Abstract
The phycobilisome (PBS) is an antenna protein complex in cyanobacteria, Glaucocystophytes, and red algae. In the standard PBS, the rod-core PBS, the rods are connected to the core by the rod-core linker protein CpcG. The rod-core PBS transfers the light energy mainly to photosystem (PS) II and to a lesser extent to PSI. Cyanobacteria assemble another type of PBS, the CpcL-PBS, which consists of only one rod. This rod-type PBS is connected to the thylakoid membrane by the linker protein CpcL and is a PSI-specific antenna. In the filamentous heterocyst-forming cyanobacterium Anabaena (Nostoc) sp. PCC 7120, the CpcL-PBS forms a complex with the tetrameric PSI (PBS-PSI supercomplex). The CpcL-PBS and the rod part of the rod-core PBS are identical except for the linker proteins CpcL and CpcG. How cells control the accumulation of the two different types of PBS is unknown. Here, we analyzed two mutant strains which either lack the major rod-core linker CpcG4 or overexpress the rod-membrane linker CpcL. In both mutant strains, more and larger PBS-PSI supercomplexes accumulated compared to the wild type. Our results suggest that CpcL and CpcG4 compete for the same phycobiliprotein pool, and therefore the CpcL/CpcG4 ratio determines the levels of PBS-PSI supercomplexes. We propose that the CpcL-PBS and the rod-core PBS fulfill distinct functions in light harvesting.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Light serves as an essential energy source for life on earth. Photosynthesis enables the conversion of light energy into chemical energy to support almost all biological processes in a phototrophic cell. Light-harvesting antenna systems which efficiently collect the light energy are crucial components of the photosynthetic apparatus. Many cyanobacteria, Glaucocystophytes, and red algae use phycobilisomes (PBSs) as light-harvesting antenna complexes. The PBS absorbs light and transfers the excitation energy to the photosystem (PS) complexes, mainly to PSII [1]. Generally, the PBS is a supercomplex composed of a core and protruding rod subcomplexes (the rod-core PBS). These core and rod subcomplexes are composed of various phycobilin-binding proteins connected by several classes of colorless linker proteins [2, 3]. The phycobiliproteins can be classified into allophycocyanin (APC, λmax = 650–655 nm) which forms the core, phycocyanin (PC, λmax = 620 nm) which is the main component of the rod, phycoerythrin (PE), and phycoerythrocyanin (PEC), both of which can be assembled into the peripheral part of the rods in different species [4, 5]. The rod-core linker protein CpcG connects the rods to the core complex and is necessary for the assembly of the rod-core PBS [3]. In general, CpcG proteins contain a conserved N-terminal linker-PBS domain (Pfam00427), located within the rod disc, and a C-terminal hydrophilic linker-G domain, which interacts with the core [3, 6]. The chromosome of the filamentous, N2-fixing cyanobacterium Anabaena (Nostoc) sp. PCC 7120 (hereafter “Anabaena”) contains three copies of the rod-core linker CpcG which are designated as CpcG1, CpcG2, and CpcG4. It is known that CpcG4 is the rod-core linker that makes the connection between rods and the central core cylinders of the PBS. CpcG1 and CpcG2 linker proteins connect the rods to the so-called half-core cylinders which are composed of only one APC hexamer (Fig. 1) [6]. These half-core cylinders are located perpendicularly on top of the two basal core cylinders in the Anabaena PBS.
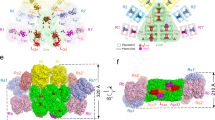
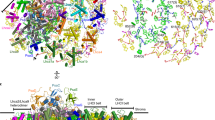
Model of the rod-core PBS (left) and the CpcL-PBS (right) from Anabaena. The rod-core linker CpcG4 connects rods with core cylinders, whereas the CpcL rod-membrane linker connects a single rod with the thylakoid membrane. The additional rod-core linkers CpcG1 and CpcG2 connect rods with the half-core cylinder (HC) [6]. In the rod-core PBS, the linker ApcE connects the core with the membrane. Anabaena PBSs contain the phycobiliproteins phycoerythrocyanin (PEC), phycocyanin (PC), and allophycocyanin (APC) and additional linkers which connect the different structures
Further, another type of PBS has been reported, which is called the CpcL-PBS [7,8,9]. The CpcL-PBS is a rod-type PBS composed of a single PBS rod and the rod-membrane linker CpcL. Previously, CpcL was called CpcG2 in Synechocystis sp. PCC 6803 and CpcG3 in Anabaena. In the following, we use only the name CpcL to designate the rod-membrane linker protein in different organisms. CpcL has an N-terminal linker-PBS domain, almost identical to that of the rod-core linker CpcG proteins, and a C-terminal hydrophobic linker-L domain [3, 9]. The hydrophobic linker-L domain enables the direct interaction of the rod with the thylakoid membrane [9]. While the rod-core PBS serves as a major antenna for PSII, CpcL-PBS mediates the energy transfer specifically to PSI [9,10,11]. The fast and efficient energy transfer from CpcL-PBS to PSI is comparable to that from the rod-core PBS to PSII [11].
In Anabaena, PSI forms a tetramer structure, which has been confirmed as the dimer-of-dimer in cryo-electron microscopy (cryo-EM) [12, 13]. The CpcL-PBS interacts with the PSI tetramer to form a PBS-PSI supercomplex [9]. The single-particle electron microscopy (EM) structure of the PBS-PSI supercomplex revealed that CpcL-PBS is attached to the interface between two monomers within the dimer of the PSI tetramer [9]. Theoretically, the dimer-of-dimer structure revealed two such binding sites within the dimers and two potential binding sites between the dimers. However, it is not yet elucidated whether more CpcL-PBS could bind to the PSI tetramer.
CpcL-PBS forms a PBS-PSI supercomplex and transfers the light energy to PSI. On the other hand, the rod-core PBS can also function as a PSI antenna. Here, we revealed the role of linker proteins on the amount of different PBS assemblies and supercomplexes. We characterized two deletion mutants lacking the linker genes for the rod-core PBS (∆cpcG4) or the CpcL-PBS (∆cpcL), and a cpcL overexpression mutant. Our results demonstrate that the linker proteins compete for the assembly of the corresponding PBS. In addition, the similarity of the PBS-PSI supercomplex assembly in the cpcL overexpression and ∆cpcG4 mutants suggests that the CpcL/CpcG4 linker ratio determines the amount of PBS-PSI supercomplexes in the cell.
2 Materials and methods
2.1 Strains and growth conditions
Wild type and mutants of Anabaena sp. PCC 7120 were grown at 31 °C in liquid BG11 medium supplemented with 20 mM HEPES KOH (pH 8.2) [14] and with bubbling with 1% CO2 under continuous illumination with white fluorescent lamps (ca. 20 μmol photons m–2 s–1). For the cultivation of mutant strains, 30 µg mL−1 neomycin or 20 µg mL−1 erythromycin were added.
2.2 Construction of mutants
All primers are listed in Table S1. For the cpcL gene deletion, upstream, downstream-1, and downstream-2 regions of the cpcL gene were amplified by PCR using primer pairs of cpcG2-1F/cpcG2-2R, cpcG4-1F/cpcG4-2R, and all0538-1F/all0538-2R, respectively. Details of the construction of the plasmids are shown in Fig. 2A. PCR products of upstream and downstream-1 were subcloned into pPCRScript separately, and the downstream-2 fragment was subcloned into pT7Blue. The respective upstream and the downstream DNA fragments were digested with BamHI/EcoRI or ClaI/XhoI, respectively, and were ligated. The neomycin resistance cassette was inserted into the EcoRI site between the downstream-1 and 2 DNA fragments. The cpcG4 gene was inactivated by the insertion of the neomycin resistance cassette into a HpaI site. The XhoI–SacI fragment from pPCRScript_∆cpcL::NmR and pT7Blue_∆cpcG4::NmR were subcloned into the sacB-vector pRL271 between the XhoI and SacI sites [15], resulting in pRL271_∆cpcL::NmR (∆cpcL) and pRL271_∆cpcG4::NmR (∆cpcG4). The overexpression plasmid for the cpcL gene was constructed as follows. The subcloned cpcL gene in the pPCRScript was digested by BglII and SalI and ligated into the BamHI/SalI digested pAM505-based shuttle vector [16, 17] pAM505::ErR creating the pAM505::ErR_cpcL plasmid. The pAM505::ErR_cpcL and the trc promoter [18,19,20] were amplified by PCR using primer pairs cpcL-6F/pAM505-6R and trc-19FpAM505/trc-20RAncpcL and combined using the In-Fusion System (Takara).
a Scheme of the cpc operon in Anabaena and a map illustrating the construction of the ∆cpcL and ∆cpcG4 mutants. The arrows show the putative transcription start sites as predicted by RNA-seq data [29]. For ∆cpcL mutant construction, to avoid polar effects on the downstream cpcG4 gene expression, the complete ORF of the cpcL gene was deleted, and the kanamycin/neomycin resistance cassette was inserted into an intergenic region after the whole cpc operon. For ∆cpcG4 construction, the kanamycin/neomycin resistance cassette was inserted into the cpcG4 gene. b Verification of complete segregation of mutant alleles in ∆cpcL and ∆cpcG4 mutants and presence of the cpcL overexpression plasmid by PCR. Wild type (WT) served as a negative control. Expected product sizes are 1755 bp or 2475 bp for ∆cpcL mutant and WT, 2281 bp or 931 bp for ∆cpcG4 mutant or WT, and 982 bp for cpcL overexpression mutant. Clone 1 was used each time for the analyses in this study
Mutation of Anabaena was achieved by triparental conjugation using strain J53 bearing RP4 and cargo strain HB101, bearing the helper plasmid pRL623 [21]. The selection of double recombinants was performed using sacB as a negative selection marker [22]. The correct genome modification and the complete segregation of mutant alleles in ∆cpcL and ∆cpcG4 mutants were verified by PCR using the primer pairs cpcG2-1F/cpcG4-2R, and cpcG4-1F/cpcG4-2R, respectively. Primer pair cpcL-4R/trc-19FpAM505 was used for PCR to verify the presence of the plasmid in the cpcL overexpression mutants.
2.3 Isolation of phycobilisomes
PBSs were isolated as previously described [9]. Briefly, cells were harvested, washed twice with 0.9 M potassium phosphate buffer (pH 7.0), and broken with zirconia beads with a bead beater (Micro Smash MS-100R; TOMY). The cell extract was treated with 2% (v/v) Triton X-100 in 0.9 M phosphate buffer (pH 7.0) for 30 min and centrifuged at 20,000×g for 20 min at 18 °C to separate the upper green Triton X-100 layer from the lower blue aqueous phase. The blue phase was loaded onto a 10–50% (w/v) linear sucrose density gradient with 0.9 M phosphate buffer and centrifuged at 130,000×g for 16 h at 18 °C.
2.4 Isolation of photosystem complexes
PS complexes were isolated as described in a previous report [9]. Cells were harvested, resuspended in buffer A containing 50 mM 4-morpholineethanesulfonic acid sodium hydroxide (pH 6.5), 10 mM MgCl2, 5 mM CaCl2, and 25% (w/v) glycerol, and disrupted with zirconia beads with a bead beater (Micro Smash MS-100R; TOMY). After removing unbroken cells, the resulting supernatant was centrifuged at 300,000×g for 30 min at 4 °C to separate thylakoid fractions and soluble proteins. Thylakoid fractions were adjusted to a concentration of 1 mg chlorophyll (Chl) mL–1 with buffer A and solubilized with 1% n-dodecyl-β-d-maltoside (DM) (Sigma-Aldrich) on ice for 30 min, followed by centrifugation at 300,000 × g for 30 min at 4 °C. The supernatant was diluted with three volumes of buffer B (50 mM 4-morpholineethanesulfonic acid sodium hydroxide (pH 6.5), 10 mM MgCl2, and 5 mM CaCl2), loaded onto a 10–30% linear sucrose density gradient in buffer B containing 0.01% DM, and centrifuged at 130,000×g for 6–8 h at 4 °C.
2.5 SDS-PAGE and Western blot analysis
Proteins were separated by SDS-urea-PAGE with a 16–22% (w/v) linear gradient of polyacrylamide containing 7.5 M urea [23], followed by Coomassie or silver staining [24]. The bilin fluorescence of the phycobiliproteins was detected through a 605 nm filter with excitation at 532 nm (FMBIO II, TaKaRa) before silver staining. Western blot analysis was performed as described before [9]. The peptide antibodies for CpcL, CpcG4, and CpcG1 were produced by Takara Bio and used for immunodetection.
2.6 Absorption and 77 K fluorescence spectroscopy
Absorption spectra were measured at room temperature (UV-2400PC; Shimadzu). Fluorescence excitation spectra were measured with a fluorescence spectrophotometer (RF-5300PC; Shimadzu) equipped with a Dewar vessel containing liquid nitrogen. Samples were excited from 400 to 700 nm and fluorescence was monitored at the PSI emission peak at 725 nm.
3 Results
3.1 Construction of Anabaena mutants lacking the linker proteins CpcL and CpcG4
In order to evaluate the assembly of PBS-PSI supercomplexes in the absence of CpcL and CpcG4, we constructed the respective mutant strains ∆cpcL and ∆cpcG4. In Anabaena, the cpcL gene locates upstream of cpcG4 in an operon including genes encoding other PBS subunits (Fig. 2a). To avoid polar effects of a cpcL mutation on the downstream cpcG4 gene expression, we deleted the complete ORF of the cpcL gene and inserted the neomycin resistance cassette into an intergenic region after the whole operon (Fig. 2a). We verified the correct genome modification and the complete segregation of mutant alleles in ∆cpcL and ∆cpcG4 mutants by PCR (Fig. 2b), and used one completely segregated clone for further analyses. In addition to the gene deletion mutants, we also created a cpcL overexpression mutant in Anabaena. The cpcL gene was inserted together with a synthetic trc promoter into a self-replicating plasmid [20]. The presence of the plasmid in conjugated cells was verified by PCR (Fig. 2b).
3.2 Isolation of PBS fractions from ∆cpcL and ∆cpcG4 mutants
To investigate the effects of the disruption of cpcL and cpcG4 genes, we isolated PBSs by linear sucrose density gradient centrifugation from wild type, ∆cpcL, and ∆cpcG4 mutants grown under standard conditions (Fig. 3a). Distributions of CpcL-PBS and the rod-core PBS in the gradients were analyzed by Western blot analysis using an anti-CpcL antibody for CpcL-PBS, anti-CpcG4, and anti-CpcG1 antibodies for the rod-core PBS, respectively (Fig. 3b). In the wild type, CpcL and CpcG4 were detected in fractions 2–10 and 5–10, respectively (Fig. 3b). Similarly to a previous study [9], we detected the rod-core PBSs in several fractions (Fig. 3a, b). Fraction 4 did not correspond to a clearly visible band but rather to a pale blue region in the sucrose gradient (Fig. 3a). This fraction mainly contained CpcL and did not show prominent APC absorption (650 nm) in the spectra (Fig. 3c), indicative of the presence of CpcL–PBS complexes only. In the ∆cpcL mutant, no colored region corresponding to fraction 4 of the wild type was observed, whereas the rod-core PBS fractions (fractions 5–10 in the wild type) are clearly visible in this mutant (Fig. 3a, fractions 15–19 in ∆cpcL). We took the non-colored region of the ∆cpcL gradient (fraction 14) and measured the absorption spectra. Fraction 14 showed almost no absorption (Fig. 3c). In the ∆cpcG4 mutant, CpcL was detected in fractions 21–25 (Fig. 3b, lower panel). In the gradient, fraction 23 was at the same position as fraction 4 of wild type and was clearly colored (Fig. 3a). This fraction did not include the rod-core linker CpcG1 (Fig. 3b, lower panel). Absorption spectra revealed that fraction 23 includes PC and APC (Fig. 3c). These results suggest that this fraction is a mixture of the CpcL-PBS and detached discs from the rod-core PBS and that the assembly of CpcL-PBS was not affected in the ∆cpcG4 mutant. These results confirm that CpcL is a specific linker of the CpcL-PBS. The rod-core linker CpcG1 was detected in fractions 24 and 25 of the ∆cpcG4 mutant preparation, indicating that these two fractions contain the rod-core PBS (Fig. 3b). Absorption spectra showed a high APC-to-PC ratio in fractions 24 and 25, suggesting that these fractions contain a low amount of PBS rods (Fig. 3c). This result is consistent with previous analyses [6, 25], which showed that CpcG4 is the central rod-core linker in the rod-core PBS, and CpcG1 or CpcG2 cannot replace CpcG4.
Isolation of CpcL-PBSs and rod-core PBSs by centrifugation with a linear sucrose gradient under high-salt conditions. a Separation patterns of PBS fractions from wild type, ∆cpcL, and ∆cpcG4 after centrifugation at 130,000×g for 18 h at 18 °C. (B) Western blot analysis of CpcL, CpcG4, and CpcG1 linker proteins with specific antibodies in the different fractions from wild type and the ∆cpcG4 mutant. The same volume of samples was applied to the SDS-PAGE gel. Lane numbers correspond to the fractions numbered in (a). c Absorption spectra of PBS fractions from wild type and mutant samples. Fractions were directly used for the measurement without further dilution. Sample numbers correspond to the numbered fractions in (A). WT, wild type
3.3 Effect of cpcL deletion on PBS-PSI supercomplex formation
Next, we separated the PS complexes from the solubilized thylakoid fraction of the ∆cpcL mutant in sucrose gradients (Fig. 4a). In the ∆cpcL mutant, a green fraction which would correspond to the PBS-PSI supercomplex was not detected (fraction 4 of wild type). Therefore, we collected the fraction from the gradient of the ∆cpcL mutant which had a similar position in the gradient as the PBS-PSI in the wild type (fraction 4* in Fig. 4a) and performed SDS-PAGE and measured absorption spectra (Fig. 4b, c). Fraction 4* from the ∆cpcL mutant contained PSI, but no CpcL and PC subunits (Fig. 4b). The absorption spectrum confirmed that the fraction 4* did not contain PC (Fig. 4c). These results indicate that the CpcL linker is crucial for the PBS-PSI supercomplex assembly.
Isolation of PBS-PSI supercomplexes of wild type, ∆cpcL, and ∆cpcG4 mutants by centrifugation using a linear sucrose gradient. a Separation patterns of thylakoid membranes from wild type, ∆cpcL, and ∆cpcG4 after centrifugation at 130,000×g for 6 h at 4 °C. b Coomassie staining following SDS-PAGE of the fractions. The samples of wild type fractions 3 and 4; 5.0 µg Chla/lane, ∆cpcL fraction 3; 5.0 µg Chla/lane, fraction 4; 3.4 µg Chla/lane, ∆cpcG4 fraction 3; 5.0 µg Chla/lane, fraction 4; 4.3 µg Chla/lane, fraction 5; 3.0 µg Chla/lane, fraction 6; 1.0 µg Chla/lane were loaded. Lane numbers correspond to the numbered fractions in (a). Bands are assigned according to a previous report based on N-terminal sequencing [9]. c Absorption spectra of fractions containing PSI tetramers and potential PBS-PSI supercomplexes. Fractions were directly used for the measurement without further dilution. Spectra were normalized relative to the Chl peak at 678 nm
3.4 Isolation of PBS-PSI supercomplexes from a cpcL overexpression mutant
Overexpression of cpcL generated two blue-green bands (fractions 5 and 6), which appeared below the typical dense bands (fractions 3 and 4) in the gradient (Fig. 5a). We analyzed the subunit composition of the larger supercomplexes (fractions 5 and 6) by SDS-PAGE. Bilin fluorescence image and silver-staining demonstrated that they include PSI as well as PC subunits, which were nearly identical to the profile of fraction 4 (Fig. 5b). Absorption spectra confirmed the presence of PC in the fractions 5 and 6. Notably, the PC content of fraction 6 was prominent compared with fraction 5.
Isolation of PBS-PSI supercomplexes from the wild type (WT) in comparison to the cpcL overexpression (cpcL ox) mutant by centrifugation using a linear sucrose gradient. a Separation patterns of thylakoid membranes from wild type and cpcL overexpression mutant after centrifugation at 130,000× g for 6 h at 4 °C. b Silver staining and bilin fluorescence image (lower panel) following SDS-PAGE of the protein fractions. The same volume of samples was applied to the SDS-PAGE gel. 3.3 µg Chla of thylakoid membranes were loaded. Lane numbers correspond to the numbered fractions in (a). Bands are assigned according to a previous report based on N-terminal sequencing [9]. c Absorption spectra of PSI tetramer and supercomplex fractions of cpcL overexpression mutant. Fractions were directly used for the measurement without further dilution. Absorption spectra were normalized relative to the Chl peak at 678 nm
The structure of Anabaena PSI tetramer is dimer-of-dimer, and CpcL-PBS is attached to the interface between the two monomers within the dimer [9, 12, 13]. Theoretically, the dimer-of-dimer structure revealed two such binding sites within the dimers and two potential binding sites between the dimers. According to the assumption, two more blue-green bands (fractions 5 and 6) may correspond to the second binding site within the dimer and a first binding site between the dimers.
Absorption spectra showed that the fractions 3 and 4 in cpcL overexpression mutant had a higher PC content than the PBS-PSI supercomplex (fraction 4) in the wild type when normalized to the Chl content (Fig. 5c, peak at 625 nm). Fractions 3 and 4 in the cpcL overexpression mutant correspond to the fractions 3 and 4 in the wild type and should have the same PC content. CpcL-PBSs which are present in fractions 5 and 6 may dissociate and therefore contaminate fractions 3 and 4 during ultracentrifugation. This could be the reason why the PC content in fractions 4 and 5 was similar in the cpcL overexpression mutant.
Our results suggest that the larger supercomplexes are PBS-PSI assemblies, which contain more CpcL-PBS, and that the cpcL expression level determines the amount and the PBS-to-PSI stoichiometry of this complex.
3.5 Isolation of PBS-PSI supercomplexes from the ∆cpcG4 mutant
Further, we investigated the effect of the cpcG4 deletion on the PBS-PSI supercomplex organization. The ∆cpcG4 mutant showed two additional larger blue-green bands below the PBS-PSI band 4 in the sucrose gradient (Fig. 4a, fractions 5 and 6). SDS-PAGE revealed that these fractions include the subunits of PSI and the CpcL-PBS (Fig. 4b). The ratio of CpcL to PsaA/B was higher in the supercomplex fractions 5 and 6 in ∆cpcG4 than in the PBS-PSI fraction 4 in the wild type (Fig. 4B). Absorption spectra confirmed that the supercomplex fractions 5 and 6 of ∆cpcG4 had a higher PC-to-Chl ratio than the PBS-PSI fraction 4 of the wild type (Fig. 6a). Fractions 4–6 of ∆cpcG4 showed almost the same absorption spectra. This may suggest a dissociation and contamination of the PC discs from the lower fractions to the upper fractions during centrifugation. Further, we measured the energy transfer from CpcL-PBS to PSI by 77 K fluorescence excitation spectroscopy (Fig. 6b). PSI fluorescence (excitation at ~ 620 nm) was increased in the supercomplex fractions 4–6 of ∆cpcG4 compared to the PBS-PSI supercomplex in the wild type. PSI fluorescence excited around 620 nm in fraction 6 was slightly higher than in fractions 4 and 5, which showed almost identical spectra. These results suggest that the PBS-PSI supercomplexes of ∆cpcG4 contain more CpcL-PBS than the ones from the wild type and that these antennae are functional. These properties of the supercomplexes are very similar between ∆cpcG4 and the cpcL overexpression mutant. Therefore, we suggest that not the CpcL protein levels but the CpcL/CpcG4 linker ratio determines the supercomplex assembly.
Absorption and 77 K fluorescence spectra of PSI tetramer and supercomplex fractions of ∆cpcG4 mutant. Fractions were directly used for the measurement without further dilution. a Absorption spectra were normalized relative to the Chl peak at 678 nm. b Fluorescence excitation spectra were measured to investigate the energy transfer efficiency from CpcL-PBS to PSI. The spectra were normalized relative to the Chl peak at 673 nm. PSI fluorescence was measured at 725 nm. Numbers relate to the fraction numbers of the gradients shown in Fig. 3a
4 Discussion
A high CpcL/CpcG4 ratio in cpcL overexpression and ∆cpcG4 mutants led to an increased level of PBS-PSI supercomplexes and also larger supercomplexes below the typical PBS-PSI supercomplex fraction in the sucrose gradient (Figs. 4 and 5). Our results suggest that these larger PSI supercomplexes contain more CpcL-PBS per PSI than typical PBS-PSI supercomplexes. The single particle EM structure of the PBS-PSI supercomplex revealed that CpcL-PBS is attached to the interface between two monomers within a dimer of the PSI tetramer [9]. According to the cryo-EM structure of the PSI tetramer, there are two types of monomer–monomer interfaces [12, 13]. There are two potential CpcL-PBS attachment sites within the dimers and two additional potential binding sites between dimers in the PSI tetramer. The two faster migrating bands below the PBS-PSI supercomplex band might represent PBS2-PSI and PBS3-PSI supercomplexes. This means that a PBS is associated with an interdimer interface in addition to the two intradimer interfaces in PBS3-PSI supercomplexes. At the moment, we could not see a band corresponding to PBS4-PSI supercomplexes in the gradient. This may be partly due to the disintegration of PBSs from PBS-PSI supercomplexes during the experiment. This might suggest that the fourth attachment site has only a low affinity for the binding of CpcL-PBS. Further, the bands in the sucrose gradients of fraction 6 of ∆cpcG4 and the cpcL overexpression mutant were less colored than the bands of the upper PBS-PSI supercomplex bands (Figs. 4a and 5a), suggesting that the third attachment site of the PSI tetramer has also a lower affinity for CpcL-PBS. We suggest that there might be two types of interaction affinities between CpcL-PBS and the two types of monomer–monomer interfaces in the PSI tetramer. In the previously published structure of the PBS-PSI supercomplex, CpcL-PBS was attached to the monomer–monomer interface within the one PSI dimer [9]. This might suggest that the affinity of CpcL-PBS binding to the position within the dimer is higher than its affinity for binding between the dimers in the PSI tetramer. In the wild type, there were no larger supercomplexes, suggesting that the binding of one CpcL-PBS to PSI tetramer occurs preferentially. The first CpcL-PBS binding to the PSI tetramer may induce the structural changes of the PSI tetramer and reduce the binding affinity of the other three potential binding sites. Structural analysis of the supercomplexes will reveal the structural changes of the PSI tetramer by the binding of the CpcL-PBS. Overexpression of CpcL and our methods improvements, such as the use of other detergents or the use of the GraFix method [26], might have led to the recovery of more CpcL-PBS attached to the supercomplex.
77 K fluorescence excitation spectra showed a greater extent of the energy transfer from CpcL-PBS to PSI in the larger supercomplexes (Fig. 6b), suggesting that there is an increased number of functional CpcL-PBS to harvest and transduce more light energy to the PSI. Further investigations on the energy transfer from CpcL-PBS to the PSI in the supercomplexes by time-resolved fluorescence spectroscopy may reveal differences in the energy transfer efficiency between the two types of CpcL-PBS and PSI interaction in the larger supercomplexes.
Here we demonstrate that the ratio of the rod-core linker CpcG4 to the rod-membrane linker CpcL determines the assembly of the PBS-PSI supercomplexes. We previously showed that the ratio of CpcL/CpcG4 proteins is higher in heterocysts than in vegetative cells in Anabaena [9]. Therefore, we speculate that heterocysts may contain an increased amount of the PBS-PSI supercomplex and also PBS2-PSI and PBS3-PSI supercomplexes. In heterocysts, which are the specialized cells for nitrogen fixation, PSI activity is necessary to generate ATP for the nitrogenase reaction. It has been reported that CpcL is important for nitrogen fixation in Anabaena [13]. PBS2-PSI and PBS3-PSI would allow an efficient nitrogen fixation activity in heterocysts.
We could detect the PSI tetramer without CpcL-PBS binding even in the sucrose gradient of ∆cpcG4 and cpcL overexpression mutants, which showed the larger PBS-PSI supercomplexes. This might suggest that the PSI tetramer without CpcL-PBS has a different function than organizing the PBS-PSI supercomplex. It is reported that PSI is assembled into a PBS-PSII-PSI supercomplex with the rod-core PBS [27]. The PSI tetramers without CpcL-PBS in the sucrose gradient might correspond to the dissociated PSI from the PBS-PSII-PSI supercomplex. This may suggest different functions of the PSI between the PBS-PSI supercomplex and the PBS-PSII-PSI supercomplex. Since PSI is involved in both the cyclic and linear electron transfer pathways, PBS-PSI supercomplex and PBS-PSII-PSI supercomplex may correspond to each function. Future physiological analysis of ∆cpcL and ∆cpcG4 may reveal differences in the functions of each PBSs as a PSI antenna.
Our results suggest that CpcL and CpcG4 compete for the same rod component pool for the assembly of the two different types of PBS structures. For optimal photosynthesis under changing environmental conditions, it is necessary to regulate the expression of the two linker genes. In Anabaena, the cpcL gene is located in the cpc operon together with cpcG4. The whole cpc operon is transcribed from the promotor in front of cpcB (Fig. 2a) [28]. RNA-seq analysis [29] suggests additional individual transcriptional start sites (TSS) for cpcL and the cpcG4 genes in the cpc gene cluster. Further, the activity of the TSS for the cpcG4 gene is repressed under nitrogen starvation conditions, whereas that for cpcL is not [29]. However, cpcL is also expressed under nitrogen repletion and in vegetative cells in which CpcL-PBS can function as a PSI antenna as well. Therefore, the ratio of CpcL to CpcG4 could also be important for regulating the energy transfer and linear and cyclic electron transfer pathways in vegetative cells. There may be some other conditions that increase the ratio of CpcL to CpcG4. Besides nitrogen starvation conditions, other conditions, such as high salt concentration, which require more cyclic electron transfer [30, 31], may also alter the ratio of CpcL to CpcG4 in vegetative cells.
In Synechocystis sp. PCC 6803, the cpcL gene expression is regulated by a two-component system consisting of the histidine kinase/response regulator pair, CcaS/CcaR, in a light color-dependent manner [7, 32, 33]. Green light activates autophosphorylation of the cyanobacteriochrome-type photoreceptor CcaS, and CcaS transfers a phosphate to the OmpR-type response regulator CcaR. The phosphorylated transcription factor CcaR induces cpcL expression. In contrast, red light converts CcaS to its ground state, and cpcL expression is not induced. It was reported that the PBS-PSI supercomplex level is increased under green light conditions in Synechocystis sp. PCC 6803 [34]. This suggests that an increased level of CpcL induces the supercomplex assembly under green light conditions in this cyanobacterium. Future work on the regulation of cpcL and cpcG4 genes in Anabaena will clarify how the photosynthetic electron transfer is optimized by employing the two different PBS antennae under changing environmental conditions.
Data availability
The data that support the findings of this study are available from the corresponding author, MW, upon reasonable request.
References
Bryant, D. A., & Canniffe, D. P. (2018). How nature designs light-harvesting antenna systems: design principles and functional realization in chlorophototrophic prokaryotes. Journal of Physics B: Atomic, Molecular and Optical Physics, 51, 033001. https://doi.org/10.1088/1361-6455/aa9c3c
Liu, L.-N., Chen, X.-L., Zhang, Y.-Z., & Zhou, B.-C. (2005). Characterization, structure and function of linker polypeptides in phycobilisomes of cyanobacteria and red algae: An overview. Biochimica et Biophysica Acta, 1708, 10. https://doi.org/10.1016/j.bbabio.2005.04.001
Watanabe, M., & Ikeuchi, M. (2013). Phycobilisome: architecture of a light-harvesting supercomplex. Photosynthesis Research, 12, 265–276.
Adir, N. (2005). Elucidation of the molecular structures of components of the phycobilisome: Reconstructing a giant. Photosynthesis Research, 85, 15–32. https://doi.org/10.1007/s11120-004-2143-y
Liu, L.-N., Chen, X.-L., Zhang, X.-Y., Zhang, Y.-Z., & Zhou, B.-C. (2005). One-step chromatography method for efficient separation and purification of R-phycoerythrin from Polysiphonia urceolata. Journal of Biotechnology, 116, 91–100. https://doi.org/10.1016/j.jbiotec.2004.09.017
Zheng, L., Zheng, Z., Li, X., Wang, G., Zhang, K., Wei, P., Zhao, J., & Gao, N. (2021). Structural insight into the mechanism of energy transfer in cyanobacterial phycobilisomes. Nature Communications, 12, 5497. https://doi.org/10.1038/s41467-021-25813-y
Hirose, Y., Chihong, S., Watanabe, M., Yonekawa, C., Murata, K., Ikeuchi, M., & Eki, T. (2019). Diverse chromatic acclimation processes regulating phycoerythrocyanin and rod-shaped phycobilisome in cyanobacteria. Molecular Plant, 12, 715–725. https://doi.org/10.1016/j.molp.2019.02.010
Liu, H., Weisz, D. A., Zhang, M. M., Cheng, M., Zhang, B., Zhang, H., Gerstenecker, G. S., Pakrasi, H. B., Gross, M. L., & Blankenship, R. E. (2019). Phycobilisomes harbor FNRL in cyanobacteria. MBio. https://doi.org/10.1128/mBio.00669-19
Watanabe, M., Semchonok, D. A., Webber-Birungi, M. T., Ehira, S., Kondo, K., Narikawa, R., Fujiwara, M., Boekema, E. J., & Ikeuchi, M. (2014). Attachment of phycobilisomes in an antenna–photosystem I supercomplex of cyanobacteria. Proceedings of the National Academy of Sciences, 111, 2512–2517. https://doi.org/10.1073/pnas.1320599111
Kondo, K., Ochiai, Y., Katayama, M., & Ikeuchi, M. (2007). The Membrane-Associated CpcG2-Phycobilisome in Synechocystis : A New Photosystem I Antenna. Plant Physiology, 144, 1200–1210. https://doi.org/10.1104/pp.107.099267
Noji, T., Watanabe, M., Dewa, T., Itoh, S., & Ikeuchi, M. (2021). Direct energy transfer from allophycocyanin-free rod-type cpcl-phycobilisome to photosystem I. The Journal of Physical Chemistry Letters, 6, 6692–6697.
Kato, K., Nagao, R., Jiang, T.-Y., Ueno, Y., Yokono, M., Chan, S. K., Watanabe, M., Ikeuchi, M., Shen, J.-R., Akimoto, S., Miyazaki, N., & Akita, F. (2019). Structure of a cyanobacterial photosystem I tetramer revealed by cryo-electron microscopy. Nature Communications, 10, 4929. https://doi.org/10.1038/s41467-019-12942-8
Zheng, L., Li, Y., Li, X., Zhong, Q., Li, N., Zhang, K., Zhang, Y., Chu, H., Ma, C., Li, G., Zhao, J., & Gao, N. (2019). Structural and functional insights into the tetrameric photosystem I from heterocyst-forming cyanobacteria. Nature Plants, 5, 1087–1097. https://doi.org/10.1038/s41477-019-0525-6
Rippka, R. (1988). Isolation and purification of cyanobacteria. In: Methods Enzymol. Elsevier, pp. 3–27. https://doi.org/10.1016/0076-6879(88)67004-2.
Black, T. A., Cai, Y., & Wolk, C. P. (1993). Spatial expression and autoregulation of hetR, a gene involved in the control of heterocyst development in Anabaena. Molecular Microbiology, 9, 77–84. https://doi.org/10.1111/j.1365-2958.1993.tb01670.x
Wei, T.-F., Ramasubramanian, T. S., & Golden, J. W. (1994). Anabaena sp. strain PCC 7120 ntcA gene required for growth on nitrate and heterocyst development. Journal of Bacteriology, 176, 4473–4482. https://doi.org/10.1128/jb.176.15.4473-4482.1994
Yoon, H.-S., & Golden, J. W. (1998). Heterocyst pattern formation controlled by a diffusible peptide. Science, 282, 935–938. https://doi.org/10.1126/science.282.5390.935
Huang, H.-H., & Lindblad, P. (2013). Wide-dynamic-range promoters engineered for cyanobacteria. Journal of Biological Engineering, 7, 10. https://doi.org/10.1186/1754-1611-7-10
Huang, H.-H., Camsund, D., Lindblad, P., & Heidorn, T. (2010). Design and characterization of molecular tools for a Synthetic Biology approach towards developing cyanobacterial biotechnology. Nucleic Acids Research, 38, 2577–2593. https://doi.org/10.1093/nar/gkq164
Olmedo-Verd, E., Flores, E., Herrero, A., & Muro-Pastor, A. M. (2005). HetR-dependent and -independent expression of heterocyst-related genes in an Anabaena strain overproducing the NtcA transcription factor. Journal of Bacteriology, 187, 1985–1991. https://doi.org/10.1128/JB.187.6.1985-1991.2005
Elhai, J., & Wolk, C. P. (1990). Developmental regulation and spatial pattern of expression of the structural genes for nitrogenase in the cyanobacterium Anabaena. EMBO Journal, 9, 3379–3388. https://doi.org/10.1002/j.1460-2075.1990.tb07539.x
Cai, Y., & Wolk, C. P. (1990). Use of a conditionally lethal gene in Anabaena sp. strain PCC 7120 to select for double recombinants and to entrap insertion sequences. Journal of Bacteriology, 172, 3138–3145. https://doi.org/10.1128/jb.172.6.3138-3145.1990
Ikeuchi, M., & Inoue, Y. (1988). Partial characterization of the iodination site in D1 protein of manganese-retaining and manganese-depleted photosystem II membranes. Plant and Cell Physiology, 29, 695–705. https://doi.org/10.1093/oxfordjournals.pcp.a077549
Aro, E.-M., Suorsa, M., Rokka, A., Allahverdiyeva, Y., Paakkarinen, V., Saleem, A., Battchikova, N., & Rintamäki, E. (2005). Dynamics of photosystem II: A proteomic approach to thylakoid protein complexes. Journal of Experimental Botany, 56, 347–356. https://doi.org/10.1093/jxb/eri041
Chang, L., Liu, X., Li, Y., Liu, C.-C., Yang, F., Zhao, J., & Sui, S.-F. (2015). Structural organization of an intact phycobilisome and its association with photosystem II. Cell Research, 25, 726–737. https://doi.org/10.1038/cr.2015.59
Kastner, B., Fischer, N., Golas, M. M., Sander, B., Dube, P., Boehringer, D., Hartmuth, K., Deckert, J., Hauer, F., Wolf, E., Uchtenhagen, H., Urlaub, H., Herzog, F., Peters, J. M., Poerschke, D., Lührmann, R., & Stark, H. (2008). GraFix: Sample preparation for single-particle electron cryomicroscopy. Nature Methods, 5, 53–55. https://doi.org/10.1038/nmeth1139
Liu, H., Zhang, H., Niedzwiedzki, D. M., Prado, M., He, G., Gross, M. L., & Blankenship, R. E. (2013). Phycobilisomes supply excitations to both photosystems in a megacomplex in cyanobacteria. Science, 342, 1104–1107. https://doi.org/10.1126/science.1242321
Bryant, D. A., Stirewalt, V. L., Glauser, M., Frank, G., Sidler, W., & Zuber, H. (1991). A small multigene family encodes the rod-core linker polypeptides of Anabaena sp. PCC7120 phycobilisomes. Gene, 107, 91–99. https://doi.org/10.1016/0378-1119(91)90301-Q
Mitschke, J., Vioque, A., Haas, F., Hess, W. R., & Muro-Pastor, A. M. (2011). Dynamics of transcriptional start site selection during nitrogen stress-induced cell differentiation in Anabaena sp. PCC7120. Proceedings of the National Academy of Sciences, 108, 20130–20135. https://doi.org/10.1073/pnas.1112724108
Jeanjean, R., Bédu, S., Havaux, M., Matthijs, H. C. P., & Joset, F. (1998). Salt-induced photosystem I cyclic electron transfer restores growth on low inorganic carbon in a type 1 NAD(P)H dehydrogenase deficient mutant of Synechocystis PCC6803. FEMS Microbiology Letters, 167, 131–137. https://doi.org/10.1111/j.1574-6968.1998.tb13218.x
Hagemann, M., Jeanjean, R., Fulda, S., Havaux, M., Joset, F., & Erdmann, N. (1999). Flavodoxin accumulation contributes to enhanced cyclic electron flow around photosystem I in salt-stressed cells of Synechocystis sp. strain PCC 6803. Physiologia Plantarum, 105, 670–678. https://doi.org/10.1034/j.1399-3054.1999.105411.x
Hirose, Y., Narikawa, R., Katayama, M., & Ikeuchi, M. (2010). Cyanobacteriochrome CcaS regulates phycoerythrin accumulation in Nostoc punctiforme, a group II chromatic adapter. Proceedings of the National Academy of Sciences, 107, 8854–8859. https://doi.org/10.1073/pnas.1000177107
Hirose, Y., Shimada, T., Narikawa, R., Katayama, M., & Ikeuchi, M. (2008). Cyanobacteriochrome CcaS is the green light receptor that induces the expression of phycobilisome linker protein. Proceedings of the National Academy of Sciences, 105, 9528–9533. https://doi.org/10.1073/pnas.0801826105
Gao, F., Ogawa, T., & Ma, W. (2018). Effect of green light on the amount and activity of NDH-1-PSI supercomplex in Synechocystis sp. strain PCC 6803. Photosynthetica, 56, 316–321. https://doi.org/10.1007/s11099-018-0790-z
Acknowledgements
This work was supported in part by the JST CREST program (JPMJCR11B1 to M.I. and M.W.) and KAKENHI (Grant 17H05716 and 20J40231 to M.W., and 16H06558 to M.I.). The authors are grateful to Prof. Conrad W. Mullineaux for his helpful comments on this manuscript.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
This publication is dedicated to Prof. Silvia E. Braslavsky, a pioneer in photobiology and photobiophysics, on the occasion of her 80th birthday.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Watanabe, M., Ikeuchi, M. & Wilde, A. The organization of the phycobilisome-photosystem I supercomplex depends on the ratio between two different phycobilisome linker proteins. Photochem Photobiol Sci 22, 1561–1572 (2023). https://doi.org/10.1007/s43630-023-00397-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43630-023-00397-2