Abstract
Hepatitis B is caused by hepatitis B virus (HBV), and persistent HBV infection is a global public health problem, with 257 million people as HBV chronic carriers. Viral covalently closed circular DNA (cccDNA) is a key factor to establish persistent infection in infected hepatocytes. Current antiviral therapies have no direct impact on pre-existing cccDNA reservoir, which can be assembled into minichromosome by hijacking host factors. Understanding the mechanisms of epigenetic regulation in cccDNA minichromosome is crucial to develop new therapy on cccDNA, an attractive target for HBV cure. This review summarizes the current advances in epigenetic regulation of cccDNA minichromosome, which might provide clues to novel druggable targets to cure hepatitis B by either silencing or eliminating cccDNA reservoir.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
HBV infection remains a global health problem and results in approximately a million death annually. Although infection rate has decreased significantly due to effective vaccines, there are more than 257 million people worldwide suffering from HBV chronic infection, with high risk of developing to liver fibrosis, cirrhosis, and hepatocellular carcinoma (WHO 2017). Large number of HBV carriers can be asymptomatic for decades, because there is no therapy available to thoroughly eliminate HBV genome in patients. It has been reported that HBV chronic infection is dependent on the persistence of covalently closed circular DNA (cccDNA) in infected cellular nucleus (Köck et al. 2010). Currently, there are two categories of approved drugs for hepatitis B treatment. First, the immune modulator Peg-IFN can epigenetically control cccDNA minichromosome and regulate host antiviral immune responses (Belloni et al. 2012; Shi et al. 2018). Second, nucleos(t)ide analogues (NAs) exhibit the repression of HBV polymerase (Lai et al. 2017; Papatheodoridis et al. 2002; TAK et al. 2016; Wong et al. 2013). Unfortunately, cccDNA is insensitive to antiviral therapy, leading to rapid reoccurrence of HBV replication in most patients once drug withdrawal (Hu et al. 2019). Besides potential serious side effects, long-term treatment with NAs can lead to drug resistance (Aspinall and Pockros 2004; Fontana 2009), although tenofovir alafenamide approved in 2016 has an excellent resistance profile in about 2-year test period (Agarwal et al. 2018). Therefore, it is urgent to develop novel therapies.
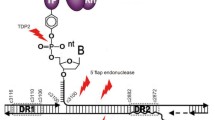
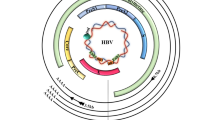
The stable episomal cccDNA minichromosome in nucleus is one of the key obstacles for cure (Fig. 1). HBV is an enveloped DNA virus of the hepadnaviridae family, whose genetic information resides in a 3.2 kb partially double-stranded relaxed circular DNA (rcDNA). After entry to hepatocyte, rcDNA is transformed into cccDNA in nucleus (Dezhbord et al. 2019; Guo et al. 2007, 2010; Yeh et al. 1998). cccDNA serves as transcriptional template for all viral RNAs, among which pre-genomic RNA (pgRNA) is reverse-transcribed into rcDNA to be further processed to replenish cccDNA reservoir (Beck and Nassal 2007). In nucleus, cccDNA takes a chromatin-like conformation, known as cccDNA minichromosome, which consists of both histones and non-histone host factors (Bock et al. 2001). cccDNA minichromosome has been considered under the regulation of nuclear transcription factors, transcriptional coactivators, chromatin-modifying enzymes, etc. However, the detailed regulatory mechanisms remain unclear. It has been proposed that rcDNA is recognized by host factors and transformed into cccDNA, which may be closely related to host DNA-damage-repair pathway (Gómez-Moreno and Garaigorta 2017; Königer et al. 2014). Results of CsCl density gradient ultracentrifugation and electron microscope observation show that HBV nucleoprotein complex displays a typical “beads-on-string” model (Bock et al. 1994). In conjunction with the results of nucleosome positioning, the evidence shows that the average number of nucleosomes is 18 and repeat unit comprises 180 bp in HBV minichromosome, which is smaller than host chromosome (Bock et al. 1994; Shi et al. 2012). That means cccDNA minichromosome may have a little more compact conformation. Therefore, it is reasonable to suppose that HBV cccDNA minichromosome might be transformed between closed conformation (inactive) and open conformation (active) to manipulate viral transcription and replication (Fig. 2). In this review, we summarize the advances of epigenetic regulation of HBV cccDNA minichromosome and provide a distinct perspective through a structural view, which might provide clues to novel druggable targets to pave the way to cure hepatitis B.
cccDNA minichromosome—the key obstacle to cure Hepatitis B. HBV binds NTCP receptor and followed by entry to the hepatocyte. Upon uncoating, rcDNA is released into nucleus and repaired to form cccDNA. Then cccDNA is packaged into minichromosome and serves as template for viral RNAs, which are transported to cytoplasm and some of them are translated into viral proteins (i.e., HBcAg, HBV polymerase, HBsAg). HBV polymerase and pre-genomic RNA (pgRNA) are packaged into HBcAg-formed capsid, where pgRNA is reverse-transcribed to form rcDNA. HBsAg proteins surround the capsid to form new virions. The clearance of cccDNA and HBsAg is rarely achieved with current therapies (in red frame), while the cccDNA minichromosome as the viral reservoir (in red) is the key obstacle to finally cure Hepatitis B
Schematic representation of HBV cccDNA minichromosome under different epigenetic regulation. In the presence of host restriction factors and inhibitive modifiers, cccDNA minichromosome is in a closed configuration, and viral genes are transcriptionally repressed (upper). In the presence of HBx and other activators, cccDNA minichromosome is in an open configuration, and viral transcription is activated and leads to a high level of virus replication (down)
Epigenetic regulation by HBV encoded proteins
HBV core protein (HBc) forms viral nucleocapsid and belongs to non-histone proteins associated with minichromosome (Bock et al. 2001). Scientists revealed that HBc may promote transcription due to its preferentially binding to CpG island 2 on minichromosome, where HBc associates with HDAC1 and CBP to regulate histone acetylation and DNA methylation (Chong et al. 2017; Guo et al. 2011). Moreover, HBc recruits DNA polymerase coordinator PCNA to minichromosome to promote viral proliferation (Feng et al. 2019). HBc also facilitates APOBEC3 family members A3A and A3B to colocalize with cccDNA minichromosome to induce deamination and subsequently Apurinic/Apyrimidinic (AP) sites for degradation, which is, respectively, upregulated by IFN-α and LTβR activation (Lucifora et al. 2014). Furthermore, core protein allosteric modulators (CpAMs) can regulate viral nucleocapsid assembly or disassembly. For instance, Bay 41-4109 and GLS4 are representative of heteroaryldihydropyrimidines (HAPs) and can introduce significant conformational changes to core protein subunits to inhibit de novo cccDNA synthesis and transcription, while ENAN-34017 of sulfamoylbenzamide (SBA) can only induce a subtle conformational change that renders viral particle susceptible to DNase degradation (Guo et al. 2017; Zhou et al. 2017). However, these CpAMs can accelerate intracellular cccDNA formation from existed progeny rcDNA.
HBV X protein (HBx) is a regulatory protein (Ramakrishnan et al. 2019). It is essential for transcription initiation, maintenance, and epigenetic regulation of cccDNA by interacting with various factors including histone acetyltransferases p300, CBP and PCAF, as well as histone deacetylases HDAC and hSirt1 (Cougot et al. 2007; Guerrieri et al. 2017; Lucifora et al. 2011). Recently, it has been shown that HBV minichromosome associated with HBx can be enriched around host transcriptionally active chromatin with host RNA Polymerase II (Pol II) and other hallmarks such as H3K36me3, H3K36ac, and H3K4me3, while silencing of HBx can decrease viral minichromosome stability (Hensel et al. 2018; Jin et al. 2019). Besides, nuclear HBx can bind plasmid-encoded HBV minichromosome through the C-terminal domain (Hensel et al. 2018), which may imply that HBx is involved in nuclear localization of viral minichromosome. Meanwhile, HBx can recruit coactivators (CBP, p300, and PCAF), and transcription factors (ATF/CREB, ATF3, c/EBP, NF-IL-6, Ets, Egr, SMAD4, Oct1, RXR receptor, p53) to HBV minichromosome to regulate cccDNA epigenetically (Belloni et al. 2009; Levrero et al. 2009). HBx is also able to bind CUL4-DDB1 ubiquitin ligase through its H-box motif (Landsberg et al. 2018; Li et al. 2010), which facilitates degradation of chromosome structure maintenance complex 5/6 (Smc5/6) (Rivière et al. 2019). In absence of HBx, Smc5/6 functions as a restriction factor to inhibit HBV transcription (Decorsière et al. 2016). However, the transcription is resumed in the presence of functional HBx (Abdul et al. 2018; Murphy et al. 2016). Additionally, viral replication and transcription are highly repressed by utilizing anti-HBx 2A7 epitope antibody, which specifically blocks the interface between HBx and DDB1 (Tao et al. 2019). Moreover, IFN-induced TRIM interacts with the C-terminal of HBx and inhibits the formation of Smc5/6-HBx–DDB1 complex, leading to suppressive HBV replication (Tan et al. 2018). Therefore, HBx-DDB1 interaction interface becomes an attractive target for drug development to silence HBV transcription (Sekiba et al. 2019). HBx-targeted siRNA can also inhibit viral replication at both mRNA and protein level (Xie et al. 2012). Besides Smc5/6, HBx interacts with other host restriction factors to regulate minichromosome. For instance, HBx is responsible for the upregulation of E3 ubiquitin ligase MSL2, which contributes to HBV cccDNA activation by inducing APOBEC3B degradation via ubiquitylation of Lys320 site of APOBEC3B (Gao et al. 2017). Similarly, HBx/STAT3 signaling is stimulated by lncRNA HULC to facilitate production of miR-539 to downregulate cytidine deaminase APOBEC3B to maintain cccDNA stability (Liu et al. 2019). Additionally, HBx can facilitate viral DNA replication through restraining MDM2-mediated ubiquitination and degradation of RNA helicase DHX9 (Shen et al. 2020). Moreover, HBx can activate Notch-CREB signaling to facilitate cccDNA replication, but it is restricted by E3 ubiquitin ligase (Gao et al. 2016; Wang et al. 2010). In addition, HBx binds lncRNA DLEU2 to modulate minichromosome transcription and relieve repression induced by chromatin-modifying enzymes like EZH2 and PRC2 (Salerno et al. 2020). Furthermore, HBx can colocalize with host transcription factor Sp110 and drive it out of ND10 complex via Sp110 deSUMOylation, which facilitates viral persistence by downregulating factors (IRF9, STAT1, and STAT2) in IFN-I-response pathway and altering epigenetic landscape via coactivator p300 (Sengupta et al. 2017). Another study revealed host pre-mRNA processing factor PRPF31 in spliceosome associates with HBx to promote cccDNA formation (Kinoshita et al. 2017). In hepatocellular carcinoma cells, 14-3-3ζ protein binds to Akt-induced RPLpS31GP motif of HBx and significantly inhibits ubiquitination and degradation of HBx, while 14-3-3ζ itself can activate Akt pathway, indicating the altered structure and biological function of phosphorylated HBx (Tang et al. 2018). In summary, both HBc and HBx are essential in cccDNA accumulation and multiple epigenetic regulatory pathways in cccDNA minichromosome. Thus, they are becoming potential virus-encoded drug targets for silencing viral transcription or eliminating the cccDNA reservoir to achieve a cure eventually.
Epigenetic regulation by histone modifications
Histones with various modifications are associated with transcriptionally active or repressive HBV minichromosome to regulate genomic transcription as found in host chromosome. Current studies have shown that HBV minichromosome is regulated by histone post-translational modifiers, which have a significant impact on viral infection, replication, and maturation (Gong et al. 2011; Tropberger et al. 2015). Histone modifications are generally reversible, including acetylation, methylation, phosphorylation, SUMOylation, ubiquitylation, ADP-ribosylation, etc. Here, we emphasize histone acetylation and methylation due to recent booming studies related to HBV.
Histone acetyltransferases (HATs) and histone deacetylases (HDACs) manipulate histone acetylation to regulate minichromosome epigenetically. HAT can transfer acetyl to lysine, which is advantageous to dissociate histone octamers from loose chromatin. Previous studies revealed that HBV replication parallels acetylation of cccDNA-associated histone H3/H4 (Pollicino et al. 2006; Wei et al. 2017). Inhibitors of histone deacetylases can promote viral replication, while H3/H4 is highly acetylated under HBx activation. In a human liver-chimeric mouse model, HAT1 can be activated by the HBx coactivating transcription factor Sp1 and recruited to minichromosome through lncRNA HULC-mediated interaction with HBc (Yang et al. 2019). The overexpression of HAT1 can promote the acetylation of H3K27, H4K5 and H4K12 in minichromosome, while downregulation of HAT1 can impair the assembly of histone H3/H4 and recruitment of HBx and p300 to impede the formation of minichromosome (Yang et al. 2019). On the contrary, HDAC can remove acetyl from specific site, causing histone positively recharged to strengthen interaction between histones and DNA, which promotes chromatin to be condensed. Acetylation of H3K9 and H3K27 is specifically downregulated by HDAC11 to limit viral replication, while acetylated H4 is not affected (Yuan et al. 2019). Moreover, HDACs can also decrease the occupancy of Pol II in transcribing minichromosome (Balakrishnan and Milavetz 2008). The relationship between HDAC and antiviral therapy has been clarified that the inhibition of HBV replication and transcription is associated with histone deacetylation of H3K9/H3K27 and recruitment of inhibitors to cccDNA in the IFN treatment (Liu et al. 2013; Zhang et al. 2019). Moreover, IL6 inhibits cccDNA transcription by enhancing the recruitment of HDAC to render hypoacetylation of cccDNA-associated histone and reducing the binding of essential transcriptional factors (HNF1α, HNF4α, and STAT3) to cccDNA (Palumbo et al. 2015). In addition, a novel E3 ubiquitination ligase NIRF reduces acetylation of H3 and acts as a negative regulator of HBc to inhibit viral replication (Qian et al. 2015). Furthermore, Retinoid X receptor α (RXRα) can increase acetylation of histones H4 and H3 to promote viral replication and transcription by recruiting p300 to cccDNA minichromosome (Nkongolo et al. 2019; Zhang et al. 2017b).
Beyond acetylation, the methylation of H3/H4 is associated with chromatin structure, which regulates HBV transcription (Kallestad et al. 2013; Peng and Karpen 2007). The Sirtuin family members (SIRT1 and SIRT3) can deacetylate histones of minichromosome and regulate the recruitment of histone methyltransferase suppressor of variegation 3–9 homolog 1 (SUV39H1) to facilitate formation of heterochromatin by increasing the chromatin repressive marker H3K9me3 and reducing the chromatin active marker H3K4me3 (Peng and Karpen 2007, 2009; Ren et al. 2014, 2018; Vaquero et al. 2007), while HBx can relieve inhibition of viral transcription by not only impairing expression and recruitment of SIRTs (Deng et al. 2017), but also recruiting LSD1 and Set1A to establish active chromatin (Alarcon et al. 2016). Recent research suggested that HBx colocalizes with the core subunit WDR5 of SET domain containing 1 (SET1)/mixed lineage leukemia (MLL) histone methyltransferase complex and inhibits DDB1-induced degradation of WDR5 to promote viral transcription by H3K4me3 modification on minichromosome (Gao et al. 2019). Moreover, protein arginine methyltransferase 5 (PRMT5) may preferentially bind to cccDNA through interaction with HBc to elevate H4R3me2 on minichromosome and disrupt HBV Pol–pgRNA interaction to abrogate pgRNA encapsidation to inhibit viral replication (Zhang et al. 2017a). It has been reported that SETDB1-mediated H3K9me2/H3K9me3 and heterochromatin protein factor 1 (HP1) can induce viral transcriptional silence by rearranging chromatin structure, but HBx can antagonize this process to allow synthesis of active chromatin (Rivière et al. 2015). Histone methylation is involved in formation of heterochromatin by recruiting HP1, which may also recruit DNA methyltransferases to methylate DNA. Modified histones can not only regulate chromatin structure directly, but also serve as binding sites for other regulatory proteins to function indirectly. Besides acetylation and methylation, the roles of various histone modifications in epigenetic regulation of HBV minichromosome remain unclear and are urgent to be fully addressed in the future.
Epigenetic regulation by DNA methylation
DNA methylation is a major epigenetic regulation on gene activities and introduced by DNA methyltransferases (DNMTs) responsible for addition of methyl groups to the CpG islands of DNA. Methylated DNA may serve as a signal recognition site to specifically recruit corresponding factors to cccDNA minichromosome and result in allosteric effects. HBV cccDNA can be methylated to various extent, which is mostly associated with the replicative repression of cccDNA (Kim et al. 2011; Zhang et al. 2014). Generally, HBV DNA methylation by DNMTs is closely related to transcriptional silence as in mammalian cells (Guo et al. 2009; Vivekanandan et al. 2009). DNMTs (DNMT1, DNMT2, DNMT3a, DNMT3b) upregulated by HBV can promote viral genome-wide methylation and reduce pgRNA production to inhibit HBV replication (Vivekanandan et al. 2010). However, further investigation indicated that DNMT inhibitors can activate host innate immune response through IFN signaling pathway, and thus inhibit both viral replication and transcription (Chiappinelli et al. 2015). Evidence also revealed that DNA methylation alone may not be efficient for inhibition, while methylated and condensed chromatin is required to repress gene transcription (Deuschle et al. 2016). DNA methylation on cccDNA seems to participate in inhibition of HBV, but meanwhile the host genes can be also methylated by elevated expression of DNMTs. Consequently, the detailed mechanism of DNA methylation during viral defense needs further exploration.
Epigenetic regulation by chromatin remodeling
Chromatin remodelers play significant roles in regulating viral transcription in the context of minichromosome. Some chromatin remodelers do regulate HBV minichromosome in the similar ways as for host chromosome, which involves sliding, replacing, reassembling, or exchanging nucleosomes. Members of human SWI/SNF family are recognized as chromatin remodelers (such as BAF and PBAF), displaying an essential role in transcriptional regulation. For example, the core ATPase subunit Brg1 of the PBAF complex can antagonize the suppression induced by PRMT5 (Zhang et al. 2017a). Meanwhile, the core ATPase subunit Brm of the BAF complex also has a promotion on viral transcription (Chen et al. 2016). In addition, HBx-associated protein HBXAP/RSF1, a component of a ISWI chromatin remodeling complex, interacts with HBx as a transcription coactivator (Shamay et al. 2002). Furthermore, it was shown that inactivating mutation of ARID2 from human SWI/SNF family is closely related to cancer genesis through genomic analysis of hepatocellular carcinoma (Li et al. 2011).
DNA topoisomerases (TOPs) can modulate chromatin structure and catalyze distinct steps of cccDNA formation (Halmer et al. 1998; Sheraz et al. 2019), and meanwhile DNA topoisomerases are believed to function in PJA1-mediated viral inhibition (Xu et al. 2018). Both TOP1 and TOP2 are involved in the repair of negative-strand DNA gap, while TOP2 also participates in the repair of positive-strand DNA gap. In addition, it was shown that human minichromosome maintenance (MCM) protein heterocomplexes (MCM2, MCM4, MCM6, and MCM7) with high affinity to histone H3 play essential roles in the replication initiation, which may make structural change once replication initiates (Ishimi et al. 1996, 1998; Méndez and Stillman 2000). Moreover, researchers found that MCMs can initiate transcription by recruiting RNA Pol II holoenzyme to minichromosome (Holland et al. 2002). Furthermore, MCM7 can be inhibited by simvastatin (SIM) to downregulate HBV replication (Li et al. 2016). Therefore, MCMs might be a potential target for novel antiviral treatment. Besides acetylation regulation, SIRT3 can also restrain the binding of host Pol II and transcription factor YY1 to cccDNA, indicating that SIRT3 participates in establishment of repressive chromatin structure and transcriptional silencing of cccDNA (Ren et al. 2018). Additionally, Parvulin (Par14 and Par17) can bind and stabilize HBx through HBx RP motif, and may bind cccDNA minichromosome through S19/44, respectively, to upregulate HBV replication in a chromatin remodeling way (Saeed et al. 2019).
Host Smc5/6 suppresses HBV transcription when localized to nuclear domain 10 (ND10) without inducing a detectable innate immune response (Niu et al. 2017). Moreover, it has been reported that PJA1 can promote Nse4 to bind viral or episomal DNA in a synergistic way through competitive substitution of Nse1 in Smc5/6 complex, and thus represses HBV proliferation (Xu et al. 2018). Another SMC family member cohesin is highly affiliated with minichromosome and severing its SMC ring domain causes cohesin dissociating from minichromosome (Ivanov and Nasmyth 2005), which indicates a topological association between cohesion and minichromosome. However, whether there is a direct link between cohesion and HBV minichromosome needs to be further investigated. Recent studies of the mechanisms of DNA compaction by cohesion provide us a new insight into the formation of cccDNA minichromosome (Davidson et al. 2019; Kim et al. 2019, 2020). In general, non-histone proteins can modulate transcription factor’s accessibility to cccDNA in either transcriptional repressive or active states. The mechanisms of some host canonical chromatin remodelers have been elucidated. However, how these remodelers participate in anti-HBV defense remains poorly understood.
Potential for the development of novel therapies
Since both Southern blot and cccDNA-specific PCR have their limitations for the detection of HBV cccDNA, different methods for quantification of cccDNA vary considerably. Therefore, it raises the possibility of using cccDNA surrogates to develop novel detection methods (Zhou et al. 2006). HBx can recruit transcription factors to transcriptionally active domain of cccDNA minichromosome and promote transcription of viral episome as well as transiently transfected plasmid (Reeves et al. 1985; van Breugel et al. 2012). However, HBx has no regulatory impact on HBV genes integrated into host chromosome (van Breugel et al. 2012). It suggests that HBx may apply a special mechanism to specifically activate expression of episome. Interestingly, researchers recently revealed that the expression of mitotic Aurora kinase A enhances viral replication in an Akt-dependent but HBx-independent manner, and DDB1 can also stimulate viral transcription via HBx-independent mechanism (Jeong and Ahn 2019; Kim et al. 2016), indicating that Aurora kinase A may be a potential substitution of HBx that would allow transcriptional stimulating of the CUL4/DDB1 complex. This property of Aurora kinase A elucidates that it might be a potential HBx surrogate and share similar signal transduction pathway as well as similar structural conformation. cccDNA is not naked but wrapped with large number of histones and non-histone proteins, which protect cccDNA from destruction by other factors, giving rise to its high stability and a long life-span. HBx or Aurora kinse A may be good potential targets for developing not only episomal DNA-targeted detection methods to improve the sensitivity and accuracy, but also for new antiviral therapies.
Gene editing enzymes comprising TALEN (Bloom et al. 2013), ZFN (Cradick et al. 2010), CRISPR/Cas9 (Moyo et al. 2018), and APOBEC (Lucifora et al. 2014) have been applied to reduce cccDNA stability to achieve the therapeutic eradication. According to the accuracy and efficiency among these gene therapies, rapidly-updated CRISPR/Cas9 tools would come to the frontline of antiviral therapeutic combat. With the use of combinations of HBV-targeting nucleases, cccDNA can be cleaved at more than one site and thus become unstable. However, in vivo precise delivery challenge and off-target effects of CRISPR/Cas9 system remain to be solved. In addition, RNA interference (RNAi) is an alternative gene therapy, including microRNAs (miRNAs), short hairpin RNAs (shRNAs), and small interfering RNAs (siRNAs) (Ely and Arbuthnot 2015; Moyo et al. 2018). Although RNAi can achieve a sustained HBV inhibition by knocking down viral transcripts, the major drawback of RNAi therapy for HBV is the failure to eliminate established cccDNA leading to HBV reoccurrence after withdrawal of gene inhibitors similar to current therapies with IFN or NAs. Hence the combination of unique viral epigenetic traits can be utilized to improve accuracy and efficiency of gene therapy tools.
Summary and perspectives
HBV cccDNA minichromosome may utilize similar epigenetic regulative mechanism as the host chromatin. Various types of histone modifications may rearrange the charge of histones to affect interactions among chromatin constituents. Moreover, multiple sites in one histone can be modified, while the same residue can be modified in various types, which dominates the intricate regulative network through antagonism or synergism. Chromatin remodeling generally results from minichromosome-associated non-histones. Due to the similarities and the differences in the catalytic ATPases, chromatin remodelers can be divided into four subfamilies: ISWI, CHD, INO80 and SWI/SNF. Besides, there are some non-canonical host remodelers such as ATRX, CSB, etc. Generally, chromatin remodelers directly bind to nucleoprotein complexes to slide, exchange, or replace nucleosome along the DNA string in an ATP-dependent manner, causing rearrangement of the relative position of histone octamer to DNA, which in consequence regulates transcription of relevant genes (Sundaramoorthy 2019). Unfortunately, there are limited studies on the mechanism of host chromatin remodelers regulating HBV genomic transcription, because the HBV episome is less abundant in infected cells and the episomal structure is quite dynamic.
Intriguingly, the recent technique advances of structural biology provide a major boost in determination of the structures of multinucleosomal complexes with linear dsDNA, which makes it feasible to determine the structures of HBV cccDNA minichromosome or other episomes in various states as well. Schalch et al. reported the 9-Å resolution crystal structure of tetranucleosomal chromatin fiber and Garcia-Saez et al. reported the 9.7-Å resolution crystal structure of hexanucleosomal chromatin fiber, which showed the advanced chromatin structure is arranged into two-start nucleosome stacks in a zigzag helix (Garcia-Saez et al. 2018; Schalch et al. 2005). Song et al. reported the 11-Å resolution cryo-EM structure of the dodecanucleosomal 30-nm chromatin fiber and ~25-Å resolution cryo-EM structure of tetracosanucleosomal 30-nm chromatin fiber, which confirmed that higher-order chromatin fibers apply a left-handed twist of the repeating tetranucleosomal units (Song et al. 2014). As expected, we could stabilize the HBV minichromosome via diverse epigenetic regulations to capture the high-resolution structures of certain conformational states to uncover the intricate regulatory mechanisms.
Novel perspective links HBV cccDNA with extrachromosomal circular DNA (ecDNA) and other viral episomal DNA. Recently, ecDNA (size range from 1 to 3 Mb or larger) found in eukaryotic species has been redefined in intimate relation to cancer pathogenesis (Verhaak et al. 2019; Wu et al. 2019). Although ecDNA can be packaged into chromatin, ecDNA chromosome lacks higher-order compaction and displays significantly enhanced chromatin accessibility compared to canonical chromatin (Wu et al. 2019). Despite intensive research concerning cccDNA formation, the mechanisms of cccDNA formation remain unclear. But there is no doubt that rcDNA would fail to be transformed into cccDNA without host DNA repair system (Guo et al. 2012), as ecDNA formation may also rely on the canonical homologous recombination (HR) or nonhomologous end joining (NHEJ)-like pathway (van Loon et al. 1994). But it can be reasonably assumed that DNA repair mechanism can be used to form higher-order chromatin due to the topological change of chromatin during rcDNA/cccDNA transformation, which can provide a novel insight into the establishment of stable minichromosome. Although cccDNA minichromosome is smaller than ecDNA chromatin or other viral episomes, the similar chromatin-like composition may indicate that the current epigenetic regulation for HBV cccDNA minichromosome might also be applied to the regulation of cancer-related ecDNA chromatin and other viral episomes in CMV (Olszewski et al. 1982), MVM (Ben-Asher et al. 1982), SV40 (Crémisi et al. 1978; Varshavsky et al. 1977), and EBV (Castán et al. 2017; Kumala et al. 2012), etc.
References
Abdul F, Filleton F, Gerossier L, Paturel A, Hall J, Strubin M, Etienne L (2018) Smc5/6 antagonism by HBx is an evolutionarily conserved function of hepatitis B virus infection in mammals. J Virol 92(16):e00769
Agarwal K, Brunetto M, Seto WK, Lim Y-S, Fung S, Marcellin P, Ahn SH, Izumi N, Chuang WL, Bae H, Sharma M, Janssen HLA, Pan CQ, Çelen MK, Furusyo N, Shalimar D, Yoon KT, Trinh H, Flaherty JF, Gaggar A, Lau AH, Cathcart AL, Lin L, Bhardwaj N, Suri V, Mani Subramanian G, Gane EJ, Buti M, Chan HLY (2018) 96 weeks treatment of tenofovir alafenamide vs. tenofovir disoproxil fumarate for hepatitis B virus infection. J Hepatol 68(4):672–681
Alarcon V, Hernández S, Rubio L, Alvarez F, Flores Y, Varas-Godoy M, De Ferrari GV, Kann M, Villanueva RA, Loyola A (2016) The enzymes LSD1 and Set1A cooperate with the viral protein HBx to establish an active hepatitis B viral chromatin state. Sci Rep 6(1):25901
Aspinall RJ, Pockros PJ (2004) The management of side-effects during therapy for hepatitis C. Aliment Pharmacol Ther 20(9):917–929
Balakrishnan L, Milavetz B (2008) HDAC inhibitors stimulate viral transcription by multiple mechanisms. Virol J 5(1):43
Beck J, Nassal M (2007) Hepatitis B virus replication. World J Gastroenterol 13(1):48–64
Belloni L, Pollicino T, De Nicola F, Guerrieri F, Raffa G, Fanciulli M, Raimondo G, Levrero M (2009) Nuclear HBx binds the HBV minichromosome and modifies the epigenetic regulation of cccDNA function. Proc Natl Acad Sci 106(47):19975
Belloni L, Allweiss L, Guerrieri F, Pediconi N, Volz T, Pollicino T, Petersen J, Raimondo G, Dandri M, Levrero M (2012) IFN-α inhibits HBV transcription and replication in cell culture and in humanized mice by targeting the epigenetic regulation of the nuclear cccDNA minichromosome. J Clin Investig 122(2):529–537
Ben-Asher E, Bratosin S, Aloni Y (1982) Intracellular DNA of the parvovirus minute virus of mice is organized in a minichromosome structure. J Virol 41(3):1044
Bloom K, Ely A, Mussolino C, Cathomen T, Arbuthnot P (2013) Inactivation of hepatitis B virus replication in cultured cells and in vivo with engineered transcription activator-like effector nucleases. Mol Ther 21(10):1889–1897
Bock C-T, Schranz P, Schröder CH, Zentgraf H (1994) Hepatitis B virus genome is organized into nucleosomes in the nucleus of the infected cell. Virus Genes 8(2):215–229
Bock CT, Schwinn S, Locarnini S, Fyfe J, Manns MP, Trautwein C, Zentgraf H (2001) Structural organization of the hepatitis B virus minichromosome11Edited by M. Yaniv. J Mol Biol 307(1):183–196
Castán A, Fernández-Calleja V, Hernández P, Krimer DB, Schvartzman JB, Fernández-Nestosa M-J (2017) Analysis of DNA topology of EBV minichromosomes in HEK 293 cells. PLoS ONE 12(11):e0188172
Chen C, Wu M, Zhang W, Lu W, Zhang M, Zhang Z, Zhang X, Yuan Z (2016) MicroRNA-939 restricts Hepatitis B virus by targeting Jmjd3-mediated and C/EBPα-coordinated chromatin remodeling. Sci Rep 6(1):35974
Chiappinelli KB, Strissel Pamela L, Desrichard A, Li H, Henke C, Akman B, Hein A, Rote Neal S, Cope Leslie M, Snyder A, Makarov V, Buhu S, Slamon Dennis J, Wolchok Jedd D, Pardoll Drew M, Beckmann Matthias W, Zahnow Cynthia A, Merghoub T, Chan Timothy A, Baylin Stephen B, Strick R (2015) Inhibiting DNA methylation causes an interferon response in cancer via dsRNA including endogenous retroviruses. Cell 162(5):974–986
Chong CK, Cheng CYS, Tsoi SYJ, Huang F-Y, Liu F, Seto W-K, Lai C-L, Yuen M-F, Wong DK-H (2017) Role of hepatitis B core protein in HBV transcription and recruitment of histone acetyltransferases to cccDNA minichromosome. Antiviral Res 144:1–7
Cougot D, Wu Y, Cairo S, Caramel J, Renard C-A, Lévy L, Buendia MA, Neuveut C (2007) The hepatitis B virus X protein functionally interacts with CREB-binding protein/p300 in the regulation of CREB-mediated transcription. J Biol Chem 282(7):4277–4287
Cradick TJ, Keck K, Bradshaw S, Jamieson AC, McCaffrey AP (2010) Zinc-finger nucleases as a novel therapeutic strategy for targeting hepatitis B virus DNAs. Mol Ther 18(5):947–954
Crémisi C, Chestier A, Yaniv M (1978) Assembly of SV40 and polyoma minichromosomes during replication. Cold Spring Harb Symp Quant Biol 42:409–416
Davidson IF, Bauer B, Goetz D, Tang W, Wutz G, Peters J-M (2019) DNA loop extrusion by human cohesin. Science 366(6471):1338
Decorsière A, Mueller H, van Breugel PC, Abdul F, Gerossier L, Beran RK, Livingston CM, Niu C, Fletcher SP, Hantz O, Strubin M (2016) Hepatitis B virus X protein identifies the Smc5/6 complex as a host restriction factor. Nature 531(7594):386–389
Deng J-J, Kong K-YE, Gao W-W, Tang H-MV, Chaudhary V, Cheng Y, Zhou J, Chan C-P, Wong DK-H, Yuen MF, Jin D-Y (2017) Interplay between SIRT1 and hepatitis B virus X protein in the activation of viral transcription. Biochimica et Biophysica Acta (BBA) 1860(4):491–501
Deuschle K, Kepp G, Jeske H (2016) Differential methylation of the circular DNA in geminiviral minichromosomes. Virology 499:243–258
Dezhbord M, Lee S, Kim W, Seong BL, Ryu W-S (2019) Characterization of the molecular events of covalently closed circular DNA synthesis in de novo Hepatitis B virus infection of human hepatoma cells. Antiviral Res 163:11–18
Ely A, Arbuthnot P (2015) Differing prospects for the future of using gene therapy to treat infections with hepatitis B virus and hepatitis C virus. Discov Med 20(109):137–143
Feng J, Yang G, Liu Y, Gao Y, Zhao M, Bu Y, Yuan H, Yuan Y, Yun H, Sun M, Gao H, Zhang S, Liu Z, Yin M, Song X, Miao Z, Lin Z, Zhang X (2019) LncRNA PCNAP1 modulates hepatitis B virus replication and enhances tumor growth of liver cancer. Theranostics 9(18):5227–5245
Fontana RJ (2009) Side effects of long-term oral antiviral therapy for hepatitis B. Hepatology 49(S5):S185–S195
Gao J, Xiong Y, Wang Y, Wang Y, Zheng G, Xu H (2016) Hepatitis B virus X protein activates Notch signaling by its effects on Notch1 and Notch4 in human hepatocellular carcinoma. Int J Oncol 48(1):329–337
Gao Y, Feng J, Yang G, Zhang S, Liu Y, Bu Y, Sun M, Zhao M, Chen F, Zhang W, Ye L, Zhang X (2017) Hepatitis B virus X protein–elevated MSL2 modulates hepatitis B virus covalently closed circular DNA by inducing degradation of APOBEC3B to enhance hepatocarcinogenesis. Hepatology 66(5):1413–1429
Gao W, Jia Z, Tian Y, Yang P, Sun H, Wang C, Ding Y, Zhang M, Zhang Y, Yang D, Tian Z, Zhou J, Ruan Z, Wu Y, Ni B (2019) HBx protein contributes to liver carcinogenesis by H3K4me3 modification through stabilizing WD repeat domain 5 protein. Hepatology 71(5):1678–1695
Garcia-Saez I, Menoni H, Boopathi R, Shukla MS, Soueidan L, Noirclerc-Savoye M, Le Roy A, Skoufias DA, Bednar J, Hamiche A (2018) Structure of an H1-bound 6-nucleosome array reveals an untwisted two-start chromatin fiber conformation. Mol Cell 72(5):902–915
Gómez-Moreno A, Garaigorta U (2017) Hepatitis B virus and DNA damage response: interactions and consequences for the infection. Viruses 9(10):304
Gong Q, Chen S, Guo J, Sun H, Zheng G, Liu Q, Ren H, He S (2011) Chromosome remodeling related to hepatitis B virus replication in HepG2 cells. DNA Cell Biol 30(6):347–354
Guerrieri F, Belloni L, D’Andrea D, Pediconi N, Le Pera L, Testoni B, Scisciani C, Floriot O, Zoulim F, Tramontano A, Levrero M (2017) Genome-wide identification of direct HBx genomic targets. BMC Genomics 18(1):184
Guo H, Jiang D, Zhou T, Cuconati A, Block TM, Guo J-T (2007) Characterization of the intracellular deproteinized relaxed circular DNA of hepatitis B virus: an intermediate of covalently closed circular DNA formation. J Virol 81(22):12472
Guo Y, Li Y, Mu S, Zhang J, Yan Z (2009) Evidence that methylation of hepatitis B virus covalently closed circular DNA in liver tissues of patients with chronic hepatitis B modulates HBV replication. J Med Virol 81(7):1177–1183
Guo H, Mao R, Block TM, Guo J-T (2010) Production and function of the cytoplasmic deproteinized relaxed circular DNA of hepadnaviruses. J Virol 84(1):387
Guo Y-H, Li Y-N, Zhao J-R, Zhang J, Yan Z (2011) HBc binds to the CpG islands of HBV cccDNA and promotes an epigenetic permissive state. Epigenetics 6(6):720–726
Guo H, Xu C, Zhou T, Block TM, Guo J-T (2012) Characterization of the host factors required for hepadnavirus covalently closed circular (ccc) DNA formation. PLoS ONE 7(8):e43270
Guo F, Zhao Q, Sheraz M, Cheng J, Qi Y, Su Q, Cuconati A, Wei L, Du Y, Li W, Chang J, Guo J-T (2017) HBV core protein allosteric modulators differentially alter cccDNA biosynthesis from de novo infection and intracellular amplification pathways. PLoS Pathog 13(9):e1006658
Halmer L, Vestner B, Gruss C (1998) Involvement of topoisomerases in the initiation of simian virus 40 minichromosome replication. J Biol Chem 273(52):34792–34798
Hensel KO, Cantner F, Bangert F, Wirth S, Postberg J (2018) Episomal HBV persistence within transcribed host nuclear chromatin compartments involves HBx. Epigenet Chromatin 11(1):34
Holland L, Downey M, Song X, Gauthier L, Bell-Rogers P, Yankulov K (2002) Distinct parts of minichromosome maintenance protein 2 associate with histone H3/H4 and RNA polymerase II holoenzyme. Eur J Biochem 269(21):5192–5202
Hu J, Protzer U, Siddiqui A (2019) Revisiting hepatitis B virus: challenges of curative therapies. J Virol 93(20):e01032
Ishimi Y, Ichinose S, Omori A, Sato K, Kimura H (1996) Binding of human minichromosome maintenance proteins with histone H3. J Biol Chem 271(39):24115–24122
Ishimi Y, Komamura Y, You Z, Kimura H (1998) Biochemical function of mouse minichromosome maintenance 2 protein. J Biol Chem 273(14):8369–8375
Ivanov D, Nasmyth K (2005) A topological interaction between cohesin rings and a circular minichromosome. Cell 122(6):849–860
Jeong GU, Ahn B-Y (2019) Aurora kinase A promotes hepatitis B virus replication and expression. Antiviral Res 170:104572
Jin X-L, Hong SK, Kim H, Lee S-K, Yi N-J, Lee K-W, Suh K-S (2019) Antiviral therapy may decrease HBx, affecting cccDNA and MSL2 in hepatocarcinogenesis. Oncol Lett 18(5):4984–4991
Kallestad L, Woods E, Christensen K, Gefroh A, Balakrishnan L, Milavetz B (2013) Transcription and replication result in distinct epigenetic marks following repression of early gene expression. Front Genet 4:140
Kim JW, Lee SH, Park YS, Hwang JH, Jeong SH, Kim N, Lee DH (2011) Replicative activity of hepatitis B virus is negatively associated with methylation of covalently closed circular DNA in advanced hepatitis B virus infection. Intervirology 54(6):316–325
Kim W, Lee S, Son Y, Ko C, Ryu W-S (2016) DDB1 stimulates viral transcription of hepatitis B virus via HBx-independent mechanisms. J Virol 90(21):9644
Kim Y, Shi Z, Zhang H, Finkelstein IJ, Yu H (2019) Human cohesin compacts DNA by loop extrusion. Science 366(6471):1345
Kim E, Kerssemakers J, Shaltiel IA, Haering CH, Dekker C (2020) DNA-loop extruding condensin complexes can traverse one another. Nature. https://doi.org/10.1038/s41586-020-2067-5
Kinoshita W, Ogura N, Watashi K, Wakita T (2017) Host factor PRPF31 is involved in cccDNA production in HBV-replicating cells. Biochem Biophys Res Commun 482(4):638–644
Köck J, Rösler C, Zhang J-J, Blum HE, Nassal M, Thoma C (2010) Generation of covalently closed circular DNA of hepatitis B viruses via intracellular recycling is regulated in a virus specific manner. PLoS Pathog 6(9):e1001082
Königer C, Wingert I, Marsmann M, Rösler C, Beck J, Nassal M (2014) Involvement of the host DNA-repair enzyme TDP2 in formation of the covalently closed circular DNA persistence reservoir of hepatitis B viruses. Proc Natl Acad Sci 111(40):E4244
Kumala S, Hadj-Sahraoui Y, Rzeszowska-Wolny J, Hancock R (2012) DNA of a circular minichromosome linearized by restriction enzymes or other reagents is resistant to further cleavage: an influence of chromatin topology on the accessibility of DNA. Nucleic Acids Res 40(19):9417–9428
Lai C-L, Wong D, Ip P, Kopaniszen M, Seto W-K, Fung J, Huang F-Y, Lee B, Cullaro G, Chong CK, Wu R, Cheng C, Yuen J, Ngai V, Yuen M-F (2017) Reduction of covalently closed circular DNA with long-term nucleos(t)ide analogue treatment in chronic hepatitis B. J Hepatol 66(2):275–281
Landsberg CD, Megger DA, Hotter D, Rückborn MU, Eilbrecht M, Rashidi-Alavijeh J, Howe S, Heinrichs S, Sauter D, Sitek B, Le-Trilling VTK, Trilling M (2018) A mass spectrometry-based profiling of interactomes of viral DDB1- and cullin ubiquitin ligase-binding proteins reveals NF-κB inhibitory activity of the HIV-2-encoded Vpx. Front Immunol 9:2978
Levrero M, Pollicino T, Petersen J, Belloni L, Raimondo G, Dandri M (2009) Control of cccDNA function in hepatitis B virus infection. J Hepatol 51(3):581–592
Li T, Robert EI, van Breugel PC, Strubin M, Zheng N (2010) A promiscuous alpha-helical motif anchors viral hijackers and substrate receptors to the CUL4-DDB1 ubiquitin ligase machinery. Nat Struct Mol Biol 17(1):105–111
Li M, Zhao H, Zhang X, Wood LD, Anders RA, Choti MA, Pawlik TM, Daniel HD, Kannangai R, Offerhaus GJA, Velculescu VE, Wang L, Zhou S, Vogelstein B, Hruban RH, Papadopoulos N, Cai J, Torbenson MS, Kinzler KW (2011) Inactivating mutations of the chromatin remodeling gene ARID2 in hepatocellular carcinoma. Nat Genet 43(9):828–829
Li W, Cao F, Li J, Wang Z, Ren Y, Liang Z, Liu P (2016) Simvastatin exerts anti-hepatitis B virus activity by inhibiting expression of minichromosome maintenance protein 7 in HepG2.2.15 cells. Mol Med Rep 14(6):5334–5342
Liu F, Campagna M, Qi Y, Zhao X, Guo F, Xu C, Li S, Li W, Block TM, Chang J (2013) Alpha-interferon suppresses hepadnavirus transcription by altering epigenetic modification of cccDNA minichromosomes. PLoS Pathog 9(9):e1003613
Liu Y, Feng J, Sun M, Yang G, Yuan H, Wang Y, Bu Y, Zhao M, Zhang S, Zhang X (2019) Long non-coding RNA HULC activates HBV by modulating HBx/STAT3/miR-539/APOBEC3B signaling in HBV-related hepatocellular carcinoma. Cancer Lett 454:158–170
Lucifora J, Arzberger S, Durantel D, Belloni L, Strubin M, Levrero M, Zoulim F, Hantz O, Protzer U (2011) Hepatitis B virus X protein is essential to initiate and maintain virus replication after infection. J Hepatol 55(5):996–1003
Lucifora J, Xia Y, Reisinger F, Zhang K, Stadler D, Cheng X, Sprinzl MF, Koppensteiner H, Makowska Z, Volz T, Remouchamps C, Chou W-M, Thasler WE, Hüser N, Durantel D, Liang TJ, Münk C, Heim MH, Browning JL, Dejardin E, Dandri M, Schindler M, Heikenwalder M, Protzer U (2014) Specific and nonhepatotoxic degradation of nuclear hepatitis B virus cccDNA. Science 343(6176):1221
Méndez J, Stillman B (2000) Chromatin association of human origin recognition complex, Cdc6, and minichromosome maintenance proteins during the cell cycle: assembly of prereplication complexes in late mitosis. Mol Cell Biol 20(22):8602
Moyo B, Bloom K, Scott T, Ely A, Arbuthnot P (2018) Advances with using CRISPR/Cas-mediated gene editing to treat infections with hepatitis B virus and hepatitis C virus. Virus Research 244:311–320
Murphy CM, Xu Y, Li F, Nio K, Reszka-Blanco N, Li X, Wu Y, Yu Y, Xiong Y, Su L (2016) Hepatitis B virus X protein promotes degradation of SMC5/6 to enhance HBV replication. Cell Rep 16(11):2846–2854
Niu C, Livingston CM, Li L, Beran RK, Daffis S, Ramakrishnan D, Burdette D, Peiser L, Salas E, Ramos H, Yu M, Cheng G, Strubin M, Delaney WEIV, Fletcher SP (2017) The Smc5/6 complex restricts HBV when Localized to ND10 without inducing an innate immune response and is counteracted by the HBV X protein shortly after infection. PLoS ONE 12(1):e0169648
Nkongolo S, Nußbaum L, Lempp FA, Wodrich H, Urban S, Ni Y (2019) The retinoic acid receptor (RAR) α-specific agonist Am 80 (tamibarotene) and other RAR agonists potently inhibit hepatitis B virus transcription from cccDNA. Antiviral Res 168:146–155
Olszewski N, Hagen G, Guilfoyle TJ (1982) A transcriptionally active, covalently closed minichromosome of cauliflower mosaic virus DNA isolated from infected turnip leaves. Cell 29(2):395–402
Palumbo GA, Scisciani C, Pediconi N, Lupacchini L, Alfalate D, Guerrieri F, Calvo L, Salerno D, Di Cocco S, Levrero M, Belloni L (2015) IL6 inhibits HBV transcription by targeting the epigenetic control of the nuclear cccDNA minichromosome. PLoS ONE 10(11):1–14
Papatheodoridis GV, Dimou E, Papadimitropoulos V (2002) Nucleoside analogues for chronic hepatitis B: antiviral efficacy and viral resistance. Am J Gastroenterol 97(7):1618–1628
Peng JC, Karpen GH (2007) H3K9 methylation and RNA interference regulate nucleolar organization and repeated DNA stability. Nat Cell Biol 9(1):25–35
Peng JC, Karpen GH (2009) Heterochromatic genome stability requires regulators of histone H3 K9 methylation. PLoS Genet 5(3):e1000435
Pollicino T, Belloni L, Raffa G, Pediconi N, Squadrito G, Raimondo G, Levrero M (2006) Hepatitis B virus replication is regulated by the Acetylation status of hepatitis B virus cccDNA-bound H3 and H4 histones. Gastroenterology 130(3):823–837
Qian G, Hu B, Zhou D, Xuan Y, Bai L, Duan C (2015) NIRF, a novel ubiquitin ligase, inhibits hepatitis B virus replication through effect on HBV core protein and H3 histones. DNA Cell Biol 34(5):327–332
Ramakrishnan D, Xing W, Beran RK, Chemuru S, Rohrs H, Niedziela-Majka A, Marchand B, Mehra U, Zábranský A, Doležal M, Hubálek M, Pichová I, Gross ML, Kwon HJ, Fletcher SP (2019) Hepatitis B virus X protein function requires zinc binding. J Virol 93(16):e00250
Reeves R, Gorman CM, Howard B (1985) Minichromosome assembly of non-integrated plasmid DNA transfected into mammalian cells. Nucleic Acids Res 13(10):3599–3615
Ren J-H, Tao Y, Zhang Z-Z, Chen W-X, Cai X-F, Chen K, Ko BCB, Song C-L, Ran L-K, Li W-Y, Huang A-L, Chen J (2014) Sirtuin 1 regulates hepatitis B virus transcription and replication by targeting transcription factor AP-1. J Virol 88(5):2442
Ren J-H, Hu J-L, Cheng S-T, Yu H-B, Wong VKW, Law BYK, Yang Y-F, Huang Y, Liu Y, Chen W-X, Cai X-F, Tang H, Hu Y, Zhang W-L, Liu X, Long Q-X, Zhou L, Tao N-N, Zhou H-Z, Yang Q-X, Ren F, He L, Gong R, Huang A-L, Chen J (2018) SIRT3 restricts hepatitis B virus transcription and replication through epigenetic regulation of covalently closed circular DNA involving suppressor of variegation 3-9 homolog 1 and SET domain containing 1A histone methyltransferases. Hepatology 68(4):1260–1276
Rivière L, Gerossier L, Ducroux A, Dion S, Deng Q, Michel M-L, Buendia M-A, Hantz O, Neuveut C (2015) HBx relieves chromatin-mediated transcriptional repression of hepatitis B viral cccDNA involving SETDB1 histone methyltransferase. J Hepatol 63(5):1093–1102
Rivière L, Quioc-Salomon B, Fallot G, Halgand B, Féray C, Buendia M-A, Neuveut C (2019) Hepatitis B virus replicating in hepatocellular carcinoma encodes HBx variants with preserved ability to antagonize restriction by Smc5/6. Antiviral Res 172:104618
Saeed U, Kim J, Piracha ZZ, Kwon H, Jung J, Chwae Y-J, Park S, Shin H-J, Kim K (2019) Parvulin 14 and parvulin 17 bind to HBx and cccDNA and upregulate hepatitis B virus replication from cccDNA to virion in an HBx-dependent manner. J Virol 93(6):e01840
Salerno D, Chiodo L, Alfano V, Floriot O, Cottone G, Paturel A, Pallocca M, Plissonnier M-L, Jeddari S, Belloni L, Zeisel M, Levrero M, Guerrieri F (2020) Hepatitis B protein HBx binds the DLEU2 lncRNA to sustain cccDNA and host cancer-related gene transcription. Gut. https://doi.org/10.1136/gutjnl-2019-319637:gutjnl-2019-319637
Schalch T, Duda S, Sargent DF, Richmond TJ (2005) X-ray structure of a tetranucleosome and its implications for the chromatin fibre. Nature 436(7047):138–141
Sekiba K, Otsuka M, Ohno M, Yamagami M, Kishikawa T, Suzuki T, Ishibashi R, Seimiya T, Tanaka E, Koike K (2019) Inhibition of HBV transcription from cccDNA With Nitazoxanide by targeting the HBx–DDB1 interaction. Cell Mol Gastroenterol Hepatol 7(2):297–312
Sengupta I, Das D, Singh SP, Chakravarty R, Das C (2017) Host transcription factor Speckled 110 kDa (Sp110), a nuclear body protein, is hijacked by hepatitis B virus protein X for viral persistence. J Biol Chem 292(50):20379–20393
Shamay M, Barak O, Doitsh G, Ben-Dor I, Shaul Y (2002) Hepatitis B virus pX interacts with HBXAP, a PHD finger protein to coactivate transcription. J Biol Chem 277(12):9982–9988
Shen B, Chen Y, Hu J, Qiao M, Ren J, Hu J, Chen J, Tang N, Huang A, Hu Y (2020) Hepatitis B virus X protein modulates upregulation of DHX9 to promote viral DNA replication. Cell Microbiol 22(3):e13148
Sheraz M, Cheng J, Tang L, Chang J, Guo J-T (2019) Cellular DNA topoisomerases are required for the synthesis of hepatitis B virus covalently closed circular DNA. J Virol 93(11):e02230
Shi L, Li S, Shen F, Li H, Qian S, Lee DHS, Wu JZ, Yang W (2012) Characterization of nucleosome positioning in hepadnaviral covalently closed circular DNA minichromosomes. J Virol 86(18):10059
Shi A, Zhang X, Xiao F, Zhu L, Yan W, Han M, Luo X, Chen T, Ning Q (2018) CD56bright natural killer cells induce HBsAg reduction via cytolysis and cccDNA decay in long-term entecavir-treated patients switching to peginterferon alfa-2a. J Viral Hepatitis 25(11):1352–1362
Song F, Chen P, Sun D, Wang M, Dong L, Liang D, Xu R-M, Zhu P, Li G (2014) Cryo-EM study of the chromatin fiber reveals a double helix twisted by tetranucleosomal units. Science 344(6182):376–380
Sundaramoorthy R (2019) Nucleosome remodelling: structural insights into ATP-dependent remodelling enzymes. Essays Biochem 63(1):45–58
Tak E, Hwang S, Lee HC, Ko G-Y, Ahn C-S, Yoon Y-I, Lim Y-S, Jun D-Y, Kim K-H, Song G-W, Moon D-B, Ryoo B-Y, Kim N, Lee S-G (2016) Apoptosis of hepatitis B virus-expressing liver tumor cells induced by a high concentration of nucleos(t)ide analogue. Anticancer Res 36(11):6059–6069
Tan G, Xu F, Song H, Yuan Y, Xiao Q, Ma F, Qin FX-F, Cheng G (2018) Identification of TRIM14 as a type I IFN-stimulated gene controlling hepatitis B virus replication by targeting HBx. Front Immunol 9:1872
Tang Y, Zhang Y, Wang C, Sun Z, Li L, Dong J, Zhou W (2018) 14-3-3ζ binds to hepatitis B virus protein X and maintains its protein stability in hepatocellular carcinoma cells. Cancer Med 7(11):5543–5553
Tao S, Pan S, Gu C, Wei L, Kang N, Xie Y, Liu J (2019) Characterization and engineering of broadly reactive monoclonal antibody against hepatitis B virus X protein that blocks its interaction with DDB1. Sci Rep 9(1):20323
Tropberger P, Mercier A, Robinson M, Zhong W, Ganem DE, Holdorf M (2015) Mapping of histone modifications in episomal HBV cccDNA uncovers an unusual chromatin organization amenable to epigenetic manipulation. Proc Natl Acad Sci USA 112(42):E5715
van Breugel PC, Robert EI, Mueller H, Decorsiere A, Zoulim F, Hantz O, Strubin M (2012) Hepatitis B virus X protein stimulates gene expression selectively from extrachromosomal DNA templates. Hepatology 56(6):2116–2124
van Loon N, Miller D, Murnane JP (1994) Formation of extrachromosomal circular DNA in HeLa cells by nonhomologous recombination. Nucleic Acids Res 22(13):2447–2452
Vaquero A, Scher M, Erdjument-Bromage H, Tempst P, Serrano L, Reinberg D (2007) SIRT1 regulates the histone methyl-transferase SUV39H1 during heterochromatin formation. Nature 450(7168):440–444
Varshavsky AJ, Nedospasov SA, Schmatchenko VV, Bakayev VV, Chumackov PM, Georgiev GP (1977) Compact form of SV40 viral minichromosome is resistant to nuclease: possible implications for chromatin structure. Nucleic Acids Res 4(10):3303–3325
Verhaak RGW, Bafna V, Mischel PS (2019) Extrachromosomal oncogene amplification in tumour pathogenesis and evolution. Nat Rev Cancer 19(5):283–288
Vivekanandan P, Thomas D, Torbenson M (2009) Methylation regulates hepatitis B viral protein expression. J Infect Dis 199(9):1286–1291
Vivekanandan P, Daniel HD-J, Kannangai R, Martinez-Murillo F, Torbenson M (2010) Hepatitis B virus replication induces methylation of both host and viral DNA. J Virol 84(9):4321
Wang F, Zhou H, Xia X, Sun Q, Wang Y, Cheng B (2010) Activated Notch signaling is required for hepatitis B virus X protein to promote proliferation and survival of human hepatic cells. Cancer Lett 298(1):64–73
Wei Z-Q, Zhang Y-H, Ke C-Z, Chen H-X, Ren P, He Y-L, Hu P, Ma D-Q, Luo J, Meng Z-J (2017) Curcumin inhibits hepatitis B virus infection by down-regulating cccDNA-bound histone acetylation. World J Gastroenterol 23(34):6252–6260
WHO (2017) Global hepatitis report
Wong DK-H, Seto W-K, Fung J, Ip P, Huang F-Y, Lai C-L, Yuen M-F (2013) Reduction of hepatitis B surface antigen and covalently closed circular DNA by nucleos(t)ide analogues of different potency. Clin Gastroenterol Hepatol 11(8):1004
Wu S, Turner KM, Nguyen N, Raviram R, Erb M, Santini J, Luebeck J, Rajkumar U, Diao Y, Li B, Zhang W, Jameson N, Corces MR, Granja JM, Chen X, Coruh C, Abnousi A, Houston J, Ye Z, Hu R, Yu M, Kim H, Law JA, Verhaak RGW, Hu M, Furnari FB, Chang HY, Ren B, Bafna V, Mischel PS (2019) Circular ecDNA promotes accessible chromatin and high oncogene expression. Nature 575(7784):699–703
Xie Q, Zhang S, Wang W, Li YM, Du T, Su XL, Wei YQ, Deng HX (2012) Inhibition of hepatitis B virus gene expression by small interfering RNAs targeting cccDNA and X antigen. Acta Virol 56(01):49–55
Xu W, Ma C, Zhang Q, Zhao R, Hu D, Zhang X, Chen J, Liu F, Wu K, Liu Y, Wu J (2018) PJA1 coordinates with the SMC5/6 complex to restrict DNA viruses and episomal genes in an interferon-independent manner. J Virol 92(22):e00825
Yang G, Feng J, Liu Y, Zhao M, Yuan Y, Yuan H, Yun H, Sun M, Bu Y, Liu L, Liu Z, Niu J-Q, Yin M, Song X, Miao Z, Lin Z, Zhang X (2019) HAT1 signaling confers to assembly and epigenetic regulation of HBV cccDNA minichromosome. Theranostics 9(24):7345–7358
Yeh C-T, Chiu H-T, Chu C-M, Liaw Y-F (1998) G1 phase dependent nuclear localization of relaxed-circular hepatitis B virus DNA and aphidicolin-induced accumulation of covalently closed circular DNA. J Med Virol 55(1):42–50
Yuan Y, Zhao K, Yao Y, Liu C, Chen Y, Li J, Wang Y, Pei R, Chen J, Hu X, Zhou Y, Wu C, Chen X (2019) HDAC11 restricts HBV replication through epigenetic repression of cccDNA transcription. Antiviral Res 172:104619
Zhang Y, Mao R, Yan R, Cai D, Zhang Y, Zhu H, Kang Y, Liu H, Wang J, Qin Y, Huang Y, Guo H, Zhang J (2014) Transcription of hepatitis B virus covalently closed circular DNA is regulated by CpG methylation during chronic infection. PLoS ONE 9(10):e110442
Zhang W, Chen J, Wu M, Zhang X, Zhang M, Yue L, Li Y, Liu J, Li B, Shen F, Wang Y, Bai L, Protzer U, Levrero M, Yuan Z (2017a) PRMT5 restricts hepatitis B virus replication through epigenetic repression of covalently closed circular DNA transcription and interference with pregenomic RNA encapsidation. Hepatology 66(2):398–415
Zhang Y, He S, Guo J-J, Peng H, Fan J-H, Li Q-L (2017b) Retinoid X receptor α-dependent HBV minichromosome remodeling and viral replication. Ann Hepatol 16(4):501–509
Zhang D, Wang Y, Zhang H-Y, Jiao F-Z, Zhang W-B, Wang L-W, Zhang H, Gong Z-J (2019) Histone deacetylases and acetylated histone H3 are involved in the process of hepatitis B virus DNA replication. Life Sci 223:1–8
Zhou T, Guo H, Guo J-T, Cuconati A, Mehta A, Block TM (2006) Hepatitis B virus e antigen production is dependent upon covalently closed circular (ccc) DNA in HepAD38 cell cultures and may serve as a cccDNA surrogate in antiviral screening assays. Antiviral Res 72(2):116–124
Zhou Z, Hu T, Zhou X, Wildum S, Garcia-Alcalde F, Xu Z, Wu D, Mao Y, Tian X, Zhou Y, Shen F, Zhang Z, Tang G, Najera I, Yang G, Shen HC, Young JAT, Qin N (2017) Heteroaryldihydropyrimidine (HAP) and sulfamoylbenzamide (SBA) inhibit hepatitis B Virus replication by different molecular mechanisms. Sci Rep 7(1):42374
Acknowledgements
We thank Prof. Yu Wei (Institut Pasteur of Shanghai, CAS) for her constructive comment. We apologize to scientists in this field, whose publications were not cited due to space limit. This work was supported by the Strategic Priority Research Program of CAS (XDB29010205), the National Key R&D Program of China (2018YFA0507303, 2018YFC1200701), and National Natural Science Foundation of China (31770816).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Lanfeng Wang, Weiwei Wang, and Zhaoning Wang declare that they have no conflict of interest.
Human and animal rights and informed consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, Z., Wang, W. & Wang, L. Epigenetic regulation of covalently closed circular DNA minichromosome in hepatitis B virus infection. Biophys Rep 6, 115–126 (2020). https://doi.org/10.1007/s41048-020-00112-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41048-020-00112-z