Abstract
Background
The use of plant extracts as antibiotics is gaining more attention because bacteria are one of the biggest threats to global health and the resistance of bacteria to antibiotics in humans and animals is increasing. Antibacterial susceptibility is used to determine which specific antibiotics a particular bacterium is sensitive to. This research is focused on the phytochemical, in vitro antibacterial susceptibility, and in silico analysis of Morinda lucida root extracts against gram-negative Pseudomonas aeruginosa and gram-positive Staphylococcus aureus. The root of M. lucida was extracted separately with ethanol, chloroform, and ethyl acetate.
Results
Crude extracts of M. lucida had active antibacterial activity and were a promising natural antibiotic when compared to gentamicin. The in vitro results showed that the extracts of M. lucida had good susceptibility properties against the two drug-resistant bacteria while the in silico showed that 2-hydroxy-1-methoxy anthraquinone is predicted to have a better susceptibility with S. aureus while alizarin has better susceptibility properties against P. aeruginosa. Finally, the MD simulation studies of Alizarin and 9,10-Anthracenedione 2-hydroxy-1- methoxy- define the stability of protein–ligand complexes within a 50 ns time scale.
Article Highlights
The research article studied the way selected compounds from the Morinda lucida plant affects selected bacteria. This study was carried our in vitro and in silico so as to identify why anthraquinone poses the best potential for novel drugs. The alizarin compound and the compound 2-hydroxyl-1-methoxyanthraquinone have the best properties.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Morinda lucida is a plant species that is widely distributed in the tropics, especially in West Africa. The plant has numerous medicinal properties. It is used traditionally in various communities for the treatment of different ailments [1]. One of the reported applications of M. lucida is the treatment of bacterial infections [2, 3]. In recent years, the use of plant extracts for the development of new antibacterial agents has been on the increase, particularly in the face of the increasing prevalence of antibiotic-resistant bacteria [4, 5]. Approximately 80% of people worldwide still rely on plant medicines for relief from illnesses, according to the World Health Organization (WHO). Since they are an essential source of physiologically active compounds, medicinal plants have a big impact on human health. [6,7,8]The study of M. lucida root extracts by in vitro and in silico antibacterial analysis is highly intriguing as it offers important information about the plant's potential for producing new antibacterial chemicals. Previous studies have reported the antibacterial potential of the leaves and stem bark of M. lucida against a range of bacterial strains and its use in traditional medicine [2, 5, 9,10,11]. The M. lucida leaf extracts have been found to have the potential to inhibit the growth of E. coli and Salmonella species, indicating its potential in the treatment of infections involving these bacteria [6]. Morinda lucida leaf aqueous extract has strong immunostimulatory and immunorestorative qualities, which may account for why it works so well for treating infections and immuno-inflammatory conditions [7]. The ethanol extract of M. lucida leaves possesses antiemetic properties, suggesting its potential for use in the development of new antiemetic drugs [8] and the leaves may have potential applications in anticoccidial therapy [12].
A different study discovered that the M. lucida leaf aqueous extract had the greatest inhibitory effect on two enzymes (α-Amylase and α-Glucosidase), and this activity might be because of phytochemicals like tannins, flavonoids, and saponins [13]. Importantly, research indicates that consuming M. lucida leaf extract has no negative effects on liver or kidney function, indicating that it might be a safe therapeutic herb [14]. According to Saganuwan et al., the kidney, lung, liver, heart, and spleen may have mild to moderate adverse effects at higher dosages. However, the aqueous extract of M. lucida is safe when administered for 28 days at test dosages of 16.26 and 162.65 mg/kg body weight. [15]The Staphylococcus aureus (a Gram-positive multidrug resistant [16, 17] facultative anaerobe bacteria typically causing a variety of clinical diseases [18]) and Pseudomonas aeruginosa (a ubiquitous facultatively anaerobic, toxin-producing gram-negative bacterium [19] associated with serious infections in immunocompromised hosts [20]). The GC–MS analysis of M. lucida was discussed by Adekunle et al., [21] indicating that the root of M. lucida is very rich in phytocompounds. The information regarding the phytochemical profile and biological functions of M. lucida roots has not yet been fully utilized. As a result, the study sought to qualitatively examine the phytochemical components and assess the antibacterial properties of the chloroform extract of M. lucida root on several chosen clinical bacterial pathogens. However, the root extract of M. lucida has not been extensively studied for its antibacterial potential, especially against antibiotic-resistant strains.
Therefore, this study investigates the antibacterial activity of M. lucida root extracts against selected two bacterial strains using in vitro and in silico assays.
2 Methodology
2.1 Materials and Methods
2.1.1 Bacteria
Selected gram-positive and gram-negative bacterial strains include; Staphylococcus aureus and Pseudomonas aeruginosa used in the in vitro antibacterial susceptibility study of M. lucida root crude extracts on the bacterial species. These isolates were subcultured onto sterile nutrient agar plates and incubated for 24 h at 37ºC, and the pure cultures were used for the antibacterial analysis.
2.1.2 Plant Sample
The fresh roots of M. lucida were carefully harvested from the Ebualawe Area, Iwo, Osun State, Nigeria in July 2021. Plant specimens were authenticated and a voucher specimen number was given (BUH036). The roots were washed and air dried for two weeks after cutting into small chunks then homogenized to fine powder which was then stored in airtight containers for further analysis.
2.1.3 Extraction
After macerating 150 g of powdered M. lucida root in 1 L of chloroform for 48 h at room temperature with gentle shaking, the material was filtered through filter paper (Whatman, No. 42). The process was repeated three times to ensure adequate extraction. The extracts were combined and concentrated using a rotary evaporator. Phytochemicals were screened using the procedure outlined by Harborne (1984) [22] before being analyzed, the concentrated extracts were sealed and kept in a refrigerator. The yield of the extract was determined and noted.
2.1.4 Antibacterial assay
The agar diffusion method was used to evaluate the in vitro antibacterial activities of the extracts [23] using a modification of Adeleke and Owoseni [24] protocol and the guidelines of the Clinical Laboratory Standards Institute [25] with slight modifications. 10% DMSO served as the negative control and gentamicin as the positive reference standard medication for the test bacterium. The agar plates were inoculated with the bacteria suspension using a sterile swab on the surface of a Mueller–Hinton agar plate. After applying uniformly, they were allowed to dry for a few minutes. Using sterile forceps, the gentamicin-impregnated disks were placed onto the inoculated agar surface, Precautions was taken to ensure that there was sufficient spacing between the disks and a complete contact between the disk and the agar surface. The inoculated agar plates were incubated at 35 °C for a period of 24 h.
The average zone of susceptibility and average deviation for the test substances against the bacteria were recorded during the duplicate experimentation.
2.1.5 Data organization and statistical analysis
With the aid of Microsoft Word 2016 and Microsoft Excel 2016, the data was arranged and tallied. The studies were performed in triplicate, and the mean and standard deviation (MSD) were determined as the average of the clear zone and standard deviations (SD).
2.2 Molecular docking of Compounds from Plant Extracts of Morinda lucida against selected Microorganisms.
2.2.1 Preparation of compounds
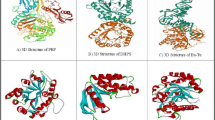
The compounds used for molecular docking studies were retrieved from a previously published work [21]. The Ligprep module of the Schrodinger suit was used to prepare the compounds. The OPLS4 (optimized potentials for liquid simulations) was utilized for force field and energy minimization [26]. The ionization state was established using the Epik ionization tool in ligprep at a pH range of 7.0 ± 1.0 [26]. The 3D conformer structure of the ligands was downloaded from PubChem (https://pubchem.ncbi.nlm.nih.gov/). To prepare the receptor, crystal structures of the different proteins, S. aureus GyrB ATPase, and Pseudomonas aeruginosa DNA gyrase B 24 kDa ATPase subdomain complexed with EBL2888, were retrieved from the protein data bank (PDB) [3U2D and 7PTG] (https://www.rcsb.org).
A grid box was generated around the centroid of the co-crystallized ligand on the prepared protein using the Glide Receptor Grid Generation module of the Schrodinger suit. The grid center coordinates on the receptor were set at 7PTG (x = 18.805600541666667, y = -17.01892358333333, z = -0.7439418333333333) and 3u2d (x = 0.2916116222222224, y = 2.6178757555555547, z = 24.38558124444444) and the inner box defines the diameter midpoint restrain of each ligand docked was set at 12 Å × 12 Å × 12 Å.
2.3 Molecular docking
The prepared compounds were docked using extra precision (XP) glide docking into the produced receptor grid. The force field of choice in the docking computation was the OPLS4 force field.
2.4 Molecular dynamics simulation
The two protein–ligand complexes that showed the best binding affinity for both strains of bacteria studied were selected for molecular dynamics simulation. 2-hydroxy-1-methoxy anthraquinone for S. aureus while alizarin for P. aeruginosa. In the molecular dynamics simulation studies, parameters such as root mean square deviation (RMSD), root mean square fluctuation (RMSF), and Intermolecular hydrogen bonding were taken into consideration.
3 Results and discussion
3.1 In vitro analysis
Figure 1 shows the antibacterial activities of the three extracts tested against two organisms. These were compared with the antibacterial activity of Gentamicin.
The extract showed different antibacterial activities against S. aureus and P. aeruginosa. The hexane extract had the weakest activity for S. aureus when compared to other extracts while the other extracts showed excellent susceptibility with chloroform extract having the best susceptibility against S. aureus. Hexane extract has better activity against P. aeruginosa while ethanol extract has a more promising activity against S. aureus.
These results are in agreement with the earlier published reports [2], which confirmed the bioactivities of the aqueous and ethanol extracts of M. lucida against Salmonella typhi, Salmonella paratyphi, Salmonella typhorium, and Escherichia coli. In addition, several parts of M. lucida have proven to possess antibacterial potential. [27]
P. aeruginosa, which is also resistant to different antibiotics [5] had its growth inhibited by ethanol and chloroform extracts of M. lucida.
Burns and wound infections are caused by P. aeruginosa; these infections frequently occur in diabetic patients and those who have recently undergone surgery. Hospital-acquired wound infections are most frequently brought about by S. aureus. [27, 28]
3.2 In silico
The in silico analysis was used to screen and identify the bioactive compounds with the potential to influence the microbial activity noted in the in vitro analysis. The binding affinity, amino residues, interactions, and bond distances for Pseudomonas aeruginosa (7PTG) and Staphylococcus aureus (5TW8).
3.2.1 Pseudomonas aeruginosa (7PTG)
It is evident from the interaction of the quinones with the 7PTG protein as seen in Table 1 that gentamicin (Fig. 2) had the highest binding affinity with a docking score of − 7.79 kcal/Mol which indicates that it has the best binding to the receptor. Gentamicin forms multiple hydrogen bonds with the following amino acid residues: Glu44, Asp51, Asn48, Ile98, Val99, Leu100, Ile96, and Gly121. It also shows electrostatic interactions with negatively charged Asp residues, forming salt bridges (Fig. 2). The variety of hydrogen bonds and electrostatic attractions with key binding site residues lead to its tight binding. [28]
From the quinones identified in M. lucida root extracts, all the compounds with the hydroxyl and methoxy functional groups had bonding affinity ranging from − 5.65 to – 3–85 kcal/mol (Figs. 3, 4). 1-Hydroxy-4-methylanthraquinone (− 5.47 kcal/mol) (Fig. 5) and 1-Hydroxy-2-methylanthraquinone (− 5.59 kcal/mol) (Fig. 6) show affinity with amino acid residues such as Thr167 and other hydrophobic interactions. The interaction of the methyl derivatives 9,10-Anthracenedione, 2-methyl- (Fig. 3) and 9,10-Anthracenedione, 1-methyl- (Fig. 4) interacting with amino acid residues indicates that the presence of the methyl group did not effectively increase the interaction as the hydroxyl groups with amino acid residues of the 7PTG protein.
Others such as 3-Hydroxyl-1-methoxyanthraquinone (Fig. 9) had an affinity of -3.85 kcal/mol, 2-hydroxyl-1-methoxyanthraquinone (Fig. 7) − 5.65 kcal/mol, and 9,10-Anthracenedione, 1,5-dimethoxy- (Fig. 8) − 4.54 kcal/mol.
The methoxy substituted anthraquinones like 1,5-dimethoxy anthraquinone (− 4.54 kcal/mol) and 1,8-dimethoxy anthraquinone (− 5.56 kcal/mol) (Figs. 9, 10 and 11 respectively) had an affinity with similar amino acids except for VAL 73. The methyl-substituted anthraquinones have varied affinities which that the position of the methyl group influences that binding affinity.
Naturally occurring alizarin (Fig. 11) shows the highest binding energy among the natural anthraquinone compounds from M. lucida (− 6.2 kcal/mol) owing to multiple hydrogen bonds, consistent with its antibiotic potential as described by Khan et al. [29]. Thus it can be predicted that alizarin contributes strongly to the susceptibility properties of M. lucida against Pseudomonas aeruginosa.
3.2.2 Staphylococcus aureus (5TW8)
Table 2 presents the results of the docking for S. aureus showing the affinities, distance, and natures of bonds. Gentamicin (Fig. 12) shows the highest binding affinity of − 7.04 kcal/mol, consistent with its potent antibacterial activity against S. aureus. [30]
2-Methyl (Fig. 13) and 1-methyl (Fig. 14) substituted anthraquinones have weaker affinities than Gentamicin with − 3.07 and − 3.67 kcal/mol respectively due to limited hydrophobic contacts.
Alizarin (Fig. 21), 1-Hydroxy-4-methylanthraquinone (Fig. 15), and 1-hydroxy-2-methylanthraquinone (Fig. 16) showed improved binding affinities (− 3.7, − 3.8, and − 4.36 kcal/mol respectively) over the methyl anthraquinones, attributed to additional hydrogen bonds with binding site residues like Tyr291. The hydroxy group enhances their antibacterial potential as noted experimentally. [31]
9,10-Anthracenedione-2-hydroxy-1-methoxy (Fig. 17) also shows a higher binding energy (− 5.5 kcal/mol) than the other hydroxyl-methyl derivative of anthraquinone, facilitated by hydrogen bonding interactions. The dimethoxy anthraquinones 1,5-dimethoxyanthraquinone (Fig. 18), 3-Hydroxyl-1-methoxyanthraquinone (Fig. 19), and 1,8-dimethoxyanthraquinone (Fig. 20) have lower affinities than the other anthraquinone derivatives potentially due to steric effects.
Naturally occurring 2-hydroxy-1-methoxyanthraquinone (Fig. 17) shows the highest binding energy among the natural anthraquinone compounds from M. lucida (− 5.5 kcal/mol) owing to multiple hydrogen bonds with amino acids such as ASN 141, ALA 74, LEU 115, SER 116.
2-hydroxyl-1-methoxyanthraquinone isolated from the stem bark of M. lucida has been reported to possess antibacterial activities. [32]. Figure 21, shows the binding interractions of alizarin with amino acids in the 5tw8 protein. Alizarin (Fig. 21)showed improved binding affinities (− 3.7 kcal/mol ) attributed to additional hydrogen bonds.
3.3 Molecular dynamics (MD) simulations
3.3.1 The root mean square deviation (RMSD) of protein–ligand
RMSD of S. aureus (5TW8) in complex with 2-hydroxy-1-methoxy anthraquinone and P. aeruginosa (7tgp) in complex with alizarin is shown in Figs. 22 and 23. The RMSD of protein–ligand was done for 50 ns. The RMSD value of 2-hydroxy-1-methoxy anthraquinone with 5TW8 is stabilized at 1.5 Å and 20 ns and remains stabilized for 50 ns (Fig. 22). Similarly, in the case of alizarin complex with 7tgp is stabilized at 5 Å and 5 ns and remains unstable for 50 ns (Fig. 23). The RMSD of the ligand with respect to the alpha carbon (prot_CA) of the protein with the reference frame shows that in Fig. 22 the protein is relatively stable and is not undergoing major conformational changes which could disrupt activities such as catalysis. It is expected that a protein being in the range of 3 Å as expected for non-globular proteins [33]. The RMSD explains how the ligand interacts with the receptor and the stability over time [34]. The alizarin ligand in the binding site of the 7tgp protein is not very stable as seen in Fig. 23.
3.3.2 The root mean square fluctuation (RMSF) of protein–ligand
To determine the flexibility of protein structures and determine the deviation of the particles from the original position of macromolecules (3-D structure of the protein) the RMSF is used [35]. As seen in Fig. 24, low fluctuations are observed for 5TW8 in a complex with 2-hydroxy-1-methoxy anthraquinone between 0.5 and 2.0 Å. Similarly, 7tgp in complex with alizarin also showed low fluctuation between 0.5 and 2.5 Å (Fig. 25). Thus, the above results revealed that RMSF plots for the two complexes and ligand RMSF have stability within the binding pocket of the target RMSF. [35]
Figures 26 and 27 further confirm the stability of the proteins by showing interaction fractions of the various residues of the protein by which the amino acids bond to the various ligands. SER 116 had the highest interaction fraction for 5TW8. SER had 87% interaction with the hydroxyl group of 2-hydroxy-1-methoxy anthraquinone (the ligand) while the oxygen on the methoxy group of the ligand had a 60% interaction (Fig. 28). ASP 75 shows the highest interaction fraction for 7PTG with water bridges. The most common type of interaction between alizarin ligand and the 7PTG protein is hydrophobic interactions. Figures 28 and 29 buttresses the nature of the most important bonding. The presence of intramolecular hydrogen bonding as seen in Fig. 29.
4 Conclusion
Since these bacteria are of public health concern and can produce toxins that are stable under heat [36], utmost care must be taken to properly address them. One of which is the identification of compounds from plants with antibacterial properties. As seen in the in vitro results, the extracts of M. lucida had good susceptibility properties against the two drug-resistant bacteria when compared to gentamicin. From the anthraquinones identified in M. lucida, 2-hydroxy-1-methoxy anthraquinone is predicted to have a better susceptibility with S. aureus while alizarin has better susceptibility properties against P. aeruginosa. The MD shows a notable level of stability between the protein and the ligands in this study.
Thus anthraquinone compounds from M. lucida show that they have bioactive potentials as antibacterial drugs. This provides a justifiable reason for further identification and characterization of all the compounds that can serve as enzyme inhibitors with potential medicinal applications. Therefore future research would be on the isolation and characterization of secondary metabolites which have been identified in the study to be responsible for the antibacterial properties .
Data Availability
The authors declare that the data supporting the findings of this study are available within the paper and its Supplementary Information files. Should any raw data files be needed in another format they are available from the corresponding author upon reasonable request. Source data are provided with this paper.
References
Adewole KE, Attah AF, Adebayo JO. M. lucida Benth (Rubiaceae): a review of its ethnomedicine, phytochemistry and pharmacology. J Ethnopharmacol. 2021;276(2021):114055.
Fakoya A, Owojuyigbe OS, Fakoya S, Adeoye SO. Possible antimicrobial activity of M. lucida stem bark, leaf and root extracts. Afr J Biotechnol. 2014;13(3):471–5.
Umeh E, Oluma H, Igoli J. Antibacterial screening of four local plants using an indicator-based microdilution technique. Afr J Trad Comp Alt Med. 2005;2(3):238–43.
Santos PRV, Oliveira ACX, Tomassini TCB. Controle microbiógico de produtos fitoterápicos. Rev Farm Bioquím. 1995;31:35–8.
Nascimento GGF, Locatelli J, Freitas PC, Silva GL. Antibacterial activity of plant extracts and phytochemicals on antibiotic-resistant bacteria. Braz J Microbiol. 2000;31(4):247–56.
Fakoya A, Owojuyigbe OS, Fakoya S, Adeoye SO. Possible antimicrobial activity of Morinda lucida stem bark, leaf and root extracts. Afr J Biotech. 2014;13:471–5.
Nworu CS, Akah PA, Okoye FB, Onwuakagba CJ, Okorafor UO, Esimone CO. Supplementation with aqueous leaf extract of Morinda lucida enhances immunorestoration and upregulates the expression of cytokines and immunostimulatory markers. Immunol Invest. 2012;41:799–819.
Osuala-Felix N, Amarachi IJ, Odoh UE. Pharmacognostic screening and antiemetic evaluation of the ethanol extract of the leaves of Morinda lucida benth. (rubiaceae). World J Biol Pharm Health Sci. 2021;07(03):001–14.
Osuntokun OT. Evaluation of antimicrobial activities of crude ethyl acetate and ethanol extracts of M. lucida leaf, stem, and bark growing in Adekunle Ajasin University Botanical Garden against selected clinical isolates. World J Biomed Pharm Sci. 2015;1(1):6–14.
Olawuwo OS, Aro A, Kobus E, McGaw L. The in vitro antibacterial activity of M. lucida and Acalypha wilkesiana leaf extracts against Campylo bacter and Escherichia coli relevant in poultry infections. The FASEB J. 2019;33(S1):672.5.
Temitope O, Oluduro A, Odubanjo O, Dada T. Evaluation of antibacterial potency of crude ethyl acetate and ethanol extracts of M. lucida leaf, stem, and bark on Mycobacterium species, isolated from the Chest Hospital, Obafemi Awolowo Teaching University, Ile-Ife, Osun State, Nigeria. J Adv Med Pharm Sci. 2016;5(3):1–8.
Ola-Fadunsin SD, Ademola IO. Anticoccidial effects of Morinda lucida acetone extracts on broiler chickens naturally infected with Eimeria species. Pharm Biol. 2014;52:330–4.
Kazeem MI, Adamson J, Ogunwande IA (2013). Modes of Inhibition of α-Amylase and α-glucosidase by aqueous extract of Morinda lucida Benth Leaf. BioMed Res Int, Volume 2013, Article ID 527570
Oduola T, Bello IO, Adeosun G, Ademosun A, Raheem G, Avwioro GO. Hepatotoxicity and nephrotoxicity evaluation in Wistar albino rats exposed to Morinda lucida leaf extract. N Am J Med Sci. 2010;2:230–3.
Saganuwan SA, Aondoaver A, Roman IT. Reassessment of acute and chronic toxicity effects of aqueous leaf extract of Morinda Lucida in Rattus norvegicus. J Hematol Res. 2014;1:36–46.
Koulenti D, Xu E, Song A, Sum Mok IY, Karageorgopoulos DE, Armaganidis A, Tsiodras S, Lipman J. Emerging treatment options for infections by multidrug-resistant gram-positive microorganisms. Microorganisms. 2020;8(2):191.
Berini F, Orlandi V, Gornati R, Bernardini G, Marinelli F. Nanoantibiotics to fight multidrug-resistant infections by gram-positive bacteria: Hope or reality? Biotechnol Adv. 2022;57: 107948.
Pal M, Shuramo MY, Tewari A, Srivastava J, Steinmetz CH. Staphylococcus aureus from a commensal to zoonotic pathogen: a critical appraisal. Int J Clin Exp Med Res. 2023;7(2):220–8. https://doi.org/10.26855/ijcemr.2023.04.023.
Khabeer BJ. Antibiogram Pattern of Pathogenic E. coli Isolates Detected in Fecal Samples of Healthy Animals (Doctoral dissertation, Tuskegee University). 2022
Magana-Arachchi D, Wanigatunge R. Ubiquitous waterborne pathogens. Waterborne Pathogens. 2020;1:15-42.
Adekunle DO, Faboro EO, Lajide L. Identification and quantification of bioactive compounds in different extracts of Morinda lucida Benth (Rubiaceae) root using GC–MS analysis. J Niger Soc Phys Sci. 2023. https://doi.org/10.46481/jnsps.2023.1534
Osuala-Felix N, Amarachi IJ, Odoh UE. Pharmacognostic screening and antiemetic evaluation of the ethanol extract of the leaves of Morinda lucida benth. (rubiaceae). World J Biol Pharm Health Sci. 2021;07(03):001–014
Adeleke OA, Owoseni AA. Antibiotic resistance and presence of plasmids in bacteria isolated from cooked street foods. Int J Environ Stud. 2022. https://doi.org/10.1080/00207233.2022.2111149.
Ojo OA, Agboola AO, Ogunro OB, Iyobhebhe M, Elebiyo TC, Rotimi DE, Ayeni JF, Ojo AB, Odugbemi AI, Egieyeh SA, Oluba OM. Beet leaf (beta Vulgaris L.) extract attenuates iron-induced testicular toxicity: experimental and computational approach. Heliyon. 2023;9(7):e17700.
Osowole AA, Ott I, Ogunlana OM. Synthesis, spectroscopic, anticancer, and antimicrobial properties of some Metal(II) complexes of (substituted) nitrophenol Schiff base. Int J Inorg Chem 2012; Article ID 206417
Shelley JC, Cholleti A, Frye LL, Greenwood JR, Timlin MR, Uchimaya M. Epik: a software program for pK a prediction and protonation state generation for drug-like molecules. J Comput Aided Mol Des. 2007;21:681–91.
Bassetti M, Vena A, Croxatto A, Righi E, Guery B. How to manage Pseudomonas aeruginosa infections. Drugs Context. 2018;7: 212527.
Chiang YT, Xiao YB, Hsu SH, Chang SW, Chou CC. Molecular interactions of tannic acid and matrix metalloproteinases 2 and 9. Comput Struct Biotechnol J. 2023;21:2792–800.
Khan A, Ezati P, Rhim JW. Alizarin: prospects and sustainability for food safety and quality monitoring applications. Colloids Surf B Biointerfaces. 2023;223:113169.
Moniruzzaman M, Jinnah MM, Islam S, Biswas J, Pramanik MJ, Uddin MS, Saleh MA, Zaman S. Biological activity of Cucurbita maxima and Momordica charantia seed extracts against the biofilm-associated protein of Staphylococcus aureus: an in vitro and in silico studies. Inf Med Unlocked. 2022;33: 101089.
Boy FR, Casquete R, Martínez A, Córdoba MG, Ruíz-Moyano S, Benito MJ. Antioxidant, antihypertensive and antimicrobial properties of phenolic compounds obtained from native plants by different extraction methods. Int J Environ Res Public Health. 2021;18(5):2475. https://doi.org/10.3390/ijerph18052475.
Mfonku NA, Mbah JA, Kodjio N, Gatsing D, Zhan J. Isolation and selective glycosylation of antisalmonellal anthraquinones from the stem bark of Morinda lucida Benth. (Rubiaceae). Phytochem Lett. 2020;37:80–4.
Adhikari B, Cheng J. Improved protein structure reconstruction using secondary structures, contacts at higher distance thresholds, and non-contacts. BMC Bioinf. 2017;18(1):1–13.
da Fonseca AM, Caluaco BJ, Madureira JMC, Cabongo SQ, Gaieta EM, Djata F, Colares RP, Neto MM, Fernandes CFC, Marinho GS, Dos Santos HS. Screening of potential inhibitors targeting the main protease structure of SARS-CoV-2 via molecular docking, and approach with molecular dynamics, RMSD, RMSF, H-bond, SASA and MMGBSA. Mol Biotechnol. 2023;1–15
Rolta R, Salaria D, Kumar V, Patel CN, Sourirajan A, Baumler DJ, Dev K. Molecular docking studies of phytocompounds of Rheum emodi Wall with proteins responsible for antibiotic resistance in bacterial and fungal pathogens: in silico approach to enhance the bio-availability of antibiotics. J Biomol Struct Dyn. 2020;40:1–15.
Mahros MA, Abd-Elghany SM, Sallam KI. Multidrug-, methicillin-, and vancomycin-resistant Staphylococcus aureus isolated from ready-to-eat meat sandwiches: an ongoing food and public health concern. Int J Food Microbiol. 2021;346: 109165.
Funding
Therefore future research would be on the isolation and characterization of secondary metabolites which have been identified in the study to be responsible for the antibacterial properties.
Author information
Authors and Affiliations
Contributions
AD and AO- carried out the invitro studies AD and OA - worked on the insilico analysis FO and LL- designed the work and worked on the manuscript. AD wrote the manuscript
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Adekunle, O.D., Adeleke, O.A., Odugbemi, A.I. et al. In vitro and in silico screening and identification of potential bioactive anthraquinones of Morinda lucida benth against pathogenic bacterial target proteins. Discov Appl Sci 6, 295 (2024). https://doi.org/10.1007/s42452-024-05832-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42452-024-05832-2