Abstract
In pears, the presence of stone cells adversely affects fruit quality. Pectin methylesterase (PME) plays various roles in plant biology, including lignin biosynthesis. However, only a limited fraction has been functionally characterized, and the distribution and function of PME in many Rosaceae trees remain unexplored. In this study, we identified 396 putative PME family candidate genes, with 81 in Pyrus bretschneideri, 92 in Malus domestica, 62 in Fragaria vesca, 65 in Prunus mume, 15 in Pyrus communis, and 81 in Pyrus pyrifolia. Leveraging insights from model plants, we categorized PME family genes into four groups. Additionally, the evolution of the PME gene family was shaped by various gene duplication events, primarily dispersed duplication, influenced by purifying selection. A specific gene, Pbr031522.1, designated PbPME35, emerged as a candidate associated with lignin biosynthesis in pear fruits, supported by RNA-seq data. The role of PbPME35 in repressing lignification was validated through its overexpression in pear callus and Arabidopsis. Overall, our findings highlight the ability of PbPME35 to reduce lignin content in pear fruit by downregulating the expression levels of lignin biosynthesis genes. These findings provide new insights into the characteristics of PME genes and their role in regulating lignification in pear fruits.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The pear, belonging to the Rosaceae family, has a rich history of global cultivation (Wu et al. 2013). Stone cells in pears significantly affect fruit quality due to their lignified cell walls (Gong et al. 2020a), characterized by sclerenchyma cells with hardened secondary cell walls (SCWs) due to lignin deposition (Tao et al. 2009).
Plant cell walls are complex structures mainly composed of lignin, cellulose, hemicellulose and pectin (Fry 2004). Pectin methylesterase (PME, EC 3.1.1.11) is an enzyme modifying pectin by removing methyl groups, leaving carboxyl groups on pectin chains (Röckel et al. 2008). PME can be classified based on the presence or absence of the PME inhibitor (PMEI) domain. Most PME proteins have both PME and PMEI domains, though some have only the PME domain (Jolie et al. 2010). Moreover, PME influences the hardness of plant cell walls, with increased activity decreasing hardness and vice versa (Pelletier et al. 2010; Levesque-Tremblay et al. 2015; Wen et al. 2020). Furthermore, PME plays a crucial role in pectin modifications during the deposition of secondary walls in xylem cells (Hertzberg et al. 2001; Geisler-Lee et al. 2006). The significance of these modifications lies in the role of de-esterified homogalacturonan, a type of pectic polysaccharide domain, in xylem lignification (Pelloux et al. 2007). When bridged by Ca2+, de-esterified homogalacturonan binds peroxidases, potentially initiating lignin polymerization (Carpin et al. 2001; Dunand et al. 2002). Supporting this, colocalization of PME, de-esterified homogalacturonan, peroxidase, and lignification initiation occurs at cell junctions in woody tissues (Guglielmino et al. 1997; Wi et al. 2005). Furthermore, the interaction between pectin and lignin monomers influences lignin polymerization (Lairez et al. 2005). In summary, these insights provide novel perspectives on the roles of PME and pectin in lignification.
PMEs are associated with different developmental processes, including lignin biosynthesis (Sexton et al. 2012), fruit ripening and softening (Wen et al. 2020), hypocotyl growth (Pelletier et al. 2010), embryo development, seed germination (Levesque-Tremblay et al. 2015), and pollen tube elongation (Tang et al. 2020). Eucalyptus pilularis pectin methylesterase 7 (EpPME7) can negatively regulate cellulose and lignin contents (Sexton et al. 2012). Studies also indicate a close relationship between pectin metabolism and post-harvest loquat fruit cellular lignification (Huang et al. 2022). In strawberries and tomatoes, specific PME gene family members play essential roles in regulating fruit firmness (Wen et al. 2020; Xue et al. 2020). However, the relationships between lignification, PME, and the role of PME in stone cell formation in pear fruits remain unclear. Elucidating this relationship could offer a novel perspective on stone cell formation mechanisms.
Functional characterization of PMEs has been limited to a subset, including those in Arabidopsis (Louvet et al. 2006), tomato (Wen et al. 2020), and strawberry (Xue et al. 2020). The distribution of PMEs in pears and many other Rosaceae species is not fully understood. Additionally, limited research on the pear PME family leaves the relationship between these genes and pear fruit lignification unclear. This study identified PME family genes from five Rosaceae species, evaluating their gene structure, duplication modes, location, collinearity, and the ratio of the nonsynonymous substitutions per nonsynonymous site (Ka) to the synonymous substitutions per synonymous site (Ks) (Ka/Ks values). Transcriptome analysis revealed that PbPME35 is correlated with stone cell formation and lignin biosynthesis, with its biological function of suppressing lignification confirmed through homologous and allogenic transformation. These findings provide new insights into the mechanism of lignification in pear fruit stone cell formation.
Materials and methods
Plant materials
Pear fruit samples from two cultivars, Pyrus bretschneideri ‘Dangshan Su’ (DS) and Pyrus pyrifolia ‘Cuiguan’ (CG), were collected at three developmental stages (15, 35, and 55 days after flowering [DAF]) from Gaoyou Orchard in Yangzhou, Jiangsu, China. Ten trees (five for each cultivar), grown under healthy and uniform conditions without disease or insect infection, were randomly selected. Upon returning to the laboratory, the pear fruit samples were immediately frozen in liquid nitrogen and stored in an ultralow-temperature refrigerator at –80°C for subsequent experiments.
Identification and phylogenetic analysis of PME genes
PME genes from different species, including Arabidopsis thaliana, Pyrus bretschneideri, Pyrus pyrifolia, Malus domestica, Fragaria vesca, Prunus mume, and Pyrus communis, were identified using the methods outlined by Qiao et al. (2015). An evolutionary tree was constructed using Orthofinder (v2.2.7), MAFFT (v7.475), TrimAI (v1.4.1), Raxml (v8.2.12), Astral (v5.7.8), and EasySpeciesTree (v1.0). The TimeTree database (http://timetree.org/home) was referenced for mapping species divergence time (Fig. S1a). Genome files from various databases reported previously were collected (Table 1). HMMER (v3.3.2) and BLAST (v2.11.0) were used to identify candidate genes of each species. The Hidden Markov Model profile (PF01095) was obtained from the Pfam database, and CDD/SPARCLE was used to confirm PME domain accuracy in the candidate proteins (Lu et al. 2020). Multiple sequence alignments were performed using MAFFT (v7.475), and maximum likelihood (ML) phylogenetic trees were constructed using IQ-TREE (v2.1.2) (Katoh et al. 2019; Minh et al. 2020). Nomenclature guidelines and details for candidate PME genes from five Rosaceae species are provided in Table S1.
Chromosomal location and gene structure analysis of PME genes
Chromosomal location information for PME genes was extracted using genomic annotation data from various species, with details presented in Table S1. MEME (v5.0.5) was used to identify conserved protein motifs (Bailey et al. 2010) with the maximum value set to 10. The motif length was adjusted within the range of 6 to 50 during motif analysis.
Gene duplication modes, collinearity analysis, and Ka/Ks calculation
To ascertain gene duplication modes in the five Rosaceae species, we employed DupGen Finder, a specialized software package (Qiao et al. 2019). TBtools (v1.108) was used to determine the localization and synteny of the PME genes and calculate the Ka/Ks value (Chen et al. 2020). MCscan (Python version) was used to analyze the collinearity relationships among the five Rosaceae species (Tang et al. 2008).
Expression pattern of PME genes in pear
RNA-seq data from two regions, one rich in stone cells content and the other with few stone cells, were obtained for three stages (15, 35, and 55 DAF) of CG and DS fruits from the NCBI bioProject PRJNA825067. The expression patterns of PbPME genes were extracted from these data, and the results are shown in Table S6.
Arabidopsis plant and pear fruit callus transformation
Following the methods outlined by Gong et al. (2022), PbPME35 was overexpressed in the Arabidopsis Columbia ecotype through the floral dip assay. T1-generation seeds were screened on media supplemented with sucrose (30 g/L), agar (0.75%), hygromycin (20 mg/L), timentin (100 mg/L), and carbenicillin (100 mg/L). Transgenic lines were identified using PCR. Positive transgenic Arabidopsis plants from the T3 generation and wild-type (WT) plants were grown on MS media without growth regulators for functional verification.
For pear callus transformation, callus tissues were added to LB cultures under aseptic conditions and placed on a shaker at 120 r·min−1 for 30 min. Infected calli were evenly spread on acetosyringone (AS) plates for 24 h after draining the liquid. Subsequently, the calli were transferred to hygromycin-containing MS solid media to screen for transgenic calli, which were then cultured at 24°C under continuous dark conditions (Bai et al. 2019).
RT-qPCR analysis
To validate the expression levels of the selected PbPME genes, RNA was extracted from pear pulp samples using a Fuji total RNA purification kit. The extracted RNA was then reverse transcribed using a TransGen Biotech reverse transcription kit. Primers were designed using Primer (v5.0) software for specificity (Table S8). The PCR mixture comprised premix (water:fluorescent = 1:2; 15 µL), template cDNA (200 ng), and forward (2.5 µL) and reverse (2.5 µL) primers. The PCR procedure involved preincubation for 5 min at 95°C, followed by 55 cycles of denaturation at 95°C for 3 s, annealing at 60°C for 10 s, and extension at 72°C for 30 s. P. bretschneideri ubiquitin (PbUBQ) served as the internal reference gene, and the relative gene expression level was calculated using the 2−ΔΔct method.
Subcellular localization of PbPME35
The transient transformation of tobacco leaves was conducted following the protocol from a previous study (Yang et al. 2000). An Agrobacterium suspension transfected with 35S:PbPME35-GFP, cultured in LB liquid media supplemented with kanamycin and rifampicin, was injected into the lower surface of 5-week-old Nicotiana benthamiana leaves. After 48 h of growth, live cell imaging of the transformed leaves was conducted using an inverted confocal microscope (Zeiss LSM 780, Carl Zeiss AG, Oberkochen, Germany).
Lignin and stone cell content determination
Lignin content was assessed using the acetyl bromide method outlined by Tao et al. (2009), utilizing a standard sample curve (Sigma-Aldrich, St. Louis, MO, USA) for calculation. Stone cell content was quantified using the Frozen-HCL method outlined by Gong et al. (2020).
Anatomical structure observation
To examine the anatomical structures of Arabidopsis stems and pear pulp, paraffin sectioning was performed (Zhang et al. 2020). Sections were cut using a microtome (Leica, Biosystems, Wetzlar, Germany), stained with safranin and fast green or toluidine blue, and observed under a microscope (Zeiss LSM 780).
Statistical analysis
The data were subjected to analysis of variance using IBM SPSS (v19.0) for significance assessment. Microsoft Office Excel 2013 was used for data processing. Plots were generated using Origin (v9.0), RStudio, Hiplot (https://hiplot.com.cn), and TBtools (Chen et al. 2020).
Results
Identification, phylogenetic analysis, and conserved motif patterns of PME family members
In this study, we identified 396 putative PME family candidate genes distributed across various species, including 81 in P. bretschneideri, 92 in M. domestica, 62 in F. vesca, 65 in P. mume, 15 in P. communis, and 81 in P. pyrifolia (Table S1). Additionally, 66 PME family genes were identified in A. thaliana, consistent with the TAIR database and previous research (Louvet et al. 2006).
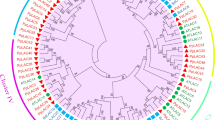
A phylogenetic tree, constructed using PME proteins from P. bretschneideri, P. pyrifolia, and A. thaliana through the ML method, revealed four clades (I, II, III, and IV) indicating an evolutionary divergence approximately 3.54 million years ago (MYA) (Fig. 1a, Fig. S1a). Further, a phylogenetic tree encompassing PME genes from Arabidopsis, along with those from P. bretschneideri, M. domestica, F. vesca, P. mume, and P. communis, showcased the same four clades (I, II, III, and IV), indicating the conserved evolutionary relationships within the PME family (Fig. S1b).
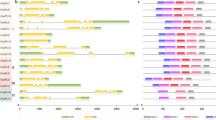
Phylogenetic and conserved motif analyses of Pectin methylesterase (PME) family members. (a) Phylogenetic analysis depicting the relationship among PMEs from Arabidopsis, Pyrus bretschneideri, and P. pyrifolia. (b) Phylogeny, conserved motif distribution, and coding sequence (CDS)–untranslated region (UTR) structure of PME genes in pear
Conserved motif analysis revealed variability in motif composition among P. bretschneideri PME genes, with motif 6 being predominant in most clades, except for PbPME17 in clade I and PbPME74 in clade IV (Fig. 1b, Fig, S2a). Exon–intron analysis revealed notable patterns, such as 64.10% of clade I PbPME genes having only two exons, consistent with similar trends observed in clades II and III. There were seven PbPME genes with two exons in clades II and III, accounting for 77.78% and 87.50%, respectively. Clade IV exhibited a higher proportion (19 PbPME genes) with more than three exons, with only two genes having two exons (Fig. 1b).
Gene duplication, evolution, and synteny analysis of PME family
Using DupGen Finder to evaluate gene duplication modes within the PME family, we identified five modes—whole-genome duplication (WGD), tandem duplication (TD), proximal duplication (PD), transpose duplication (TRD), and dispersed duplication (DSD) as potential drivers (Qiao et al. 2019). All PME genes across the studied species showed involvement in WGD, TD, PD, TRD, or DSD. DSD played a substantial role in evolution, with proportions of 51.41%, 56.49%, 65.12%, 75.68%, and 57.89% in P. bretschneideri, M. domestica, F. vesca, P. mume, and P. communis, respectively. However, the second significant duplication mode differed among these species, with WGD being the second most significant mode in P. bretschneideri and M. domestica, TRD in F. vesca and P. communis, and PD in P. mume (Fig. 2a, Table S2).
Gene-duplication modes, Ka/Ks values, gene locations, and collinearity analysis of the Pectin methylesterase (PME) gene family. (a) Various modes of gene duplication, with the x-axis representing the number of duplicated gene pairs and the y-axis denoting the species. (b) Ka/Ks values across five Rosaceae species. (c) Ka/Ks values corresponding to different duplication modes in Pyrus bretschneideri. (d) Gene locations in P. bretschneideri and collinearity analysis, displaying genes distributed across different chromosomes. Red lines represent syntenic gene pairs. (e) Collinearity analysis comparing P. bretschneideri with four Rosaceae species. DSD, dispersed duplication; TRD, transpose duplication; PD, proximal duplication; TD, tandem duplication; WGD, whole genome duplication; Ka, nonsynonymous substitutions per nonsynonymous site; Ks, synonymous substitutions per synonymous site
To evaluate selection pressure, we calculated Ka/Ks values for PME genes in the five species using TBtools (Table S3). Results indicated Ka/Ks values below 1 for almost all gene pairs in P. bretschneideri and all gene pairs in other species, signifying purifying selection. However, two gene pairs (PbPME67–PbPME68 and PbPME20–PbPME21) in P. bretschneideri underwent positive selection (Ka/Ks > 1; Fig. 2b). These gene pairs evolved through PD (PbPME67–PbPME68) and DSD (PbPME20–PbPME21) (Table S1, Fig. 2c and d).
In P. bretschneideri, 62 PbPME genes were unevenly distributed across 16 chromosomes, except for chromosome 14, and 19 PbPME genes were located on unanchored scaffolds (Fig. 2d). Chromosome 2 had the most PME genes, accounting for 17.28% of all PME genes (Fig. 2d and Table S1). Additionally, intragenomic synteny blocks identified 69 syntenic gene pairs, with P. bretschneideri and M. domestica having the highest number (37 and 26 pairs, respectively). F. vesca, P. mume, and P. communis had fewer syntenic pairs (3, 1, and 2 pairs, respectively; Fig. 2d and Table S4). Furthermore, a comparative syntenic graph revealed orthologous relationships between PME genes in P. bretschneideri and other species (M. domestica, F. vesca, P. mume, and P. communis), showing 81 syntenic gene pairs with M. domestica, 20 with P. communis, 32 with F. vesca, and 45 with P. mume (Fig. 2e and Table S5).
Expression analysis of PME genes
Transcriptome data from two pear varieties with varying stone cell contents identified 19 PbPME genes expressed in pear fruits (Fig. 3a and Table S6). RT-qPCR analysis was performed to validate expression patterns for four randomly selected genes (PbPME13, PbPME29, PbPME35, and PbPME55), highly consistent with RNA-seq data (Fig. 3b). Critical stone cell formation periods in pear fruits were identified as 15, 35, and 55 DAF. Additionally, although stone cells are distributed near the fruit cores, they can be categorized into different regions with few (L) or rich (R) stone cells (Gong et al. 2022).
Expression analysis of Pectin methylesterase (PME) genes in pear. (a) Heatmap analysis presenting the expression patterns of PME genes. (b) Correlation analysis between the expression levels and RNA-seq data of four PME genes. (c) Gene expression clustering trend analysis of PME genes at various fruit development stages. (d) Candidate PME genes were screened through gene expression trend analysis and Venn analysis. CG and DS indicate different cultivars (CG, Pyrus pyrifolia ‘Cuiguan’; DS, Pyrus bretschneideri ‘Dangshan Su’). L and R indicate different fruit pulp parts (L, Regions with few stone cells in pear fruits; R, Regions with rich stone cells in pear fruits). The numbers 15, 35, and 55 indicate different pear fruit development stages concerning the number of days after flowering (DAF). PbPME, Pyrus bretschneideri Pectin methylesterase
To ensure consistency of PbPME gene expression in cultivars or stone cell regions, we categorized expression levels of the PbPME genes extracted from RNA-seq data into four groups: regions in CG with few stone cells (CG_L), regions in CG with rich stone cells (CG_R), and the same for DS (DS_L) and (DS_R) (Fig. 3c). Our results revealed two distinct expression patterns (Trend 1 and Trend 2) shared across all groups, each containing two common genes (Trend 1: PbPME34 and PbPME35; Trend 2: PbPME20 and PbPME75). In all groups, PbPME34 and PbPME35 exhibited an initial increase followed by a decrease, whereas PbPME20 and PbPME75 showed a decrease, followed by stabilization. RT-qPCR data for these four genes confirmed these patterns (Fig. 3c and d). Notably, PbPME34 and PbPME35 exhibited high expression levels in RNA-seq data, whereas PbPME20 and PbPME75 exhibited low expression (Fig. 3d). Moreover, the gene pair PbPME34–PbPME35, originating from WGD (Table S2) and displaying synteny (Table S4) with an identity of up to 99.19% (Table S7), led us to select PbPME35 as a candidate gene influencing lignin biosynthesis due to its high homology with PbPME34.
Analysis of phenotypic traits, PbPME35 expression in mature pear fruit and subcellular localization of PbPME35
Utilizing the phloroglucinol-HCl method to stain cross sections of mature fruits revealed a higher concentration of stone cells near the fruit core. Further observation of stone cell clusters in different pear fruit parts through paraffin sectioning indicated larger stone cell clusters and greater stone cell density near the core regions compared to those near the peel (Fig. 4a). Consequently, we categorized pear fruits into two regions: the stone cell enrichment area (near the core) and the non-enrichment area (near the peel). Results from fruit firmness, stone cell content, and lignin content determination demonstrated higher expression of these phenotypic traits in the stone cell enrichment area than in the non-enrichment area (Fig. 4b). However, the expression level of PbPME35 in the stone cell enrichment area was lower than that in non-enrichment area (Fig. 4c). Furthermore, the expression levels of lignin biosynthetic genes (P. bretschneideri 4-coumarate: CoA ligase 1 [Pb4CL1], P. bretschneideri p-coumarate 3-hydroxylase 1 [PbC3H1], PbC3H2, P. bretschneideri cinnamyl alcohol dehydrogenase 2 [PbCAD2], P. bretschneideri Caffeoyl coenzyme A O-methyltransferase [PbCCOAOMT], P. bretschneideri cinnamoyl-CoA reductase 1 [PbCCR1], P. bretschneideri caffeoyl shikimate esterase 1 [PbCSE1], P. bretschneideri hydroxycinnamoyl CoA:shikimate/quinate hydroxycinnamoyl transferase 17 [PbHCT17], PbPAL1: P. bretschneideri phenylalanine ammonia-lyase 1 [PbPAL1], P. bretschneideri peroxidase [PbPOD1], and PbPOD2) were significantly higher in regions with greater stone cell counts (Fig. 4d). Additionally, correlation analysis was used to evaluate the relationships between PbPME35 and other stone cell-related traits (or genes). The expression of PbPME35 was significantly negatively correlated with stone cell-related traits (including stone cell content, lignin content, and fruit firmness) and stone cell-related genes (all mentioned lignin biosynthetic genes), highlighting the inhibitory role of PbPME35 in stone cell formation (Fig. 4e).
Phenotypic trait analysis and Pyrus bretschneideri Pectin methylesterase 35 (PbPME35) expression levels in mature pear fruits. (a) Phloroglucinol staining and cross-sectional staining of mature pear fruit. (b) Analysis of fruit firmness, stone cell content, and lignin content in different regions of mature pear fruit. (c) PbPME35 expression levels in different regions of mature pear fruit. (d) Relative expression patterns of lignin biosynthetic genes in pear fruits enriched (or not) with stone cells. (e) Correlation analysis among phenotypic traits, expression levels of lignin biosynthetic genes, and PbPME35 expression levels. (f) Subcellular localization of PbPME35. * P < 0.05, ** P < 0.01. Pb4CL1, P. bretschneideri 4-coumarate: CoA ligase 1; PbC3H1, P. bretschneideri p-coumarate 3-hydroxylase 1; PbCAD2, P. bretschneideri cinnamyl alcohol dehydrogenase 2; PbCCOAOMT, P. bretschneideri Caffeoyl coenzyme A O-methyltransferase; PbCCR1, P. bretschneideri cinnamoyl-CoA reductase 1; PbCSE1, P. bretschneideri caffeoyl shikimate esterase 1; PbHCT17, P. bretschneideri hydroxycinnamoyl CoA:shikimate/quinate hydroxycinnamoyl transferase 17; PbPAL1, P. bretschneideri phenylalanine ammonia-lyase 1; PbPOD1, P. bretschneideri peroxidase 1; GFP, green fluorescence
To investigate subcellular localization, we employed Agrobacterium-mediated transformation, introducing the 35S:PbPME35-GFP fusion construct (treatment) and 35S:GFP (control) into tobacco leaves. While the positive control expressed a green fluorescence signal in the cell membrane and nucleus, leaves injected with 35S:PbPME35-GFP expressed this signal solely in the cell membrane (Fig. 4f).
Overexpression of PbPME35 in Arabidopsis
Arabidopsis thaliana (Col-0) was infected with inflorescences to generate overexpression (OE) lines of PbPME35 (Fig. 5a and b). Three independent transgenic lines, exhibiting significantly higher PbPME35 expression levels, were selected for further analysis (Fig. 5c). After 10 d of growth, both root and hypocotyl lengths were significantly extended in the transgenic plants compared to the WT plants (Fig. 5a, d, and e). At 5 weeks of age, the transgenic plants displayed increased growth rates and taller heights compared to the controls (Fig. 5b). Additionally, lignin content in the stems of the transgenic Arabidopsis plants significantly decreased after PbPME35 OE (Fig. 5f). Paraffin sectioning further revealed less pronounced phenotypic traits in the interfascicular fibers and xylem cells of transgenic plants (Fig. 5i). Furthermore, RT-qPCR quantified the downregulation of six lignin biosynthetic genes (4CL1, CAD4, CAD9, CCOAOMT, ferulate 5-hydroxylase 1 [F5H1], and hydroxycinnamoyl CoA:shikimate/quinate hydroxycinnamoyl transferase [HCT]) in the transgenic plants (Fig. 5g). Additionally, six transcription factors involved in lignin biosynthesis (NST1, MYB85, MYB103, MYB58, MYB83, and MYB46) were also downregulated in the transgenic plants (Fig. 5h), indicating the negative regulatory effect of PbPME35 on lignin biosynthesis.
Overexpression (OE) of Pyrus bretschneideri Pectin methylesterase 35 (PbPME35) in Arabidopsis. (a) Ten-day-old plants. (b) Five-week-old plants. (c) Expression patterns of PbPME35. (d) Root length. (e) Hypocotyl length. (f) Lignin content. (g) Expression patterns of lignin biosynthetic genes. (h) Expression patterns of transcription factors regulating lignin biosynthesis and secondary wall synthesis. (i) Stem anatomy of wild-type (WT) and transgenic Arabidopsis plants. Toluidine blue-stained sections illustrate variations in cell walls in Arabidopsis plants overexpressing PbPME35. Black arrows highlight the lignification of the xylem and interfascicular fiber cells. Different lowercase letters in a column indicate significant differences at the 5% level. * P < 0.05, ** P < 0.01
Overexpression of PbPME35 in pear fruit callus
We generated a homologous stable OE system in pear fruit callus to elucidate the biological function of PbPME35. The expression level of PbPME35 significantly increased in OE calli compared to control calli (Fig. 6c). Phloroglucinol-HCl staining revealed a lighter red stain in control, whereas the transgenic line exhibited minimal red staining, suggesting a reduction in lignin accumulation upon PbPME35 OE (Fig. 6a). This observation aligned with the lignin determination results (Fig. 6b). Additionally, the expression levels of five lignin biosynthetic genes (Pb4CL1, PbC3H1, PbCCR1, PbPAL1, and PbPOD1) were significantly downregulated in the OE callus (Fig. 6d). These findings provide an explanation for the decreased lignin content and underscore the inhibitory role of PbPME35 in lignin biosynthesis.
Discussion
The PME family, recognized for its significant role in plant development, particularly in lignin biosynthesis (Sexton et al. 2012), was investigated in the context of stone cell formation, a process tightly linked to lignin accumulation. Five Rosaceae plants and pear species were analyzed, leading to the identification of 396 PME genes across six species, namely P. bretschneideri, M. domestica, F. vesca, P. mume, P. communis, and P. pyrifolia. The distribution of PME genes varied among species, with P. bretschneideri hosting 81, and M. domestica, F. vesca, P. mume, P. communis, and P. pyrifolia harboring 92, 62, 65, 15, and 81 PME genes, respectively. The classification of PME genes into four clades in P. bretschneideri and P. pyrifolia exhibited high consistency (Louvet et al. 2006).
Gene duplication, a prominent force in gene family evolution (Lynch and Force 2000), was evident in these Rosaceae species, indicating a shared WGD event from a common Rosaceae ancestor (Jiao et al. 2011). Despite a more recent WGD event estimated at approximately 35–45 MYA in pear (Huang et al. 2015), our analysis indicated that P. bretschneideri and P. pyrifolia diverged approximately 3.54 MYA (Fig. S1a). This discrepancy in divergence times may contribute to the high homology observed in the PME genes of these two species. The conservation of motifs within the same subfamilies supported the credibility of our classification, as closely related genes in the phylogenetic tree shared motifs, suggesting functional similarity among PbPME proteins within the same clade.
Previous studies have classified gene duplication patterns into five modes: WGD, PD, TD, DSD, and TRD, with each mode contributing uniquely to gene family expansion (Freeling 2009; Qiao et al. 2019). WGD, TD, and DSD are prominent features in eukaryotic genome evolution (Fawcett et al. 2009), a trend consistent with our findings, where DSD emerged as the primary force, aligning with previous research. Moreover, the Ka/Ks values for almost all PME genes were < 1, indicating robust purifying selection (Qiao et al. 2015). However, two gene pairs (PbPME67–PbPME68 and PbPME20–PbPME21) in P. bretschneideri exhibited Ka/Ks > 1.2, suggesting positive selection driving their evolution.
Mapping 62 PbPME genes to 16 chromosomes and locating 19 other PbPME genes on various scaffolds revealed an uneven distribution across chromosomes. To gain further insights into the evolutionary history and identify orthologous genes in the P. bretschneideri PME family, we constructed comparative syntenic maps (Fig. 2e). Interspecific collinear analysis highlighted a higher number of orthologous genes (81) between P. bretschneideri and M. domestica than between P. bretschneideri and P. communis (20). This suggests a significant divergence in the PME gene family post-split between P. bretschneideri and P. communis, with a relatively smaller divergence between P. bretschneideri and M. domestica after their divergence.
PMEs are associated with cell lignification and fruit softening (Sexton et al. 2012; Wen et al. 2020). Therefore, we hypothesized that PbPME35 plays a critical role in regulating fruit firmness, lignin biosynthesis, and stone cell formation in pears. Our findings revealed a significant negative correlation between PbPME35 expression and stone cell content, lignin content, fruit firmness, and lignin biosynthetic genes. Furthermore, the overexpressing of PbPME35 in transgenic Arabidopsis and transgenic pear fruit calli exhibited decreased lignin content and tissue lignification. This reduction may arise from the downregulation of genes related to lignin biosynthesis, aligning with previous research (Sexton et al. 2012). Our results suggest that PbPME35 reduces lignin content by suppressing the expression of lignin biosynthesis-related genes, contributing to a decline in stone cell content and fruit firmness, similar to mechanisms identified in previous studies on stone cells (Gong et al. 2020b).
To explore whether PbPME35 influences transcription factor expression, we investigated six transcription factors associated with lignin biosynthesis and secondary cell wall biosynthesis (NST1, MYB85, MYB103, MYB58, MYB83, and MYB46). These transcription factors reportedly promote the expression of lignin-related genes and improve lignin biosynthesis (Mitsuda et al. 2005; Zhong et al. 2008; McCarthy et al. 2009; Zhou et al. 2009; Öhman et al. 2013). Notably, P. bretschneideri NAC stone cell promoting factor (PbrNSC) and P. bretschneideri myeloblastosis 169 (PbrMYB169), homologous to A. thaliana NAC secondary wall thickening promoting factor 1 (NST1) and A. thaliana myeloblastosis 85 (MYB85), respectively, regulate lignin biosynthesis genes and influence lignin formation (Xue et al. 2019; Wang et al. 2021). The expression levels of all investigated transcription factors were lower in transgenic Arabidopsis plants than in WT plants, indicating that PbPME35 could modulate lignification during stone cell development by regulating transcription factor expression. Therefore, we speculate that PbPME35 might interact with lignin biosynthetic proteins or critical switches, leading to the downregulation of lignin biosynthetic genes and, consequently, a reduction in lignin content and inhibition of cell lignification.
The quantity, distribution, and cell wall characteristics of stone cells significantly influence pear fruit texture and quality (Tao et al. 2009; Zhang et al. 2021). Lignin is a major component of stone cells (Gong et al. 2022). Our exploration of the function of PbPME35 provides new insights into lignin biosynthesis and stone cell formation in pear fruit, laying a theoretical groundwork for cultivating pear cultivars with reduced stone cell counts. Notably, we identified high homology between PbPME35 and MdPME42, MdPME43, and FvPME14 (Table S7 and Fig. S3). Consequently, we propose that MdPME42 and MdPME43 may regulate anthocyanin biosynthesis and fruit color in apples through substrate competition with lignin biosynthesis (Hu et al. 2021). Similarly, inhibiting the expression of FvPME14, assuming it functions similarly to PbPME35, may increase the lignin content and firmness of strawberry fruits, potentially extending their shelf life (He et al. 2018). However, these genes functions require further study to clarify their effects on lignin deposition or fruit firmness. Moreover, lignin deposition and stone cell formation in pear fruit is a complex biological process that is regulated by various developmental or environmental signals (Wang et al. 2021). Although this study clarifies the role of PbPME35 in suppressing fruit lignification, the lignin biosynthetic proteins or critical switches that might interact with PbPME35 have not been revealed. The molecular hubs connecting PME and lignification needs further study.
Conclusions
In this study, we identified 396 PME genes across six species (P. bretschneideri, M. domestica, F. vesca, P. mume, P. communis, and P. pyrifolia), with 81 genes belonging to P. bretschneideri. Notably, PbPME35 was identified as a key regulator, suppressing lignification in pear fruit. Specifically, our findings demonstrate that PbPME35 reduces pear fruit lignin content by downregulating the expression of transcription factors and lignin biosynthesis genes.
Availability of data and materials
The datasets used in this study have been uploaded to the NCBI (https://www.ncbi.nlm.nih.gov/) and the BioProject number is PRJNA825067. Anyone who wants to obtain the public data in the research can contact us at taost@njau.edu.cn.
References
Bai S, Tao R, Tang Y, Yin L, Ma Y, Ni J, et al. BBX16, a B-box protein, positively regulates light-induced anthocyanin accumulation by activating MYB10 in red pear. Plant Biotechnol J. 2019;17:1985–97. https://doi.org/10.1111/pbi.13114.
Bailey TL, Bodén M, Whitington T, Machanick P. The value of position-specific priors in motif discovery using MEME. BMC Bioinf. 2010;11:179. https://doi.org/10.1186/1471-2105-11-179.
Carpin S, Crèvecoeur M, de Meyer M, Simon P, Greppin H, Penel C. Identification of a Ca2+-pectate binding site on an apoplastic peroxidase. Plant Cell. 2001;13:511–20. https://doi.org/10.1105/tpc.13.3.511.
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant. 2020;13:1194–202. https://doi.org/10.1016/j.molp.2020.06.009.
Daccord N, Celton JM, Linsmith G, Becker C, Choisne N, Schijlen E, et al. High-quality de novo assembly of the apple genome and methylome dynamics of early fruit development. Nature Genet. 2017;49:1099–106. https://doi.org/10.1038/ng.3886.
Dunand C, Tognolli M, Overney S, von Tobel L, de Meyer M, Simon P, et al. Identification and characterisation of Ca2+-pectate binding peroxidases in Arabidopsis thaliana. J Plant Physiol. 2002;159:1165–71. https://doi.org/10.1078/0176-1617-00768.
Fawcett JA, Maere S, Van De Peer Y. Plants with double genomes might have had a better chance to survive the Cretaceous-Tertiary extinction event. Proc Natl Acad Sci USA. 2009;106:5737–42. https://doi.org/10.1073/pnas.0900906106.
Freeling M. Bias in plant gene content following different sorts of duplication: tandem, whole-genome, segmental, or by transposition. Annu Rev Plant Biol. 2009;60:433–53. https://doi.org/10.1146/annurev.arplant.043008.092122.
Fry SC. Primary cell wall metabolism: tracking the careers of wall polymers in living plant cells. New Phytol. 2004;161:641–75. https://doi.org/10.1111/j.1469-8137.2004.00980.x.
Gao Y, Yang Q, Yan X, Wu X, Yang F, Li J, et al. High-quality genome assembly of “Cuiguan” pear (Pyrus pyrifolia) as a reference genome for identifying regulatory genes and epigenetic modifications responsible for bud dormancy. Hortic Res. 2021;8:197. https://doi.org/10.1038/s41438-021-00632-w.
Geisler-Lee J, Geisler M, Coutinho PM, Segerman B, Nishikubo N, Takahashi J, et al. Poplar carbohydrate-active enzymes. Gene identification and expression analyses. Plant Physiol. 2006;140:946–62. https://doi.org/10.1104/pp.105.072652.
Gong X, Qi K, Chen J, Zhao L, Xie Z, Yan X, et al. Multi-omics analyses reveal the difference of stone cell distribution in pear fruit. Plant J. 2022;113:626–42. https://doi.org/10.1111/tpj.16073.
Gong X, Xie Z, Qi K, Zhao L, Yuan Y, Xu J, et al. PbMC1a/1b regulates lignification during stone cell development in pear (Pyrus bretschneideri) fruit. Hortic Res. 2020;7:59. https://doi.org/10.1038/s41438-020-0280-x.
Guglielmino N, Liberman M, Catesson A, Mareck A, Prat R, Mutaftschiev S, et al. Pectin methylesterases from poplar cambium and inner bark: localization, properties and seasonal changes. Planta. 1997;202:70–5. https://doi.org/10.1007/s004250050104.
He Y, Bose SK, Wang W, Jia X, Lu H, Yin H. Pre-harvest treatment of chitosan oligosaccharides improved strawberry fruit quality. Int J Mol Sci. 2018;19:2194. https://doi.org/10.3390/ijms19082194.
Hertzberg M, Aspeborg H, Schrader J, Andersson A, Erlandsson R, Blomqvist K, et al. A transcriptional roadmap to wood formation. Proc Natl Acad Sci USA. 2001;98:14732–7. https://doi.org/10.1073/pnas.261293398.
Hu Y, Cheng H, Zhang Y, Zhang J, Niu S, Wang X, et al. The MdMYB16/MdMYB1-miR7125-MdCCR module regulates the homeostasis between anthocyanin and lignin biosynthesis during light induction in apple. New Phytol. 2021;231:1105–22. https://doi.org/10.1111/nph.17431.
Huang W, Shi Y, Yan H, Wang H, Wu D, Grierson D, et al. The calcium-mediated homogalacturonan pectin complexation in cell walls contributes the firmness increase in loquat fruit during postharvest storage. J Adv Res. 2022;49:47–62. https://doi.org/10.1016/j.jare.2022.09.009.
Huang X, Li K, Xu X, Yao Z, Jin C, Zhang S. Genome-wide analysis of WRKY transcription factors in white pear (Pyrus bretschneideri) reveals evolution and patterns under drought stress. BMC Genomics. 2015;16:1–14. https://doi.org/10.1186/s12864-015-2233-6.
Jiao Y, Wickett NJ, Ayyampalayam S, Chanderbali AS, Landherr L, Ralph PE, et al. Ancestral polyploidy in seed plants and angiosperms. Nature. 2011;473:97–100. https://doi.org/10.1038/nature09916.
Jolie RP, Duvetter T, Van Loey AM, Hendrickx ME. Pectin methylesterase and its proteinaceous inhibitor: a review. Carbohydr Res. 2010;345:2583–95. https://doi.org/10.1016/j.carres.2010.10.002.
Katoh K, Rozewicki J, Yamada KD. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Briefings Bioinf. 2019;20:1160–6. https://doi.org/10.1093/bib/bbx108.
Lairez D, Cathala B, Monties B, Bedos-Belval F, Duran H, Gorrichon L. Aggregation during coniferyl alcohol polymerization in pectin solution: a biomimetic approach of the first steps of lignification. Biomacromol. 2005;6:763–74. https://doi.org/10.1021/bm049390y.
Levesque-Tremblay G, Müller K, Mansfield SD, Haughn GW. Highly methyl esterified seeds is a pectin methyl esterase involved in embryo development. Plant Physiol. 2015;167:725–37. https://doi.org/10.1104/pp.114.255604.
Li Y, Pi M, Gao Q, Liu Z, Kang C. Updated annotation of the wild strawberry Fragaria vesca V4 genome. Hortic Res. 2019;6:61. https://doi.org/10.1038/s41438-019-0142-6.
Linsmith G, Rombauts S, Montanari S, Deng CH, Celton J-M, Guérif P, et al. Pseudo-chromosome–length genome assembly of a double haploid “Bartlett” pear (Pyrus communis L.). Gigascience. 2019;8:giz138. https://doi.org/10.1093/gigascience/giz138.
Louvet R, Cavel E, Gutierrez L, Guénin S, Roger D, Gillet F, et al. Comprehensive expression profiling of the pectin methylesterase gene family during silique development in Arabidopsis thaliana. Planta. 2006;224:782–91. https://doi.org/10.1007/s00425-006-0261-9.
Lu S, Wang J, Chitsaz F, Derbyshire MK, Geer RC, Gonzales NR, et al. CDD/SPARCLE: the conserved domain database in 2020. Nucleic Acids Res. 2020;48:D265–8. https://doi.org/10.1093/nar/gkz991.
Lynch M, Force A. The probability of duplicate gene preservation by subfunctionalization. Genetics. 2000;154:459–73. https://doi.org/10.1093/genetics/154.1.459.
McCarthy RL, Zhong R, Ye ZH. MYB83 is a direct target of SND1 and acts redundantly with MYB46 in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell Physiol. 2009;50:1950–64. https://doi.org/10.1093/pcp/pcp139.
Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, Von Haeseler A, et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol. 2020;37:1530–4. https://doi.org/10.1093/molbev/msaa015.
Mitsuda N, Seki M, Shinozaki K, Ohme-Takagi M. The NAC transcription factors NST1 and NST2 of Arabidopsis regulate secondary wall thickenings and are required for anther dehiscence. Plant Cell. 2005;17:2993–3006. https://doi.org/10.1105/tpc.105.036004.
Öhman D, Demedts B, Kumar M, Gerber L, Gorzsás A, Goeminne G, et al. MYB103 is required for FERULATE-5-HYDROXYLASE expression and syringyl lignin biosynthesis in Arabidopsis stems. Plant J. 2013;73:63–76. https://doi.org/10.1111/tpj.12018.
Pelletier S, Van Orden J, Wolf S, Vissenberg K, Delacourt J, Ndong YA, et al. A role for pectin de-methylesterification in a developmentally regulated growth acceleration in dark-grown Arabidopsis hypocotyls. New Phytol. 2010;188:726–39. https://doi.org/10.1111/j.1469-8137.2010.03409.x.
Pelloux J, Rusterucci C, Mellerowicz EJ. New insights into pectin methylesterase structure and function. Trends Plant Sci. 2007;12:267–77. https://doi.org/10.1016/j.tplants.2007.04.001.
Qiao X, Li M, Li L, Yin H, Wu J, Zhang S. Genome-wide identification and comparative analysis of the heat shock transcription factor family in Chinese white pear (Pyrus bretschneideri) and five other Rosaceae species. BMC Plant Biol. 2015;15:12. https://doi.org/10.1186/s12870-014-0401-5.
Qiao X, Li Q, Yin H, Qi K, Li L, Wang R, et al. Gene duplication and evolution in recurring polyploidization–diploidization cycles in plants. Genome Biol. 2019;20:38. https://doi.org/10.1186/s13059-019-1650-2.
Röckel N, Wolf S, Kost B, Rausch T, Greiner S. Elaborate spatial patterning of cell-wall PME and PMEI at the pollen tube tip involves PMEI endocytosis, and reflects the distribution of esterified and de-esterified pectins. Plant J. 2008;53:133–43. https://doi.org/10.1111/j.1365-313X.2007.03325.x.
Sexton TR, Henry RJ, Harwood CE, Thomas DS, McManus LJ, Raymond C, et al. Pectin methylesterase genes influence solid wood properties of Eucalyptus pilularis. Plant Physiol. 2012;158:531–41. https://doi.org/10.1104/pp.111.181602.
Tang C, Zhu X, Qiao X, Gao H, Li Q, Wang P, et al. Characterization of the pectin methyl-esterase gene family and its function in controlling pollen tube growth in pear (Pyrus bretschneideri). Genomics. 2020;112:2467–77. https://doi.org/10.1016/j.ygeno.2020.01.021.
Tang H, Bowers JE, Wang X, Ming R, Alam M, Paterson AH. Synteny and collinearity in plant genomes. Science. 2008;320:486–8. https://doi.org/10.1126/science.1153917.
Tao S, Khanizadeh S, Zhang H, Zhang S. Anatomy, ultrastructure and lignin distribution of stone cells in two Pyrus species. Plant Sci. 2009;176:413–9. https://doi.org/10.1016/j.plantsci.2008.12.011.
Wang R, Xue Y, Fan J, Yao JL, Qin M, Lin T, et al. A systems genetics approach reveals PbrNSC as a regulator of lignin and cellulose biosynthesis in stone cells of pear fruit. Genome Biol. 2021;22:313. https://doi.org/10.1186/s13059-021-02531-8.
Wen B, Zhang F, Wu X, Li H. Characterization of the tomato (Solanum lycopersicum) pectin methylesterases: evolution, activity of isoforms and expression during fruit ripening. Front Plant Sci. 2020;11:238. https://doi.org/10.3389/fpls.2020.00238.
Wi S, Singh A, Lee K, Kim Y. The pattern of distribution of pectin, peroxidase and lignin in the middle lamella of secondary xylem fibres in alfalfa (Medicago sativa). Ann Bot. 2005;95:863–8. https://doi.org/10.1093/aob/mci092.
Wu J, Wang Z, Shi Z, Zhang S, Ming R, Zhu S, et al. The genome of the pear (Pyrus bretschneideri Rehd.). Genome Res. 2013;23:396–408. https://doi.org/10.1101/gr.144311.112.
Xue C, Guan SC, Chen JQ, Wen CJ, Cai JF, Chen X. Genome wide identification and functional characterization of strawberry pectin methylesterases related to fruit softening. BMC Plant Biol. 2020;20:13. https://doi.org/10.1186/s12870-019-2225-9.
Xue C, Yao JL, Xue YS, Su GQ, Wang L, Lin LK, et al. PbrMYB169 positively regulates lignification of stone cells in pear fruit. J Exp Bot. 2019;70:1801–14. https://doi.org/10.1093/jxb/erz039.
Yang Y, Li R, Qi M. In vivo analysis of plant promoters and transcription factors by agroinfiltration of tobacco leaves. Plant J. 2000;22:543–51. https://doi.org/10.1046/j.1365-313x.2000.00760.x.
Zhang F, Lu K, Gu Y, Zhang L, Li W, Li Z. Effects of low-temperature stress and brassinolide application on the photosynthesis and leaf structure of tung tree seedlings. Front Plant Sci. 2020;10:1767. https://doi.org/10.3389/fpls.2019.01767.
Zhang MY, Xue C, Hu H, Li J, Xue Y, Wang R, et al. Genome-wide association studies provide insights into the genetic determination of fruit traits of pear. Nature Commun. 2021;12:1144. https://doi.org/10.1038/s41467-021-21378-y.
Zhang Q, Chen W, Sun L, Zhao F, Huang B, Yang W, et al. The genome of Prunus mume. Nat Commun. 2012;3:1318. https://doi.org/10.1038/ncomms2290.
Zhong R, Lee C, Zhou J, McCarthy RL, Ye ZH. A battery of transcription factors involved in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell. 2008;20:2763–82. https://doi.org/10.1105/tpc.108.061325.
Zhou J, Lee C, Zhong R, Ye ZH. MYB58 and MYB63 are transcriptional activators of the lignin biosynthetic pathway during secondary cell wall formation in Arabidopsis. Plant Cell. 2009;21:248–66. https://doi.org/10.1105/tpc.108.063321.
Acknowledgements
We appreciate the contributions of Yiming Shen and Xuebing Zhang (technicians of Orchard in Baoying) to managing the fruit trees. We appreciate the contribution of Textcheck (http://www.textcheck.com; Florida, USA) to providing service in language help.
Funding
This work was financially supported by the Jiangsu Provincial Key Research and Development Program (BE2023365), the National Natural Science Foundation of China (31972361 and U2003121), the Jiangsu Agriculture Science and Technology Innovation Fund (CX(22)2025), and the Achievement Transformation Fund Project of Sanya Institute of Nanjing Agricultural University (NAUSY-CG-YB06).
Author information
Authors and Affiliations
Contributions
F.Z., Q.W., K.Y. and S.T. designed the entire experiment. K.Q., Z.X., X.G. and S.Z. prepared the plant materials and collected the samples. F.Z., Q.W., K.Y., and W.L. carried out the experiment. F.Z. analyzed the results. F.Z., Q.W., and S.T. wrote the main text of this manuscript. All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing financial interests or personal relationships that may influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Fig. S1.
(a) Species evolution tree of seven species. (b) Phylogenetic analysis of Pectin methylesterases (PMEs) from Arabidopsis, Pyrus bretschneideri, Malus domestica, Fragaria vesca, Prunus mume, and P. communis. Fig. S2. Logo of motifs. Fig. S3. Homologous genes of PbPME35 in Pyrus bretschneideri, P. pyrifolia, Malus domestica, Fragaria vesca, Prunus mume, and P. communis, and phylogeny, conserved motifs, and coding sequence (CDS)–untranslated region (UTR) structure of these genes.
Additional file 2: Table S1.
Candidate PME genes. Table S2. Duplication gene pairs of different Rosaceae species in PME family. Table S3. Ka and Ka/Ks values in different Rosaceae species. Table S4. Syntenic pairs information of different Rosaceae species. Table S5. Syntenic pairs between Pyrus bretschneideri and other Rosaceae species. Table S6. RNA-seq data of PME genes in Pyrus bretschneideri. Table S7. Homologous genes identification of PbPME35 by BLAST. Table S8. Sequences for the gene-specific primers used in this work.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, F., Wang, Q., Yuan, K. et al. Genome-wide analysis of the PME gene family reveals its role in suppressing fruit lignification in pear. HORTIC. ADV. 2, 9 (2024). https://doi.org/10.1007/s44281-024-00033-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44281-024-00033-8