Abstract
Parkinson’s disease (PD) is a neurological condition characterized by the disruption of both motor and non-motor functions. Given the absence of a definitive diagnostic method, it is crucial to uncover its root causes. Consequently, individuals displaying symptoms of Parkinson’s disease can promptly receive treatment and comprehensive care. To address this, our study aims to develop an AI-powered system capable of detecting Parkinson’s disease and subsequently evaluating the primary factors influencing its development. We collected 12 distinct datasets from the well-known PPMI database, covering various medical assessments such as motor abilities, olfaction, cognition, sleep patterns, and depressive symptoms. Subsequently, we refined this raw data using advanced search techniques to tailor it to our model’s requirements. Moreover, we introduced a novel labeling approach known as the majority voting algorithm. Following data preparation, we conducted Single and Multi-Modality analyses, focusing on single-treatment approaches and integrating multiple treatments for a comprehensive therapeutic strategy. To analyze these both, we employed five distinct Machine Learning algorithms. Notably, the Support Vector Machine (linear) emerged as the top performer, reaching an accuracy of 100% in both single and multimodality analysis. Furthermore, we employed four tree-based models for feature selection, with the Gradient Boosted Decision Tree excels in identifying the most significant features. Finally, we employed an Artificial Neural Network utilizing these key features, achieving the highest accuracy of 91.41%.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Parkinson’s disease is a progressive neurological condition caused by the loss of dopamine-producing brain neurons, crucial for controlling movement [1]. Essentially, it disrupts the nervous system’s functioning, affecting organs regulated by these nerves. James Parkinson first described this disease in 1817, leading to its eponymous designation [2]. Over time, his contribution gained significant recognition in the medical community, driving increased interest and research. According to the World Health Organization (WHO), the prevalence of PD has doubled in the last 25 years. In 1990, 2.5 million individuals worldwide were affected, a number that soared to 6.1 million by 2016 [3]. This figure has continued to rise, with over 8.5 million estimated cases globally in 2019 [4].
Typically, Parkinson’s manifests around the age of 70, with a small percentage experiencing it before 50 [5]. Its incidence increases significantly after 60 [6]. However, it turns out that its prevalence rises dramatically over the age of 60 [6]. The prevalence of Parkinson’s disease varies among populations in Europe and Asia, with higher rates in North America, Europe, and Australia, and especially among people between the ages of 70 and 79 [7]. The impact of Parkinson’s extends beyond the individual, affecting families, caregivers, and healthcare systems. As the population continues to age, understanding and addressing Parkinson’s disease remains an essential field of research and a matter of public health concern.
Parkinson’s disease currently has no known cure, although available drugs and exercise, which may have few side effects, can manage symptoms and slow down the disease’s progression [8]. Thus, early Parkinson’s disease detection becomes essential for timely medical intervention to improve the quality of life. Currently, medical professionals around the world are actively investigating a range of diagnostic procedures to precisely diagnose Parkinson’s disease, placing significant emphasis on both single and multi-modality analysis. Single modality treatment entails selecting one approach from a range of available options to address a specific disease. It offers simplicity but may lack accuracy in capturing all influencing factors. Multi-modality treatments, on the flip side, combine various treatment approaches, either sequentially or simultaneously, to comprehensively address the disease process. This analysis provides a comprehensive understanding, though it involves increased complexity and requires advanced techniques. However, many previous studies have tended to narrow their focus on multimodality analyses, such as aggregating clinical assessments or audio tests, rather than single modality analysis. As a consequence, it creates limitations by conducting comprehensive and detailed investigations of various aspects that are important for understanding the causes of Parkinson’s disease. Furthermore, the limited focus hinders the ability to find the major factors involved in the development of Parkinson’s disease. Therefore, this study addresses this gap by employing a machine learning approach to analyze both single and multi-modalities concurrently to develop an automated system that can determine if a person has Parkinson’s disease along with assessing the vital factors that lead to the disorder. Moreover, the rationale for incorporating both single and multi-modal analysis techniques lies in the desire to comprehensively explore and compare outcomes. Furthermore, our research employs both singular and multimodal analytic methods to offer a thorough comprehension of the topic being studied. To provide a broader interpretation of the results, this approach looks at how many data sources contribute to the overall study, allowing for comparisons and the discovery of subtle trends across multiple dimensions. Thus, a brief overview of the major contributions of this study is listed below:
-
To attain our goal, 12 distinct sets of recent samples focusing on different medical assessments from the year between 2010 to 2023 are collected from a reputable database named Parkinson’s Progression Marker Initiative (PPMI). This repository is a global study collaborating with researchers, medical professionals, and patients launched in 2010, to create a data set and bio-sample library for Parkinson’s research. Afterward, a series of techniques are applied for Dataset Creation that effectively fits our model. Notably, a novel method named as majority voting technique is then applied to accurately label every sample.
-
To detect Parkinson’s disease, Single and Multi-Modality analyses are performed using 5 different ML algorithms. In single-modality analysis, each test is considered individually for analysis, while multi-modality analysis involves merging all tests for comprehensive analysis. So in our study, in single-modality analysis, the model is trained separately using 12 different datasets, whereas in multi-modality analysis, these individual datasets are combined to form a unified dataset for training. Hence, the Support Vector Machine (linear) outperforms the others in terms of accuracy (reaching 100%).
-
The study identified the top 10 important features that contribute most to Parkinson’s disease using 4 tree-based ML models. In this study, we have adopted tree-based models because Tree-based models are highly effective for structured/tabular data with a moderate to large number of features by making nonlinear relationships between features and the target variable. Additionally, It reduces the risk of overfitting by filtering out less important variables during the training process. Therefore, in our study, the Gradient Boosted Decision Tree outperformed the others in determining the most important features. The most effective feature selection method is then determined by using the same models for training and testing. Finally, we apply an Artificial Neural Network using top important features to evaluate the performance and achieve the highest accuracy of 91.41%
As of our knowledge, an entirely new set of data from 2010 to the present day is collected from the well-known database Parkinson’s Progression Marker Initiative (PPMI) repository. Following that, we propose innovative techniques for data creation, notably a unique Majority Voting Algorithm for precise class labeling. Subsequently, both single- and multi-modality approaches are conducted, and identified the key factors that lead to Parkinson’s disease.
The rest of the paper is documented as follows. The background research on our subject is covered in Sect. 2, which also outlines the development of the field’s expertise over time. Our overall system’s architecture is presented in Sect. 3, along with the selection of the data processing pipeline, machine learning models, and assessment metrics details. Following that, an exhaustive analysis of the experiment with the algorithms, testing results, and identification of key features are explained in Sect. 4. After discussing the essential aspects of this study in Sect. 5, the paper concludes with an outline of the future scope in Sect. 6.
2 Related Work
The development of non-intrusive techniques for patient monitoring has been aided by the last ten years’ improvement in the identification of Parkinson’s disease symptoms. Parallel to this, the creation of reliable solutions for patients with diverse medical diseases has been sped up thanks to the rapid advancement of machine learning and data analytics techniques. This confluence of developments highlights the possibility of improving healthcare monitoring and diagnostics in a more user-friendly and effective way.
C. Xiao and his colleagues [9] applied a tree-based, important-driven model to find key features to classify Parkinson’s disease. A customized dataset with 751 samples is collected from a well-known database name as PPMI. Afterwards, the Gradient Boosting Decision Tree Model is applied to find the topmost features. Finally, an neural network with 2 hidden layers (30 and 40 neurons on each hidden layer) is applied by varying the number of features for PD-Classification. Their research established that a highly accurate Parkinson’s Disease classification model can be developed applying just a few key characteristics. The tests on Hopkins verbal learning, motor functions, cognitive assessment, symbol digit modalities, olfactory function, and autonomic function are the most significant of the top 12 characteristics that contribute to high performance. Finally, considering the top 28 important features, an optimal accuracy of 99.74% and F1-score of 99.86% are achieved.
Tyler D. Alexander and his allies [10] develop a machine learning algorithm that predicts changes in Parkinson’s patients’ quality of life (QoL). To accomplish their goal, they collected at least 12 months identified follow-up patients records of 630 from the Parkinson’s Progression Markers Initiative database. Utilizing UPDRS-II scores, an artificial neural network with sigmoidal activation function with 10 hidden neurons is created to forecast whether or not a patient would experience clinical worsening at 12 months of follow-up. Additionally,Twelve unique divisions of data were generated, and for each subdivision, six unique models were developed to predict UPDRS-II score worsening at 12 months. The algorithm achieved a 90% specificity and 56% sensitivity, which suggests that a learning model may rule out patients whose QoL had significantly deteriorated, necessitating additional research and attention.
Alex, John Sahaya Rani [11] created a PPMI Data Repository for Parkinson’s Disease Imaging and Modeling to aid in early detection. They explores the feasibility of creating a cost-effective, robust wearable device for early detection and real-time data collection of Parkinson’s Disease. In order to attain goal, A gait dataset is created by collecting 168 records by monitoring 81 patients with 56 features, derived from six experimental tasks from PPMI database. Data analysis is done using a variety of categorization algorithms. Afterwards, A Mutual-Information algorithm is employed as features selection techniques to check for high correlation between individual features and the predictor variable. After identifying top features, different ML algorithm are applied. An 91.90% which is highest accuracy is achieved from KNN model. With the help of this technology, it may be possible to identify motor changes that occur before a Parkinson’s diagnosis and possibly create more precise biomarkers for the condition.
Michela Russo and his fellow [12] mates talk about Gait analysis which can be an useful technique for early identification and assessment of distinguish both PD patients with and without mild cognitive impairment (MCI). To carry out, a custom dataset created with 80 participants were selected between January 2018 and March 2022 of the University of Salerno, Italy. To study PD-MCI utilizing spatial and temporal gait data, the researchers applied Uni-variate statistical analysis applied on clinical features and spatial and temporal parameters. Afterwards, machine learning algorithms are utilized to analyze model performances. The results demonstrated that PD-MCI patients had reduced gait performance and impaired dynamic balance, which increased the chance of falling. The ML algorithms successfully recognized PD-MCI more than 80% of the time. This study supports the use of gait analysis in the early diagnosis of Parkinson’s disease.
Haewon Byeon [13] used the Parkinson’s Dementia Clinical Epidemiology Data for develop a random forest-based prediction model for Parkinson’s disease with mild cognitive impairment (PD-MCI). In order to develop model, a total of 96 records are collected. The model took into account health-related behaviors, environmental factors, medical history, bodily functions, depression, and cognitive processes. Random forest is used in different ways is applied for attain the goal. Based on the PD-MCI prediction model, the study suggests creating a methodology to quickly identify early-stage PDD for specialized monitoring and following high-risk populations.
Maitane Martinez–Eguiluz and her colleagues [14] developed a technique that permitted them to choose the most discriminative non-motor factors to make possible tools for PD screening. They Collected data from Biocruces (96 subjects) and PPMI (687 subjects). Due to the small data in Biocruces, they added PPMI data with Biocruces and called the BIO-PPMI dataset. To improve the result, they combined both datasets due to a large number of missing datasets, and they used questions 1 to 21 from the SCOPA-AUT test. In the PPMI dataset, they dropped more than 20% values. In feature selection, they applied filter methods on both datasets’ FPI and FPT versions. They have used nine algorithms to identify PD patients. Most of the algorithms could perform high accuracy which is more than 80%. In Support Vector Machine and Multi-Layer Perceptron, they got the best result, which is 86.3% and 84.7%.
Aditi Govindu and her colleagues [15] aims to develop a PD detection technique through telemedicine. They have collected 195 audio samples from PPMI and UCI between the ages of 46 and 85. Following that, 3 different scenarios are implemented using 4 different machine learning (ML) techniques, including support vector machine, logistic regression, random forest, and k closest neighbor models. First, each of the 4 models are applied to 22 attributes, Second, principal component analysis is applied to identify 5 key attributes with the 4 models, Finally, the entire dataset is balanced by sampling the minority class and applied the 4 algorithms. Therefore, Results are compared for each case using the confusion matrix, ROC-AUC curve, and accuracy. Thus, the best result is achieved for SVM model with accuracy of 91.836%.
Kamal and Venkata [16] discover an ML forecasting model for Parkinson’s disease (PD) is a major public health problem, with one million people suffering in the US and 5 million worldwide.Predicting Parkinson’s disease (PD) early stages is essential for treatment. Machine learning algorithms can improve model performance and predict disease progression, aiding neuroprotective therapies. PPMI, an observational study, collected data from 184 normal patients and 402 early PD subjects. The study used features from the University of Pennsylvania Smell Identification Test, RBD screening questionnaire, CSF markers, and SPECT measurements of striatal binding ratio (SBR). Boosted Logistic Regression showed the best performance, with an impressive accuracy of 97.159 % and a 98.9 % area under the ROC curve. This research demonstrates the potential of machine learning algorithms for better decision-making and diagnosis in nuclear experts.
Arti Rana and her fellow mates [17] focus on Neurodegenerative disease which affects the neural, physiological, and behavioral systems of the brain. Slow movements, also known as ”bradykinesia,” are among the early symptoms. They start to show up in middle age and get worse as one age. The study suggests using supervised classification techniques like SVM, naive Bayes, K-NN, and ANN to identify subjective Parkinson’s disease. The suggested approach involves selecting features based on filtering, wrapping, and classification techniques. The duration and cost of PD screening might be decreased with this strategy. They uses the voice feature of 195 Parkinson’s disease patients which are identified in the Max Little dataset from Oxford University, and a ”status” column is used to contrast healthy and unwell people. According to the results of the experiment, ANN has an accuracy of 96.7%. Which is the highest accuracy. So, ANN is the most accurate classifier.
Marimuthu et al. [18] proposed a system that uses the UCI ML Parkinson’s dataset and the XGB Classifier algorithm to predict Parkinson’s disease. The dataset has 197 records and 24 attributes, including noise-to-tonal component ratio, minimum vocal fundamental frequency, average vocal fundamental frequency, maximum vocal fundamental frequency, and various measures of fundamental frequency variation. According to the results, XGB Classifier shows the highest accuracy which is 94.87%.
Khushal Thakur and his fellow mates [19] do this research to create a machine-learning model that can effectively identify Parkinson’s disease based on speech characteristics. The Random Forest Classifier, Decision Trees Classifier, SVM, and Extra Trees Classifier are among the models that are employed. Their research-trained model indicates that there are 106 patients in total, of which 92 have Parkinson’s disease and 14 are healthy. Their trained model then predicts that 7 subjects will be healthy and 99 will have Parkinson’s disease. The accuracy of the Extra Trees Classifier is 94.339% when used in conjunction with the SelectFromModel feature selection strategy. With an accuracy of 91.57%, the Random Forest Classifier likewise operates effectively.
Pankaj Kumar Keserwani and his colleagues [20] emphasize Artificial Intelligence (AI) as a possible path for better medical care and early detection. AI is a crucial tool for PD detection because it can process massive datasets and produce precise statistical predictions. This article examines the state of the art in AI research as well as several AI-based techniques, such as meta-heuristic algorithms, deep learning, and machine learning.
A concise overview of related comparative works is discussed in Table 1.
3 Methodology and Model Architecture
This section presents comprehensive descriptions of different components of our methodology. Fig 1 shows the overview of the steps that we have applied in our study.
3.1 Data Process Pipeline
A comprehensive overview of the data collection, dataset creation, and data preprocessing procedures given in this section.
3.1.1 Data Collection
This subsection provides comprehensive details about our data source and the records we have collected.
-
Parkinson’s Progression Marker Initiative:PPMI database serves as the primary repository for our study. The Parkinson’s Progression Marker Initiative (PPMI) is a novel research program aims to advance our understanding of Parkinson’s disease. It is a collaborative study aimed to bring together a diverse group of researchers, medical professionals, and patients, was founded by the Michael J. Fox Foundation and and supported by a variety of industry partners for Parkinson’s Research in 2010. PPMI includes diverse data types including clinical data and imaging information. It also includes genetic data, surveys, as well as biological samples from more than 1500 patients at 33 clinical locations throughout the world to provide a comprehensive understanding of Parkinson’s disease for improved diagnosis and treatment.
-
Data: To achieve our goal, we collect a wide array of samples from 2010 to 2023. This data consists of two groups of patient records for single-modality and multi-modality analysis: those with Parkinson’s disease and those who do not have Parkinson’s disease. The collected records are associated with a variety of assessments, including movement behavior, speech, sense of smell, writing tests, etc. These individual tests consist of multiple sets of questions from various subjects that are assigned specific scores. In total, 12 tests are collected, each with a significant number of features. Here, Table 2 illustrates the overall breakdown of our collected samples.
3.1.2 Data Preprocessing
Several features are eliminated during the pre-processing stage that is irrelevant to the training model, i.e., ”Rec id”, ”Origin entry”, and ”Last update”. A significant percentage of collected records were found to be null and duplicate, which were removed as well. However. in certain cases, null entries are filled with the min and average values. Besides, feature renaming is carried out along with standardizing features to ensure all numerical attributes fall into a similar range. Noted that the null and duplicate records checking is carried out twice: once before creating the dataset and then again after it is created.
3.1.3 Dataset Creation
Our study focuses on single and multi-modality analysis as two primary areas and to carry it out, unlabeled samples and a mix of patient records from individuals who do not undergo all 12 tests are collected. While conducting a single modality analysis, 12 tests are investigated one by one which we are referring to as individual datasets for our convenience. Since a single test is considered at a time, therefore, we do not require those patients who undergo of all tests. However, to conduct a multi-modality analysis, these 12 tests need to be combined into a single one. To accomplish that, we require only those patients who undergo all 12 tests which we are referring to as common patients to ease our work. However, data analysis finds that the majority of patients have not undergone all 12 tests. Therefore, it is necessary to identify common records among all tests and label them to train our model. To label records, we thoroughly reviewed plenty of related studies. Here, Fig 2 illustrates the key steps that are carried out for dataset creation.
-
1.
Common Patients Dataset Creation: To conduct this study, a search algorithm is designed based on a tree-styled technique, as shown in Fig 3. Fig 3 illustrates that two datasets are used in each stage to apply the searching algorithm while taking account of two parameters, namely Event Id and Patient No. Following the identification of common records, the findings are merged and forwarded to the next stage. It should be noted that, Event Id and Patient No represent patient sectors by treatment duration and unique event, respectively. After completing all iterations, 3,509 patients are determined to undergo all 12 tests.
-
2.
Individual Dataset Creation: To carry out this research, we are required to create 12 individual datasets containing the patients who undergo all of the tests. Simply put, while making T1, we must include records of those patients who completed Test 1 as well as the other 11 tests. Therefore, each of the 12 datasets is taken one by one and matched up with the common patient dataset, considering two attributes, i.e., ’Patient No. and ’Event ID.
-
3.
Individual Dataset Labeling: To label data, each record is assigned a specific value, following which the sum of these values is calculated to determine the final score. Two ranges for binary classification labels, PD and NON-PD, are determined with respect to the final score. Patients fall into the PD and NON-PD groups, respectively, having Parkinson’s disease and do not show any signs of the disorder. Additionally, as shown in Table 3, each test has a unique range for diagnosing Parkinson’s disease.
-
4.
Combined Datasets Creations: Following the individual dataset labeling process, we create two datasets, namely Dataset 1and Dataset 2, by combining multiple individual tests. Dataset 1 contains Tests 1 to 6, while Dataset 2 includes all 12 tests.
-
5.
Majority Voting Algorithm Application: A majority voting technique is implemented to determine the most accurate class for both combined datasets i.e., Dataset 1 and Dataset 2. The voting mechanism takes into account all labels for all tests (6 labels for dataset 1 and 12 labels for dataset 2). Following that, it checks each tuple and counts the total number of PD and NON-PD records before determining the final class based on the three scenarios stated in Table 4. In addition, algorithm 1 represents the pseudo code of majority voting algorithm implemented in this research.
3.1.4 Final Dataset Description
The overview of a list of all datasets after pre-processing is discussed in Table 5. In addition, Table 6 illustrates the specific datasets that are applied to specific methods.
3.2 Single and Multi-Modality analysis
A single modality treatment is a focused strategy used in the field of medical diagnostics, where a particular disease process is examined and treated using one modality or method that has been carefully selected from a variety of alternatives. On the contrary, A comprehensive and integrated approach to medical therapy known as multi-modality treatment combines various approaches, modalities, or methods to treat a particular condition or disease process. In this study, while single modality analysis, each of the 19 diagnostics tested used to train and evaluate with various ML algorithms individually. whereas, Multi-Modality analysis combined all 19 diagnostic tests altogether and applied different ML algorithms.
3.2.1 Algorithm Application
After pre-processing, all of the datasets are split for training (80%) and testing (20%) to perform single and multi-modality analysis. Afterward, five machine learning classifiers i.e., (i) Support Vector Machine (ii) Random Forest, (iii) Adaboost Classifier, (iv) Gradient Boosting Decision Tree, and (v) Extra Trees Classifier are applied for the binary classification of Parkinson’s disease.
These Ml models are adopted in this study because tree-based models are ideal for structured data due to their collaboration effectively in tabular or structured datasets that have a moderate to large number of features. In binary classification tasks, Support Vector Machines (SVM) are also very effective at determining the best decision boundaries and differentiating between different categories. They are highly effective at classifying data and are chosen for tasks that call for accurate binary outcome classification. Support Vector Machine predicts classes using hyperplane, margin, and kernel characteristics, plotting data on an n-dimensional graph to identify the optimal hyperplane for each end class. In our study, 3 different kernels i.e., (i) Rbf, (ii) Linear and (ii) Sigmoid were used. Random Forest generates decision trees using the bagging technique for reliable results, selecting the highest-voted output class. It involves random sample selection, tree construction, voting technique, and highest-voted class selection. AdaBoost is a boosting technique that assigns a sample weight to all records, creating a sequential base learner using a decision tree with a single depth. The model is trained, and the total error is calculated by summarizing all weights. Gradient Boosting is an ensemble algorithm for minimizing bias error and adjusting initial assumptions in continuous and categorical target variables. It uses mean squared error as a cost function in classification problems. Extra Trees builds multiple decision trees by selecting split points at random, lowering variance, avoiding over-fitting, and producing reliable predictions.
3.3 Identifying Important Features
The key to early diagnosis of Parkinson’s disease involves understanding the primary factors contributing to Parkinson’s disease. Hence, this study employs tree-based machine learning models as feature extraction techniques. These models are chosen due to their capacity to represent complex nonlinear correlations between the target variable and the input features. One notable advantage is their innate capability to filter out less relevant variables during the training phase, thereby enhancing the model’s efficiency. In addition to that, tree-based models maintain interpretability, allowing for easy visualization and comprehension of each decision made by the model. Besides, Ensemble methods like random forests and gradient-boosting machines combine multiple decision trees to enhance predictive accuracy, reduce overfitting risk, and enhance model robustness and efficacy. Therefore, our study identifies, the top 10 features that cause Parkinson’s Disease by applying 4 different tree-based machine learning models, including (i) Random Forest, (ii) Adaboost Classifier, (iii) Gradient Boosting Decision Tree, and (iv) Extra Trees Classifier, which we discussed in the previous section of 3.2.1. Afterward, The same tree-based model that we applied for feature selection techniques was used to train and test our data for the top 10 features after identifying them. This way, the top 10 features for each tree-based technique were identified and the most accurate feature selection method was determined. Lastly, The Artificial Neural Network (ANN) was applied to the features that provided the best accuracy.
Table 7 illustrates the parameters of the ANN model that were used in this research.
4 Experiment and Result Analysis
This section illustrates the experiment and results of our adopted methodology.
4.1 Performance Evaluation
The performance of various machine learning models is evaluated using different metrics including accuracy, precision, recall, and F1 score. The equations to calculate these metrics are shown in (1), (2) and (3).
Where, TP and TN indicates true positives and true negatives, respectively, whereas FP and FN indicates false positives and false negatives.
4.2 Performance Analysis of Different Algorithms for Single and Multi-modality Data
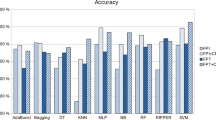
Table 8 displays the accuracy values obtained by various machine learning algorithms on all 12 datasets in single modality analysis.It demonstrates that in the majority of datasets, SVM using a linear kernel consistently attained 100% accuracy. In contrast, the Sigmoid kernel performs poorly across all datasets. While the accuracy values of the Random Forest, AdaBoost, and Gradient Boosting algorithms are competitive, Extra Trees excelled effectively across all datasets, with accuracy scores ranging from 81.18% to 95.43%.
The precision, recall, and f1-score values for various algorithms among 12 different datasets are compared in Tables 9, 10 and 11, respectively for single modality analysis. After analyzing all three table, it is noticeable that, the SVM (Linear) technique constantly achieves a perfect score of 100% on the majority of datasets. In contrast, SVM(Sigmoid) exhibits the poorest performance across most datasets. Aside from that, from Table 9, Extra Trees algorithm produces relatively consistent precision values, with scores ranging from 54.55% to 99.48% in contrast to others.
Table 12 compares the performance of various machine learning algorithms in multi-modality analysis. The findings suggest that, it seems that SVM (Linear) consistently outperforms SVM (Sigmoid) in accuracy, precision, and recall, while SVM (Linear) outperforms SVM (Sigmoid) in accuracy. Data on accuracy show that GBDT has the highest performance of 93.35%, followed by ADA at 92.96 percent. In addition, SVM with Kernal RBF and Sigmoid reach 100% precision, indicating no false positives. Aside from that, recall results are analyzed, and it is discovered that SVM (Linear) has the higjest recall (94.66%) with no false negatives.
Figure 4 illustrates the accuracy performance of single modality analysis. The SVM (Linear) consistently outperforms competing algorithms and achieves accuracy levels close to 100%. In contrast, ADA performs equally well. However, in every test case, Extra Tree performs worse than the other methods.
Figure 5 illustrates the performance of different matrices on various algorithms for Multi-Modality analysis. In terms of accuracy, GBDT has the highest accuracy 93.35% while SVM(RBF) and SVM(Sigmoid) perform better in terms of precision. However, the GBDT algorithm outperforms all others in terms of overall performance. SVM (sigmoid), on the other hand, performs the poorest.
4.3 Performance Analysis for Top Important Features
The factors that are most likely to cause Parkinson’s disease are identified by applying four deferent tree-based model. In this dedicated section, a thorough assessment is done to determine how well the models performed by looking at various measures like accuracy, precision, recall and f1-score.
Figure 6 shows the distribution of the percentages of the top 10 important features which is ranked by ADA, RF, ET and GBDT In Dataset-1. Here, only ADAboost model is identifying important features by providing same importance value. The most significant tests in dataset 1 are T1-NP1COG and T2-NP2DRES because they consistently perform well across all models and rank in the top 10 features. Despite having superior performance and continuing to be among the top features, T2-NP2EAT and T3-NP3RISNG. When it is come to ADA, we can see from figure that, Test 6’s feature ToTAL_CORRECT is the most important feature. In Random Forest, We can see that the feature NP2RISE from Test-2 is the most significant feature, and NP3RISNG from Test-3 is the second-most significant feature. According Extra Tress In Dataset-1’s top 10 most significant features NP3RISNG from Test-3 is the most essential feature. Applying GBDT the feature MCACITY is the most vital feature for Test-4, as shown in the Fig. 6. Furthermore, according to Test-2, the feature NP2RISE is the second-most necessary feature, whereas in the random forest, it was the essential feature.
The distribution of the percentages of the top 10 features, as determined by ADA, RF, ET, and GBDT in Dataset-2, is shown in Fig 7. T12-GDSDROPD is the most important test in dataset 2 since it consistently outperforms all other tests across all models and ranks among the top 10 features. T11-ESS1 and T2-NP2HOBB continue to rank among the top features despite their higher performance. Since Test 2 performs better almost always, the majority of the features are found in Test 2. In Test-3 NP3RTARU and VLTANIM in Test-16 are the most essential features when it comes to ADA. The feature NP2HOBB from Test-2 is the most important feature in Random Forest, and the feature NP2HYGN from Test-2 is the least important feature. The most important feature, as listed in Extra Tress In Dataset-2’s top 10, is GDSDROPD from Test-12. The most important feature for Test-2, as depicted in the 7, is applying GBDT to the feature NP2HOBB. The remainder of the feature performs poorly.
Table 13 shows the performance of the tree-based model for the top 10 features After applying the same model as feature selection. This table indicates that, among all models, GBDT has the highest accuracy for Dataset 1 at 91%. Additionally, both RF and GBDT provide the highest accuracy for Dataset 2 at 92%. As an outcome, it can be stated that since these models perform the best, the top features we identified after using them are most important to the detection of Parkinson’s disease.
Figure 8 and Fig. 9 the performance of the tree-based model when applied to Datasets 1 and 2. Figure 8 shows that the GBDT has the highest accuracy (91%), as well as the best Precision, Recall, and F1-Score. However ADA outperforms in each section. Moving on to Fig. 9, we found that for Dataset 2, GBDT acchives an accuracy of 92%. However, Random Forest (RF) did even better than GBDT in all the measurements we looked at. This means RF is be the better choice for Dataset 2.
Figures 10 and 11 show the NN application for the top 10 important features using GBDT for both datasets 1 and 2. For Dataset 1, after 10 epochs, GBDT provides 91.41% accuracy. On the other hand, for Dataset 2, RF and GBDT provide 89.06% and 89.84%, respectively. Thus, in all cases, GBDT showed the best performance. Additionally, it found the top features that are most important in both datasets.
While reviewing related studies of Parkinson’s disease, we identified similarities between our research and Xiao Chun’s study. Both our and Xiao Chun’s [9] research propose a key feature-based machine learning approach to identify key features for PD classification using different tree-based models and then apply ANN to compare model performance. However, our research also thoroughly analyzed each sample and developed a unique strategy for data labeling known as the majority voting technique. Additionally, this study deeply explores Single and Multi-Modality treatment by applying machine learning approaches along with multiple tree-based models to identify top features. Furthermore, it is found that ANN achieves 91.41% accuracy for dataset 1 when comparing the outcomes of the top 10 features using GBDT. However, before the ANN application, we also analyzed using SVM (SVM with a linear kernel offers the maximum accuracy for single and multi-modality analysis), and it was discovered that SVM offers 87.7% accuracy for the same scenario. However, it is worth noting that ANN’s training time is minimal at 3.70 s, despite potential higher computation costs associated with using all features. Here, Table 14 below provides a summary of the models’ running times for different no of features while training Dataset 1.
5 Discussion
This study aims to develop an AI-based system that can detect Parkinson’s disease and the factors that contribute to it. Our study conducted a series of analyses to identify the underlying factors. As an outcome, our analysis found that older people over the age of 60 are more affected by disease. Furthermore, the main factors causing this illness are identified in several tests, including motor function, autonomic function, the Epworth Sleepiness Scale, semantic fluency, etc. Thus, people who have issues with body position, movement, and muscle strength, as well as breathing, heartbeat, and digestion are more likely to develop PD.
Our study deeply explores Single and Multi-Modality treatment by applying machine learning approaches along with multiple tree-based models to identify top features. In both single and multi-modality analysis, the SVM (linear) outperforms other algorithms in accuracy achieving 100%. One significant factor contributing to achieving high performance is the quality of our dataset. We took several steps to ensure the dataset’s integrity, including using a majority voting algorithm for precise data labeling and implementing rigorous preprocessing techniques such as removing null and duplicated entries, eliminating irrelevant features, and standardizing features. These efforts were instrumental in improving the performance of our models.
Apart from that, we attain this remarkable accuracy through the implementation of feature selection techniques. After applying feature selection, the Neural Network (NN) was trained and achieved the highest accuracy of 91.41%. Therefore, this feature selection technique plays a crucial role in achieving high accuracy by addressing the curse of dimensionality and pinpointing the most relevant aspects of the data, which can lead to better generalization of unseen data. Moreover, by training models on a reduced set of features, feature selection simplifies the models and makes them more interpretable. This simplification enables the model to understand the underlying relationships between input features and the target variable more effectively. Additionally, by focusing on the most informative features, feature selection techniques facilitate the model in distinguishing between signal and noise in the data more accurately.
It is noteworthy that GBDT, RF, ANN, and SVM stand out as the algorithms delivering the highest results. Tree-based models like GBDT and RF are highly effective for structured/tabular data with a moderate to large number of features, contributing significantly to our dataset’s high performance. Conversely, SVM and ANN excel in binary classification by identifying optimal decision boundaries and learning complex nonlinear relationships between input features and target labels, respectively. Hence, we obtained satisfactory results using these models.
Throughout the course of our research, we encountered numerous challenges. Managing diverse datasets and unlabeled data posed significant hurdles, making the task quite challenging. Additionally, to gain a thorough comprehension of disease symptoms and determine the most effective strategy for class creation, we found it necessary to extensively review numerous related studies.
6 Limitations
This study delves into an extensive exploration of Parkinson’s disease and the critical factors underlying this condition using machine learning algorithms. However, we encountered several limitations in our research. One significant hurdle arose from the substantial volume of irrelevant data, which restricted our capacity to build a larger and more representative dataset for our studies. Furthermore, our research is confined to analyzing only tabular data
7 Conclusion and Future Scope
This study introduces machine learning and neural network techniques to identify Parkinson’s disease and analyze various aspects to pinpoint key features associated with the condition. To carry out this investigation, data from 12 different tests from the well-known site PPMI are collected and a custom dataset is constructed using the majority voting method. Subsequently, five distinct machine learning algorithms are employed to conduct Single and Multi-Modality analyses. Notably, in both single-modality analyses, SVM (Linear) consistently outperforms other algorithms, exhibiting higher accuracy, precision, recall, and F1 Score metrics. Similarly, in multi-modality analysis, SVM (Linear) also surpasses other algorithms by achieving a remarkable accuracy of 100%. Afterward, four tree-based models are utilized as feature selection techniques. For Dataset 1, Gradient Boosted Decision Tree (GBDT) emerges as the top feature selection technique with 91.41% accuracy, while Random Forest (RF) and GBDT excel for Dataset 2, achieving accuracies of 89.06% and 89.84%, respectively. Additionally, Neural Network (NN) is applied to the top 10 features, achieving 91.41% accuracy for Dataset 1, while RF and GBDT yield accuracies of 89.06% and 89.84% for Dataset 2, respectively. In the future, more data can be collected to conduct in-depth research on the disease. Moreover, besides the tree-based strategy, other feature selection techniques may be explored for comparative analysis. Additionally, adopting more advanced artificial intelligence methods, particularly explainable AI, holds promise for enhancing the study’s outcomes.
Data and Code Availability
A comprehensive overview of the PPMI Data source is provided in the Sect. 3. The Data is made accessible through a formal data sharing procedures to author Nusrat Islam upon a valid and reasonable request. The data is available at Parkinson’s Progression Markers Initiative (PPMI): https://www.ppmi-info.org/access-data-specimens/download-data. In addition, others materials will be made available on request.
Abbreviations
- AI:
-
Artificial intelligence
- ML:
-
Machine learning
- PPMI:
-
Parkinson’s progression Markers Initiative
- PD:
-
Parkinson’s disease
- ADA:
-
Adaptive boosting
- RF:
-
Random forest
- ET:
-
Extra trees
- GBDT:
-
Gradient-boosted decision trees
- SVM:
-
Support vector machine
- ANN:
-
Artificial neural network
References
Bereczki Dániel. The description of all four cardinal signs of Parkinson’s disease in a Hungarian medical text published in 1690. Parkinsonism & Related Disorders. 2010;16(4):290–3.
Parkinson James. An essay on the shaking palsy. J Neuropsychiatry Clin Neurosci. 2002;14(2):223–36.
GBD 2016 Parkinson’s Disease Collaborators. “Global, regional, and national burden of Parkinson’s disease, 1990-2016: a systematic analysis for the Global Burden of Disease Study 2016.” The Lancet. Neurology .2018; vol. 17(11): 939-953. https://doi.org/10.1016/S1474-4422(18)30295-3
Ternák G, et al. Antibiotic consumption patterns in European countries are associated with the prevalence of Parkinson’s sisease; the possible augmenting role of the narrow-Spectrum Penicillin. Antibiotics. 2022;11(9):1145.
Abdullah R et al. (2015) ’Parkinson’s disease and age: the obvious but largely unexplored link. Exp Gerontol 68 : 33-38.
Gazewood JD, Roxanne Richards D, Clebak K. Parkinson disease: an update. Am Family Phys. 2013;87(4):267–73.
Santos GD, et al. Present and future of parkinson’s disease in Spain: Parkinson-2030 delphi project. Brain Sci. 2021;11(8):1027.
Toulouse André, Sullivan Aideen M. Progress in Parkinson’s disease-where do we stand? Progress Neurobiol. 2008;85(4):376–92.
Xiao C. et al. Key Marker Selection for the Detection of Early Parkinson’s Disease using Importance-Driven Models. In: 2018 40th annual international conference of the IEEE engineering in medicine and biology society (EMBC). IEEE. 2018;
Alexander Tyler D, Nataraj Chandrasekhar, Chengyuan Wu. A machine learning approach to predict quality of life changes in patients with Parkinson’s Disease. Ann Clin Trans Neurol. 2023;10(3):312–20.
Alex JSR. ’Early Detection of Parkinson’s Disease using Motor Symptoms and Machine Learning. arXiv preprint arXiv:2304.09245(2023).
Russo M et al. (2023) Identification of a Gait pattern for detecting mild cognitive impairment in Parkinson’s disease. Sensors. 23(4): 1985.
Byeon Haewon. Is the random forest algorithm suitable for predicting parkinson’s disease with mild cognitive impairment out of parkinson’s disease with normal cognition? Int J Environ Res Publ Health. 2020;17(7):2594.
Martinez-Eguiluz M, et al. Diagnostic classification of Parkinson’s disease based on non-motor manifestations and machine learning strategies. Neural Comput Appl. 2023;35(8):5603–17.
Govindu Aditi, Palwe Sushila. Early detection of Parkinson’s disease using machine learning. Proc Comput Sci. 2023;218:249–61.
Challa KNR et al. An improved approach for prediction of Parkinson’s disease using machine learning techniques. In: 2016 international conference on signal processing, communication, power and embedded system (SCOPES). IEEE. 2016
Rana A, et al. An Efficient Machine Learning Approach for Diagnosing Parkinson’s Disease by Utilizing Voice Features. Electronics. 2022;11(22):3782.
Marimuthu M et al. (2021) Detection of Parkinson’s disease using machine learning approach. Ann Romanian Soc Cell Biol 2544-2550.
Thakur K et al. Diagnosis of Parkinson’s Disease Using Machine Learning Algorithms. Congress on Intelligent Systems. Singapore: Springer Nature Singapore. 2022
Keserwani PK, Das S, Sarkar N. A comparative study: prediction of parkinson’s disease using machine learning, deep learning and nature inspired algorithm. Multimedia Tools Appl 2024; 1-49.
Goetz CG et al. Movement Disorder Society-sponsored revision of the Unified Parkinson’s Disease Rating Scale (MDS-UPDRS): scale presentation and clinimetric testing results. Movement disorders. 2008 ;23(15): 2129-2170.
Hirschauer Thomas J, Adeli Hojjat, Buford John A. Computer-aided diagnosis of Parkinson’s disease using enhanced probabilistic neural network. J Med Syst. 2015;39:1–12.
Visser M et al. Assessment of autonomic dysfunction in Parkinson’s disease: the SCOPA-AUT. Movement disorders. 2004;19(11) : 1306-1312.
Johns MW A new method for measuring daytime sleepiness: the Epworth sleepiness scale. sleep. 1991; 14(6) : 540-545.
Lopez FV et al. Does the Geriatric Depression Scale measure depression in Parkinson’s disease?. Int J Geriatric Psychiatry. 2018; 33.12 : 1662-1670.
Stiasny-Kolster K et al. The REM sleep behavior disorder screening questionnaire–a new diagnostic instrument. Movement Disorders. 2007; 22(16) : 2386-2393.
Obeso I et al. Semantic and phonemic verbal fluency in Parkinson’s disease: Influence of clinical and demographic variables. Behavioural Neurol 2012; 25(2): 111-118.
Yang H-J et al. Measuring anxiety in patients with early-stage Parkinson’s disease: Rasch analysis of the State-Trait Anxiety Inventory. Front Neurol. 2019;10: 49.
Funding
The authors of this work did not receive any financial support for their research and writing.
Author information
Authors and Affiliations
Contributions
NI: conceptualization, investigation, methodology development, formal analysis, data collection & curation, visualization, writing & drafting, editing & proofreading. MSAT: conceptualization, investigation, formal analysis, visualization, figures & illustrations, writing & drafting, editing & proofreading. SIF: Conceptualization, Investigation, Formal analysis, Visualization, Writing & Drafting, Editing & Proofreading. RMR: Conceptualization, Investigation, Visualization, Formal analysis, Editing & Proofreading, Review & Feedback, Supervision.
Corresponding author
Ethics declarations
Conflict of Interest
The authors confirm that they have no potential Conflict of interest that might uphold the integrity of their research.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Islam, N., Turza, M.S.A., Fahim, S.I. et al. Single and Multi-modal Analysis for Parkinson’s Disease to Detect Its Underlying Factors. Hum-Cent Intell Syst 4, 316–334 (2024). https://doi.org/10.1007/s44230-024-00069-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s44230-024-00069-z