Abstract
Direct reprogramming of differentiated cells to pluripotent stem cells has great potential to improve our understanding of developmental biology and disorders such as cancers, and has implications for regenerative medicine. In general, the effects of transcription factors (TFs) that are transduced into cells can be influenced by pre-existing transcriptional networks and epigenetic modifications. However, previous work has identified four key TFs, Oct4, Sox2, Klf4 and c-Myc, which can reprogram various differentiated cells to generate induced pluripotent stem cells. Here, we show that in the heart, the transduction of cardiac mesenchymal progenitors (CMPs) with Klf4 and c-Myc (KM) was sufficient to drive the differentiation of these cells into adipocytes without the use of adipogenic stimulation cocktail, that is, insulin, 3-isobutyl-1-methylxanthine (IBMX) and dexamethasone. KM-transduced CMPs exhibited a gradually increased expression of adipogenic-related genes, such as C/Ebpα, Pparγ and Fabp4, activation of the peroxisome proliferator-activated receptor (PPAR) signaling pathway, inactivation of the cell cycle-related pathway and formation of cytoplasmic lipid droplets within 10 days. In contrast, NIH3T3 fibroblasts, 3T3-L1 preadipocytes, and bone marrow-derived mesenchymal stem cells transduced with KM did not differentiate into adipocytes. Both in vitro and in vivo cardiac ischemia reperfusion injury models demonstrated that the expression of KM genes sharply increased following a reperfusion insult. These results suggest that ectopic adipose tissue formation in the heart following myocardial infarction results from CMPs that express KM following a stress response.
Similar content being viewed by others
Main
Adipocyte differentiation, that is, adipogenesis, has been extensively investigated, and its regulation via transcriptional cascades has been described for in vitro model systems.1 The adipogenic transcriptional cascade consists of two waves. The first wave converges at the CCAAT/enhancer-binding protein (C/Ebp)β/γ, which induces the second wave consisting of nuclear receptor peroxisome proliferator-activated receptor (Ppar)γ and C/EBPα activity. In addition, c-Myc is periodically expressed during the early phase of adipogenesis.2 Krüppel-like factor (Klf) family members include both repressors and activators of adipogenesis, and are activated during the first wave.3 KLF4 and c-MYC (KM) coordinately bind the promoters of genes that are activated during the reprogramming of differentiated cells to pluripotency.4 Whether KM work together in adipogenesis has not been examined.
Mesenchymal stem cells (MSCs) are multipotent cells with a capacity to differentiate to mesodermal lineages and show a vigorous proliferation capacity under conventional culture conditions.5 The criteria for identifying MSCs include adherence to a plastic dish, a characteristic surface profile and differentiation capacity in vitro.6 Although most prior reports have identified bone marrow as the origin for MSCs, other organs including adipose tissue7 and the heart8, 9 also harbor fibroblasts that fulfill the criteria for MSCs. MSCs derived from different organs demonstrate varying capacities for proliferation and differentiation.10 Although several reports have demonstrated adipose tissue formation in the myocardium following reperfusion therapy for ischemic heart diseases,11, 12, 13, 14 it is unclear how fat depositions in the heart are generated.
Direct reprogramming of differentiated cells using specific transcription factors (TFs) opens the door to understanding the mechanisms underlying development and the pathogenesis of various disorders, and has applications in regenerative medicine.15, 16 Transdifferentiation or direct conversion, which occurs when a differentiated cell type is reprogrammed to another cell type, could be implemented via the same strategy of using a set of TFs to generate cardiomyocytes, neurons and so on.17, 18 Moreover, there are similarities and overlaps between the pathways for the generation of induced pluripotent stem cells (iPSCs) and tumorigenesis, such as a mesenchymal-to-epithelial transition.19 Recently, it was reported that partial reprogramming of differentiated cells using four reprogramming TFs (Oct4 , Sox2, Klf4 and c-Myc (OSKM)) in vivo could generate tumors via epigenetic reprogramming.20 Direct reprogramming can shed light on cancer biology, and vice versa.
Transcriptional cascades definitively determine cell fate during reprogramming to pluripotency and normal differentiation. We examined whether murine cardiac mesenchymal progenitors (CMPs)9 expressing Sca-1 antigen and TFs associated with cardiomyocytes can differentiate into adipocytes and how the process is regulated. Elucidating the mechanisms underlying adipose tissue generation in the heart should help us to understand the pathophysiologies of ischemic reperfusion injury (IRI) myocardial infarction (MI).
Results
Transduction of OSKM into CMPs is sufficient to induce their differentiation into adipocytes
To test our hypothesis that reprogrammed CMPs can differentiate into other cell types, we transduced CMPs with Sendai virus encoding OSKM. We modified standard reprogramming medium by removing leukemia inhibitory factor to avoid the generation of iPSCs, following a previous report (Figure 1a).21 Nine days after infection (day 8), OSKM-transduced CMPs (OSKM-CMPs) formed cytoplasmic lipid droplets, which were not formed by untreated CMPs (CMP control) or CMPs treated with adipogenic differentiation cocktails (CMP with adipogenic cocktails) (Figure 1b). The lipid droplets in OSKM-CMPs were clearly stained by Oil Red O (Figure 1b). Next, to identify gene expression in reprogrammed CMPs, we performed quantitative reverse transcription polymerase chain reaction (qRT-PCR) analysis (Figure 1c). The expression levels of Oct4 and Sox2 in OSKM-CMPs decreased at day 2 and were maintained at a low level thereafter. Klf4 and c-Myc expression in OSKM-CMPs also decreased at day 2. The expression levels of the adipogenic-related genes C/Ebpα and Fabp4 in OSKM-CMPs increased at day 4. The expression levels of Fas, Pparγ1 and Pparγ2 in OSKM-CMPs were higher than those in untreated CMPs at day 6. Moreover, the expression levels of the cardiac-related genes Mef2c, Gata4 and Tbx5 in CMP controls increased, but these genes were not expressed in OSKM-CMPs.
OSKM-transduced CMPs differentiated into adipocytes. (a) Schematic representation of the adipocyte differentiation method. MC, medium change. Growth medium indicates the basal medium for each cell line, and reprogramming medium indicates KO-DMEM-based medium. (b) Phase contrast microscope images. CMPs treated with OSKM Sendai virus (CMP with OSKM) clearly accumulated large cytosolic lipid droplets at day 8. These droplets were stained with Oil Red O. Untreated CMPs (CMP cntrl) and those treated with adipogenic stimulation cocktails (CMP with adipogenic cocktails) did not form cytosolic lipid droplets at day 8 and were not stained with Oil Red O. The white bar indicates 50 μm. (c) qRT-PCR analysis of the expression of each gene in CMPs on each day. Individual RNA expression levels were normalized to Gapdh expression. Error bars indicate S.E. (n=3). * and ** indicate significant changes compared with untreated controls at day 8 (P<0.05 and 0.01, respectively)
Transduction of OSKM into NIH3T3 fibroblasts is insufficient to induce their differentiation into adipocytes
Next, we transduced OSKM into NIH3T3 fibroblasts. At day 8, NIH3T3 fibroblasts changed in shape from fibroblast-like cells to round cells; however, there were no iPSC-like colonies or Oil Red O-positive cells (Figure 2a). Expression of OSKM genes at day 2 increased rapidly; however, the expression levels of the adipogenic genes Fas, C/Ebpα and Pparγ2 decreased steeply from day 2 (Figure 2b). In particular, C/Ebpα and Fas expression did not differ from that of the control (without OKSM). These results showed that OSKM-transduced NIH3T3 fibroblasts did not differentiate into adipocytes.
OSKM-transduced NIH3T3 fibroblasts did not differentiate into adipocytes. (a) Phase contrast microscope images. NIH3T3 fibroblasts treated with OSKM Sendai virus (NIH3T3 fibroblasts with OSKM) did not form cytosolic lipid droplets at day 8 and were not stained with Oil Red O. White bar indicates 50 μm. PhC, phase contrast. (b) qRT-PCR analysis of the expression of each gene in NIH3T3 fibroblasts on each day. Individual RNA expression levels were normalized to Gapdh expression. Error bars indicate S.E. (n=3). * and ** indicate significant changes compared with untreated controls at day 8 (P<0.05 and 0.01, respectively)
Microarray analysis of OSKM-CMPs
To analyze global gene expression in OSKM-CMPs, we performed microarray analysis using an Agilent mouse microarray chip and the NIA Array Analysis website.22 Based on hierarchical clustering analysis of gene expression, OSKM-CMPs could be clearly discriminated from CMP controls (Figure 3a). In addition, principal component analysis (PCA) of gene expression showed that the OSKM-CMPs were different from the CMP controls and gradually shifted from right to left on the PC1 axis in a time-dependent manner (Figure 3b). Furthermore, a group of genes with decreasing expression over time (positive direction along PC1, 4577 probes) and a group with increasing expression over time (negative direction along PC1, 5314 probes) were observed (Figure 3b). These genes were categorized based on gene ontology (GO) annotations and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways (Figure 3c). Many genes showing decreasing expression over time (PC1-positive direction) were assigned to functional categories related to cell cycling and cell division. In addition, many of the genes showing decreasing expression over time were assigned to functions related to focal adhesion and regulation of the actin cytoskeleton. Otherwise, the genes showing increasing expression over time (PC1-negative direction) were functionally related to adipocyte differentiation, including saturated and unsaturated fatty acid metabolism, fat cell differentiation and the PPAR signaling pathway. These results strongly indicated that OSKM-CMPs differentiated into adipocytes.
Global gene expression of OSKM-transduced CMPs. (a) Hierarchical clustering analysis of OSKM-transduced CMPs on each day by NIA array analysis. CMP cntrl indicates untreated CMPs. (b) PCA by NIA array analysis. CMPs are categorized based on the principal component 1 (PC1) direction (left). A total of 4577 probes were in the positive direction on PC1, indicating decreased expression over time, and 5314 probes were in the negative direction of PC1, indicating increased expression over time. (c) Genes in the PC1-positive and -negative directions were categorized based on biological processes using GO annotations (white bars) and KEGG pathways (gray bars)
Klf4 and c-Myc have important roles in the differentiation of CMPs into adipocytes
To determine which of the reprogramming factors among OSKM were critical for CMP differentiation into adipocytes, we examined the effect of removing each factor. We searched genetic databases for information regarding gene expression during adipogenesis in 3T3-L1 cells. The available information from previous studies indicated that Klf4 was expressed before adipogenic stimulation and that c-Myc sharply rose up in response, then the expression of both genes decreased to become undetectable at 1 week, at which time lipid-laden adipocytes were macroscopically recognized using the standard protocol (GSE34150). Both Klf4 and c-Myc have been reported to be adipocyte differentiation-related factors.3, 23 We withdrew OSKM sequentially. Neither the withdrawal of Oct4 nor that of Sox2 influenced adipogenic differentiation based on Oil Red O staining. The withdrawal of either Klf4 or c-Myc decreased the percentage of Oil Red O-positive cells 9 days after infection, compared with that observed when these factors were present (Figure 4a). The expression levels of genes related to adipogenesis including C/Ebpα, Fas and Pparγ2 in TF(s)-transduced CMPs at day 8 showed a similar pattern, regardless of the TFs used except OKM-infected CMP. In OKM-infected CMP cells, the expression of both C/Ebpα and Pparγ1 was upregulated, but Fas, which is involved in lipogenesis,24 was downregulated. C/Ebpα is mainly attributed to a role in insulin sensitivity in adipocyte differentiation,25 and Pparγ2, but not Pparγ1, has an essential role in adipogenic differentiation in vitro.26 Overexpression of c-MYC elicits p53-dependent apoptosis in primary fibroblasts.27, 28 Infection of CMP cells might result in apoptosis, therefore the expression level of c-MYC was almost the same as that of the negative control. KLF4 might be required to suppress p53 and c-Myc-induced apoptosis. Fabp4, which is expressed during terminal differentiation,29 was not expressed in K- and M-transduced CMPs (Figure 4b). This observation might be ascribed to the inability of those CMPs to differentiate into mature lipid-laden adipocytes. Furthermore, KM genes in CMPs resulted in a greater relative area of Oil Red O-positive cells than that observed for the transduction of OSKM, OKM and SKM (Figure 4c). These results showed that the combination of Klf4 and c-Myc is indispensable for the differentiation of CMPs into adipocytes.
Adipocyte differentiation properties of TF-transduced CMPs. (a) Phase contrast microscope images. CMPs were transduced using a combination of OSKM, SKM, KM, K and M Sendai virus. The white bar indicates 200 μm. (b) qRT-PCR analysis of the expression of each gene in CMPs at day 8. Individual RNA expression levels were normalized to Gapdh expression. Error bars indicate S.E. (n=3). * and ** indicate significant changes compared with KM-treated CMPs (white box, P<0.05 and 0.01, respectively). (c) Calculation of Oil Red O staining area. Each well image was captured using a Keyence BZ-X700 digital microscope. The black bar indicates 5 mm (left). The graph shows the percentages of the total area that were positive for Oil Red O staining (right). Error bars indicate S.E.; * and ** indicate significant changes (P<0.05 and 0.01, respectively). OKM: Oct4, Klf4 and c-Myc; SKM: Sox2, Klf4 and c-Myc; KM: Klf4 and c-Myc; K: Klf4; M: c-Myc. Negative cntrl: untreated CMPs
Klf4 and c-Myc do not induce adipogenic differentiation of MSCs
To test the ability of other cell types to differentiate into adipocytes via KM transduction, we used the same methods to induce adipocyte differentiation (Figure 1a). KM genes did not increase the frequency of adipocyte differentiation in 3T3-L1 preadipocytes (Figure 5a). All of the adipogenic genes exhibited a similar expression pattern in both non-treated and KM-transduced 3T3-L1 preadipocytes (Figure 5b). Furthermore, we examined whether other mouse multipotent MSCs derived from bone marrow (KUSA-A1, KUM5 and KUM9 cells) could be induced to undergo differentiation to adipocytes by KM transduction. Cells treated with KM did not show any formation of lipid droplets in the cytosol (Figure 5c). The cells exhibited high expression levels of c-Myc and Klf4, and low or unchanged expression levels of adipogenic genes such as C/Ebpα, Fas, Pparγ2 and Fabp4 at day 8 (Figure 5d). Interestingly, the expression of all adipogenic genes in these MSCs decreased at day 8. These results showed that these cells were not able to differentiate into adipocytes via KM. The expression levels of a set of TFs in bone marrow-derived mesenchymal cell lines, including KUSA-A1, KUM5 and KUM9 cells, were determined relative to the expression of Gapdh from cardiac tissues. All TFs related to adipogenesis did not increase following exogenous KM gene transfer, suggesting that the induction of adipogenesis by KM gene transfer could be specific to CMPs.
Adipocyte differentiation properties of KM-transduced 3T3-L1 preadipocytes and MSCs derived from bone marrow (KUSA-A1, KUM5 and KUM9 cells). (a) Phase contrast microscope images. 3T3-L1 preadipocytes were transduced with KM Sendai virus. One day after infection, cells were cultured in reprogramming medium for 8 days. At day 8, cells were fixed and stained with Oil Red O. The white bar indicates 50 μm. Cntrl indicates untreated 3T3-L1 preadipocytes. (b) qRT-PCR analysis of the expression of each gene in 3T3-L1 preadipocytes on each day. Individual RNA expression levels were normalized to Gapdh expression. Error bars indicate S.E. (n=3). * and ** indicate significant changes compared with KM-treated 3T3-L1 cells at day 8 (P<0.05 and 0.01, respectively). (c) Phase contrast microscope images. MSCs derived from mouse bone marrow were transduced with KM Sendai virus. One day after infection, cells were cultured in reprogramming medium for 8 days. At day 8, cells were fixed and stained with Oil Red O. The white bar indicates 50 μm. Cntrl indicates untreated MSCs. (d) qRT-PCR analysis of the expression of each gene in MSCs KUSA-A1 (black bars), KUM5 (gray bars) and KUM9 (white bars) at day 8. Individual RNA expression levels were normalized to Gapdh expression. Error bars indicate S.E. (n=3). * and ** indicate significant changes from untreated control cells (KM–, day –2) (P<0.05 and 0.01, respectively)
Klf4 and c-Myc were induced by ischemia reperfusion injury in vitro and in vivo
CMPs were exposed to IRI model culture conditions in vitro (Figure 6a). CMPs immediately detached from the tissue culture dish under hypoxic conditions, and detached CMPs re-attached to the culture dish under normoxic conditions (Figure 6b). CMPs expressed the hypoxia-induced gene Hif1α at 3 h, and expression returned to baseline levels at 24 h (Supplementary Figure 1), indicating that the culture system could successfully mimic IRI. The expression level of Klf4 increased at 3 h after hypoxia, remained high until 6 h under normoxia, then returned to the baseline level, whereas the expression of c-Myc increased sharply just after normoxic conditions were applied. HIF1, the abundance of which indicates the extent of the ischemic insult, is upregulated by the inhibition of proteolysis in cardiac IRI,30 and its abundance can be measured by western blotting. To assess the link between a set of TFs and IRI using a consistent methodological approach, qRT-PCR was chosen. As a surrogate marker for IRI, instead of Hif-1, the gene expression levels of c-Fos and c-Jun, the protein products of which form the TF AP1, were examined. The expression of both c-Fos and c-Jun drastically and temporarily increased just after exposure to normoxic conditions.
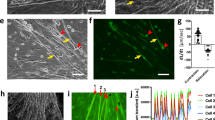
In vitro and in vivo IRI model. (a) Schematic representation of the in vitro IRI method. PBS, phosphate-buffered saline. Growth medium indicates the basal medium for CMP. Hypoxia indicates 1% O2, 5% CO2, balance N2 conditions in a hypoxic chamber, and normoxia indicates 21% O2, 5% CO2 conditions. (b) Phase contrast microscope images of CMPs under hypoxic conditions. (c) qRT-PCR analysis of the expression of each gene in CMPs. Error bars indicate S.E. (n=3). * and ** indicate significant changes compared with CMPs at 0 h (white box, P<0.05 and 0.01, respectively). H indicates hypoxia condition; N indicates hormoxia condition. (d) Schematic representation of the in vivo IRI method. (e) Photograph of open-chest mouse with 8-0 Prolene suture thread on the LAD. (f) qRT-PCR analysis of the expression of each gene in an IRI model LV heart. Error bars indicate S.E. (n=3). * and ** indicate significance (P<0.05 and 0.01, respectively). ‘I’ indicates ischemic condition; ‘R’ indicates reperfusion condition. (g) Western blotting of IRI model LV heart. Error bars indicate S.E. (n=3). * and ** indicates statistically significant differences at 0 h (white box, P<0.05 and 0.01, in the order described)
Moreover, we determined that KM was involved in in vivo murine cardiac IRI (Figures 6d–f). Injured murine ventricles acutely and temporarily expressed Klf4 and c-Myc in IRI, similar to the pattern observed for in vitro IRI. The expression levels of both c-Fos and c-Jun were transiently increased 1 h after ischemia, and increased after reperfusion, indicating that Klf4 might be involved in the cellular response following the insult as early as c-Fos and c-Jun. Platelet-derived growth factor, which is involved in cardiac IRI, induces c-Myc expression via an AP1-dependent signaling pathway under in vitro culture conditions.31 The kinetics of c-Fos and c-Jun expression showed an earlier response to IRI than was found for c-Myc, suggesting that AP1, which is composed of c-Fos and c-Jun, might be an upstream regulator of c-Myc. The expression of C/Ebpβ and C/Ebpδ gradually increased; however, at 24 h the gene expression levels of both C/Ebpβ and C/Ebpδ returned to baseline, whereas C/Ebpα gene expression was increased and maintained at a higher level 3 h after left anterior descending artery (LAD) ligation. The expression levels of Pparγ1 and Pparγ2 transiently increased at 1 h (Figure 6f), and the expression of PPARγ protein transiently increased at days 1 and 2 (Figure 6g). However, reperfused hearts were not stained by Oil Red O staining (Supplementary Figure 2).
Regional expression of adipogenic-related genes in a RA, an AAR and the AON
To evaluate RNA expression in the LV wall of the IRI model in detail at 2 h after LAD ligation, a set of TFs related to adipogenesis was examined by qRT-PCR for each area consisting of the area of necrosis (AON), an area at risk (AAR) and a remote area (RA), which were defined by double staining with 2,3,5-triphenyltetrazolium chloride (TTC) and Evans blue (Figure 7a). Klf4 expression significantly increased in the AON, and c-Myc expression significantly increased in all areas over time. The first wave of TFs (C/Ebpβ and C/Ebpδ) for adipogenesis was significantly raised in all areas. In contrast, expression of the second wave of TFs (Pparγ2, and C/Ebpα) did not differ in AON and AAR compared with that in RA. Comparison of gene expression among areas revealed that Klf4 expression in AON was significantly higher than that in RA, and c-Fos, c-Jun, and c-Myc expression levels in AON and AAR were significantly higher than those in RA. The gene expression of the first wave of TFs in AON was significantly higher than that in RA, and the gene expression of the second wave of TFs was not significantly different among the three areas. The surrogate TFs (c-Fos and c-Jun) showed significantly increased expression in AON and AAR compared with that in RA, which was the same tendency as that observed for Klf4, c-Myc, C/Ebpβ and C/Ebpδ, suggesting that adipogenesis might be initiated via KM induction in this cardiac IRI model (Figure 7b).
Gene expression in tested areas of ischemia reperfusion injury hearts. (a) TTC staining and Oil red O staining of ischemic reperfusion heart. RA, AOR and AAR indicate RA, AON and AAR, respectively. Black bar: 1 mm. (b) qRT-PCR analysis for the expression of genes in the ischemic reperfusion mouse heart on each day. Individual RNA expression levels were normalized to the respective mouse Gapdh expression levels. Error bars indicate S.E. (n=3). * and ** indicates statistically significant differences in each color box. (P<0.05 and 0.01, in the order described). † also indicates statistically significant difference from IRI non-treatment heart (0 h) (P<0.05 and 0.01, in the order described). Relative gene expression in the RA at 0 h is regarded as 1
Discussion
We demonstrated that CMPs could effectively differentiate into adipocytes via two TFs, Klf4 and c-Myc, without adipogenic stimulation. Neither Klf4 nor c-Myc transduction alone resulted in CMPs differentiating into adipocytes. These results suggested that KM proteins in CMPs coordinately regulate adipogenesis. Interestingly, this new protocol was only effective in CMPs, but not in MSCs from bone marrow and 3T3-L1 preadipocytes. CMPs did not differentiate into adipocytes when treated with adipogenic stimulation cocktails, contradicting the hypothesis that CMPs contain adipogenic progenitors or stem cells. These results might indicate that the route to adipocyte differentiation is not uniform as defined by the in vitro cellular model, but rather that it depends upon the cell type and environment. The increased expression levels of both Klf4 and c-Myc in IRI models might be due to intracardiac fatty degeneration following MI.
Previous in vitro studies of the molecular pathways underlying adipogenesis are based on limited adipogenic cell lines.32, 33, 34, 35 Adipogenesis involves two distinct waves of TF expression and six defined differentiation stages: mesenchymal precursors, committed preadipocytes, growth-arrested preadipocytes, mitotic clonal expansion, terminal differentiation and mature adipocytes.36, 37, 38 Preadipocytes differentiate into lipid-laden and insulin-sensitive adipocytes upon the addition of exogenous adipogenic stimulation cocktails in confluent culture growth conditions. Klf4 is one of the earliest TFs in the first wave and is regulated by cAMP3 and the JAK-STAT pathway, mechanisms that maintain the pluripotency of embryonic stem cells.39 As the Jak-Stat pathway is activated by external stimuli such as reactive oxygen species (ROS),40 Klf4 mediates the response to external stress.41 KLF4 directly transactivates the C/EBPβ gene by binding to the promoter region, and is a key TF in the first wave that relays the signal to PPARγ, a central factor in adipogenesis.3 However, within 1 h of adipogenic stimulation of confluent 3T3-L1 cells, c-Myc is rapidly and highly expressed, along with c-Fos and c-Jun.23 Constitutively overexpressed c-Myc inhibits the differentiation of 3T3-L1 cells, possibly by precluding the entry of cells to a distinct predifferentiation stage in G0/G1.42 The peak in the expression of c-Myc might function as an amplifier of the expression of other genes to surpass the threshold from a stable, low-level position in adipocytes and not as an activator of the cell cycle.43 In this experiment, neither Klf4 nor c-Myc transduction alone induced adipogenesis from CMPs, suggesting that KM cooperatively function to induce adipogenesis.
Little is known about in vivo adipogenesis or de novo adipocyte generation, which is referred to as hyperplasia in terms of tissue growth, owing to the post-mitotic nature of mature adipocytes. In adipose tissue, resident MSCs are considered to be a major source for adipocyte generation.36 Some studies have reported in vivo adipocyte differentiation from MSCs, which expressed similar cell surface antigens to those expressed by the CMPs in this study.44, 45 Recently, myocardium-derived stem/progenitor cells such as cardiac stem cells and CMPs have been reported by several institutes.9, 46, 47, 48 CMPs, which we isolated from murine hearts and defined as a Sca-1-positive population, expressed a similar surface antigen profile to that of MSCs,6 except for CD73 and CD34. The origin of each MSC influences its molecular phenotype, including the transcriptional network, epigenetic landscape and subsequent differentiation potential.10, 38 CMPs are a distinct population from bone marrow, adipose tissue or skin-derived MSCs, with an expression profile of TFs characteristic of the heart. Only KM induced the differentiation of CMPs into adipocytes, potentially owing to the default settings of the TF network.
Baroldi et al.11 reported adipose tissue formation in the excised heart during transplant surgery, and this was termed lipomatous metaplasia. Another study using the recipient heart in transplantation showed consistent ectopic fat formation, representing 84% of healed MI.12 Imaging analyses in patients with a history of MI using either computed tomography13 or magnetic resonance imaging14 have demonstrated a similar prevalence of ectopic fat formation, which was found in approximately 65% of individuals. These reports suggest that adipose formation in the myocardium should be a more common pathology than is currently recognized. A new mechanism of arrhythmogenesis in ventricular tachycardia proposes that intramyocardial adipose tissue hinders myocardial conduction and worsens local electrophysiological properties, which in turn results in an increased propensity for ventricular tachycardia.49 The KM genes transduced into CMP maintained high expression for about 1 week, resulting in differentiation of the CMPs to lipid-laden adipocytes and activation of the second wave of TFs for adipogenesis. In contrast, cardiac IRI temporarily induced Klf4 and c-Myc expression, which sharply fell and disappeared after only a few days, resulting in failure to maintain the second wave and generate adipocytes. During left ventricular (LV) remodeling post MI, the renin–angiotensin–aldosterone system (RAAS) is activated, which leads to AP1 activation50 and might result in c-Myc induction.51 Angiotensin II can induce Klf4 expression in cardiac fibroblasts including CMPs.52 Although RAAS does not induce a high expression of KM such as that which we observed during in vitro adipogenesis in this study, the low level of KM expression over a long period induced by RAAS might slowly form adipogenic enhanceosomes at enhancer regions for late-acting TFs in adipogenesis, such as Pparγ.37 Inhibition of either Klf4 or c-Myc induction might be a novel strategy to treat LV remodeling post MI.
Global mRNA profiling of the myocardium after IRI has revealed that Klf family members, including Klf4 and c-Myc, exhibit significantly increased expression following ischemia and additional increases after reperfusion.53 Ischemic events generate interleukin 6 in the heart, activating STAT3,54 which is linked to Klf4 expression.39 However, ROS, which have been characterized as negative factors in reperfusion injuries, are involved in signal transduction in many biological processes, including inflammation, stemness and differentiation, cancer, and aging.55 The thioredoxin family member nucleoredoxin (NRX), which is a redox sensor regulated by ROS, interacts with dishevelled (Dvl) under a reduction state. The oxidized form of NRX liberates Dvl, which in turn stabilizes β-catenin, leading to the transcription of WNT target genes including c-Myc.56 Consistent with the aforementioned studies on signal transduction, myocardial ischemia led to increased expression of Klf4 and reperfusion stimulated c-Myc expression. These results strongly suggest that the two TFs KLF4 and c-MYC in CMPs are causative factors for intracardiac adipogenesis following myocardial reperfusion. The regional assessment revealed that the expression levels of Klf4, c-Myc, c-Fos and C/Ebpδ in AON and AAR were raised more than those in RA, suggesting that the adipogenic differentiation process had already been launched in the area directly affected by the insult of ischemia and reperfusion at 2 h.
MSCs can be isolated from various tissue types including bone marrow, adipose tissue, heart and skeletal muscle. However, the characteristics and epigenetic background of these MSCs differ.10 Transduction of CMPs with KM genes was highly effective in inducing their differentiation into adipocytes, whereas transducing the same genes into MSCs derived from other tissues did not induce them to differentiate. Furthermore, these phenomena might provide a basis for ectopic fat formation in ischemic hearts. Understanding CMP adipogenesis should shed light on post-MI and IRI pathophysiology and facilitate the development of better treatments for these disorders.
Materials and Methods
Materials
Geltrex and basic fibroblast growth factor (bFGF) were purchased from Life Technologies (Carlsbad, CA, USA). The CytoTune-iPS ver. 1.0 Sendai Reprogramming Kit was purchased from DNAVEC (Ibaraki, Japan). Oil Red O powder was purchased from Nacalai Tesque, Inc. (Kyoto, Japan). Percoll Plus was purchased from GE Healthcare UK (Buckinghamshire, England). The adipogenic stimulation cocktail ingredients insulin, IBMX, and dexamethasone were purchased from Sigma-Aldrich (St. Louis, MO, USA).
Cell preparation
Experimental procedures and protocols were approved by the Animal Experiment Ethics Committee of the Kyoto Prefectural University of Medicine. Murine CMPs were isolated from wild-type C57BL/6 mouse hearts (10- to 16-week-old).9 Briefly, the mice were killed by deep anesthesia with pentobarbital. The hearts were excised, and atria were used in this study. The minced tissue fragments were digested twice for 30 min at 37 °C with 0.2% (w/v) type II collagenase and 0.01% (w/v) DNase I (Worthington Biochemical, Lakewood, NJ, USA). After digestion, cells were passed through a 70-μm filter to remove debris and transferred to Dulbecco's modified Eagle’s medium (DMEM)/F12 supplemented with 10% (v/v) fetal bovine serum (FBS) (Life Technologies). The cells were collected and size fractionated on a 30–70% Percoll gradient to obtain CMPs expressing the Sca-1 antigen. CMPs were seeded on 60-mm collagen I-coated dishes (Asahi Glass, Tokyo, Japan) in DMEM/F12 supplemented with 10% (v/v) FBS and 20 ng/ml bFGF. The medium was changed every 3 days.
Cell culture and adipocyte differentiation
CMPs were cultured in DMEM/F12 supplemented with 10% (v/v) FBS and 20 ng/ml bFGF in a humidified atmosphere containing 5% CO2. NIH3T3 fibroblasts and MSCs derived from bone marrow KUM5, KUM9 and KUSA-A133 were cultured in DMEM (Wako Chemical Co., Osaka, Japan) supplemented with 10% (v/v) FBS in a humidified atmosphere containing 5% CO2. The 3T3-L1 preadipocytes were cultured in minimum essential media (MEM) (Life Technologies) supplemented with 10% (v/v) FBS in a humidified atmosphere containing 5% CO2. For adipocyte differentiation, before viral transduction, cells were seeded at 0.5 × 105 per well on Geltrex-coated six-well plates (1:40, Life Technologies) in growth medium (day –2). On the next day (day –1), cells were transduced using the CytoTune-iPS ver. 1.0 Sendai Reprogramming Kit according to the manufacturer’s recommendations. At 24 h after transduction (day 0), cells were transferred to reprogramming media, that is, knockout DMEM (KO-DMEM) with 5% (v/v) knockout serum replacement, 15% (v/v) FBS, 1% (v/v) GlutaMAX solution, 1% (v/v) nonessential amino acids solution and 0.1 mM β-mercaptoethanol (all components obtained from Life Technologies). Using another conventional method for adipocyte differentiation, cells were exposed to adipogenic differentiation cocktails containing dexamethasone (1 μM), IBMX (0.5 mM), insulin (5 μg/ml) and 10% (v/v) FBS. The cells were maintained in reprogramming medium for 8 days beginning at day 0, and the media was exchanged every 48 h throughout all experiments (Figure 1a).
Total RNA extraction and qRT-PCR analysis
Total RNAs from cells were extracted using TRIzol (Life Technologies) and a Direct-zol RNA MiniPrep Kit (Zymo Research, Irvine, CA, USA) with DNase I according to the manufacturer’s recommendations. To perform the qRT-PCR assay, 400 ng of total RNAs was reverse-transcribed using the PrimeScript RT Reagent Kit and SYBR Premix Ex Taq (Takara Bio, Shiga, Japan) according to the manufacturer’s recommendations. qRT-PCR was performed using a Thermal Cycler Dice Real Time System using the default cycling program (Takara Bio). The primers used in this experiment are listed in Supplementary Table 2. The relative gene expression levels of mouse total heart RNAs (Takara Bio) or human iPSC RNAs were normalized to Gapdh expression.
Tissue preparation
Ten- to 12-week-old C57BL/6 mice were anesthetized and killed, and their hearts were removed at indicated time points. For total RNA and proteins extraction, the walls of the LV were dissociated from the whole heart. For total RNA isolation, the samples were cut into small pieces and homogenized with TRIzol using Bio Masher II (Nippi, Tokyo, Japan).
To isolate whole proteins, the samples were cut into small pieces and homogenized with lysis buffer (20 mM Tris-HCl (pH7.5), 137 mM NaCl, 10% glycerol (vol/vol), 1% NP-40 (vol/vol) (Wako Chemical Co.)), subsequently the lysates were sonicated with Bioruptor (CosmoBio Co. Ltd, Tokyo, Japan) for 4 min (30-s ON/30-s OFF) in ice-water.
For frozen sections, the kidneys were fixed with 4% paraformaldehyde (PFA; Wako Chemical Co.) for 2 h on ice, incubated overnight in 30% (vol/vol) sucrose in phosphate-buffered saline (PBS) at 4 °C and embedded in optimum cutting temperature compound (Sakura FineTek Japan Co., Ltd, Tokyo, Japan). Subsequently, 5-μm thick sections were cut.
Oil Red O staining and area calculation
Oil Red O powder (75 mg) was dissolved in 25 ml of 100% isopropyl alcohol and the solution was filtered to remove undissolved powder. PFA-fixed samples were washed with PBS and 60% (v/v) isopropyl alcohol. The samples were stained with 60% (v/v) Oil Red O solution for 15 min. Fat droplets in adipocytes were stained. Oil Red O-stained cells and frozen section samples were observed and images were captured with an IX71 inverted microscope (Olympus, Tokyo, Japan) or a BZ-X700 digital microscope (Keyence, Osaka, Japan). The percentage of total cell culture area positive for Oil Red O staining was calculated using ImageJ software (National Institutes of Health, Bethesda, MD, USA). At least three different wells were measured for each condition. For frozen sections, Mayer's hematoxylin was used as a counter-stain.
Hierarchical clustering, PC and GO analyses
Gene expression analysis was performed using a SurePrint G3 Mouse GE Microarray Kit 8 × 60 K (Agilent Technologies, Santa Clara, CA, USA). Raw data were normalized and analyzed using GeneSpring GX11 software (Agilent Technologies). These normalized data were analyzed using the NIA (National Institute on Aging) Array Analysis website (http://lgsun. grc.nia.nih.gov/ANOVA/),22 a web-based tool for microarray data analysis using hierarchical clustering of averages and PCA. A hierarchical clustering analysis was performed using a minimum distance value of 0.001, a separation ratio of 0.5, and the standard definition of the correlation distance. GO and KEGG pathway enrichments were evaluated statistically following the instructions provided by the Database for Annotation, Visualization and Integrated Discovery (DAVID) 6.7.57 The gene expression microarray data have been submitted to the GEO (Gene Expression Omnibus) online database (http://www.ncbi.nlm.nih.gov/geo/) under accession number GSE70088.
In vitro and in vivo IRI models
For the in vitro IRI model, CMPs were grown to 80% confluence and incubated in PBS for 3 h under hypoxic (1% O2, 5% CO2, balanced N2) conditions at 37 °C in a hypoxic chamber (ASTEC, Fukuoka, Japan), and subsequently incubated in growth media for 21 h under normoxic conditions (21% O2, 5% CO2). CMPs were collected at various time points (0, 3, 4, 6 and 24 h).58 Ten- to 12-week-old C57BL/6 mice were anesthetized by intraperitoneal injection of pentobarbital (50 mg/kg body weight) (Kyoritsu Seiyaku, Tokyo, Japan), and were intubated and ventilated under a respirator (SN-480-7, Shinano Manufacturing, Tokyo, Japan). General anesthesia was maintained by isoflurane. Following left thoracotomy, 7-0 Prolene suture thread was passed beneath the LAD just distal to the main trunk. The threads were tied transiently over a polyethylene tube for 60 min for ischemia, and were thereafter released for reperfusion. The LVs, including the areas at risk of IRI, were collected at various time points (0, 1, 1.5, 2 and 24 h) to examine gene profiles. At the end of the 24- h reperfusion period, Evans blue and TTC double staining was performed to verify IRI.59
Western blotting
Samples (50 μg) were mixed with bromophenol blue and 2-mercaptoethanol, boiled for 10 min, electrophoresed on 10% SDS polyacrylamide gel and electroblotted onto a PVDF transfer membrane (Millipore, Billerica, MA, USA). The membrane was blocked with PBS containing 5% skimmed milk, 0.05% Tween 20 and then incubated for 1 h with rabbit polyclonal antibodies to PPAR-γ (sc-7196; Santa Cruz Biotechnologies, Inc., Dallas, TX, USA), and mouse monoclonal antibodies to GAPDH (MAB374; Millipore), which were diluted to 1 : 500 with blocking buffer. After washing, the membrane was incubated with 1 : 5000 dilution of horseradish peroxidase (HRP)-conjugated donkey anti-rabbit IgG or HRP-conjugated donkey anti-mouse IgG (GE Healthcare, Little Chalfont, UK) in blocking buffer. Subsequently, the blots were developed using the ECL detection kit (GE Healthcare) and protein bands were visualized using the VersaDoc system (Bio-Rad Laboratories, Inc., Hercules, CA, USA).
Statistical analysis
Results are expressed as mean values±S.E. The statistical significance of differences between groups was evaluated using Student’s t-test, and P-values <0.05 were considered significant.
Accession codes
Abbreviations
- C/EBP:
-
CCAAT/enhancer-binding protein
- PPAR:
-
peroxisome proliferator-activated receptor
- Klf:
-
Krüppel-like factor
- KM:
-
KLF4 and c-MYC
- MSC:
-
mesenchymal stem cell
- TF:
-
transcription factor
- iPSC:
-
induced pluripotent stem cell
- OSKM:
-
Oct4, Sox2, Klf4 and c-Myc
- CMP:
-
cardiac mesenchymal progenitor
- IRI:
-
ischemic reperfusion injury
- MI:
-
myocardial infarction
- qRT-PCR:
-
quantitative reverse transcription polymerase chain reaction
- PCA:
-
principal component analysis
- PC:
-
principal component
- GO:
-
gene ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- LAD:
-
left anterior descending artery
- AON:
-
area of necrosis
- AAR:
-
area at risk
- RA:
-
remote area
- ROS:
-
reactive oxygen species
- LV:
-
left ventricular
- RAAS:
-
renin–angiotensin–aldosterone system
- NRX:
-
nucleoredoxin
- Dvl:
-
dishevelled
- bFGF:
-
basic fibroblast growth factor
- IBMX:
-
3-isobutyl-1-methylxanthine
- DMEM:
-
Dulbecco's modified Eagle’s medium
- FBS:
-
fetal bovine serum
- MEM:
-
minimum essential media
- KO-DMEM:
-
knockout DMEM
- PFA:
-
paraformaldehyde
- NIA array analysis:
-
National Institute on Aging array analysis
- DAVID:
-
Database for Annotation, Visualization and Integrated Discovery
- GEO:
-
Gene Expression Omnibus
- TTC:
-
2,3,5-triphenyltetrazolium chloride
References
Lefterova MI, Lazar MA . New developments in adipogenesis. Trends Endocrinol Metab 2009; 20: 107–114.
Pulverer B, Sommer A, McArthur GA, Eisenman RN, Luscher B . Analysis of Myc/Max/Mad network members in adipogenesis: inhibition of the proliferative burst and differentiation by ectopically expressed Mad1. J Cell Physiol 2000; 183: 399–410.
Birsoy K, Chen Z, Friedman J . Transcriptional regulation of adipogenesis by KLF4. Cell Metab 2008; 7: 339–347.
Buganim Y, Faddah DA, Jaenisch R . Mechanisms and models of somatic cell reprogramming. Nat Rev Genet 2013; 14: 427–439.
Pittenger MF, Mackay AM, Beck SC, Jaiswal RK, Douglas R, Mosca JD et al. Multilineage potential of adult human mesenchymal stem cells. Science 1999; 284: 143–147.
Dominici M, Le Blanc K, Mueller I, Slaper-Cortenbach I, Marini F, Krause D et al. Minimal criteria for defining multipotent mesenchymal stromal cells. The International Society for Cellular Therapy position statement. Cytotherapy 2006; 8: 315–317.
Kern S, Eichler H, Stoeve J, Kluter H, Bieback K . Comparative analysis of mesenchymal stem cells from bone marrow, umbilical cord blood, or adipose tissue. Stem Cells 2006; 24: 1294–1301.
Goumans MJ, de Boer TP, Smits AM, van Laake LW, van Vliet P, Metz CH et al. TGF-beta1 induces efficient differentiation of human cardiomyocyte progenitor cells into functional cardiomyocytes in vitro. Stem Cell Res 2007; 1: 138–149.
Kami D, Kitani T, Nakata M, Gojo S . Cardiac mesenchymal progenitors from postmortem cardiac tissues retained cellular characterization. Transplant Proc 2014; 46: 1194–1197.
Al-Nbaheen M, Vishnubalaji R, Ali D, Bouslimi A, Al-Jassir F, Megges M et al. Human stromal (mesenchymal) stem cells from bone marrow, adipose tissue and skin exhibit differences in molecular phenotype and differentiation potential. Stem Cell Rev 2013; 9: 32–43.
Baroldi G, Silver MD, De Maria R, Parodi O, Pellegrini A . Lipomatous metaplasia in left ventricular scar. Can J Cardiol 1997; 13: 65–71.
Su L, Siegel JE, Fishbein MC . Adipose tissue in myocardial infarction. Cardiovasc Pathol 2004; 13: 98–102.
Ichikawa Y, Kitagawa K, Chino S, Ishida M, Matsuoka K, Tanigawa T et al. Adipose tissue detected by multislice computed tomography in patients after myocardial infarction. JACC Cardiovasc Imaging 2009; 2: 548–555.
Goldfarb JW, Roth M, Han J . Myocardial fat deposition after left ventricular myocardial infarction: assessment by using MR water-fat separation imaging. Radiology 2009; 253: 65–73.
Takahashi K, Yamanaka S . Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006; 126: 663–676.
Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 2007; 131: 861–872.
Vierbuchen T, Wernig M . Direct lineage conversions: unnatural but useful? Nat Biotechnol 2011; 29: 892–907.
Ieda M, Fu JD, Delgado-Olguin P, Vedantham V, Hayashi Y, Bruneau BG et al. Direct reprogramming of fibroblasts into functional cardiomyocytes by defined factors. Cell 2010; 142: 375–386.
Goding CR, Pei D, Lu X . Cancer: pathological nuclear reprogramming? Nat Rev Cancer 2014; 14: 568–573.
Ohnishi K, Semi K, Yamamoto T, Shimizu M, Tanaka A, Mitsunaga K et al. Premature termination of reprogramming in vivo leads to cancer development through altered epigenetic regulation. Cell 2014; 156: 663–677.
Efe JA, Hilcove S, Kim J, Zhou H, Ouyang K, Wang G et al. Conversion of mouse fibroblasts into cardiomyocytes using a direct reprogramming strategy. Nat Cell Biol 2011; 13: 215–222.
Sharov AA, Dudekula DB, Ko MS . A web-based tool for principal component and significance analysis of microarray data. Bioinformatics 2005; 21: 2548–2549.
Ntambi JM, Young-Cheul K . Adipocyte differentiation and gene expression. J Nutr 2000; 130: 3122S–3126S.
Jeong HJ, Park SW, Kim H, Park SK, Yoon D . Coculture with BJ fibroblast cells inhibits the adipogenesis and lipogenesis in 3T3-L1 cells. Biochem Biophys Res Commun 2010; 392: 520–525.
Rosen ED, Hsu CH, Wang X, Sakai S, Freeman MW, Gonzalez FJ et al. C/EBPalpha induces adipogenesis through PPARgamma: a unified pathway. Genes Dev 2002; 16: 22–26.
Tontonoz P, Hu E, Spiegelman BM . Stimulation of adipogenesis in fibroblasts by PPAR gamma 2, a lipid-activated transcription factor. Cell 1994; 79: 1147–1156.
Yamanaka S . Strategies and new developments in the generation of patient-specific pluripotent stem cells. Cell Stem Cell 2007; 1: 39–49.
Rowland BD, Bernards R, Peeper DS . The KLF4 tumour suppressor is a transcriptional repressor of p53 that acts as a context-dependent oncogene. Nat Cell Biol 2005; 7: 1074–1082.
Hunt CR, Ro JH, Dobson DE, Min HY, Spiegelman BM . Adipocyte P2 gene: developmental expression and homology of 5'-flanking sequences among fat cell-specific genes. Proc Natl Acad Sci USA 1986; 83: 3786–3790.
Maxwell PH, Wiesener MS, Chang G-W, Clifford SC, Vaux EC, Cockman ME et al. The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature 1999; 399: 271–275.
Iavarone C, Catania A, Marinissen MJ, Visconti R, Acunzo M, Tarantino C et al. The platelet-derived growth factor controls c-myc expression through a JNK- and AP-1-dependent signaling pathway. J Biol Chem 2003; 278: 50024–50030.
Sarjeant K, Stephens JM . Adipogenesis. Cold Spring Harb Perspect Biol 2012; 4: a008417.
Umezawa A, Maruyama T, Segawa K, Shadduck RK, Waheed A, Hata J . Multipotent marrow stromal cell line is able to induce hematopoiesis in vivo. J Cell Physiol 1992; 151: 197–205.
Kuri-Harcuch W, Green H . Adipose conversion of 3T3 cells depends on a serum factor. Proc Natl Acad Sci USA 1978; 75: 6107–6109.
Green H, Meuth M . An established pre-adipose cell line and its differentiation in culture. Cell 1974; 3: 127–133.
Cristancho AG, Lazar MA . Forming functional fat: a growing understanding of adipocyte differentiation. Nat Rev Mol Cell Biol 2011; 12: 722–734.
Siersbaek R, Nielsen R, Mandrup S . Transcriptional networks and chromatin remodeling controlling adipogenesis. Trends Endocrin Metab 2012; 23: 56–64.
Ali AT, Hochfeld WE, Myburgh R, Pepper MS . Adipocyte and adipogenesis. Eur J Cell Biol 2013; 92: 229–236.
Niwa H, Ogawa K, Shimosato D, Adachi K . A parallel circuit of LIF signalling pathways maintains pluripotency of mouse ES cells. Nature 2009; 460: 118–122.
Simon AR, Rai U, Fanburg BL, Cochran BH . Activation of the JAK-STAT pathway by reactive oxygen species. Am J Physiol Cell Physiol 1998; 275: C1640–C1652.
McConnell BB, Yang VW . Mammalian Kruppel-like factors in health and diseases. Physiol Rev 2010; 90: 1337–1381.
Freytag SO . Enforced expression of the c-myc oncogene inhibits cell differentiation by precluding entry into a distinct predifferentiation state in G0/G1. Mol Cell Biol 1988; 8: 1614–1624.
Nie Z, Hu G, Wei G, Cui K, Yamane A, Resch W et al. c-Myc is a universal amplifier of expressed genes in lymphocytes and embryonic stem cells. Cell 2012; 151: 68–79.
Rodeheffer MS, Birsoy K, Friedman JM . Identification of white adipocyte progenitor cells in vivo. Cell 2008; 135: 240–249.
Uezumi A, Fukada S, Yamamoto N, Takeda S, Tsuchida K . Mesenchymal progenitors distinct from satellite cells contribute to ectopic fat cell formation in skeletal muscle. Nat Cell Biol 2010; 12: 143–152.
Bolli R, Chugh AR, D'Amario D, Loughran JH, Stoddard MF, Ikram S et al. Cardiac stem cells in patients with ischaemic cardiomyopathy (SCIPIO): initial results of a randomised phase 1 trial. Lancet 2011; 378: 1847–1857.
Makkar RR, Smith RR, Cheng K, Malliaras K, Thomson LE, Berman D et al. Intracoronary cardiosphere-derived cells for heart regeneration after myocardial infarction (CADUCEUS): a prospective, randomised phase 1 trial. Lancet 2012; 379: 895–904.
Smits AM, van Vliet P, Metz CH, Korfage T, Sluijter JP, Doevendans PA et al. Human cardiomyocyte progenitor cells differentiate into functional mature cardiomyocytes: an in vitro model for studying human cardiac physiology and pathophysiology. Nat Protocols 2009; 4: 232–243.
Pouliopoulos J, Chik WW, Kanthan A, Sivagangabalan G, Barry MA, Fahmy PN et al. Intramyocardial adiposity post-myocardial infarction: new implications of a substrate for ventricular tachycardia. Circulation 2013; 128: 2296–2308.
Hou X, Zhang Y, Shen YH, Liu T, Song S, Cui L et al. PPAR-gamma activation by rosiglitazone suppresses angiotensin II-mediated proliferation and phenotypictransition in cardiac fibroblasts via inhibition of activation of activator protein 1. Eur J Pharmacol 2013; 715: 196–203.
Wang C, Mayer JA, Mazumdar A, Fertuck K, Kim H, Brown M et al. Estrogen induces c-myc gene expression via an upstream enhancer activated by the estrogen receptor and the AP-1 transcription factor. Mol Endocrinol 2011; 25: 1527–1538.
Zhang Y, Wang Y, Liu Y, Wang N, Qi Y, Du J . Kruppel-like factor 4 transcriptionally regulates TGF-beta1 and contributes to cardiac myofibroblast differentiation. PloS One 2013; 8: e63424.
Gabrielsen A, Lawler PR, Yongzhong W, Steinbruchel D, Blagoja D, Paulsson-Berne G et al. Gene expression signals involved in ischemic injury, extracellular matrix composition and fibrosis defined by global mRNA profiling of the human left ventricular myocardium. J Mol Cell Cardiol 2007; 42: 870–883.
Boengler K, Hilfiker-Kleiner D, Drexler H, Heusch G, Schulz R . The myocardial JAK/STAT pathway: from protection to failure. Pharmacol Ther 2008; 120: 172–185.
Holmstrom KM, Finkel T . Cellular mechanisms and physiological consequences of redox-dependent signalling. Nat Rev Mol Cell Biol 2014; 15: 411–421.
Funato Y, Michiue T, Asashima M, Miki H . The thioredoxin-related redox-regulating protein nucleoredoxin inhibits Wnt-beta-catenin signalling through dishevelled. Nat Cell Biol 2006; 8: 501–508.
Huang, da W, Sherman BT, Lempicki RA . Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Prot 2009; 4: 44–57.
Fukai K, Nakamura A, Hoshino A, Nakanishi N, Okawa Y, Ariyoshi M et al. Pyk2 aggravates hypoxia-induced pulmonary hypertension by activating HIF-1alpha. Am J Physiol Heart Circ Physiol 2015; 308: H951–H959.
Michael LH, Entman ML, Hartley CJ, Youker KA, Zhu J, Hall SR et al. Myocardial ischemia and reperfusion: a murine model. Am J Physiol 1995; 269 (6 Pt 2): H2147–H2154.
Acknowledgements
We would like to express our sincere thanks to Toyoda Masashi. (Tokyo Metropolitan Institute of Gerontology) for helpful discussions regarding the results presented in the manuscript. This study was supported by a Grant-in-Aid for Exploratory Research from JSPS KAKENHI 24659594.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by D Aberdam
Supplementary Information accompanies this paper on Cell Death and Disease website
Rights and permissions
Cell Death and Disease is an open-access journal published by Nature Publishing Group. This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kami, D., Kitani, T., Kawasaki, T. et al. Cardiac mesenchymal progenitors differentiate into adipocytes via Klf4 and c-Myc. Cell Death Dis 7, e2190 (2016). https://doi.org/10.1038/cddis.2016.31
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2016.31
- Springer Nature Limited
This article is cited by
-
Dual Specific Phosphatase 7 Exacerbates Dilated Cardiomyopathy, Heart Failure, and Cardiac Death by Inactivating the ERK1/2 Signaling Pathway
Journal of Cardiovascular Translational Research (2022)