Abstract
Induced mutations are important for genetic research and breeding. Mutations induced by physical or chemical mutagenesis are usually heterozygous during the early generations. However, mutations must be fixed prior to phenotyping or field trials, which requires additional rounds of self-pollination. Microspore culture is an effective method to produce double-haploid (DH) plants that are fixed homozygotes. In this study, we conducted ethyl methanesulfonate (EMS)-induced mutagenesis of microspore cultures of barley (Hordeum vulgare) cultivar ‘Hua30’ and landrace ‘HTX’. The EMS concentrations were negatively correlated with the efficiency of callus induction and the frequency of mutant plant regeneration. The two genotypes showed different regeneration efficiencies. The phenotypic variation of the regenerated M1 plants and the presence of genome-wide nucleotide mutations, revealed by whole-genome sequencing, highlight the utility of EMS-induced mutagenesis of isolated microspore cultures for developing DH mutants. Genome-wide analysis of the mutation frequency in the regenerated plants revealed that a considerable proportion of mutations resulted from microspore culture (somaclonal variation) rather than EMS-induced mutagenesis. In addition to producing a population of 1972 homozygous mutant lines that are available for future field trials, this study lays the foundation for optimizing the regeneration efficiency of DH plants and the richness of mutations (mainly by fine-tuning the mutagen dosage).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mutation-based breeding accelerates crop improvement by generating new genetic diversity beyond natural variation (Jankowicz-Cieslak et al. 2017). The Mutant Variety Database (MVD), which is maintained by the Joint Food and Agriculture Organization of the United Nations (FAO) and the International Atomic Energy Agency (IAEA) Centre of Nuclear Techniques in Food and Agriculture (https://nucleus.iaea.org/sites/mvd/), lists over 3000 mutant varieties from a range of plant species, including cereal (staple food) crops, economically important crops (e.g., soybean [Glycine max], cotton [Gossypium hirsutum], rapeseed [Brassica napus]), and horticultural species. The generation of mutagenized populations with uniform genetic backgrounds but enriched with genetic/phenotypic variation has greatly accelerated the modern genetics research using forward and reverse genetic approaches (Uauy et al. 2017; Jiang et al. 2022).
Radiation methods, such as X-ray and fast-neutron treatment, were commonly used for mutagenesis during the first three-quarters of the last century (Lundqvist 2014). These physical agents induce breaks in double-stranded DNA and the deletion of chromosomal segments. The iconic barley (Hordeum vulgare) cultivar (cv.) ‘Golden Promise’, which is the mainstay of the European brewing industry and is widely employed as the receptor genotype during barley transformation, was developed by γ-ray-induced mutation of the historical cv. ‘Maythorpe’, followed by selection for semi-dwarf, sturdy culms, and higher yield performance (Schreiber et al. 2020). In the recently released barley pan-genome database (Jayakodi et al. 2020), Golden Promise has the smallest genome and the fewest genes among representative cultivars/accessions, possibly due to the γ-ray-induced segmental deletions in its genome.
Since the 1970s, chemical mutagens have become increasingly popular compared to radiation (Lundqvist 2014). Commonly used chemical mutagens include sodium azide (NaN3), N-methyl-N-nitrosourea (MNU), and ethyl methanesulfonate (EMS), each of which induces nucleotide transitions (predominantly G–A, or C–T). Several chemically induced mutant populations have been developed in major crops and other economically important plants (Abe et al. 2012; Henry et al. 2014; Jiao et al. 2016; Gupta et al. 2017; Li et al. 2017; Schreiber et al. 2019; Gao et al. 2020; Sashidhar et al. 2020; Nie et al. 2021). Mutants in an elite parental cultivar/variety background with desirable performance for a specific trait (e.g., salt tolerance) can readily be used to develop new varieties via genetic improvement (Takagi et al. 2015).
Double haploid (DH) technology rapidly generates completely homozygous lines, which greatly shortens the duration of breeding (Ren et al. 2017; Ma et al. 2018). DHs are commonly produced by androgenesis, a highly efficient method involving the culture of isolated microspores (Esteves et al. 2014). In this technique, microspores are isolated from anthers prior to culture. The isolated microspores can be diverted from the normal gametophytic developmental pathway to a sporophytic pathway, subsequently producing embryogenic calli and haploid or DH plants (Lu et al. 2008; Ferrie and Caswell 2011). This procedure can be accelerated by optimizing the culture conditions, such as introducing nutrients into the culture medium (Shrivastava et al. 2021). An isolated microspore culture provides a large number of single haploid cells that can be subjected to uniform treatments. Thousands of embryogenic calli can thus be rapidly induced, many of which can later develop into regenerated plants (Li and Devaux 2005; Esteves et al. 2014). In recent years, higher frequencies of embryogenic callus formation and green plantlet regeneration have been achieved in several plant species, such as rice (Oryza sativa), wheat (Triticum aestivum), tobacco (Nicotiana tabacum), rapeseed, and eggplant (Solanum melongena) (Shariatpanahi et al. 2006; Mohammadi et al. 2012; Shariatpanahi and Ahmadi 2016; Dong et al. 2021; Calabuig-Serna et al. 2022; Rahman et al. 2022). An efficient isolated microspore culture system was established in barley (Hordeum vulgare) following the optimization of factors that affect microspore embryogenesis (Lu et al. 2008, 2016).
The heterozygosity of early-generation plant populations produced by chemically induced mutagenesis, via seed treatment, makes it difficult to identify phenotypic variation in quantitative or recessive traits. EMS mutagenesis of isolated microspore cultures can be used to produce DH lines with fixed homozygous mutations. Moreover, microspore mutagenesis is highly efficient, as millions of microspores can be treated at the same time in a small space, such as a Petri dish. This strategy is practical, as many examples of phenotypic variation have been observed in DH lines derived from mutagenesis of microspores in Ethiopian mustard (Brassica carinata) (Barro et al. 2001), Chinese cabbage (Brassica rapa spp. pekinensis) (Huang et al. 2016), and barley (Gao et al. 2018). In the current study, we developed a homozygous mutant population of cultivated barley by EMS-induced mutagenesis of microspores from two genotypes. We evaluated the effects of plant genotype and EMS dosage as well as the mutation efficiency, via whole-genome sequencing. Our findings highlight the efficiency of our technique for producing homozygous mutant populations in barley.
Materials and methods
Plant materials and growth conditions
Two barley genotypes were included in this study: the cultivar ‘Hua30’, which was developed by anther culture from a cross between the spring two-rowed cv. ‘Xiumai 1’ and the breeding line ‘82,164’; and the landrace ‘HTX’ (Jiang et al. 2022). The seeds were sown in the field at Shanghai Academy of Agricultural Science (Shanghai, China) in November 2018, and spikes were collected in March 2019 for isolated microspore culture. Green plantlets at the 4- to 5-leaf stage regenerated from microspore culture were grown for seed multiplication under normal greenhouse conditions at the field station (Kunming, Yunnan province, China). Five seedlings were planted in each pot containing 25 kg of local field soil, and 30 g NPK compound fertilizer was applied to each pot. The plants were watered once or twice per week and treated with 30 g of compound fertilizer after flowering. Aphids and powdery mildew were controlled by regular spraying with the pesticides imidacloprid and carbendazim.
Microspore culture, EMS treatment, and ploidy determination
Microspore culture was performed, as previously described (Lu et al. 2008). Briefly, the collected spikes were subjected to cold pre-treatment at 4 °C for 2 weeks. The microspores were collected by crushing the anthers from surface-sterilized spikes and incubated in extraction buffer (pH 5.8, containing 330 mM mannitol, 10 mM CaCl2, and 4.6 mM MES hydrate) at 25 °C for 2 days. The microspore titer was adjusted to a density of 5.0 × 105 microspores/mL for embryogenic callus induction and cultured at 25 °C for 19 days in the dark in callus induction medium alone or containing EMS at a concentration of 50, 100, 200, or 300 mg/L (Fig. 1); the callus induction medium was N6 medium (pH 5.8) supplemented with 4.5 μM 2,4-D, 2.3 μM KT, 0.7 mM hydrolyzed casein, 4.6 mM MES hydrate, 10 mM glutamine, and 250 mM maltose. Following induction, the embryogenic calli were transferred to differentiation medium (pH 5.8) composed of 2/3 Murashige and Skoog (MS) basal medium supplemented with 2.2 μM 6-BA, 7.0 μM KT, 0.25 μM NAA, and 83.3 mM maltose. The induced embryogenic calli were carefully removed from the medium by pipetting, weighed, and the regenerated plantlets, surviving plants, and albino plants counted.
Pipeline used to produce homozygous mutants by microspore culture. Various concentrations of the mutagen EMS (0, 50, 100, 200, 300 mg/L) were included in the induction medium at the microspore culturing step. A Collected young spikes; B sample pre-treatment; C microspore isolation; D microspore culture with EMS treatment; E Induced embryogenic calli; F regenerated M1 plantlets; G typical results of ploidy determination; H plant cultivation and seed multiplication in the greenhouse
The ploidy levels of a subset of regenerated green plantlets were assessed with a BD Accuri C6 flow cytometer (BD Biosciences, Franklin Lakes, NJ, USA). In brief, a 0.5–1 cm piece of a leaf blade was placed into a 1.5 mL tube, ground in 1 mL of liquid nitrogen, and re-suspended in 0.5 mL buffer solution (15 mM Tris–HCl, 2 mM Na2EDTA, 80 mM KCl, 20 mM NaCl, 0.1% [v/v] Triton X-100). The samples were filtered to remove debris, incubated on ice for at least 5 min, and stained by adding 0.2 mL propidium iodide (PI) in RNase staining buffer solution (BD-Pharmingen). Following incubation on ice in the dark for 30 min, the samples were analyzed on a BD Accuri C6 flow cytometer, and statistical analysis was performed using BD Accuri C6 Software. Using the position of the DNA peak from mesophyll cells of different ploidy levels in the control group as a standard, the ploidy levels of the regenerated plants were determined. Rooted green plantlets were acclimated for 1 to 2 weeks by hydroponics, transferred to soil for seed multiplication, and collected as DH lines.
DNA extraction and whole-genome sequencing
The leaves of seedling were sampled from germinated cv. ‘Hua30’ seedlings and from Hua30 and HTX seedlings generated by microspore culture. Genomic DNA was extracted from the samples and its concentration quantified following a standard procedure (Yang et al. 2013). Following a quality check, the DNA samples were used for library construction, followed by sequencing using an Illumina NovaSeq platform (Novogene, Beijing) following a standard protocol (Jiang et al. 2022).
Read mapping and identification of mutations
Raw reads were trimmed and filtered to obtain clean reads using fastp (v0.20.1) with default parameters (Chen et al. 2018). The clean reads were mapped to the assembled MorexV3 (https://doi.org/10.5447/ipk/2021/3) reference genome (Mascher et al. 2021) with bwa (v0.7.17) (Li and Durbin 2009). The mapped reads were sorted using samtools (v1.10) (Li et al. 2009), and the duplicates generated by PCR were marked and removed using the MarkDuplicates function of GATK4 (v4.2.6.1) (McKenna et al. 2010). Only reads with a mapping quality ≥ 20 were subjected to variant calling using BCFtools (v1.9) (Li 2011).
First, single-nucleotide polymorphisms (SNPs) in each sample of germinated or regenerated plants were called against the MorexV3 reference genome and subjected to the following filtration steps: (1) only bi-allelic variants were accepted; (2) genotype calls were considered valid when their read depth was ≥ 2 and ≤ 30 for each sample; (3) heterozygous SNPs were not considered. The number of mutation sites in each regenerated plant sample was then calculated as the number of mutation sites in the SNP matrix for each sample against MorexV3 compared to those in the SNP matrix against MorexV3 in plants of the corresponding genotype germinated from seeds. Only SNPs meeting the following criteria were accepted: (i) calls for HTX or Hua30 were homozygous (genotype call 0/0 or 1/1 in VCF format); (ii) the genotype calls in the mutant samples were homozygous and different from the HTX or Hua30 genotype calls; and (iii) the alternate allele was only present in a single sample. The mutation frequency for each mutant was calculated as the number of SNPs divided by the cumulative size of the whole genome sequence or gene-coding region.
Results
Different genotypes show different rates of plant regeneration following microspore mutagenesis
We subjected the two barley genotypes Hua30 and HTX to EMS-induced mutagenesis of isolated microspore cultures and investigated the rates of embryogenic callus induction and mutagenized plant regeneration. There was no significant difference in the rate of embryogenic callus induction between Hua30 and HTX at the same EMS concentration (Fig. 2A). However, the number of regenerated plants, surviving plants, and albino plants was significantly lower for HTX than Hua30, regardless of EMS concentration (Fig. 2B–D). Indeed, the number of surviving HTX plants was less than 10% that of Hua30 (Fig. 2C). These results demonstrate that the genotypes Hua30 and HTX show little difference in embryogenic callus induction but large differences in plant regeneration following microspore mutagenesis.
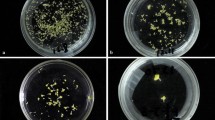
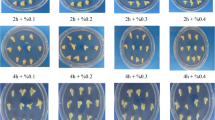
The effects of mutagen dosage and genotype on the embryogenesis and regeneration of microspore cultures. Various traits were quantified: weight of induced embryogenic calli on each plate (A), number of regenerated surviving plantlets (B), number of surviving regenerated green plantlets (C), and number of albino plantlets that eventually died (D). The number of regenerated plantlets on each plate of embryogenic callus induction medium was recorded, and three plates were quantified per treatment. Statistical analysis was conducted using Tukey’s Honest Significant Difference (P = 0.05) with three independent replicates. E Induced embryogenic calli treated with different EMS concentrations. Arrows represent microspore-derived embryogenic calli. Scale bars, 100 μm. F Regenerated plantlets from microspores treated with different concentrations of EMS. Two and five independent experiments were conducted for Hua30 and HTX, respectively; one experiment per genotype is shown in the figure
EMS concentration is negatively correlated with embryogenic callus induction and plant regeneration
We examined whether EMS treatment had dose-dependent effects on embryogenic callus induction or plant regeneration. We observed a significant reduction in callus yield with increasing EMS concentration (Fig. 2E). Compared to the control (without EMS treatment), the number and the size of calli decreased with increasing EMS concentration. When the EMS concentration increased to 300 mg/L, we observed about a 50% reduction in the yield of embryogenic calli in both Hua30 and HTX (Fig. 2A). Thus, treatment with higher concentrations of EMS significantly impairs callus induction.
In Hua30, the number of regenerated plants gradually decreased with increasing EMS concentration (Fig. 2B and F). We obtained the greatest number of surviving Hua30 plants at an EMS concentration of 50 mg/L, and the fewest at an EMS concentration of 300 mg/L (Fig. 2C). Accordingly, the number of albino plants in Hua30 significantly increased when using higher concentrations of EMS (Fig. 2D). For HTX, which had a significantly lower ratio of regenerated plants compared to Hua30, different EMS concentrations had little effect on the number of regenerated plants, surviving plants, or albino plants.
Generation of an EMS-induced homozygous mutant population
We self-pollinated the surviving plants (M1) to produce seeds from the DH homozygous mutants. Of the 2793 regenerated plants, derived from seven isolated microspore culture experiments, 1972 M2 DH lines (70.64%) were fertile and produced seeds (Table 1). These M2 lines included 99 DH HTX lines and 1873 DH Hua30 lines. For Hua30, we collected seeds from 1818 DH lines, comprising 258, 434, 688, and 438 lines that were treated with EMS concentrations of 50 mg/L, 100 mg/L, 200 mg/L, or 300 mg/L, respectively. We noticed one regenerated M1 plant lacking a wax coating (non-glaucous) on the uppermost internode and flag leaf sheath (Fig. 3). This phenotypic variation suggests that EMS treatment of isolated microspores induces mutations in the barley genome.
Examining mutation frequency by whole-genome sequencing
To evaluate the mutation frequencies in the homozygous mutant populations, we conducted whole-genome sequencing via the Illumina approach on a subset of plants regenerated from microspore culture. We obtained 25.15–47.58 Gb of clean data, representing approximately 6–11 × coverage of the barley reference genome (Morex V3, 4.2 Gb) (Mascher et al. 2021), for each Hua30 and HTX sample (Table 2).
To determine whether the mutations were induced by microspore culture (control condition, 0 mg/L) or by microspore culture in the presence of EMS (300 mg/L), we analyzed the mutation sites in the regenerated Hua30 and HTX plants. We detected a higher genome-wide mutation frequency in Hua30 than in HTX, regardless of treatment (Table 2). We detected an average of 1.84 and 1.23 mutations/Mb in Hua30 and HTX seedlings regenerated from seedlings derived from microspore culture on growth medium lacking EMS. A treatment with 300 mg/L EMS in the induction medium resulted in a significant increase in mutation frequency (2.19 mutations/Mb in Hua30, P < 0.01; 1.57 mutations/Mb in HTX, P < 0.05) compared to EMS-free medium. After subtracting the number of mutations generated by microspore culture, there were approximately 0.35 and 0.34 EMS-induced mutations/Mb in regenerated Hua30 and HTX seedlings, respectively. Specifically, for the gene-coding regions, we observed a similar pattern of induced mutations (Table 2). We also examined whole-genome sequencing datasets from seeds subjected to EMS-induced mutagenesis (Jiang et al. 2022) to compare the mutation frequencies obtained using different treatment approaches (seeds vs. isolated microspores). We determined that HTX seeds treated with 2800 mg/L and 4000 mg/L EMS accumulated 3.99 mutations/Mb and 4.51 mutations/Mb, respectively (Table 2).
In addition, we counted the numbers of nucleotide transitions (A/T to G/C; G/C to A/T) or transversions (A/T to C/G; A/T to T/A; G/C to T/A; G/C to C/G) (Fig. 4A). The G/C to A/T transition, a typical effect of EMS-induced mutation, represented the majority of mutations in HTX mutants derived from EMS treatment of seeds in a previous study (Fig. 4A). Following treatment with 300 mg/L EMS in the induction medium, the ratio of G/C to A/T transition significantly increased in both Hua30 (P < 0.05) and HTX (P < 0.005) DH mutants (Fig. 4B and C), while the number of other types of mutations also significantly increased (P < 0.05). Non-G/C to A/T mutations were detected at a ratio over 50% in the regenerated plants following microspore culture both with and without EMS treatment.
Single nucleotide transitions or transversions detected by whole-genome sequencing of mutated plants. A Overall number of single nucleotide differences detected in plants from mutagenized seeds (left, adapted from Jiang et al. 2022) or microspores (right). The G/C to A/T transition is a typical effect of EMS-induced mutagenesis, which significantly increased in regenerated plants treated with 300 mg/L EMS in both the HTX (B) and Hua30 (C) genotypes. The abbreviated name of each sample indicates the genotype, the EMS concentration, and the biological replicate. For example, HTX-2.8_1 indicates the first replicate of HTX treated with 2800 mg/L EMS
Collectively, these results demonstrate that EMS-induced mutagenesis of isolated microspore cultures can produce homozygous mutations in the regenerated DH plants.
Discussion
In this study, we developed a homozygous mutant population of cultivated barley after only one generation of seed multiplication by combining microspore culture with EMS-induced mutagenesis. We performed microspore mutagenesis in two distinct genotypes with a series of EMS concentrations together with whole genome re-sequencing of the regenerated plants, revealing the genotype- and EMS dosage-dependent effects on the regeneration efficiency of mutagenized DH lines.
Most mutant plant populations developed to date were obtained by mutagenesis of seeds (Irshad et al. 2020). Seeds before and after mutagen treatment are designated as M0 and M1 seeds, respectively (Caldwell et al. 2004). M1 plants that germinate from M1 seeds are usually heterozygous at each mutagenized locus. M2 plants and stocks of M3 seeds are normally used to construct mutant populations. For either breeding or genetic research, several rounds of self-pollination are required to obtain homozygous mutant seeds with stabilized phenotypic alterations. However, lethal or sterile effects can still occur in M3 plants or later generations (Jiang et al. 2022). In vitro culture of haploid microspores is a common approach for developing DH lines, producing regenerated plants homozygous at every genetic locus (Lu et al. 2008). Previous studies have included chemical mutagens in the microspore suspension or induction medium during microspore culture of Brassica species (Barro et al. 2001; Huang et al. 2016) and barley (Castillo et al. 2001; Gao et al. 2018). The regenerated M1 seedlings were homozygous, allowing their leaves to be sampled for DNA extraction to develop a TILLING population and their seeds (M2 seeds) to be directly sown for phenotyping in a field trial. In the current study, we obtained seeds from 1873 DH lines from the parental line Hua30 and 99 from HTX with nucleotide mutations. These DH lines with the same genetic background are valuable resources for phenotyping traits of interest.
Tissue culture-induced genomic variation (somaclonal variation) is a well-known phenomenon that has been widely studied (Bednarek et al. 2007; Zhang et al. 2014; Machczyńska et al. 2015; Bednarek et al. 2020). Previous studies have used amplification fragment length polymorphic (AFLP) markers to reveal the extent of somaclonal variations. In the present study, we utilized a new strategy to estimate gametoclonal variation. We subtracted the number of SNPs between plants obtained by seed germination vs. plants regenerated by microspore culture to calculate the genome-wide mutation frequency (1.84 and 1.23 mutations/Mb in Hua30 and HTX, respectively) that resulted from isolated microspore culture. Over 50% of the observed mutations differed from the G/C to A/T transition, the typical effect of EMS-induced mutagenesis. The mutation frequency, regeneration rate, and survival rate were shown to be genotype-dependent, as they were significantly higher in regenerated Hua30 vs. HTX plants with or without EMS treatment. For both genotypes, the mutation frequency was lower in the gene-coding regions than in the overall genome, implying that the non-coding regions have higher tolerance to mutations.
When we calculated the number of SNPs in the regenerated seedlings (0 vs. 300 mg/L EMS), we determined that ca. 0.35 mutations/Mb are induced by EMS genome-wide, accounting for 19.0% and 28.5% of that resulting from microspore culture without mutagenesis in Hua30 and HTX, respectively. Compared to the mutation frequency in the same genotype (HTX) obtained by mutagenesis of seeds with a much higher dosage of EMS (i.e., 2800 mg/L; 3.99 mutations/Mb), the mutation frequency of microspores obtained in this study was low, with only 1.57 mutations per Mb (300 mg/L EMS), i.e., 39.35% that of seed mutagenesis. The low mutation frequency in microspores treated with EMS is unexpected and is likely due to an insufficient dose of EMS. Indeed, microspores that are competent to undergo embryogenesis possess plasticity, and EMS treatment could impose stress and induce cell reprogramming (Testillano 2019). The totipotency of isolated microspores is a complicated program including dedifferentiation (embryogenic callus induction) and differentiation (plantlet regeneration). Although embryogenic calli were induced in the presence of EMS, a portion of the resulting mutations might have recovered or been removed by selection against mutations in essential genes during later development (i.e., plantlet regeneration). Therefore, more studies are needed to fully evaluate the efficiency of microspore mutagenesis.
We did not sequence regenerated plants from calli treated with a lower dosage of EMS, as a much lower mutation frequency would be expected in these lines. These DH lines with lower rates of mutation but homozygous mutations would be quite useful in future breeding programs or genetic studies due to minimized background noise. Enlarging the population size with saturated mutations, combined with a highly sensitive mutation detection method (i.e., Fast Identification of Nucleotide variants by droplet DigITal PCR [FIND-IT]) (Knudsen et al. 2022), may help identify sufficient numbers of mutants in target genes. Considering that the mutagen was included in the induction medium, followed by 19 days of culture, the main reason for the low mutation frequency is likely the low level of EMS utilized. As Hua30 microspores showed a certain level of tolerance to mutagenesis, a dosage of > 300 mg/L may be considered to increase the frequency of mutations in the barley genome.
Data availability
The whole genome re-sequencing clean reads datasets were deposited in NCBI database with the accession ID “SAMN32366957” to “SAMN32366969” under the project ID “PRJNA915006”.
References
Abe A, Kosugi S, Yoshida K, Natsume S, Takagi H, Kanzaki H, Matsumura H, Yoshida K, Mitsuoka C, Tamiru M, Innan H, Cano L, Kamoun S, Terauchi R (2012) Genome sequencing reveals agronomically important loci in rice using MutMap. Nat Biotechnol 30:174–178. https://doi.org/10.1038/nbt.2095
Barro F, Fernández-Escobar J, DeLaVega M, Martín A (2001) Doubled haploid lines of Brassica carinata with modified erucic acid content through mutagenesis by EMS treatment of isolated microspores. Plant Breed 120:262–264. https://doi.org/10.1046/j.1439-0523.2001.00602.x
Bednarek PT, Orłowska R, Koebner RM, Zimny J (2007) Quantification of the tissue-culture induced variation in barley (Hordeum vulgare L.). BMC Plant Biol 7:10. https://doi.org/10.1186/1471-2229-7-10
Bednarek PT, Zebrowski J, Orłowska R (2020) Exploring the biochemical origin of DNA sequence variation in barley plants regenerated via in vitro anther culture. Int J Mol Sci 21:5770. https://doi.org/10.3390/ijms21165770
Calabuig-Serna A, Camacho-Fernández C, Mir R, Porcel R, Carrera E, López-Díaz I, Seguí-Simarro JM (2022) Quantitative and qualitative study of endogenous and exogenous growth regulators in eggplant (Solanum melongena) microspore cultures. Plant Growth Regul 96:345–355. https://doi.org/10.1007/s10725-021-00780-y
Caldwell DG, Mccallum N, Shaw P, Muehlbauer GJ, Marshall DF, Waugh R (2004) A structured mutant population for forward and reverse genetics in barley (Hordeum vulgare L.). Plant J 40:143–150. https://doi.org/10.1111/j.1365-313X.2004.02190.x
Castillo AM, Cistué L, Vallés MP, Sanz JM, Romagosa I, Molina-Cano JL (2001) Efficient production of androgenic doubled-haploid mutants in barley by the application of sodium azide to anther and microspore cultures. Plant Cell Rep 20:105–111. https://doi.org/10.1007/s002990000289
Chen S, Zhou Y, Chen Y, Gu J (2018) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:i884–i890. https://doi.org/10.1093/bioinformatics/bty560
Dong YQ, Gao YH, Zhao T, Ren GQ, Liu YL, Guan B, Jin RX, Gao F, Zhang YL, Tan XF, Zhu HC, Zhang YH, Zhang JX, Peng D, Yan YX (2021) Influencing factors and physiochemical changes of embryogenesis through in vitro isolated microspore culture in Brassica species. Biologia 76:2629–2654. https://doi.org/10.1007/s11756-021-00721-0
Esteves P, Clermont I, Marchand S, Belzile F (2014) Improving the efficiency of isolated microspore culture in six-row spring barley: II-exploring novel growth regulators to maximize embryogenesis and reduce albinism. Plant Cell Rep 33:871–879. https://doi.org/10.1007/s00299-014-1563-1
Ferrie AMR, Caswell KL (2011) Isolated microspore culture techniques and recent progress for haploid and doubled haploid plant production. Plant Cell, Tissue Organ Cult 104:301–309. https://doi.org/10.1007/s11240-010-9800-y
Gao R, Guo G, Fang C, Huang S, Chen J, Lu R, Huang J, Fan X, Liu C (2018) Rapid generation of barley mutant lines with high nitrogen uptake efficiency by microspore mutagenesis and field screening. Front Plant Sci 9:450. https://doi.org/10.3389/fpls.2018.00450
Gao YL, Yao XF, Li WZ, Song ZB, Wang BW, Wu YP, Shi JL, Liu GS, Li YP, Liu CM (2020) An efficient TILLING platform for cultivated tobacco. J Integr Plant Biol 62:165–180. https://doi.org/10.1111/jipb.12784
Gupta P, Reddaiah B, Salava H, Upadhyaya P, Tyagi K, Sarma S, Datta S, Malhotra B, Thomas S, Sunkum A, Devulapalli S, Till BJ, Sreelakshmi Y, Sharma R (2017) Next-generation sequencing (NGS)-based identification of induced mutations in a doubly mutagenized tomato (Solanum lycopersicum) population. Plant J 92:495–508. https://doi.org/10.1111/tpj.13654
Henry IM, Nagalakshmi U, Lieberman MC, Ngo KJ, Krasileva KV, Vasquez-Gross H, Akhunova A, Akhunov E, Dubcovsky J, Tai TH, Comai L (2014) Efficient genome-wide detection and cataloging of EMS-induced mutations using exome capture and next-generation sequencing. Plant Cell 26:1382–1397. https://doi.org/10.1105/tpc.113.121590
Huang S, Liu Z, Li D, Yao R, Feng H (2016) A new method for generation and screening of Chinese cabbage mutants using isolated microspore culturing and EMS mutagenesis. Euphytica 207:23–33. https://doi.org/10.1007/s10681-015-1473-5
Irshad A, Guo H, Zhang S, Liu L (2020) TILLING in cereal crops for allele expansion and mutation detection by using modern sequencing technologies. Agronomy 10:405. https://doi.org/10.3390/agronomy10030405
Jankowicz-Cieslak J, Mba C, Till BJ (2017) Mutagenesis for crop breeding and functional genomics. Biotechnologies for plant mutation breeding: protocols. Springer International Publishing. https://doi.org/10.1007/978-3-319-45021-6_1
Jayakodi M, Padmarasu S, Haberer G, Bonthala VS, Gundlach H, Monat C et al (2020) The barley pan-genome reveals the hidden legacy of mutation breeding. Nature 588:284–289. https://doi.org/10.1038/s41586-020-2947-8
Jiang C, Lei M, Guo Y, Gao G, Shi L, Jin Y, Cai Y, Himmelbach A, Zhou S, He Q, Yao X, Kan J, Haberer G, Duan F, Li L, Liu J, Zhang J, Spannagl M, Liu C, Stein N, Feng Z, Mascher M, Yang P (2022) A reference-guided TILLING by amplicon-sequencing platform supports forward and reverse genetics in barley. Plant Commun 3:100317. https://doi.org/10.1016/j.xplc.2022.100317
Jiao Y, Burke J, Chopra R, Burow G, Chen J, Wang B, Hayes C, Emendack Y, Ware D, Xin Z (2016) A sorghum mutant resource as an efficient platform for gene discovery in grasses. Plant Cell 28:1551–1562. https://doi.org/10.1105/tpc.16.00373
Knudsen S, Wendt T, Dockter C, Thomsen HC, Rasmussen M, Egevang-Jørgensen M, Lu Q et al (2022) FIND-IT: accelerated trait development for a green evolution. Sci Adv 8:2266. https://doi.org/10.1126/sciadv.abq2266
Li H (2011) A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27:2987–2993. https://doi.org/10.1093/bioinformatics/btr509
Li H, Devaux P (2005) Isolated microspore culture overperforms anther culture for green plant regeneration in barley (Hordeum vulgare L.). Acta Physiol Plant 27:611–619. https://doi.org/10.1007/s11738-005-0065-8
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25:1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Li Z, Jiang L, Ma Y, Wei Z, Hong H, Liu Z, Lei J, Liu Y, Guan R, Guo Y et al (2017) Development and utilization of a new chemically-induced soybean library with a high mutation density. J Integr Plant Biol 59:60–74. https://doi.org/10.1111/jipb.12505
Lu R, Wang Y, Sun Y, Shan L, Chen P, Huang J (2008) Improvement of isolated microspore culture of barley (Hordeum vulgare L.): the effect of floret co-culture. Plant Cell, Tissue Organ Cult 93:21–27. https://doi.org/10.1007/s11240-008-9338-4
Lu R, Chen Z, Gao R, He T, Wang Y, Xu H, Guo G, Li Y, Liu C, Huang J (2016) Genotypes-independent optimization of nitrogen supply for isolated microspore cultures in barley. BioMed Res Int 2016:1801646. https://doi.org/10.1155/2016/1801646
Lundqvist U (2014) Scandinavian mutation research in barley—a historical review. Hereditas 151:123–131. https://doi.org/10.1111/hrd2.00077
Ma H, Li G, Würschum T, Zhang Y, Zheng D, Yang X, Li J, Liu W, Yan J, Chen S (2018) Genome-wide association study of haploid male fertility in maize (Zea mays L.). Front Plant Sci 9:974. https://doi.org/10.3389/fpls.2018.00974
Machczyńska J, Zimny J, Bednarek PT (2015) Tissue culture-induced genetic and epigenetic variation in triticale (Triticosecale spp. Wittmack ex A. Camus 1927) regenerants. Plant Mol Biol 89:279–292. https://doi.org/10.1007/s11103-015-0368-0
Mascher M, Wicker T, Jenkins J, Plott C, Lux T, Koh CS, Ens J, Gundlach H, Boston LB, Tulpová Z, Holden S, Hernández-Pinzón I, Scholz U, Mayer KFX, Spannagl M, Pozniak CJ, Sharpe AG, Šimková H, Moscou MJ, Grimwood J, Schmutz J, Stein N (2021) Long-read sequence assembly: a technical evaluation in barley. Plant Cell 33:1888–1906. https://doi.org/10.1093/plcell/koab077
Mckenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, Depristo MA (2010) The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303. https://doi.org/10.1101/gr.107524.110
Mohammadi PP, Moieni A, Ebrahimi A, Javidfar F (2012) Doubled haploid plants following colchicine treatment of microspore-derived embryos of oilseed rape (Brassica napus L.). Plant Cell, Tissue Organ Cult 108:251–256. https://doi.org/10.1007/s11240-011-0036-2
Nie S, Wang B, Ding H, Lin H, Zhang L, Li Q, Wang Y, Zhang B et al (2021) Genome assembly of the Chinese maize elite inbred line RP125 and its EMS mutant collection provide new resources for maize genetics research and crop improvement. Plant J 108:40–54. https://doi.org/10.1111/tpj.15421
Rahman ZA, Seman ZA, Othman AN, Ghaffar MBA, Razak SA, Yusof MFM, Nasir KH, Ahmad K, Chow YL, How TC, Saad NM, Subramaniam S (2022) Establishment of effective plantlets regeneration protocol via isolated microspore culture in Malaysian indica rice MR219. Plant Biotechnol Rep 16:343–355. https://doi.org/10.1007/s11816-022-00742-4
Ren J, Wu P, Trampe B, Tian X, Lübberstedt T, Chen S (2017) Novel technologies in doubled haploid line development. Plant Biotechnol J 15:1361–1370. https://doi.org/10.1111/pbi.12805
Sashidhar N, Harloff HJ, Jung C (2020) Identification of phytic acid mutants in oilseed rape (Brassica napus) by large-scale screening of mutant populations through amplicon sequencing. New Phytol 225:2022–2034. https://doi.org/10.1111/nph.16281
Schreiber M, Barakate A, Uzrek N, Macaulay M, Sourdille A, Morris J, Hedley PE, Ramsay L, Waugh R (2019) A highly mutagenised barley (cv. Golden Promise) TILLING population coupled with strategies for screening-by-sequencing. Plant Methods 15:99–99. https://doi.org/10.1186/s13007-019-0486-9
Schreiber M, Mascher M, Wright J, Padmarasu S, Himmelbach A, Heavens D, Milne L, Clavijo BJ, Stein N, Waugh R (2020) A genome assembly of the barley “transformation reference” cultivar golden promise. G3 10:1823–1827. https://doi.org/10.1534/g3.119.401010
Shariatpanahi ME, Ahmadi B (2016) Isolated microspore culture and its applications in plant breeding and genetics. Plant tissue culture: propagation conservation and crop improvement. Springer, Singapore. https://doi.org/10.1007/978-981-10-1917-3_21
Shariatpanahi ME, Belogradova K, Hessamvaziri L, Heberle-Bors E, Touraev A (2006) Efficient embryogenesis and regeneration in freshly isolated and cultured wheat (Triticum aestivum L.) microspores without stress pretreatment. Plant Cell Rep 25:1294–1299. https://doi.org/10.1007/s00299-006-0205-7
Shrivastava V, Savarimuthu A, Patil M, Sarkar P, Hadole S, Dasgupta S (2021) Gametic embryogenesis and callogenesis in isolated microspore culture of Jatropha curcas L. a recalcitrant bioenergy crop. Plant Cell, Tissue Organ Cult 144:359–370. https://doi.org/10.1007/s11240-020-01959-3
Takagi H, Oli MT, Abe A, Yoshida K, Uemura A, Yaegashi H, Obara T, Oikawa K, Utsushi H, Kanzaki E, Mitsuoka C, Natsume S, Kosugi S, Kanzaki H, Matsumura H, Urasaki N, Kamoun S, Terauchi R (2015) MutMap accelerates breeding of a salt-tolerant rice cultivar. Nat Biotechnol 33:445–449. https://doi.org/10.1038/nbt.3188
Testillano PS (2019) Microspore embryogenesis: targeting the determinant factors of stress-induced cell reprogramming for crop improvement. J Exp Bot 70:2965–2978. https://doi.org/10.1093/jxb/ery464
Uauy C, Wulff BBH, Dubcovsky J (2017) Combining traditional mutagenesis with new high-throughput sequencing and genome editing to reveal hidden variation in polyploid wheat. Annu Rev Genet 51:435–454. https://doi.org/10.1146/annurev-genet-120116-024533
Yang P, Perovic D, Habekuß A, Zhou R, Graner A, Ordon F, Stein N (2013) Gene-based high-density mapping of the gene rym7 conferring resistance to Barley mild mosaic virus (BaMMV). Mol Breed 32:27–37. https://doi.org/10.1007/s11032-013-9842-z
Zhang D, Wang Z, Wang N, Gao Y, Liu Y, Wu Y, Bai Y, Zhang Z, Lin X, Dong Y, Ou X, Xu C, Liu B (2014) Tissue culture-induced heritable genomic variation in rice, and their phenotypic implications. PLoS ONE 9:e96879. https://doi.org/10.1371/journal.pone.0096879
Acknowledgements
This study was financially supported by the earmarked fund for CARS (CARS-05-01A-02), SAAS Program for Excellent Research Team (2022(018)), Agricultural Science and Technology Innovation Program of CAAS.
Author information
Authors and Affiliations
Contributions
PY and CL designed the research. LH, GQG, PY, CL, CJ, GMG, QH, and YZ performed the experiments and data analysis. CJ, LH, and PY wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, L., Gao, G., Jiang, C. et al. Generating homozygous mutant populations of barley microspores by ethyl methanesulfonate treatment. aBIOTECH 4, 202–212 (2023). https://doi.org/10.1007/s42994-023-00108-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42994-023-00108-6