Abstract
Citrus fruit coloration is one of the vital quality traits that is determined primarily by the composition and content of carotenoids. Natural citrus fruit pigment mutants are available to study diverse and complex carotenoid metabolism. Here, ‘Jinlegan’ (MT) tangor is a spontaneous bud mutant derived from ‘Shiranuhi’ (WT) with distinctive bright yellow fruit. High performance liquid chromatography (HPLC) analysis revealed that the yellowish MT flavedo and pulp were primarily caused by the decrease in total carotenoid content. The total carotenoid content in MT flavedo was reduced by 75% (79.98 μg/g DW) compared with that in WT (318.40 μg/g DW), including approximately 84%, 80%, and 60% reductions in the contents of β-cryptoxanthin, violaxanthin and zeaxanthin, respectively. The total carotenoid content in MT pulp was 60% lower (10.09 μg/g DW) than that in WT pulp (26.61 μg/g DW), which was mainly due to a 70% and 30% decrease in the contents of β-cryptoxanthin and zeaxanthin, respectively. To explore the molecular mechanism underlying carotenoid variation in MT, RNA-seq analyses were performed on the flavedo and pulp of WT and MT at five developmental stages. The reduced expression of phytoene synthase (CrPSY) and β-carotene hydroxylase 1 (CrBCH1) in the flavedo and pulp of MT at the breaker stage might be the major cause of the reduction in carotenoids. Weighted gene co-expression network analysis (WGCNA) further identified 23 key transcription factors that are closely associated with carotenoid accumulation. This study demonstrated a comprehensive picture of the metabolic and transcriptional alterations of a unique yellowish citrus fruit mutant, which provides new insights into the molecular regulation of carotenoid accumulation in citrus fruit.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Carotenoids, a class of secondary metabolites widely distributed in nature, play an essential role in plant appearance and human health. In plants, carotenoids not only generate vibrant colors to attract insects or other animals to spread pollen and seeds but also protect the cell from damage caused by photooxidation stress during the photosynthesis progress (Holt et al. 2005; Cazzonelli 2011). Additionally, carotenoids are the precursors of volatile compounds, such as β-ionone, and the hormones abscisic acid (ABA) and strigolactones that involved in a variety of plant growth and development processes (Dudareva et al. 2006; Hou et al. 2016; Jia et al. 2018). Furthermore, as vitamin A precursors and antioxidants, carotenoids play a crucial role in human health. For example, β-cryptoxanthin, the precursor of vitamin A, and zeaxanthin, which have high antioxidant activity, effectively protect the eye from diseases and certain cancers (DellaPenna and Pogson 2006; Fiedor and Burda 2014; Kato 2016).
As one of the most important fruits in the world, citrus is enriched with carotenoids, which are crucial for both nutritional and product value. Moreover, the content and composition of carotenoids vary between cultivars and tissues (Nisar et al. 2015). In general, most mandarin cultivars mainly accumulate β-cryptoxanthin; orange cultivars are classified into the violaxanthin abundant category; and some cultivars are separate from oranges and mandarins due to their low carotenoid contents (Kato et al. 2004; Ikoma et al. 2016). Moreover, the carotenoid contents in the flavedo are generally higher than those in the pulp. The leaves are also abundant in carotenoids, especially those that respond to photosynthesis, such as lutein (Rodrigo et al. 2004). Therefore, citrus serves as a great model species for researching plant carotenoid metabolism owing to its diversified and complex carotenoid metabolism (Sun et al. 2018).
The carotenoid biosynthetic pathway has been well established in model plants and horticultural crops (Yuan et al. 2015; Wurtzel 2019). Exploiting the inherent genetic diversity of fruit colors in numerous species and spontaneous mutants with various carotenoid accumulation patterns is a crucial tactic for investigating carotenoid metabolism. For example, the tomato mutant r (yellow-flesh) with yellow-fleshed fruit showed a reduction in carotenoid accumulation as a consequence of the deletion in the phytoene synthase (PSY1) gene region (Fray and Grierson 1993; Gady et al. 2012). The tomato Beta (B) mutant is caused by increased β-carotene in the fruit, which changed from red to yellow (Ronen et al. 2000). Orange-fleshed watermelon usually containes lycopene and β/ε-carotene, whereas yellow-fleshed watermelon is predominantly enriched in violaxanthin and lutein (Lv et al. 2015). The yellow or orange fruit of chili peppers is a result of various levels of capsanthin and capsorubin accumulation (Rodriguez-Uribe et al. 2012).
Both natural and artificial selection of citrus mutants has generated many new citrus cultivars (Deng 2022). The exploitation of spontaneous mutants with elite pigment variation facilitates the development of citrus genetic breeding and enhances the understanding of carotenoid metabolism. For instance, Cara Cara, a spontaneous bud mutation of Navel orange, characterized by developing fruits with a pulp of bright red coloration due to the presence of lycopene (Alquezar et al. 2008). 'Zong Cheng', a brown flavedo navel orange mutant has a mutation of the STAY-GREEN (SGR) protein, which simultaneously regulates carotenoid biosynthesis and chlorophyll degradation (Zhu et al. 2021). Pinalate, a yellowish Navelate orange mutant, is due to a mutated allele of ζ-carotene isomerase (Z-ISO), resulting in elevated proportions of phytoene, phytofluene and ζ-carotene and reduced β, β-xanthophylls (Rodrigo et al. 2003, 2019). The spontaneous yellowish Ponkan (YP) mutant consistently shows reduced total carotenoid and xanthophylls in the pulp (Luo et al. 2015). Tamnaneunbong and Saebyeolbong were developed by embryo sac culture of Shiranui, both of which had orange pulp. However, the contents of β-cryptoxanthin and β-carotene are higher in Tamnaneunbong and the lowest in Saebyeolbong compared to Shiranui (Kim et al. 2021). Therefore, studying and utilizing natural variations effectively facilitates the understanding of the diverse and complex metabolic regulatory mechanisms of carotenoids in citrus and perhaps other fruit crops.
Shiranuhi tangor was bred by crossing Kiyomi (Citrus unshiu Marcov. × Citrus sinensis Osbeck) and Nakano No.3 Ponkan (Citrus reticulata Blanco) in Japan, which is a typical high-sugar and high-acid variety that completes coloring in early December. It was introduced to China from Japan in 1992 and mainly produced in Sichuan Province. ‘Jinlegan’ (MT) tangor, a natural mutant derived from a branch mutant of ‘Shiranuhi’ (WT) was found in 2007 in an orchard in Jintang County, Sichuan Province, China. Bright yellow instead of orange distinguished the peel of the MT from the WT. The contents of both total soluble solids (TSS) and titratable acid (TA) were higher in MT than in WT at the same stage, and the TA content decreased more slowly during the maturation. MT fruit pulp has a crispier texture than WT, indicating that MT superior fruit quality. Later, it was introduced to Danling County in Sichuan Province where the fruit matured a month later than the fruit of WT in mid to late April. Moreover, ‘Jinlegan’ tangor was highly productive, and its yield reached 45 metric tons per hectare. Its characteristics were stable and its comprehensive nature was excellent after years of observation and evaluation. MT was authorized and given new variety rights (UPOV) in June 2021 by the Ministry of Agriculture and Rural Affairs as a late-season citrus variety (Zhu et al. 2022). However, the molecular mechanism of color variation in this mutant has not been comprehensively studied.
In this study, HPLC was used to identify the composition and content of carotenoids in the flavedo, pulp and leaves of MT. In addition, RNA-seq was further used to understand the underlying molecular mechanism of carotenoid variation between the two cultivars, which provided novel insights into the molecular mechanism of carotenoid biosynthesis in citrus fruit.
Materials and methods
Plant materials
Both MT and WT were propagated by grafting in Chongqing, China. Fruit samples were collected in 2018 at different developmental stages from immature to fully colored stages: 120, 150, 170, 190, 210, and 240 DAF (days after flowering). Young leaves were sampled for further analysis. All samples were frozen in liquid nitrogen after collection and stored at -80°C for further analysis. Three biological replicates were analyzed.
Color measurement
Peel color parameters of L* (whiteness or brightness/darkness), a* (redness/greenness), and b* (yellowness/blueness) were measured at six evenly distributed equatorial sites of the fruit using a color analyzer (KONICA MINOLTA CR-400, Japan). The CCI [= 1000 × a/(L × b)] was calculated as described previously (Luo et al. 2015). Three biological replicates were analyzed.
Carotenoid extraction and HPLC analysis
For carotenoid analysis, the samples were powdered after lyophilization using a lyophilizer (FreeZone, LABCONCO, USA). Carotenoid extraction was performed according to previous studies (Zhu et al. 2022). The sample was dissolved in ethyl acetate for subsequent HPLC analysis. Carotenoid detection and analysis were performed using HPLC (e2695, Waters, USA), as previously described (Xu et al. 2006). Retention times and absorption spectra were compared to standards to determine the carotenoids. Peak areas for phytoene, phytofluene, and other compounds were measured at 286, 348, and 450 nm, respectively. The carotenoid levels were quantified using calibration curves prepared with appropriate standards. Three biological replicates were performed for each set of data.
RNA extraction and RT-qPCR
RNA extractions were performed from all samples using the TRIzol method according to previously reported. cDNA synthesis was synthesized using HiScript II Q RT SuperMix for qPCR (+ gDNA wiper; Vazyme). The RT-qPCR primers used in this study are listed in Supplemental Table S1. RT-qPCR products were quantified using a Roche LightCycler 480 system (Roche, https://www.roche.com). Three biological replicates were analyzed.
RNA-seq library construction and analysis
A total of 60 RNA-Seq libraries of flavedo (150, 170, 190, 210, and 240 DAF) and pulp (120, 150, 170, 190, and 210 DAF) samples at five developmental stages from WT and MT were constructed and sequenced. Three independent biological replicates were submitted to RNA-seq. The construction and sequencing of the RNA libraries were completed by Novogene Bioinformatics Technology (Tianjin) using Illumina HiSeq.
The raw reads were processed to remove adapter and low-quality sequences using fastp v0.22 (Chen et al. 2018). The resulting cleaned reads were mapped to the Citrus clementina v1.0 genome (GCF_000493195.1) using HISAT2 (Kim et al. 2019). Read counting was performed using featureCounts v2.0.2 (Liao et al. 2014). The RPKM (reads per kilobase per million mapped reads) was used to represent gene expression, which was calculated by StringTie v2.1.6 (Kovaka et al. 2019). Differentially expressed genes (DEGs) of each group analysis were identified using DESeq2 with a p value cutoff of 0.05 (Love et al. 2014). Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses were performed using the Cluster Profiler R package. Heatmaps were implemented using TBtools (Chen et al. 2020). Venn diagrams were implemented using Draw Venn Diagram.
Regulatory network construction by Weighted gene coexpression network analysis (WGCNA)
WGCNA was completed using the R package (Zhang and Horvath 2005). The network construction and module detection were performed using an unsigned type of topological overlap matrix (TOM), a power β of 26, a minimum module size of 20, and a branch merge cut height of 0.3.
Statistical analysis
The data were analyzed using GraphPad 8.0. Student’s t-test was used to identify whether the differences between the two datasets were statistically significant (* P < 0.05, ** P < 0.01).
Results
Characterization of MT with yellowish flavedo and pulp
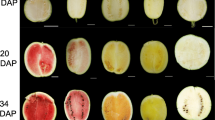
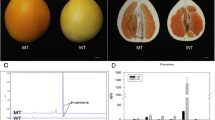
The fruits of the MT had distinctive bright yellow-colored flavedo and pulp that were distinguishable from the orange-colored fruits of the WT. To characterize the coloration variation of MT, fruits were sampled at 120, 150, 170, 190, 210, and 240 DAF. The flavedo phenotype differed after the breaker stage at 190 DAF (Fig. 1a), while the difference in pulp appeared at 150 DAF (Fig. 2a). After the breaker stage, the flavedo color of MT fruit turned bright yellow from green, and in WT fruit, the flavedo turned orange. A similar process occurred in the pulp of WT and MT. To provide quantitative results for describing color changes between MT and WT fruit, the L*, a*, b*, and CCI values were measured. The values of a* and the CCI index of MT flavedo were significantly lower than those of WT after the breaker stage (190 DAF). The value of a* changed from -0.46 at 190 DAF to 12.29 at 240 DAF in MT, while that of WT increased from 10.73 to 29.43 (Fig. 1d). The CCI index of MT increased from -0.14 (190 DAF) to 2.21 (240 DAF) compared to that of WT, which changed from 2.68 to 6.72 (Fig. 1b). However, in the pulp of MT, the values of a* and the CCI index were significantly lower than those in WT fruit at 150 DAF. The value of a* was 1.19 at 150 DAF and increased to 8.55 at 240 DAF, whereas the control increased from 4.06 to 16.04 (Fig. 2d). The CCI index of MT pulp increased from 0.80 (150 DAF) to 3.24 (240 DAF) compared to that of WT, which changed from 5.04 to 9.37 (Fig. 2b). These results indicated that the coloration of MT fruit was paler, resulting in a golden yellow color, whereas WT turned orange. In addition, it was found that the breaker stage of MT was delayed by 20 days in both the flavedo and pulp.
Phenotypes and color indices in the flavedo of WT and MT. a Phenotypes of WT (‘Shiranuhi’) and MT (‘Jinlegan’) flavedo at different developmental stages. b Citrus color index (CCI). c L* value. d a* value. e b* value. DAF, days after flowering. Three biological replicates were performed. Values were means ± SEs. Asterisks indicate statistically significant differences (Student’s t test, *, P < 0.05, **, P < 0.01)
Phenotypes and color indices in the pulp of WT and MT. a Phenotypes of WT (‘Shiranuhi’) and MT (‘Jinlegan’) pulp at different developmental stages. b Citrus color index (CCI). c L* value. d a* value. e b* value. DAF, days after flowering. Three biological replicates were performed. Values were means ± SEs. Asterisks indicate statistically significant differences (Student’s t test, **, P < 0.01)
Variations in carotenoid content in the fruits and leaves of MT and WT during fruit ripening
Five ripening stages in the flavedo (150, 170, 190, 210, and 240 DAF) and pulp (120, 150, 170, 190, and 210 DAF) of fruits were selected to evaluate the alterations in carotenoid composition and content between MT and WT.
In the flavedo, no significant difference was observed from 150 to 170 DAF. After the breaker stage (190 DAF), the total carotenoid contents in the flavedo in MT were significantly lower than those in WT, and detailed differences in individual carotenoids were also identified (Fig. 3a). At the breaker stage, the contents of β-cryptoxanthin and zeaxanthin in MT were 5.35 μg/g DW and 3.20 μg/g DW, respectively, which were reduced by approximately 75% compared to 21.70 μg/g DW and 14.40 μg/g DW in WT (Fig. 3b, c). Phytoene and phytofluene were reduced by approximately 60% and 70% respectively, compared to 3.31 and 2.22 μg/g DW in WT, while MT contained merely 1.35 and 0.60 μg/g DW (Fig. 3g, h). Furthermore, a slight reduction was also detected in the contents of violaxanthin (Fig. 3i). Nevertheless, no difference was observed in the contents of lutein and α-carotene (Fig. 3e, f). After the breaker stage, β-cryptoxanthin and zeaxanthin in MT were consistently lower than those in WT. At 240 DAF, the total amount of carotenoids in MT was 79.98 μg/g DW, while the amount in WT was more than threefold higher (318.40 μg/g DW). The content of β-cryptoxanthin in MT was 12.3 μg/g DW at 240 DAF, compared to a sixfold (77.23 μg/g DW) increase in WT. Comparable variations were also observed in the contents of zeaxanthin. Violaxanthin rapidly accumulated in WT at 240 DAF, but this process was not observed in MT. The content of β-carotene gradually decreased with fruit development. Interestingly, the content of β-carotene in MT was increased by twofold compared to that in WT at 190 DAF and 210 DAF, but no differences were observed at 240 DAF (Fig. 3d).
Variations in the carotenoid contents in the flavedo of WT and MT at five developmental stages. a Total carotenoids (μg/g DW). b β-Cryptoxanthin (μg/g DW). c Zeaxanthin (μg/g DW). d β-Carotene (μg/g DW). e Lutein (μg/g DW). f α-Carotene (μg/g DW). g Phytofluene (μg/g DW). h Phytoene (μg/g DW). i Violaxanthin (μg/g DW) of WT and MT fruit. WT, ‘Shiranuhi’. MT, ‘Jinlegan’. DAF, days after flowering. DW, dry weight. Three biological replicates were performed. Values were means ± SEs. Asterisks indicate statistically significant differences (Student’s t test, *, P < 0.05, **, P < 0.01)
In the pulp of both varieties, significant differences in carotenoid contents occurred after the breaker stage (150 DAF). Consistent with the results in flavedo, total carotenoids, particularly β-cryptoxanthin and zeaxanthin, decreased in MT compared to WT after the breaker stage (Fig. 4a). At the breaker stage, all carotenoids, including lutein, α-carotene, β-carotene, β-cryptoxanthin, and zeaxanthin, decreased in MT compared with WT; of these, β-cryptoxanthin decreased by approximately 70%, and the others decreased by 40% (Fig. 4b-f). The carotenoid content of the pulp in WT was predominantly composed of β-cryptoxanthin, which accounted for approximately 50% of the total carotenoid content and increased rapidly after the breaker stage. At 210 DAF, the content of β-cryptoxanthin in MT was 5.44 μg/g DW, while in WT, this amount was increased threefold (17.32 μg/g DW). The changing trend of zeaxanthin was consistent with that of β-cryptoxanthin. Furthermore, the total carotenoid content was less than half in MT (10.09 μg/g DW) than in WT (26.61 μg/g DW) at 210 DAF. Violaxanthin in the pulp was barely detectable and could be ignored.
Variations in the carotenoid contents in the pulp and young leaves of WT and MT. a Total carotenoids (μg/g DW). b β-Cryptoxanthin (μg/g DW). c Zeaxanthin (μg/g DW). d Lutein (μg/g DW). e β-Carotene (μg/g DW). f α-Carotene (μg g.−1 μg/g DW). g The carotenoid content in the young leaves of WT and MT. WT, ‘Shiranuhi’. MT, ‘Jinlegan’. DAF, days after flowering. DW, dry weight. Three biological replicates were performed. Values were means ± SEs. Asterisks indicate statistically significant differences (Student’s t test, **, P < 0.01)
However, no variations in the composition and content of carotenoids were detected in the young leaves between MT and WT, which primarily accumulated lutein, violaxanthin, luteoxanthin, α-carotene and β-carotene (Fig. 4g).
Transcriptomic differences in the flavedo and pulp between MT and WT during developmental stages
To further explore the underlying molecular mechanism of carotenoid variation in the flavedo and pulp, a transcriptome comparison was performed during five developmental stages in the flavedo (150, 170, 190, 210, and 240 DAF) and pulp (120, 150, 170, 190, and 210 DAF) between MT and WT. We mainly focused on the differentially expressed genes (DEGs) between MT and WT at the same developmental stage.
In the flavedo, a total of 18,698 DEGs (11,576 nonredundant) were identified, including 2406 DEGs at 150 DAF, 3283 DEGs at 170 DAF, 6029 DEGs at 190 DAF, 2368 DEGs at 210 DAF, and 4613 DEGs at 240 DAF (Fig. 5a, d). In the pulp, 19,602 DEGs (12,464 nonredundant) were identified, revealing 4244 DEGs at 120 DAF, 2896 DEGs at 150 DAF, 8460 DEGs at 170 DAF, 2058 DEGs at 190 DAF, and 1944 DEGs at 210 DAF (Fig. 5d, e).
Transcriptome analysis of the flavedo and pulp between WT and MT at five developmental stages. a Venn diagram showing the number of DEGs (differentially expressed genes) in the flavedo between the WT (‘Shiranuhi’) and MT (‘Jinlegan’) at five developmental stages. b Venn diagram showing the number of DEGs in the pulp between the WT and MT at five developmental stages. c The common and unique DEGs between 190 DAF (days after flowering) in the flavedo and 150 DAF in the pulp. d The number of upregulated and downregulated DEGs in the flavedo at five developmental stages. e The number of upregulated and downregulated DEGs in the pulp between the WT and MT at five developmental stages. f Twelve significantly enriched KEGG (Kyoto Encyclopedia of Genes and Genomes) pathways in DEGs of the flavedo. g Eight significantly enriched KEGG pathways in DEGs of the pulp
To understand the functions and metabolic pathways of DEGs between MT and WT, GO and KEGG pathways were next analyzed. DEGs in the ‘photosynthesis’, ‘carbohydrate metabolic process’, and ‘response to stimulus’ pathways were significantly enriched in the GO terms in the flavedo, and ‘cellular macromolecule biosynthetic process’, ‘intracellular organelle’, and ‘translation’ were significantly enriched in the pulp (Supplementary data Fig. S1a, b). KEGG analysis reveals additional information on the metabolic pathways of the identified DEGs. The DEGs were mainly involved in metabolic pathways associated with ‘photosynthesis’, ‘fatty acid biosynthesis’, and ‘carotenoid biosynthesis’ in the flavedo, while ‘ribosome’, ‘amino sugar and nucleotide sugar metabolism’ and ‘estrogen signaling pathway’ were the most frequently annotated pathways (Fig. 5f, g).
The difference in carotenoid accumulation between the flavedo and pulp of both cultivars appeared at their respective breaker stage, which was 190 DAF in the flavedo and 150 DAF in the pulp; therefore the DEGs of the two stages deserve further attention. Using MapMan to show the function of DEGs during this stage, it was found that the DEGs at 190 DAF in the flavedo contained 267 genes participating in secondary metabolism, of which 14 were related to carotenoid metabolism, 166 were related to protein modification, 318 were related to protein degradation, and 463 were annotated as transcription factors, including members of the bHLH, MYB, ERF, WRKY, bZIP, HB and MADS families. In addition, 368 genes were related to signaling, 114 genes were related to abiotic stress, 26 genes were related to ABA biosynthesis metabolism and 38 genes were related to ethylene biosynthesis metabolism (Supplementary Table S2).
In the DEGs of pulp at 150 DAF, 88 genes were related to secondary metabolism, 97 genes were related to protein modification, 183 genes were related to protein degradation and 296 DEGs were annotated as transcription factors, including members of the bHLH, MYB, ERF, WRKY, C2H2, bZIP, HB and MADS families. Additionally, several genes that respond to hormonal signals were also identified, of which 35 genes were related to IAA metabolism, 19 genes were related to ABA metabolism and 34 genes were related to ethylene metabolism (Supplementary Table S3).
An intersection analysis was performed between the DEGs of the flavedo at 190 DAF and pulp at 150 DAF, and 988 genes were identified to be differentially expressed (Fig. 5c), of which 599 genes showed the same expression trend in the two tissues. Four genes related to carotenoid metabolism, CrPSY, CrBCH1, carotenoid cleavage dioxygenase 4a (CrCCD4a), and 9-cis-epoxycarotenoid dioxygenase 5 (CrNCED5), were downregulated in MT. In addition, 48 transcription factors were included, of which 21 were downregulated and 27 were upregulated, such as MYB, WRKY, and ERF, which have been reported to be involved in the regulation of carotenoid metabolism, suggesting that these transcription factors might be involved in the regulation of carotenoid metabolism genes in the two cultivars (Supplementary Table S4).
Alterations in gene expression related to carotenoid metabolism in flavedo and pulp
To further investigate the underlying molecular mechanism of carotenoid metabolism between the two cultivars, heatmaps were constructed to display the expression of genes involved in carotenoid metabolism in the flavedo and pulp.
In the flavedo, most genes associated with carotenoid metabolism in DEGs, including CrPSY, phytoene desaturase (CrPDS), ζ-carotene desaturase (CrZDS), carotene isomerase (CrCRTISO), CrBCH1, zeaxanthin epoxidase (CrZEP), and CrNCED5, had lower expression levels in MT than in WT at 190 DAF, which was consistent with the observed decrease in carotenoid content. At 240 DAF, the expression levels of deoxy-D-xylulose 5-phosphate synthase (CrDXS), which is upstream of carotenoid genes, and lycopene β-cyclase 2 (CrLCYB2) and CrBCH1, which are carotenoid synthesizing genes, were significantly decreased in MT, while the expression level of carotenoid cleavage dioxygenase 1 (CrCCD1) was increased (Fig. 6a).
The expression of genes involved in carotenoid metabolism in the flavedo of WT and MT at five developmental stages. a Heatmaps showing the expression of genes involved in carotenoid metabolism at 150, 170, 190, 210, and 240 DAF (days after flowering) in the flavedo of WT (‘Shiranuhi’) and MT (‘Jinlegan’). b-e The relative expression levels of genes at 150, 170, 190, 210, and 240 DAF in the flavedo of WT and MT using RT-qPCR. CrPSY, phytoene synthase; CrPDS, phytoene desaturase; CrZDS, ζ-carotene desaturase; CrNCED5, 9-c-epoxycarotenoid dioxygenase 5. Three biological replicates were performed. Values were means ± SEs. Asterisks indicate statistically significant differences (Student’s t test, *, P < 0.05, **, P < 0.01)
In the pulp, genes related to carotenoid metabolism in DEGs, including CrPSY, CrBCH1, CrCCD4a, and CrNCED5, had lower expression levels in MT than in WT at 150 DAF. Notably, the expression of CrPSY, CrBCH1, and CrLCYB2 was higher in MT at 170 DAF, while the expression of genes related to ABA synthesis and carotenoid cleavage, such as CrZEP, CrCCD1a, CrCCD1b, and CrCCD4a was also increased (Fig. 7a). Several key genes related to carotenoid biosynthesis were further investigated by RT-qPCR analysis in WT and MT at the above five stages (Fig. 6b, c, d), and the results were consistent with the above transcriptome results.
The expression of genes involved in carotenoid metabolism in the pulp of WT and MT at five developmental stages. a Heatmaps describing the expression of genes involved in carotenoid metabolism at 120, 150, 170, 190, and 210 DAF (days after flowering) in the pulp of WT (‘Shiranuhi’) and MT (‘Jinlegan’). b-e The relative expression patterns of genes at 120, 150, 170, 190, and 20 DAF in the pulp of WT and MT using quantitative real time RT-qPCR. CrPSY, phytoene synthase; CrLCYB2, lycopene β-cyclase 2; CrBCH1, β-carotene hydroxylases 1; CrCCD1, carotenoid cleavage dioxygenase 1. Three biological replicates were performed. Values were means ± SEs. Asterisks indicate statistically significant differences (Student’s t test, **, P < 0.01)
Throughout fruit ripening, the expression levels of CrPSY, CrBCH, CrLCYB2, CrZEP, violaxanthin de-epoxidase (CrVED), CrCCD1b, and CrNCED3 increased progressively in both flavedo and pulp. However, the expression of CrDXS, 1-deoxy-D-xylulose 5-phosphate reductoisomerase (CrDXR), 1-hydroxy-2-methyl-2-(E)-butenyl 4-diphosphate synthase (CrHDS), hydroxymethylbutenyl 4-diphosphate reductase (CrHDR), and carotenoid synthesis genes carotene ε-ring hydroxylase 1 (CrLUT1) and lycopene ε-cyclase (CrLCYE), gradually decreased in the flavedo but increased in the pulp. Nevertheless, the expression of genes for carotenoid cleavage such as CrCCD4a and CrNCED5 showed the opposite expression pattern, gradually increasing in the flavedo but decreasing in the pulp (Figs. 6a, and 7a).
Identification of WGCNA modules associated with carotenoid metabolism
To characterize the candidate genes participating in the regulation of carotenoid metabolism in the pulp of the two cultivars, WGCNA was performed to evaluate the associations between DEGs and carotenoid contents and the values of CCI and a* of MT and WT. The results show that these DEGs are grouped into 14 modules, which are color coded (Fig. 8a).
Weighted gene co-expression network analysis (WGCNA) of differentially expressed genes (DEGs) identified from the pulp of WT and MT at five developmental stages. a Hierarchical cluster tree displaying fourteen modules of co-expressed genes. The lower panel shows modules in specified colors, such as ‘Red’, ‘Black’ and others. b Module-carotenoid weight correlations and corresponding P values (in parentheses). The left panel shows the fourteen modules. The color scale on the right shows module-trait correlation from -1 (blue) to 1 (red). WT, ‘Shiranuhi’. MT, ‘Jinlegan’
The MEblack and MEturquoise modules showed comparatively high positive correlation coefficients (Fig. 8b), which may contain candidate genes participating in carotenoid metabolism. The MEblack module comprised 199 genes, of which only one gene involved in carotenoid metabolism (CrLUT1) and 16 transcription factors. The MEturquoise module contained 3037 genes, including genes involved in carotenoid metabolism, such as CrPSY, CrPDS, CrLCYB2, CrBCH1, CrZEP, and CrVED. The MEturquoise module contained a large number of carotenoid metabolism genes, which had the highest correlation with β-cryptoxanthin; hence, this module was further analyzed. The MEturquoise module contained 117 genes related to protein modification, 198 genes related to protein degradation, 101 genes related to secondary metabolism, 172 genes related to signaling, 79 genes related to abiotic stress, and 235 transcription factors, including members of the bHLH, MYB, ERF, WRKY, C2H2, bZIP, HB and MADS families (Supplementary Table S5). Of these, 23 transcription factors were identified to be differentially expressed in the pulp both at 150 DAF and 170 DAF, 15 of which were expressed in a similar pattern as PSY and BCH1 and eight were expressed in the opposite pattern (Supplementary Table S6).
Discussion
The coloration of citrus, one of the main quality attributes that has a major impact on consumers’ choices, is predominantly due to the content and composition of carotenoids. Investigation of citrus pigmentation mutants contributes to a better comprehension of carotenoid metabolism (Yuan et al. 2015).
Yellowish coloration in the flavedo and pulp of the MT, was primarily associated with the reduction in the total carotenoid content. The composition of carotenoids was no differentce between WT and MT during all developmental stages. The yellowing of the flavedo in MT was attributed to the decrease in the content of β-cryptoxanthin, zeaxanthin, and violaxanthin, while in the pulp, it was primarily caused by the reduction in β-cryptoxanthin and zeaxanthin. To better evaluate the contribution of different carotenoids to citrus fruit coloration, we correlated the value of CCI and a* with the content of each carotenoid (Supplementary Table S7). The results showed that the content of β-cryptoxanthin had the highest correlation with CCI and a* in the flavedo, while violaxanthin and zeaxanthin had slightly lower correlation coefficients with CCI and a*. In the pulp, β-cryptoxanthin content correlated the highest with CCI and a* values. These results better indicated that the reduction in β-cryptoxanthin content had a significant contribution to the yellowish color of the MT fruit. However, young leaves from WT and MT displayed identical carotenoid types and contents, indicating that MT was a fruit-specific mutant. In addition, we observed that the total carotenoid content in the flavedo was more than tenfold higher than that in the pulp, as well as differences in the accumulation pattern. Substantial amounts of violaxanthin were detected in the flavedo, while they were almost undetectable in the pulp. The contents of lutein and β-carotene progressively decreased with the developmental stage of the fruit in the flavedo but increased in the pulp. In our study, the trends in the contents of β-cryptoxanthin and β-carotene were consistent with the results of the former research, but there were discrepancies in the other substances, probably due to the different detection and calculation methods (Wang et al. 2023). We calculated the specific carotenoid content based on the carotenoid standard curve, whereas they used the proportion of individual carotenoid components as measured by dividing the peak area of individual carotenoids by the sum of the peak areas of all carotenoids detected.
The phenotype of MT was distinct from that of other yellow citrus fruit mutants that had already been identified. In a previous study, YP demonstrated that the content of different carotenoids decreased in the flavedo and pulp of the fruit. The total carotenoids in the flavedo were decreased, including β-cryptoxanthin, violaxanthin, α-carotene, β-carotene, zeaxanthin, and lutein, whereas in the pulp, only the contents of β-cryptoxanthin, lutein, and β-carotene decreased (Luo et al. 2015). However, Pinalate, a yellowish mutant of Navelate orange was caused by the elevated proportions of upstream carotenoids, such as phytoene, phytofluene, and ζ-carotene, with reduced β, β-xanthophylls. Moreover, a decline in carotenoids was detected in its young leaves (Rodrigo et al. 2003, 2019). The alterations in carotenoids in the flavedo, pulp, and leaves of MT were distinct from those of other citrus fruit mutants, although all were yellow mutants. Thus, each mutant has a unique phenotype, and it would be of interest to investigate the mechanism of mutation in each mutant.
The accumulation pattern of carotenoids in the flavedo and pulp is regulated by key genes involved in the carotenoid metabolic pathway. PSY is the primary rate-determining enzyme of the carotenoid pathway, which determines the carotenoid content in the fruit during development. Silencing SlPSY1 reduced the accumulation of total carotenoids, resulting in tomato fruits with yellow flesh (Fantini et al. 2013). Silencing of β-carotene hydroxylase (Csβ-CHX) induced an elevation of β-carotene in the pulp, leading to the production of fruit with a golden coloration in orange (Pons et al. 2014). In this study, we observed that the expression levels of CrPSY and CrBCH1 were lower in the flavedo and pulp of MT at the breaking stage, which may be the reason for the lower carotenoid content in MT, especially β-cryptoxanthin. At 240 DAF, the decreased contents of β-cryptoxanthin, zeaxanthin, and violaxanthin in the flavedo may be due to the decreased expression of CrBCH1 and increased expression of CrCCD1, which caused the catabolism of carotenoids to outpace biosynthesis. This phenomenon was also observed in YP), where the content of most carotenoids decreased in the flavedo during coloring, while the expression of most carotenoid related genes increased. In the pulp of YP, the contents of total carotenoid contents decreased steadily; however, the dominantly expressed PSY1 showed threefold higher expression in YP pulp than in PK (Luo et al. 2015). In the pulp, β-cryptoxanthin and zeaxanthin were still declining in MT at 170 DAF; however, the expression levels of most genes involved in carotenoid synthesis and catabolism were higher than those of WT at this stage. It was possible that the pulp color of WT began to alter, along with numerous genes associated with carotenoid synthesis beginning to function.
Gene mutation is the most direct factor leading to the differences in gene expression, which results in the generation of phenotypes. For instance, Pinalate produces yellowish fruit as a result of the Z-ISO mutant allele (Rodrigo et al. 2019). The fruit with yellow flesh of tomato mutant r (yellow-flesh) is caused by the absence of PSY1, which reduces the accumulation of carotenoids (Fray and Grierson 1993; Gady et al. 2012). Due to an underlying lesion in the gene encoding the SGR protein, ‘Zong Cheng’ cannot degrade chlorophyll, resulting in the formation of brown fruit (Zhu et al. 2021). In the present study, we found that the changes in carotenoids in the flavedo and pulp were predominantly a reduction in the content of β-cryptoxanthin and zeaxanthin, while CrBCH1 was the essential enzyme controlling the conversion of β-carotene to β-cryptoxanthin and zeaxanthin. Therefore, we cloned the CDS and promoter region for CrBCH1 from MT and WT, and no sequence differences were observed between MT and WT, indicating that there might be alternative transcription regulators responsible for the suppression of CrBCH1 mRNAs in MT in addition to genetic variance.
Transcriptional regulation also plays an essential role in mutant variability. Previous studies have identified several transcription factors that are involved in the color variation of mutants. For example, the expression of CrMYB68 was found to be significantly higher in the flavedo of Green Ougan (MT) compared to Ougan (WT), which negatively regulated the expression of CrBCH2 and CrNCED5 and consequently resulted in reduced conversion of α- and β-branched carotenoids (Zhu et al. 2017). The R2R3-type MYB transcription factor MdMYB90-like regulates anthocyanin biosynthesis directly by activating anthocyanin biosynthesis genes, which is responsible for the appearance of enhanced peel color in the apple bud sport mutant Fuji (Sun et al. 2021). In this study, RNA-seq and WGCNA identified 23 transcription factors that may be involved in the regulation of carotenoid metabolism in the pulp, such as bZIP5 (ciclev10027584m), bHLH (ciclev10005233m), WRKY (ciclev10019820m) and MYB (ciclev10012089m). They belonged to the same module (MEturquoise) and displayed complementary or opposing patterns of expression with CrPSY1 and CrBCH1. Hence, the underlying regulatory mechanisms of these transcription factors are worth exploring in the future.
Conclusions
In summary, our study identified the variations in carotenoid contents in the flavedo and pulp of MT, providing insight into the transcriptional regulatory mechanism underlying fruit coloration, which can be valuable in improving fruit breeding strategies.
Availability of data and materials
The datasets generated and/or analysed during the current study available from the corresponding author on reasonable request.
References
Alquezar B, Rodrigo MJ, Zacarias L. Regulation of carotenoid biosynthesis during fruit maturation in the red-fleshed orange mutant Cara Cara. Phytochemistry. 2008;69(10):1997–2007. https://doi.org/10.1016/j.phytochem.2008.04.020.
Cazzonelli CI. Carotenoids in nature: insights from plants and beyond. Funct Plant Biolo. 2011;38(11):833–47. https://doi.org/10.1071/FP11192.
Chen S, Zhou Y, Chen Y, Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34(17):884–90. https://doi.org/10.1093/bioinformatics/bty560.
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant. 2020;13(8):1194–202. https://doi.org/10.1016/j.molp.2020.06.009.
DellaPenna D, Pogson BJ. Vitamin synthesis in plants: tocopherols and carotenoids. Annu Rev Plant Biol. 2006;57:711–38. https://doi.org/10.1146/annurev.arplant.56.032604.144301.
Deng X. A Review and Perspective for Citrus Breeding in China During the Last Six Decades. Acta Horticulturae Sinica. 2022;49(10):2063–74. https://doi.org/10.16420/j.issn.0513-353x.2021-0701.
Dudareva N, Negre F, Nagegowda DA, Orlova I. Plant volatiles: recent advances and future perspectives. CRC Crit Rev Plant Sci. 2006;25(5):417–40. https://doi.org/10.1080/07352680600899973.
Fantini E, Falcone G, Frusciante S, Giliberto L, Giuliano G. Dissection of tomato lycopene biosynthesis through Virus-induced gene silencing. Plant Physiol. 2013;163(2):986–98. https://doi.org/10.1104/pp.113.224733.
Fiedor J, Burda K. Potential role of carotenoids as antioxidants in human health and disease. Nutrients. 2014;6(2):466–88. https://doi.org/10.3390/nu6020466.
Fray RG, Grierson D. Identification and genetic analysis of normal and mutant phytoene synthase genes of tomato by sequencing, complementation and co-suppression. Plant Mol Biol. 1993;22(4):589–602. https://doi.org/10.1007/BF00047400.
Gady ALF, Vriezen WH, Van de Wal MHBJ, Huang P, Bovy AG, Visser RGF, et al. Induced point mutations in the phytoene synthase 1 gene cause differences in carotenoid content during tomato fruit ripening. Mol Breed. 2012;29(3):801–12. https://doi.org/10.1007/s11032-011-9591-9.
Holt NE, Zigmantas D, Valkunas L, Li XP, Niyogi KK, Fleming GR. Carotenoid cation formation and the regulation of photosynthetic light harvesting. Science. 2005;307(5708):433–6. https://doi.org/10.1126/science.1105833.
Hou X, Rivers J, Leon P, McQuinn RP, Pogson BJ. Synthesis and function of apocarotenoid signals in plants. Trends Plant Sci. 2016;21(9):792–803. https://doi.org/10.1016/j.tplants.2016.06.001.
Ikoma Y, Matsumoto H, Kato M. Diversity in the carotenoid profiles and the expression of genes related to carotenoid accumulation among citrus genotypes. Breed Sci. 2016;66(1):139–47. https://doi.org/10.1270/jsbbs.66.139.
Jia K-P, Baz L, Al-Babili S. From carotenoids to strigolactones. J Exp Bot. 2018;69(9):2189–204. https://doi.org/10.1093/jxb/erx476.
Kato M, Ikoma Y, Matsumoto H, Sugiura M, Hyodo H, Yano M. Accumulation of carotenoids and expression of carotenoid biosynthetic genes during maturation in citrus fruit. Plant Physiol. 2004;134(2):824–37. https://doi.org/10.1104/pp.103.031104.
Kato M. Mechanism of β-cryptoxanthin accumulation in citrus fruits. Acta Hortic.2016; 1135,1–10. https://doi.org/10.17660/ActaHortic.2016.1135.1.
Kim D, Paggi JM, Park C, Bennett C, Salzberg SL. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotechnol. 2019;37(8):907–15. https://doi.org/10.1038/s41587-019-0201-4.
Kim D-S, Lee S, Park SM, Yun SH, Gab H-S, Kim SS, et al. Comparative metabolomics analysis of citrus varieties. Foods. 2021; 10(11). https://doi.org/10.3390/foods10112826.
Kovaka S, Zimin AV, Pertea GM, Razaghi R, Salzberg SL, Pertea M. Transcriptome assembly from long-read RNA-seq alignments with StringTie2. Genome Biol. 2019; 20(1). https://doi.org/10.1186/s13059-019-1910-1.
Liao Y, Smyth GK, Shi W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014;30(7):923–30. https://doi.org/10.1093/bioinformatics/btt656.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014; 15(12). https://doi.org/10.1186/s13059-014-0550-8.
Luo T, Xu K, Luo Y, Chen J, Sheng L, Wang J, et al. Distinct carotenoid and flavonoid accumulation in a spontaneous mutant of Ponkan (Citrus reticulata Blanco) results in yellowish fruit and enhanced postharvest resistance. J Agric Food Chem. 2015;63(38):8601–14. https://doi.org/10.1021/acs.jafc.5b02807.
Lv P, Li N, Liu H, Gu H, Zhao W. Changes in carotenoid profiles and in the expression pattern of the genes in carotenoid metabolisms during fruit development and ripening in four watermelon cultivars. Food Chem. 2015;174:52–9. https://doi.org/10.1016/j.foodchem.2014.11.022.
Nisar N, Li L, Lu S, Khin NC, Pogson BJ. Carotenoid metabolism in plants. Mol Plant. 2015;8(1):68–82. https://doi.org/10.1016/j.molp.2014.12.007.
Pons E, Alquezar B, Rodriguez A, Martorell P, Genoves S, Ramon D, et al. Metabolic engineering of β-carotene in orange fruit increases its in vivo antioxidant properties. Plant Biotechnol J. 2014;12(1):17–27. https://doi.org/10.1111/pbi.12112.
Rodrigo MJ, Marcos JF, Alferez F, Mallent MD, Zacarias L. Characterization of Pinalate, a novel Citrus sinensis mutant with a fruit-specific alteration that results in yellow pigmentation and decreased ABA content. J Exp Bot. 2003;54(383):727–38. https://doi.org/10.1093/jxb/erg083.
Rodrigo MJ, Marcos JF, Zacarias L. Biochemical and molecular analysis of carotenoid biosynthesis in flavedo of Orange (Citrus sinensis L.) during fruit development and maturation. J Agric Food Chem. 2004;52(22):6724–31. https://doi.org/10.1021/jf049607f.
Rodrigo MJ, Lado J, Alos E, Alquezar B, Dery O, Hirschberg J, et al. A mutant allele of ζ-carotene isomerase (Z-ISO) is associated with the yellow pigmentation of the “Pinalate” sweet orange mutant and reveals new insights into its role in fruit carotenogenesis. BMC Plant Biol. 2019; 19(1). https://doi.org/10.1186/s12870-019-2078-2.
Rodriguez-Uribe L, Guzman I, Rajapakse W, Richins RD, O’Connell MA. Carotenoid accumulation in orange-pigmented Capsicum annuum fruit, regulated at multiple levels. J Exp Bot. 2012;63(1):517–26. https://doi.org/10.1093/jxb/err302.
Ronen G, Carmel-Goren L, Zamir D, Hirschberg J. An alternative pathway to β -carotene formation in plant chromoplasts discovered by Map-Based cloning of beta and old-gold color mutations in tomato. Proc Natl Acad Sci U S A. 2000;97(20):11102–7. https://doi.org/10.1073/pnas.190177497.
Sun T, Yuan H, Cao H, Yazdani M, Tadmor Y, Li L. Carotenoid metabolism in plants: the role of plastids. Mol Plant. 2018;11(1):58–74. https://doi.org/10.1016/j.molp.2017.09.010.
Sun C, Wang C, Zhang W, Liu S, Wang W, Yu X, et al. The R2R3-type MYB transcription factor MdMYB90-like is responsible for the enhanced skin color of an apple bud sport mutant. Hortic Res. 2021; 8(1). https://doi.org/10.1038/s41438-021-00590-3.
Wang X, Huang J, Yin Z, Xu K, Jiang D, Lin L, et al. Carotenoid components and their biosynthesis in a bud mutant of Shiranui mandarin (Citrus reticulata Blanco) with citrine flavedo. J Zhejiang Univ Sci B. 2023;24(1):94–100. https://doi.org/10.1631/jzus.B2200431.
Wurtzel ET. Changing form and function through carotenoids and synthetic biology. Plant Physiol. 2019;179(3):830–43. https://doi.org/10.1104/pp.18.01122.
Xu C, Fraser PD, Wang W, Bramley PM. Differences in the carotenoid content of ordinary citrus and lycopene-accumulating mutants. J Agric Food Chem. 2006;54(15):5474–81. https://doi.org/10.1021/jf060702t.
Yuan H, Zhang J, Nageswaran D, Li L. Carotenoid metabolism and regulation in horticultural crops. Hortic Res. 2015; 2. https://doi.org/10.1038/hortres.2015.36.
Zhang B, Horvath S. A general framework for weighted gene co-expression network analysis. Stat Appl Genet Mol Biol. 2005; 4. https://doi.org/10.2202/1544-6115.1128.
Zhu F, Luo T, Liu C, Wang Y, Yang H, Yang W, et al. An R2R3-MYB transcription factor represses the transformation of α- and β-branch carotenoids by negatively regulating expression of CrBCH2 and CrNCED5 in flavedo of Citrus reticulate. New Phytol. 2017;216(1):178–92. https://doi.org/10.1111/nph.14684.
Zhu K, Zheng X, Ye J, Huang Y, Chen H, Mei X, et al. Regulation of carotenoid and chlorophyll pools in hesperidia, anatomically unique fruits found only in Citrus. Plant Physiol. 2021;187(2):829–45. https://doi.org/10.1093/plphys/kiab291.
Zhu S, Wen R, Wang Y, Zeng Y. A new very late ripening citrus variety ‘Jinlegan.’ Acta Horticulturae Sinica. 2022;49(S1)43–44. https://doi.org/10.16420/j.issn.0513-353x.2022-0027.
Zhu K, Chen H, Zhang Y, Liu Y, Zheng X, Xu J, et al. Chapter Six - Carotenoid extraction, detection, and analysis in citrus, in: Wurtzel ET, editors. Methods in Enzymology, Carotenoids: Carotenoid and Apocarotenoid Analysis. Holland: Elsevier Inc; 2022. p. 179–212. https://doi.org/10.1016/bs.mie.2022.01.006.
Acknowledgements
Not applicable.
Funding
This work was supported by the National Key Research and Development Program of China (2022YFF1003100) and the National Natural Science Foundation of China (No. 31930095).
Author information
Authors and Affiliations
Contributions
XD conceived and coordinated this project; HC performed the experiments with contributions from HJ, SZ, and KZ; HC wrote the manuscript, under the supervision of and with contributions from XD and JY. The authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests. Xiuxin Deng is the Editor-in-Chief of Horticulture Advances and was not involved in the journal’s review or decisions related to this manuscript.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Figure S1.
GO terms enriched in DEGs of the flavedo and pulp.
Additional file 2:
Table S1. Primers used in this study.
Additional file 3: Table S2.
Functional annotation of DEGs in the flavedo of WT and MT at 190 DAF.
Additional file 4: Table S3.
Functional annotation of DEGs in the pulp of WT and MT at 150 DAF.
Additional file 5: Table S4.
Common transcription factors in the DEGs between the flavedo at 190 DAF and pulp at 150 DAF.
Additional file 6: Table S5.
The transcription factor genes in the WGCNA modules were highly ‘black’ and ‘turquoise’.
Additional file 7: Table S6.
The 23 transcription factors were highly associated with carotenoid content in the pulp.
Additional file 8: Table S7.
Correlation coefficients of CCI and a with the content of each carotenoid type in the flavedo and pulp.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, H., Ji, H., Zhu, S. et al. Carotenoid and transcriptome profiles of a novel citrus cultivar ‘Jinlegan’ reveal mechanisms of yellowish fruit formation. HORTIC. ADV. 1, 5 (2023). https://doi.org/10.1007/s44281-023-00005-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44281-023-00005-4