Abstract
The blockade of inhibitory receptors such as CTLA-4 (CD152) is being used as immune-checkpoint therapy, offering a powerful strategy to restore effective immune responses against tumors. To determine signal components that are induced under the control of CTLA-4 we analyzed activated murine CD8+ T cells by quantitative proteomics. Accurate mass spectrometry revealed that CTLA-4 engagement led to central changes in the phosphorylation of proteins involved in T-cell differentiation. Beside other targets, we discovered a CTLA-4-mediated induction of the translational inhibitor programmed cell death-4 (PDCD4) as a result of FoxO1 nuclear re-localization. PDCD4 further bound a distinct set of mRNAs including Glutaminase, which points out a critical role for CTLA-4 in CD8+ T-cell metabolism. Consequently, PDCD4-deficient cytotoxic T-lymphocytes (CTLs) expressed increased amounts of otherwise repressed effector molecules and ultimately led to superior control of tumor growth in vivo. These findings reveal a novel CTLA-4-mediated pathway to attenuate CTLs and indicate the importance of post-transcriptional mechanisms in the regulation of anti-tumor immune responses.
Similar content being viewed by others
Main
Cytotoxic T-lymphocytes (CTLs) are the effector cells of the adaptive immune system that exclusively recognize MHC-I presented antigens, thus having a central role in the recognition and clearance of malignant cells.1, 2 A complex interplay of stimulatory and inhibitory receptor-ligand interactions, as well as inflammatory cytokines orchestrate the activation of CD8+ T cells and their differentiation into CTLs.3
To control the magnitude of T-cell responses, the inhibitory surface receptor CTLA-4 has been identified as a primary attenuator of T cells.4, 5 Furthermore, CTLA-4 is overexpressed in exhausted CTLs during chronic diseases alongside other inhibitory receptors such as PD-1 (ref. 6). Its blockade during immune-checkpoint therapy promisingly restores anti-tumor immunity in mice and men.7, 8, 9 CTLA-4 has already been shown to be involved in the regulation of CD8+ T cells;10, 11 however, the mechanisms by which CTLA-4 controls CTL responses still remain incompletely understood. Because of the temporary delayed increase of receptor expression, CTLA-4 strongly impacts on highly activated cells with already established transcription profiles.12 We therefore hypothesize that CTLA-4 could exploit posttranscriptional or -translational mechanisms to modulate CD8+ T-cell differentiation.
In this study we characterized the phosphoproteome response that resulted from CTLA-4 engagement in activated CD8+ T cells by using iTRAQ quantitative mass spectrometry. The analysis identified previously unknown targets and mechanisms how CTLA-4 affects CD8+ T-cell differentiation. We confirmed that CTLA-4-mediated mechanisms abrogated the phosphorylation of the AP-1 family transcription factor Fos-related antigen 2 (Fra-2) and led to a nuclear re-localization of the central transcription factor FoxO1, which caused a strong induction of the translational inhibitor programmed cell death-4 (PDCD4). Strikingly, PDCD4-deficient CTLs showed enhanced production of the otherwise repressed effector molecule IFN-γ and loss of PDCD4 ultimately resulted in superior control of tumor growth in vivo. This novel pathway delineates how CTLA-4 is able to regulate CD8+ T-cell differentiation and the identified mechanisms further provide new strategies to improve anti-tumor immune responses.
Results
CTLA-4 modulates central CD8+ T-cell processes
To identify novel proteins and signaling mechanisms exclusively targeted by CTLA-4 in CD8+ T cells, we performed a comparative mass spectrometry analysis of phosphorylated proteins from cells that were differentiated in vitro with or without CTLA-4 engagement concomitant with α CD3 and α CD28 activation (Supplementary Figure S1a upper).4 To control the effectiveness of CTLA-4-mediated signals we monitored CD8+ T cells by flow cytometry. The cells showed equal activation on day 1 as controlled by proliferation, CD62L downregulation, CD44 and T-bet expression; however, CD8+ T cells that received a CTLA-4 stimulus had 55% less IFN-γ producers on day 2 and less than one-fifth on day 3 (Supplementary Figures S1b-d), which proved a strong impact of CTLA-4-mediated effects.13 Interestingly, CTLA-4-triggered CTLs showed a pronounced re-expression of CD62L on day 2 (Supplementary Figure S1d).
After 48 h of stimulation, which marked the time-point of maximal CTLA-4 expression (Supplementary Figure S2a), the phosphorylated proteins were isolated, digested and the resulting phosphopeptides were measured for their abundance in two independent biological replicates. These analyses led to the detection of 89 phosphopeptides belonging to 74 proteins that were differentially regulated upon CTLA-4 engagement. Sixty-three of 89 peptides showed enhanced phosphorylated residues while 26 peptides were less phosphorylated. Among these proteins, PKC-η and VAV-1 have already been connected to CTLA-4.14, 15 As targets with multiple affected phosphopeptides, NUCKS and PDCD4 were found to be the most upregulated ones, whereas Fra-2 was the strongest dephosphorylated protein (Figure 1a and Supplementary Table S1). The analysis of phosphorylation motifs in the CTLA-4-regulated phosphopeptides revealed specific but also common patterns like RxxS of overrepresented amino acid residues in down- and upregulated sites (Figure 1b). The RxxS motif could be recognized by PKA or CaMKII.16
CTLA-4 modulates the phosphoproteome in differentiating CD8+ T cells. (a) Comparative phosphorylation profile of significantly (P<0.05) regulated proteins in CD8+ T cells from day 2 after differentiation with α CD3, α CD28 and additional CTLA-4 engagement or not, acquired by iTRAQ mass spectrometry in two independent experiments (E1, E2). Proteins were aligned according to GO:BP semantic relations.54 Blue and red represent low and high relative phosphorylation, respectively. Refer to Supplementary Table S1 for complete data. (b) Significantly (P<0.001) overrepresented amino acid residues in motifs containing dephosphorylated (left panel; blue) or stronger phosphorylated (right panel; red) serine residues in CD8+ T cells upon CTLA-4 engagement. Sequences (±7 residues) of the identified sites in (a) were analyzed with the PhosphoSitePlus motif analysis generator using the exhaustive 1–2 AA algorithm.55 (c) Possible interactions of the CTLA-4-affected proteins from a analyzed by the NetworkAnalyst software.18 A GO:BP enrichment analysis was performed to highlight functional subsets significantly (P<0.001) affected by CTLA-4 engagement. See also Supplementary Table S2
Furthermore, by using BABELOMICS functional annotation analysis software17 we could allocate these proteins to six major GO:BP categories namely cytokine production, T-cell activation, RNA processing, mRNA metabolism, DNA replication and regulation of microtubule-based processes (Supplementary Table S2). These specifically enriched clusters further suggest a role for CTLA-4 in modulating central processes of CD8+ T-cell differentiation. With the NetworkAnalyst software18 we created a interaction-network of 67 detected or predicted proteins with 81 connections. Proteins involved in the processes of T-cell activation and cytokine production are more closely connected than the proteins of the other functional clusters, rendering the former as primary CTLA-4 targets. Furthermore, this analysis revealed the central transcription factor FoxP3 as a central interaction hub for CTLA-4-regulated proteins (Figure 1c).19
To analyze the extend of CTLA-4-mediated posttranslational modifications we further characterized the hypophosphorylation of the AP-1 family transcription factor Fra-2, which functions as a key regulator of T-cell differentiation.20 As Fra-2 forms several bands in mobility gel shift assays upon phosphorylation we analyzed nuclear extracts of CD8+ T cells by immunoblotting.21 On day 1, Fra-2 showed an equal phosphorylation pattern of three mobility gel shift bands in both CTLA-4-triggered and control cells, which was maintained in the latter over the following 2 days. CTLA-4 engagement however consistently led to a more than sevenfold reduction of Fra-2 phosphorylation forms in CD8+ T cells on day 2 and 3 (Figure 2a). To confirm that Fra-2 regulation is attributed to posttranslational effects, Fra-2 (Fosl2) mRNA was quantified and showed similar amounts in all stimulated samples (Figure 2b). Among the kinases that are able to phosphorylate Fra-2 we tested the involvement of PKA. The application of the specific PKA inhibitor 14-22 amide led to a more than 30% decreased formation of the slow (upper) and fast (lower) migrating Fra-2 phosphorylation forms in control cells, whereas increased PKA activity due to incubation with the cAMP elevator Forskolin specifically intensified those phosphorylation forms more than six times in CTLA-4-triggered cells (Figure 2c).
CTLA-4 mediates Fra-2 dephosphorylation during CD8+ T-cell differentiation. (a) Immunoblot analysis of total and phosphorylated Fra-2 in nuclear extracts of CD8+ T cells from day 1, 2, and 3 after differentiation with α CD3, α CD28, and additional CTLA-4 engagement or not. (b) Fosl2 (Fra-2) mRNA expression profile of CD8+ T cells after differentiation as described in (a). (c) Immunoblot analysis of phosphorylated Fra-2 (p-Fra-2) in nuclear extracts from day 2 of CD8+ T cells after differentiation as described in (a), treated for 60 min with vehicle, 14–22 amid or Forskolin. Numbers represent relative protein amounts of the phosphorylated gel mobility shifts normalized as indicated. Data are representative of n=2–3 independent experiments. Data points represent individual mice with mean+S.D. ns, not significant, calculated by Mann–Whitney test
Collectively, these findings substantiate the significance of the mass spectrometry dataset for the identification of CTLA-4-mediated signaling effects and further revealed that hyperphosphorylation of Fra-2 integrates PKA signaling in differentiating CD8+ T cells.
CTLA-4 interferes with translation initiation via PDCD4 induction
The iTRAQ mass spectrometry data revealed that CTLA-4 mediated a strong phosphorylation of the translational inhibitor PDCD4 at S94 and S457 (Figure 1a and Supplementary Table S1), which could function as a capable post-transcriptional regulator.22 First, we complemented our proteome results and analyzed PDCD4 S457 phosphorylation as well as total abundance by immunoblotting. Consistent with temporal CTLA-4 expression, we detected on day 2 and 3 a four- to fivefold increase of PDCD4 levels in CTLA-4-triggered CD8+ T cells. The protein was continuously phosphorylated and exclusively distributed in the cytoplasm of CD8+ T cells, where it can interfere with translation initiation (Figure 3a). To fulfill this function, PDCD4 has been shown to interact with the RNA helicase eIF4A to prevent the association of eIF4G.23 To proof this relationship in CD8+ T cells, we conducted eIF4A IPs and quantified the amounts of co-immunoprecipitated eIF4G by immunoblotting.24 Thereby, we detected threefold lower eIF4G protein amounts in eIF4A precipitates from CD8+ T cells differentiated with additional CTLA-4 engagement when compared with precipitates from control cells (Figure 3b).
CTLA-4 induces PDCD4 to restrict protein translation and IFN-γ production. (a) Immunoblot analysis of PDCD4 (total and S457-phosphorylated) in subcellular extracts of CD8+ T cells from day 1, 2 and 3 after differentiation with α CD3, α CD28 and additional CTLA-4 engagement or not. (b) Immunoblot analysis of eIF4A immunoprecipitates (IP) or whole cell lysates (WCL) of CD8+ T cells from day 2 after differentiation as described in a treated with Cycloheximide prior protein extraction. The co-immunoprecipitated eIF4G was quantified to eIf4A (right panel). (c) Immunoblot for PDCD4 (total and S457-phosphorylated) in whole-cell lysates of TCR-transgenic CTLA-4 WT or deficient OT-I CD8+ T cells from day 2 and 3 after differentiation with APCs and OVA. (d) IFN-γ production of TCR-transgenic CTLA-4 and PDCD4 WT or CTLA-4- or PDCD4-deficient OT-I CTLs differentiated as described in (c). IFN-γ-positive cells were determined by flow cytometry and normalized to WT. Numbers represent relative protein amounts normalized as indicated. Data are representative of n=2–4 independent experiments. Data points represent individual mice with mean+S.D. *P<0.05; ns, not significant, calculated by Mann–Whitney test for (b) or by Kruskal–Wallis test with Dunn’s correction for (d)
In conclusion, we revealed a CTLA-4-mediated induction of the translational inhibitor PDCD4 that is localized in the cytoplasm where it interferes with protein translation initiation during CD8+ T-cell differentiation.
PDCD4 attenuates IFN-γ and anti-tumor responses of CTLs
PDCD4 has originally been connected to apoptosis.25 However, in this regard the analysis of PI and Annexin V staining showed no difference between WT and PDCD4-deficient CD8+ T cells (Supplementary Figure S3). Furthermore, PDCD4 has been shown to regulate the cytokine production of activated splenocytes and to be involved in autoimmune inflammation.26 To exclusively determine the function of PDCD4 in physiologically activated CTLs, we cultured TCR transgenic CD8+ T cells from OT-I mice together with Ovalbumin-pulsed APCs (Supplementary Figure S1a lower) and controlled PDCD4 expression by immunoblotting as well as IFN-γ production by flow cytometry. Within the first 3 days, there was no detectable up-regulation of PDCD4 in CD8+ T cells that lack CTLA-4 expression, whereas CTLA-4 sufficiency led to threefold increased PDCD4 amounts at 72 h after T-cell activation (Figure 3c). Moreover, TCR transgenic PDCD4-deficient CD8+ T cells consistently showed 3-fold higher IFN-γ levels on day 3 in comparison to WT cells (Figure 3d). Notably, the absence of PDCD4 neither impaired CTLA-4 expression nor led to changes in the proliferative capacity of the cells (Supplementary Figures S2b and c).
CTLA-4 has been proven to restrict immune responses against malignant cells.7, 8 To initially assess a functional relevance of CTLA-4-induced PDCD4 expression for anti-tumoral CTL responses we compared the ability of TCR transgenic PDCD4- or CTLA-4-deficient CTLs to control the growth of Ovalbumin expressing B16 melanoma cells in an adoptive transfer model. On day 8 after T cell transfer, the mean tumor volume was more than fivefold smaller in mice that received PDCD4−/− CD8+ T cells when compared with control mice (Figures 4a and b). We next analyzed the activated transferred peripheral CD8+ T cells regarding their relative abundance and capacity to produce IFN-γ. Consistently, PDCD4 deficiency led to significantly increased IFN-γ levels in the in vivo activated PDCD4−/− CD8+ T cells whereas the numbers of antigen-specific CD8+ T cells did not significantly differ (Figure 4c).
PDCD4 deficiency leads to enhanced control of experimental tumors. (a) Tumor growth curves of mice s.c. inoculated with B16-OVA melanoma cells, followed 5 days later by i.v. transfer of no T cells (gray) or OT-I TCR-transgenic CTLA-4 and PDCD4 WT (black) or CTLA-4- (red) or PDCD4-deficient (blue) CD8+ T cells. Thin lines represent individual tumors and bold lines the mean tumor growth (n=4–8). (b) Tumor volume of (a) on day 8 after transfer. (c) Frequencies of total CD8+ T cells (left) and IFN-γ producers (right) of i.v. injected peripheral CD62Llo CD8+ T cells on day 6 after transfer as described in (a) determined by flow cytometry. (d) Bioluminescence images of PDCD4 WT and deficient mice from day 5 or day 9 after s.c. inoculation of TRAMP-C1 Luc prostate cancer cells. (e) Frequency of tumor bearing WT (solid black line, n=9), WT CTLA-4-blocked (dashed line, n=3) or PDCD4-deficient (solid blue line, n=6) mice as described in (d). Data points represent individual mice with mean+S.D. *P<0.05, ***P<0.001; ns - not significant, calculated by Kruskal–Wallis test with Dunn’s correction for (b and c) or by Log rank (Mantel–Cox) test for (d)
Similar to the systemic effect of CTLA-4-blocking antibodies, we finally tested mice that completely lacked PDCD4 expression for their capacity to control tumor implantation. We therefore inoculated luciferase-expressing tumor cells in a prostate cancer model that had been shown to be susceptible for immune-checkpoint therapy.9 The PDCD4-deficient as well as CTLA-4-blocked WT mice showed enhanced regression of implanted tumors when compared with untreated PDCD4-sufficient mice. Moreover, all PDCD4−/− mice showed complete tumor rejection within two weeks, whereas only three of nine WT mice were tumor-free until day 14 (Figures 4d and e).
Taken together, these data demonstrated an important role for the translational inhibitor PDCD4 in the downregulation of the IFN-γ production in CTLs and implicated this mechanism of restricted protein expression to be critical for anti-tumor responses.
CTLA-4-mediated nuclear relocalization of FoxO1 promotes the expression of PDCD4
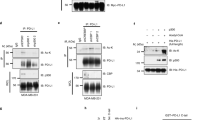
To clarify a possible regulation of PDCD4 due to ubiquitin ligase mediated protein degradation,24, 27 we applied a proteasomal inhibitor during stimulation of CD8+ T cells. Despite a strong accumulation of ubiquitinated proteins, the immunoblot analysis showed no alteration in the relative PDCD4 abundance and therefore no involvement of this mechanism in the CTLA-4-mediated regulation of PDCD4 (Figure 5a). We further ruled out an implication of the PDCD4-targeting miR-21,28 as evenly distributed micro-RNA levels could be detected (Figure 5b). The quantification of Pdcd4 transcripts revealed a consistent abundance of mRNA levels in comparison with the respective protein amounts, suggesting a regulation of PDCD4 through mRNA expression (Figures 5c and 3a). PDCD4 has been shown to be induced by the Foxo transcription factor.29 In murine regulatory T cells, FoxO1 has been confirmed to bind within the 5′-UTR of the Pdcd4 gene and loss of FoxO1 leads to decreased Pdcd4 mRNA levels.30 To assess the impact of FoxO1 on PDCD4 expression in CD8+ T cells, we applied an inhibitor of FoxO1 transactivation 24 h after beginning of T cell activation, which completely prevented the induction of PDCD4 (Figure 5d). Furthermore, we performed a Foxo1 deletion through transduction of Foxo1fl/fl CD8+ T cells with TAT-Cre recombinase.31 This treatment led to both reduced FoxO1 and decreased PDCD4 protein levels with a concomitantly elevated production of IFN-γ (Figure 5e). Thus, PDCD4 upregulation is primarily dependent on FoxO1.
CTLA-4 mediates PDCD4 expression via FoxO1 reactivation. (a) Immunoblot analysis of PDCD4 and ubiquitinated proteins in CD8+ T cells from day 2 after differentiation with α CD3, α CD28 and additional CTLA-4 engagement or not, treated for 2 h with vehicle or MG132 prior cell lysis. (b) miR-21 or (c) Pdcd4 mRNA expression profile of CD8+ T cells after differentiation as described in a. (d) Immunoblot analysis of PDCD4 in CD8+ T cells from day 1 and 2 after differentiation as described in a, treated on day 1 for 24 h with vehicle or AS1842856. (e) Immunoblot analysis of whole-cell lysates of Foxo1fl/fl CD8+ T cells from 60 h after treatment with TAT-Cre recombinase or not (Mock) followed by differentiation with α CD3, α CD28 and additional CTLA-4 engagement. Cells were further analyzed by flow cytometry for IFN-γ production (lower panel). (f) Immunoblot analysis of FoxO1 (total and S253-phosphorylated) and Akt (pan and T308- or S473-phosphorylated) in subcellular extracts of CD8+ T cells from 6 and 48 h after differentiation as described in (a). Data are representative of n=2–4 independent experiments. Numbers represent relative protein amounts normalized as indicated. Data points represent individual mice with mean+S.D. *P<0.05; ns, not significant, calculated by Mann–Whitney test
Upon T-cell activation, FoxO1 is regulated by a highly conserved mechanism that involves phosphorylation by Akt, leading to its cytoplasmic retention and subsequent inactivation.32, 33 Hence, we analyzed the cellular distribution of FoxO1 as well as phosphorylation events by immunoblotting to confirm CTLA-4-mediated FoxO1 activity. After 6 h of stimulation we consistently detected a strong Akt activation in all cells as determined by T308 and S473 phosphorylation. At the same time, FoxO1 was equally phosphorylated at the Akt site S253 and consequently localized in the cytoplasm. However, at 48 h FoxO1 was strongly enriched in the nuclei of CD8+ T cells that were differentiated with additional CTLA-4 engagement despite an equal reduction of Akt and FoxO1 phosphorylation in both CTLA-4-triggered and control cells (Figure 5f).
We therefore conclude that CTLA-4-mediated mechanisms led to an Akt-independent nuclear relocalization of the transcription factor FoxO1, resulting in augmented gene expression of the translational inhibitor PDCD4.
Targets of PDCD4 are critical for CTL responses
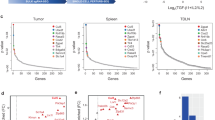
PDCD4 directly associates with mRNAs to repress the translation of the corresponding proteins.34 To identify PDCD4 targets in CTLs we immunoprecipitated PDCD4 and subsequently sequenced the co-precipitated RNA fraction. The sequencing data revealed mRNA molecules derived from 21 different genes to be significantly enriched in the PDCD4 precipitates (Figure 6a and Supplementary Table S3). Among these targets, Glutaminase (Gls) and SENP3 have been shown to be essential for cellular metabolism and biosynthesis, thus playing a critical role in CTL responses.35, 36 We finally sought to compare the in situ mRNA levels of Gls and Senp3 with the respective protein amounts to prove a direct effect of PDCD4 on the translation of these targets. Despite similar mRNA amounts, the Glutaminase and SENP3 protein levels were more than 50% lower in WT CD8+ T cells than in their PDCD4-deficient counterparts (Figure 6b and Supplementary Figure S2d). To further asses the influence of PDCD4-mediated effects on cellular glutaminolysis of CTLs we supplemented activated CD8+ T cells with glutamate. This treatment resulted in a significant increase in the IFN-γ production of WT cells (Figure 6c).
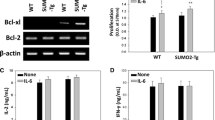
PDCD4-targeted protein translation. (a) Significantly enriched mRNAs in PDCD4 immunoprecipitates of CD8+ T cells from day 2 after differentiation with α CD3, α CD28 and additional CTLA-4 engagement. Data were aligned according to GO:BP semantic relations.54 See also Supplementary Table S3. (b) Immunoblot analysis of Glutaminase, SENP3 and PDCD4 in subcellular extracts of OT-I TCR-transgenic PDCD4 WT or deficient CD8+ T cells from day 5 after differentiation with OVA-peptide pulsed microspheres loaded with DimerX I, B7.1-Fc and α CTLA-4. Numbers represent relative protein amounts normalized as indicated. (c) Flow cytometric analysis (left) and cumulative graph (right) of glutamate-induced IFN-γ in OT-I TCR-transgenic PDCD4 WT or deficient CD8+ T cells from day 5 after differentiation as described in (b), supplemented with glutamate 60 h prior measurement. Data are representative of n=2–4 independent experiments. Data points represent individual mice with mean+S.D. *P<0.05, calculated by Mann–Whitney test
Together, these data show how CTLA-4-induced expression of PDCD4 might attenuate CD8+ T cell effector functions, leading to abrogated anti-tumor responses.
Discussion
In this study we conducted a comprehensive analysis of CTLA-4-mediated intracellular changes in differentiating CTLs, which are a central part of the effector response towards antigenic tumors. The differentiation of naïve CD8+ T cells is characterized by an early commitment to an autonomous developmental program.37 After 48 h, CTLA-4 is strongly expressed on the surface of CD8+ T cells, that continue to differentiate in absence of further activation and which already peak in gene transcription.12, 38 Consequently, CTLA-4 needs to act in a modulatory manner to attenuate CTLs responses rather than to solely ablate their stimulation.39 In this study, we identified 89 CTLA-4-responding phosphorylation sites at 74 different proteins supporting the notion of CTLA-4-mediated posttranscriptional or -translational modifications of already established signaling circuits. Importantly, the applied differentiation system4 enabled similar initial stimulation of all CD8+ T cells as verified by uniform values for every analyzed parameter 24 h after beginning of the stimulation. On day two, an inverse relation between IFN-γ and CTLA-4 was detectable, which has been previously reported for CD8+ T cells.11, 13
The mass spectrometric analysis provided unprecedented details on the signal network downstream of CTLA-4 and focusing on phosphorylated components significantly facilitated the detection of novel mechanisms. However, a further analysis of the phospho-proteins is required to characterize the identified regulation as a direct phosphorylation event. Interestingly, the protein interaction approach contributed several CTLA-4-regulated proteins capable to associate with the regulatory T-cell factor FoxP3.19 Furthermore, the discovered CTLA-4-dependent phosphorylation of PKC-η at S675 adds valuable information to the previously for Treg cells described interaction.14 Therefore, the findings in this study might also give indications for CTLA-4-mediated functions in other lymphocyte subpopulations.
The identification of common phosphorylation motifs implies the involvement of multiple kinases. In this regard, we characterized the phosphorylation of the AP-1 family transcription factor Fra-2 (Fosl2) which has been implicated in T-helper cell differentiation.20 Consistently, we demonstrated in activated CD8+ T cells an initial strong phosphorylation leading to the formation of several gel mobility band shifts and that CTLA-4 engagement completely abrogated these pos-translational modifications.21 This could be due to a regulation of the activity or the nuclear shuttling of the responsible kinases. Notably, Fosl2 gene expression has recently been identified to be regulated in exhausted CD8+ T cells.40
One of the most powerful CTLA-4-controlled proteins was the translational inhibitor PDCD4 that functions as a crucial posttranscriptional regulator.22 The central impact of the CTLA-4-PDCD4 axis could be unambiguously demonstrated in several independent in vitro and in vivo settings, delineating the cell- and context-specific role of PDCD4 in attenuating effector responses of CTLs. This appears to be unlike to B lymphocytes, were PDCD4 is primarily involved in maintaining cell quiescence.26 As the blockade of CTLA-4 is applied in tumor therapy, we initially assessed the role of its newly identified downstream target PDCD4 in the impairment of anti-tumor CTL responses. Our data demonstrate that a general and a CD8+ T-cell-specific deficiency of PDCD4 led to improved control of tumor growth. We consistently observed increased IFN-γ levels in PDCD4-deficient CTLs that could be responsible for an enhanced anti-tumor activity.41 Although CD8+ T cells are the main effectors of the adaptive immune system that lyse malignant cells, a concomitant suppression through CTLA-4-expressing regulatory T cells further accounts for ineffective immune responses against tumors.42 It is tempting to speculate that PDCD4 as a FoxP3 interacting and FoxO1-induced protein has likely a critical role in regulatory T cells as well, strengthening PDCD4 as a promising target for therapeutic interventions.19, 30
Furthermore, the identified main target of PDCD4 in CTLs, the rate-limiting enzyme Glutaminase, serves as a metabolic checkpoint during cell differentiation and has an important role in the anti-tumor response, as its substrate glutamine provides an essential source for the cellular metabolism of CTLs in the glucose-deprived tumor microenvironment.35, 43 In line with the function as a translational inhibitor, we revealed that PDCD4 expression caused a downregulation of the SUMO-specific protease SENP3, which has been shown to act as a crucial factor in ribosome biogenesis.36 In conclusion, CTLA-4-mediated PDCD4 expression induced a defined restriction of protein translation that critically shaped the quality of CTLs leading to impaired anti-tumor responses.
The global blockade of CTLA-4 impinges on more than one pathway, which is also reflected by the detected positive feedback of CTLA-4 on FoxO1. This transcription factor not only regulated PDCD4 expression but furthermore controls several proteins critical for CD8+ T-cell differentiation-like CD62L or TCF-1.44, 45, 46 Thus, dissecting CTLA-4-initiated pathways might identify targets which primarily control single aspects of anti-tumor immune responses.
Interestingly, the CTLA-4-mediated FoxO1 regulation occurred independently of Akt, indicating a central cellular role of FoxO1 downstream of multiple pathways. In this regard, FoxO1 has recently been shown to be activated by PD-1 and to promote CD8+ T-cell exhaustion.47 Moreover, increased PDCD4 mRNA levels have been detected in chronically activated CD8+ T cells.48 Thus, the induction of PDCD4 expression by FoxO1 could be a part of a redundant mechanism that is addressed by both CTLA-4 and PD-1. In T-helper cells, PDCD4 has already been confirmed as a target of PD-1 (ref. 49). As a consequence, PDCD4 could be upregulated even in CTLs that do not express CTLA-4 at a given time and the same relation could account for PD-1. This notion would further support the observed superior anti-tumor response of PDCD4-deficient CD8+ T cells being similar to synergistic effects of a combinatory CTLA-4 and PD-1 blockade.50
Taken together, the control of protein biosynthesis through the induction of the translational inhibitor PDCD4 marks an important immune-checkpoint in CTL differentiation. Our data further suggests novel targets on CTLs that evoke posttranscriptional strategies for improving anti-tumor immune-checkpoint therapy.
Materials and Methods
Mice
OVA-specific TCRtg mice (OT-I) both CTLA-4+/+ and PDCD4+/+ or CTLA-4−/− or PDCD4−/−, as well as Ly5.1, C57BL/6 and PDCD4−/− mice were bred under specific pathogen-free conditions in the central animal facility of the University of Magdeburg Medical School (Germany). OT-I TCRtg was controlled by flow cytometry and both CTLA-4−/− and PDCD4−/− mice were genotyped as described elsewhere.13, 26 All animal experiments were performed in accordance with institutional, state and federal guidelines.
Antibodies
The following antibodies against murine antigens were used: α CD152 (UC10-4F10), isotype control antibody (A19-3), α Vα2-TCR (B20.1), α T-bet (O4-46), α CD44 (IM7), α CD45.2 (104) (BD Biosciences, Franklin Lakes, NJ, USA); α CD3 (145-2C11), α CD28 (37.51), α CD8a (53-6.7), α CD62L (MEL-14), α IFN-γ (XMG1.2) (BioLegend, Koblenz, Germany); α Fra-2 (Q20), α GAPDH (A3), α eIF4A (yN-20) (Santa Cruz Biotechnology, Dallas, TX, USA); α PDCD4 (D29C6), α eIF4G (C45A4), α eIF4A (C32B4), α FoxO1 (C29H4), α phospho-FoxO1S253, α phospho-AKTS473, α phospho-AKTT308, α Akt (pan) (C67E7), α SENP3 (D20A10), α Aldolase A (D73H4), α Lamin B1, α Ubiquitin (P4D1) (Cell Signaling Technology, Danvers, MA, USA); α phospho-PDCD4S457 (9G6) (Rockland Immunochemicals, Limerick, PA, USA); α Glutaminase (EP7212) (Abcam, Cambridge, UK). For in vivo antibody treatment of mice α CTLA-4 (UC10-4F10) were purified from hybridoma supernatants and controlled by flow cytometry. α DEC-OVA conjugates were produced as previously described.51
Cell isolation and stimulation
Isolation of APCs or naïve CD8+ CD44low CD62Lhigh T cells from spleens, inguinal- and axillary-lymph nodes was performed with α CD90 microbeads or α CD8a-FITC and α FITC microbeads (Miltenyi Biotec, Bergisch Gladbach, Germany) according to the manufacturer’s instructions. Isolated CD8+ T cells from C57BL/6 mice were stimulated with antibodies immobilized on microspheres (Thermo Fisher Scientific, Waltham, MA, USA). Microspheres were coated as described elsewhere.4 The following concentrations were used: 0.75 μg/ml α CD3, 2 μg/ml α CD28 and 7 μg/ml α CTLA-4 or isotype control antibody. Cells were cultured with complete medium (RPMI, with 100 U/ml Penicillin, 100 μg/ml Streptomycin (Thermo Fisher Scientific), and 10% FCS (Biochrom, Berlin, Germany)) supplemented with 10 ng/ml of recombinant IL-2 (Miltenyi Biotec) and 10 ng/ml of recombinant IL-12 (BioLegend). CD8+ T cells from TCR-transgenic OT-I mice were stimulated with 100 μg/ml OVA and 0.2 μg/ml α DEC-OVA in complete medium supplemented with 1 ng/ml IL-2 and 1 ng/ml IL-12 and CD90 depleted splenocytes from congenic C57BL/6 mice or with OVA257-264 peptide-pulsed microspheres loaded with 1 μg/ml DimerX I fusion protein (BD Biosciences), 5 μg/ml B7.1-Fc (BioLegend) and 8 μg/ml α CTLA-4. In some experiments, CD8+ T cells were treated with 4 mM L-glutamic acid (Sigma-Aldrich, Munich, Germany) for 60 h prior analysis.
TAT-Cre transduction
Foxo1fl/fl cells and TAT-Cre recombinase were kindly provided by K. Rajewsky (Max Delbrück Center for Molecular Medicine, Berlin, Germany). Purified Foxo1fl/fl CD8+ T cells were washed in serum-free medium (HyClone Laboratories, Logan, UT, USA), prewarmed and transduced with sterile-filtrated TAT-Cre dilution31 at a density of 5 × 106 cells/ml for 45 min at 37 °C. Cells were then washed in complete medium and differentiated with 0.75 μg/ml α CD3, 2 μg/ml α CD28 and 7 μg/ml α CTLA-4 antibodies attached to microspheres in complete medium supplemented with 10 ng/ml of recombinant IL-2 and 10 ng/ml of recombinant IL-12. Mock-transduced CD8+ T cells were identically treated except of omitted recombinase.
Flow cytometry
For intracellular IFN-γ staining α CD3/α CD28/α CTLA-4 or APC-activated cells were treated with 5 μg/ml Brefeldin A for 3 h, DimerX I/B7.1-Fc/α CTLA-4 activated or ex vivo analyzed cells with 10 μg/ml PMA, 1 μg/ml Ionomycin and 5 μg/ml Brefeldin A for 4 h, fixed and permeabilized with 0.5% saponin (Sigma-Aldrich). In some experiments, cells were loaded with CFSE (Thermo Fisher Scientific) prior activation. Intracellular T-bet staining was performed using a FoxP3 buffer set (Thermo Fisher Scientific). Cytometric measurements were performed on a FACS-Canto II (BD Biosciences) and analyzed with FlowJo (Tree Star, Ashland, OR, USA) software.
iTRAQ mass spectrometric analysis of phosphorylated proteins
Isolated phosphorylated proteins of CD8+ T cells from day 2 after differentiation with or without additional CTLA-4 engagement were comparatively analyzed by iTRAQ mass spectrometry. Fifteen minutes before analysis, cells were thoroughly re-suspended, shortly centrifuged and incubated at 37 °C to ensure active contacts. Detailed proteomic methods and phospho-peptide sequencing are described in Supplementary Materials and Methods.
Immunoprecipitation and immunoblotting
Immunoprecipitation was performed with Protein G Microbeads (Miltenyi Biotec) according to the manufacturer’s instructions. In brief, CD8+ T cells were lysed and incubated in NP-40 lysis buffer with α eIF4A (yN-20) and 10 mM NaCl or α PDCD4 or α Aldolase A antibodies with 100 mM NaCl and Protein G microbeads, and subsequently purified by positive selection. Immunoprecipitation of PDCD4 or Aldolase A was conducted with extracts of CD8+ T cells on day 2 after differentiation with α CD3, α CD28 and additional CTLA-4 engagement. APC-activated CD8+ T cells were isolated with α CD8a-FITC and α FITC microbeads before protein extraction. Cellular extracts or immunoprecipitates were separated on 10% SDS-PAGE gels and transferred onto nitrocellulose membranes. Blots were probed with antibodies, visualized and quantified using the Odyssey scanner and software (LI-COR, Lincoln, NE, USA). In some experiments the CD8+ T cells were treated with 100 μg/ml Cycloheximide (AppliChem, Darmstadt, Germany) or 40 μM Forskolin (Sigma-Aldrich) or 10 μM 14-22 amid for 60 min or 5 μM MG132 for 2 h or 0.5 μM AS1842856 (Merck Millipore, Darmstadt, Germany) for 24 h before protein extraction.
Next-generation sequencing and qPCR measurement of mRNA or microRNA
Small and large RNAs from CD8+ T cells or immunoprecipitates were extracted by using NucleoSpin miRNA Kit and RNA Kit (Macherey-Nagel, Düren, Germany), respectively. Detailed methods for preparation of cDNA libraries, as well as sequencing and detection of RNA levels see Supplementary Materials and Methods.
Animal procedures
Ly5.1 recipient mice were s.c. inoculated with 2 × 105 B16-OVA cells. B16-OVA cells were kindly provided by J. Huehn (Helmholtz Centre for Infection Research, Braunschweig, Germany).52 Congenic naive CD8+ OT-I T cells (3.3 × 106) both CTLA-4+/+ and PDCD4+/+ or CTLA-4−/− or PDCD4−/− were i.v. injected 5 days later. Tumor size was monitored by measurement in three dimensions with a caliper and calculated as hemi-elliptical volume. Blood samples were taken on day 6 after CD8+ T-cell transfer. In the tumor model with prostate cancer cells,9 2 × 106 Luciferase-expressing TRAMP-C1 cells (Applied Biological Materials, Richmond, BC, Canada) were s.c. inoculated into PDCD4 WT or deficient mice. Part of the WT mice additionally received 100 μg i.p. injections of α CTLA-4 antibody on day 6, 10 and 13. Complete eradication of TRAMP-C1 Luc tumors was controlled by in vivo luminescence imaging. In brief, luciferin (150 mg/kg) was injected i.v. and mice were anesthetized with isoflurane and measured by IVIS 200 in vivo imaging system (PerkinElmer, Waltham, MA, USA).
Statistical analysis
P-values<0.05 were considered statistical significant and calculated with a two-sided Mann–Whitney rank sum test for the comparison of two groups; for more than two groups a Kruskal–Wallis test with Dunn’s correction was used. All determined significances remained by using an equivalent parametric test. The percentage of tumor-bearing mice was calculated with a Log rank (Mantel–Cox) test. The determination of significantly different phosphopeptides was performed by the discrimination of regulated from randomly fluctuating values in a large number of small samples.53 Statistical significance is indicated as followed: ns, not significant, *P<0.05 and ***P<0.001.
References
Kilinc MO, Gu T, Harden JL, Virtuoso LP, Egilmez NK . Central role of tumor-associated CD8+ T effector/memory cells in restoring systemic antitumor immunity. J Immunol 2009; 182: 4217–4225.
Hadrup S, Donia M, Thor SP . Effector CD4 and CD8 T cells and their role in the tumor microenvironment. Cancer Microenviron 2013; 6: 123–133.
Haring JS, Badovinac VP, Harty JT . Inflaming the CD8+ T cell response. Immunity 2006; 25: 19–29.
Krummel MF, Allison JP . CD28 and CTLA-4 have opposing effects on the response of T cells to stimulation. J Exp Med 1995; 182: 459–465.
Walunas TL, Lenschow DJ, Bakker CY, Linsley PS, Freeman GJ, Green JM et al. CTLA-4 can function as a negative regulator of T cell activation. Immunity 1994; 1: 405–413.
Wherry EJ, Ha SJ, Kaech SM, Haining WN, Sarkar S, Kalia V et al. Molecular signature of CD8+ T cell exhaustion during chronic viral infection. Immunity 2007; 27: 670–684.
Leach DR, Krummel MF, Allison JP . Enhancement of antitumor immunity by CTLA-4 blockade. Science 1996; 271: 1734–1736.
Phan GQ, Yang JC, Sherry RM, Hwu P, Topalian SL, Schwartzentruber DJ et al. Cancer regression and autoimmunity induced by cytotoxic T lymphocyte-associated antigen 4 blockade in patients with metastatic melanoma. Proc Natl Acad Sci USA 2003; 100: 8372–8377.
Kwon ED, Hurwitz AA, Foster BA, Madias C, Feldhaus AL, Greenberg NM et al. Manipulation of T cell costimulatory and inhibitory signals for immunotherapy of prostate cancer. Proc Natl Acad Sci USA 1997; 94: 8099–8103.
Thaventhiran JE, Hoffmann A, Magiera L, La Roche Mde, Lingel H, Brunner-Weinzierl M et al. Activation of the Hippo pathway by CTLA-4 regulates the expression of Blimp-1 in the CD8+ T cell. Proc Natl Acad Sci USA 2012; 109: E2223–E2229.
Zumwalde NA, Domae E, Mescher MF, Shimizu Y . ICAM-1-dependent homotypic aggregates regulate CD8 T cell effector function and differentiation during T cell activation. J Immunol 2013; 191: 3681–3693.
Best JA, Blair DA, Knell J, Yang E, Mayya V, Doedens A et al. Transcriptional insights into the CD8(+) T cell response to infection and memory T cell formation. Nat Immunol 2013; 14: 404–412.
Pandiyan P, Hegel JKE, Krueger M, Quandt D, Brunner-Weinzierl MC . High IFN-gamma production of individual CD8 T Lymphocytes is controlled by CD152 (CTLA-4). J Immunol 2007; 178: 2132–2140.
Kong KF, Fu G, Zhang Y, Yokosuka T, Casas J, Canonigo-Balancio AJ et al. Protein kinase C-eta controls CTLA-4-mediated regulatory T cell function. Nat Immunol 2014; 15: 465–472.
Wei B, da RDS, Wang H, Rudd CE . CTL-associated antigen-4 ligation induces rapid T cell polarization that depends on phosphatidylinositol 3-kinase, Vav-1, Cdc42, and myosin light chain kinase. J Immunol 2007; 179: 400–408.
Amanchy R, Periaswamy B, Mathivanan S, Reddy R, Tattikota SG, Pandey A . A curated compendium of phosphorylation motifs. Nat Biotechnol 2007; 25: 285–286.
Al-Shahrour F, Minguez P, Vaquerizas JM, Conde L, Dopazo J . BABELOMICS: a suite of web tools for functional annotation and analysis of groups of genes in high-throughput experiments. Nucleic Acids Res 2005; 33: W460–W464.
Xia J, Benner MJ, Hancock REW . NetworkAnalyst - integrative approaches for protein-protein interaction network analysis and visual exploration. Nucleic Acids Res 2014; 42: W167–W174.
Rudra D, deRoos P, Chaudhry A, Niec RE, Arvey A, Samstein RM et al. Transcription factor Foxp3 and its protein partners form a complex regulatory network. Nat Immunol 2012; 13: 1010–1019.
Ciofani M, Madar A, Galan C, Sellars M, Mace K, Pauli F et al. A validated regulatory network for Th17 cell specification. Cell 2012; 151: 289–303.
Gruda MC, Kovary K, Metz R, Bravo R . Regulation of Fra-1 and Fra-2 phosphorylation differs during the cell cycle of fibroblasts and phosphorylation in vitro by MAP kinase affects DNA binding activity. Oncogene 1994; 9: 2537–2547.
Yang H, Jansen AP, Komar AA, Zheng X, Merrick WC, Costes S et al. The transformation suppressor Pdcd4 is a novel eukaryotic translation initiation factor 4 A binding protein that inhibits translation. Mol Cell Biol 2003; 23: 26–37.
Yang HS, Cho MH, Zakowicz H, Hegamyer G, Sonenberg N, Colburn NH . A novel function of the MA-3 domains in transformation and translation suppressor Pdcd4 is essential for its binding to eukaryotic translation initiation factor 4 A. Mol Cell Biol 2004; 24: 3894–3906.
Kroczynska B, Sharma B, Eklund EA, Fish EN, Platanias LC . Regulatory effects of programmed cell death 4 (PDCD4) protein in interferon (IFN)-stimulated gene expression and generation of type I IFN responses. Mol Cell Biol 2012; 32: 2809–2822.
Shibahara K, Asano M, Ishida Y, Aoki T, Koike T, Honjo T . Isolation of a novel mouse gene MA-3 that is induced upon programmed cell death. Gene 1995; 166: 297–301.
Hilliard A, Hilliard B, Zheng SJ, Sun H, Miwa T, Song W et al. Translational regulation of autoimmune inflammation and lymphoma genesis by programmed cell death 4. J Immunol 2006; 177: 8095–8102.
Dorrello NV, Peschiaroli A, Guardavaccaro D, Colburn NH, Sherman NE, Pagano M . S6K1- and betaTRCP-mediated degradation of PDCD4 dromotes protein translation and cell growth. Science 2006; 314: 467–471.
Davis BN, Hilyard AC, Lagna G, Hata A . SMAD proteins control DROSHA-mediated microRNA maturation. Nature 2008; 454: 56–61.
Olson CM, Donovan MR, Spellberg MJ, Marr MT . The insulin receptor cellular IRES confers resistance to eIF4A inhibition. Elife 2013; 2: e00542.
Ouyang W, Liao W, Luo CT, Yin N, Huse M, Kim MV et al. Novel Foxo1-dependent transcriptional programs control Treg cell function. Nature 2012; 491: 554–559.
Peitz M, Pfannkuche K, Rajewsky K, Edenhofer F . Ability of the hydrophobic FGF and basic TAT peptides to promote cellular uptake of recombinant Cre recombinase: a tool for efficient genetic engineering of mammalian genomes. Proc Natl Acad Sci USA 2002; 99: 4489–4494.
Biggs W, Meisenhelder J, Hunter T, Cavenee WK, Arden KC . Protein kinase B/Akt-mediated phosphorylation promotes nuclear exclusion of the winged helix transcription factor FKHR1. Proc Natl Acad Sci USA 1999; 96: 7421–7426.
Kane LP, Weiss A . The PI-3 kinase/Akt pathway and T cell activation: pleiotropic pathways downstream of PIP3. Immunol Rev 2003; 192: 7–20.
Wedeken L, Ohnheiser J, Hirschi B, Wethkamp N, Klempnauer KH . Association of tumor suppressor protein Pdcd4 with ribosomes is mediated by protein-protein and protein-RNA interactions. Genes Cancer 2010; 1: 293–301.
Wang R, Dillon CP, Shi LZ, Milasta S, Carter R, Finkelstein D et al. The transcription factor Myc controls metabolic reprogramming upon T lymphocyte activation. Immunity 2011; 35: 871–882.
Haindl M, Harasim T, Eick D, Muller S . The nucleolar SUMO-specific protease SENP3 reverses SUMO modification of nucleophosmin and is required for rRNA processing. EMBO Rep 2008; 9: 273–279.
van Stipdonk MJ, Lemmens EE, Schoenberger SP . Naive CTLs require a single brief period of antigenic stimulation for clonal expansion and differentiation. Nat Immunol 2001; 2: 423–429.
Kaech SM, Ahmed R . Memory CD8+ T cell differentiation: initial antigen encounter triggers a developmental program in naive cells. Nat Immunol 2001; 2: 415–422.
Riley JL, Mao M, Kobayashi S, Biery M, Burchard J, Cavet G et al. Modulation of TCR-induced transcriptional profiles by ligation of CD28, ICOS, and CTLA-4 receptors. Proc Natl Acad Sci USA 2002; 99: 11790–11795.
Tinoco R, Carrette F, Barraza ML, Otero DC, Magana J, Bosenberg MW et al. PSGL-1 is an immune checkpoint regulator that promotes T cell exhaustion. Immunity 2016; 44: 1190–1203.
Shankaran V, Ikeda H, Bruce AT, White JM, Swanson PE, Old LJ et al. IFNgamma and lymphocytes prevent primary tumour development and shape tumour immunogenicity. Nature 2001; 410: 1107–1111.
Peggs KS, Quezada SA, Chambers CA, Korman AJ, Allison JP . Blockade of CTLA-4 on both effector and regulatory T cell compartments contributes to the antitumor activity of anti-CTLA-4 antibodies. J Exp Med 2009; 206: 1717–1725.
Siska PJ, Rathmell JC . T cell metabolic fitness in antitumor immunity. Trends Immunol 2015; 36: 257–264.
Fabre S, Carrette F, Chen J, Lang V, Semichon M, Denoyelle C et al. FOXO1 regulates L-selectin and a network of human T cell homing molecules downstream of phosphatidylinositol 3-kinase. J Immunol 2008; 181: 2980–2989.
Kim MV, Ouyang W, Liao W, Zhang MQ, Li MO . The transcription factor Foxo1 controls central-memory CD8+ T cell responses to infection. Immunity 2013; 39: 286–297.
Zhou X, Yu S, Zhao D, Harty JT, Badovinac VP, Xue H . Differentiation and persistence of memory CD8(+) T cells depend on T cell factor 1. Immunity 2010; 33: 229–240.
Staron MM, Gray SM, Marshall HD, Parish IA, Chen JH, Perry CJ et al. The transcription factor FoxO1 sustains expression of the inhibitory receptor PD-1 and survival of antiviral CD8+ T cells during chronic infection. Immunity 2014; 41: 802–814.
Doering TA, Crawford A, Angelosanto JM, Paley MA, Ziegler CG, Wherry EJ . Network analysis reveals centrally connected genes and pathways involved in CD8+ T cell exhaustion versus memory. Immunity 2012; 37: 1130–1144.
Iliopoulos D, Kavousanaki M, Ioannou M, Boumpas D, Verginis P . The negative costimulatory molecule PD-1 modulates the balance between immunity and tolerance via miR-21. Eur J Immunol 2011; 41: 1754–1763.
Curran MA, Montalvo W, Yagita H, Allison JP . PD-1 and CTLA-4 combination blockade expands infiltrating T cells and reduces regulatory T and myeloid cells within B16 melanoma tumors. Proc Natl Acad Sci USA 2010; 107: 4275–4280.
Hegel JK, Knieke K, Kolar P, Reiner SL, Brunner-Weinzierl MC . CD152 (CTLA-4) regulates effector functions of CD8+ T lymphocytes by repressing Eomesodermin. Eur J Immunol 2009; 39: 883–893.
Klages K, Mayer CT, Lahl K, Loddenkemper C, Teng MW, Ngiow SF et al. Selective depletion of Foxp3+ regulatory T cells improves effective therapeutic vaccination against established melanoma. Cancer Res 2010; 70: 7788–7799.
Klawonn F . Significance tests to identify regulated proteins based on a large number of small samples. Kybernetika 2012; 48: 478–493.
Mazandu GK, Mulder NJ . DaGO-Fun: tool for Gene Ontology-based functional analysis using term information content measures. BMC Bioinformatics 2013; 14: 284.
Hornbeck PV, Zhang B, Murray B, Kornhauser JM, Latham V, Skrzypek E . PhosphoSitePlus, 2014: mutations, PTMs and recalibrations. Nucleic Acids Res 2015; 43: D512–D520.
Acknowledgements
The study was supported by grants of the DFG (CRC854 B14 and B16, Br1860/8-1) and the Wilhelm Sander Foundation (number 2012.090.1).
Author contributions
HL designed and performed experiments, analyzed data and wrote paper. JW, DS, SL, MP and AA performed experiments. FK performed statistical analysis. LJ supervised proteomic experiments and wrote paper. MZ supervised transcriptome experiments. MBW supervised the study, designed experiments and wrote paper.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by S Martin
Supplementary Information accompanies this paper on Cell Death and Differentiation website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Lingel, H., Wissing, J., Arra, A. et al. CTLA-4-mediated posttranslational modifications direct cytotoxic T-lymphocyte differentiation. Cell Death Differ 24, 1739–1749 (2017). https://doi.org/10.1038/cdd.2017.102
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2017.102
- Springer Nature Limited
This article is cited by
-
The CUL5 E3 ligase complex negatively regulates central signaling pathways in CD8+ T cells
Nature Communications (2024)
-
Programmed cell death 4 governs NLRP3-mediated pyroptosis in septic lung disorders
Molecular Biology Reports (2024)
-
Advances in immune checkpoint-based immunotherapies for multiple sclerosis: rationale and practice
Cell Communication and Signaling (2023)
-
An integrated proteome and transcriptome of B cell maturation defines poised activation states of transitional and mature B cells
Nature Communications (2023)
-
ST3GAL1 and βII-spectrin pathways control CAR T cell migration to target tumors
Nature Immunology (2023)