). If (active) TEs residing in the supernumerary genome transpose to the core genome, they become subject to inactivation by RIP, implying that sex (still) may occur in this organism that is generally considered to be asexual.
RIP analysis of the transposable element DTF2_Fot2 in the core and the supernumerary genome. A comprehensive RIPcal analysis was performed on the core and the supernumerary genome, separately. In the core genome, DTF2_Fot2 exhibits RIPcal patterns that are typical for RIP (strong dominance of CpA → TpA mutations; red trace). In the supernumerary genome, all mutations occur at similar frequencies. (Picture adapted from Vanheule et al., 2016)
Concluding remarks
NGS technologies have generated hundreds of fungal genomes, the majority of which still are composed of large numbers of contigs. Integration of high quality short reads using HiSeq and long- read SMRT, allows the construction of assemblies that cover chromosomes from one telomere to the other. The assembly of AT rich regions such as centromeres and repeat clusters, such as the ribosomal RNA repeats remains challenging. Nevertheless, we were recently successful in assembling the genomes from F. subglutinans and F. temperatum into 12 contigs each (Zhang et al.,. manuscript in preparation). These Fusaria may represent sibling species in the Fusarium fujikuroi species complex, FFSC. All contigs in both species are telomere to telomere representations of the 12 chromosomes visualized in FFSC by both Pulsed Field Gel Electrophoresis as well as GTBM. In addition, all contigs contained long stretches of high AT content and multiple tandem repeats of the telomere repeat TTAGGG. Synteny to the chromosomal level could be demonstrated using mummer (Fig. 4). On the second largest contig in both species one of the telomere repeat is missing. This was due to the presence of the rDNA repeat, located at the end of a chromosome in most species (viz. Figure 1) as was also shown by King et al. (2015). The number of rDNA repeats was 110 in JL22, while 80 copies were observed in F. temperatum JL513.
References
Chuma I, Isobe C, Hotta Y, Ibaragi K, Futamata N, Kusaba M, Yoshida K, Terauchi R, Fujita Y, Nakayashiki H, Valent B, Tosa Y (2011) Multiple translocation of the AVR-Pita effector gene among chromosomes of the rice blast fungus Magnaporthe oryzae and related species. PLoS Pathog 7:e1002147
Connolly LR, Smith KM, Freitag M (2013) The Fusarium graminearum histone H3 K27 methyltransferase KMT6 regulates development and expression of secondary metabolite gene clusters. PLoS Genet 9:e1003916
Cuomo CA, Güldener U, Xu JR, Trail F, Turgeon BG, Di Pietro A, Walton JD, Ma LJ, Baker SE, Rep M, Adam G, Antoniw J, Baldwin T, Calvo S, Chang YL, DeCaprio D, Gale LR, Gnerre S, Goswami RS, Hammond-Kosack K, Harris LJ, Hilburn K, Kennell JC, Kroken S, Magnuson JK, Mannhaupt G, Mauceli E, Mewes HW, Mitterbauer R, Muehlbauer G, Münsterkötter M, Nelson D, O’Donnell K, Ouellette T, Qi WH, Quesneville H, Roncero MIG, Seong KY, Tetko IV, Urban M, Waalwijk C, Ward TJ, Yao J, Birren BW, Kistler HC (2007) The Fusarium graminearum genome reveals a link between localized polymorphism and pathogen specialization. Science 317:1400–1402
Dutheil JY, Mannhaupt G, Schweizer G, Sieber CMK, Munsterkotter M, Guldener U, Schirawski J, Kahmann R (2016) A tale of genome compartmentalization: the evolution of virulence clusters in smut fungi. Genome Biol Evol. doi:10.1093/gbe/evw026
Edel-Hermann V, Brenot S, Gautheron N, Aimé S, Alabouvette C, Steinberg C (2009) Ecological fitness of the biocontrol agent Fusarium oxysporum Fo47 in soil and its impact on the soil microbial communities. FEMS Microbiol Ecol 68:37–45
Galagan JE, Selker EU (2004) RIP: the evolutionary cost of genome defense. Trends Genet 20:417–23
Gale LR, Bryant JD, Calvo S, Giese H, Katan T, O’Donnell K, Suga H, Taga M, Usgaard TR, Ward TJ, Kistler HC (2005) Chromosome complement of the fungal plant pathogen Fusarium graminearum based on genetic and physical mapping and cytological observations. Genetics 171:985–1001
Geiser DM, Aoki T, Bacon CW, Baker SE, Bhattacharyya MK, Brandt ME, Brown DW, Burgess LW, Chulze S, Coleman JJ, Correll JC, Covert SF, Crous PW, Cuomo CA, De Hoog GS, Di Pietro A, Elmer WH, Epstein L, Frandsen RJN, Freeman S, Gagkaeva T, Glenn AE, Gordon TR, Gregory NF, Hammond-Kosack KE, Hanson LE, Jimenez-Gasco MM, Kang S, Kistler HC, Kuldau GA, Leslie J, Logrieco A, Lu G, Lysoe E, Ma LJ, McCormick SP, Migheli Q, Moretti A, Munaut F, O’Donnell K, Pfenning L, Ploetz RC, Proctor RH, Rehner SA, Robert VARG, Rooney AP, Salleh BD, Scandiani MM, Scauflaire J, Short DPG, Steenkamp E, Suga H, Summerell BA, Sutton DA, Thrane U, Trail F, Van Diepeningen A, VanEtten HD, Viljoen A, Waalwijk C, Ward TJ, Wingfield MJ, Xu JR, Yang XB, Yli-Mattila T, Zhang N (2013) One fungus, One name: defining the genus Fusarium in a scientifically robust way that preserves longstanding use. Phytopathology 103:400–408
Hoogendoorn K, Waalwijk C, van der Lee TAJ, Medema MH (2016) Evolution and diversity of biosynthetic gene clusters in Fusarium. In preparation
King R, Urban M, Hammond-Kosack MCU, Hassani-Pak K, Hammond-Kosack KE (2015) The completed genome sequence of the pathogenic ascomycete fungus Fusarium graminearum. BMC Genomics 16:544
Martin SH, Wingfield BD, Wingfield MJ, Steenkamp ET (2011) Structure and evolution of the Fusarium mating type locus: new insights from the Gibberella fujikuroi complex. Fungal Genet Biol 48:731–740
O’Donnell K, Cigelnik E, Nirenberg HI (1998) Molecular systematics and phylogeography of the Gibberella fujikuroi species complex. Mycologia 90:465–493
O’Donnell K, Ward TJ, Aberra D, Kistler HC, Aoki T, Orwig N, Kimura M, Bjørnstad Klemsdal SS (2008) Multilocus genotyping and molecular phylogenetics resolve a novel head blight pathogen within the Fusarium graminearum species complex from Ethiopia. Fungal Genet Biol 45:1514–1522
Proctor RH, Brown DW, Plattner RD, Desjardins AE (2003) Co-expression of 15 contiguous genes delineates a fumonisin biosynthetic gene cluster in Gibberella moniliformis. Fungal Genet Biol 38:237–249
Thomma BPHJ, Seidl MF, Shi-Kunne X, Cook DE, Bolton MD, van Kan JAL, Faino L (2016) Mind the gap; seven reasons to close fragmented genome assemblies. Fungal Gen Biol 90:24–30
Tsuchiya D, Taga M (2010) Fluorescence in situ hybridization for molecular cytogenetic analysis in filamentous fungi. In: molecular and cell biology methods for fungi (ed: Sharon, A). Methods Mol Biol 638:235–257
Van der Lee TAJ, Medema MH (2016) Computational strategies for genome-based natural product discovery and engineering in fungi. Fungal Gen Biol 90:29–36
Vanheule A, Audenaert K, Warris S, van de Geest H, Schijlen E, Höfte M, De Saeger S, Haesaert G, Waalwijk C, van der Lee T (2016) Living apart together: crosstalk between the core and supernumerary genomes in a fungal plant pathogen. BMC Genomics 17:670
Waalwijk C, van der Lee TAJ, de Vries PM, Hesselink T, Arts J, Kema GHJ (2004) Synteny in toxigenic Fusarium species: the fumonisin gene cluster and the mating type region as examples. Eur J Plant Pathol 110:533–544
Wiebe MG (2004) QuornTM Myco-protein – Overview of a successful fungal product. Mycologist 18:17–20
Wiemann P, Sieber CMK, von Bargen KW, Studt L, Niehaus EM, Espino JJ, Huss K, Michielse CB, Albermann S, Wagner D, Bergner SV, Connolly LR, Fisher A, Reuter G, Kleiwege K, Bald T, Wingfield BD, Ophir R, Freeman S, Hippler M, Smith KM, Brown DW, Proctor RH, Münsterkötter M, Freitag M, Humpf HU, Güldener U, Tydzynski B (2013) Deciphering the cryptic genome: genome-wide analyses of the rice pathogen Fusarium fujikuroi reveal complex regulation of secondary metabolism and novel metabolites. PLoS Pathog 9:e1003475
Zhao C, Waalwijk C, de Wit PJGM, Tang DZ, van der Lee TAJ (2014) Relocation of genes generates non-conserved chromosomal segments in Fusarium graminearum that show distinct and co-regulated gene expression patterns. BMC Genomics 15:191
Author information
Authors and Affiliations
Corresponding author
Additional information
Section Editor: Emerson M. Del Ponte
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Waalwijk, C., Vanheule, A., Audenaert, K. et al. Fusarium in the age of genomics. Trop. plant pathol. 42, 184–189 (2017). https://doi.org/10.1007/s40858-017-0128-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40858-017-0128-6