Abstract
The integrity of the genome is maintained by a host of surveillance and repair mechanisms that are pivotal for cellular function. The tumour suppressor protein p53 is a major component of the DNA damage response pathway and plays a vital role in the maintenance of cell-cycle checkpoints. Here we show that a microRNA, miR-486, and its host gene ankyrin-1 (ANK1) are induced by p53 following DNA damage. Strikingly, the cytoskeleton adaptor protein ankyrin-1 was induced over 80-fold following DNA damage. ANK1 is upregulated in response to a variety of DNA damage agents in a range of cell types. We demonstrate that miR-486-5p is involved in controlling G1/S transition following DNA damage, whereas the induction of the ankyrin-1 protein alters the structure of the actin cytoskeleton and sustains limited cell migration during DNA damage. Importantly, we found that higher ANK1 expression correlates with decreased survival in cancer patients. Thus, these observations highlight ANK1 as an important effector downstream of the p53 pathway.
Similar content being viewed by others
Main
Cells respond to DNA damage by orchestrating a series of events, either resulting in cell-cycle arrest and DNA repair, or elimination of the damaged cell. DNA double-strand breaks (DSB) are one of the most toxic types of DNA damage experienced by cells.1 A complex network of mechanisms has evolved to detect and repair DSBs. DNA repair is achieved either via non-homologous end-joining or in a more accurate manner by homologous recombination.2 Failure of either of these mechanisms triggers apoptosis.3
The DNA damage response pathway (DDR) involves a cascade of events with multiple effector components,3, 4, 5, 6, 7 key to which is the tumour suppressor protein p53.8 DNA damage leads to stabilisation of p53 resulting from the degradation of its ubiquitin ligase, MDM2.9 This induces the transcription of genes whose products trigger cell-cycle arrest, DNA repair or apoptosis.7 More recently, p53 has been shown to regulate certain microRNAs (miRNA) by facilitating their transcription or modulating the activity of the miRNA biogenesis machinery.10
MiRNAs are ~22 nt RNA molecules which base pair with target mRNAs resulting in translation inhibition and destabilisation of the bound mRNA.11 These small RNAs are involved in a range of biological processes and regulate more than half of all protein-coding genes.11 For example, the transcriptional activation of the miR-34 family by p53 following DNA damage12 results in the inhibition of key targets, including the transcription factor c-Myc which controls genes involved in cell-cycle progression and cell growth.13, 14
In addition to its roles in cell death, p53 has also been implicated in cell motility,15 and mutant p53 promotes tumour cell invasion and results in loss of directionality during migration.16 Cell migration is a complex process and is controlled by many proteins,17 and the specific role of p53 in this mechanism is not yet completely understood.
Here, we initially set out to identify new miRNAs associated with DDR. We found miR-486-5p levels increased ~8-fold following DNA damage, and to our surprise, found the host gene ANK1 increased ~80-fold. We show that both miR-486 and ANK1 are regulated by p53 and that miR-486-5p is involved in controlling G1/S transition following DNA damage. On the other hand, ankyrin-1 plays a role in sustaining cell motility through actin cytoskeleton remodelling upon non-apoptotic levels of DNA damage. Importantly, we found that high ANK1 levels correlate with reduced survival in cancer patients.
Results
Identification of miRNAs upregulated following DNA damage
To identify miRNAs that change following DNA damage, we treated the non-tumorigenic MCF10A cell line with the DNA topoisomerase II inhibitor doxorubicin to induce DSBs18 (Figure 1a) and subjected them to small RNA deep sequencing (Figure 1b). Induction of histone H2A.X phosphorylation (γ-H2AX, Ser139) and p53 phosphorylation (Ser15) confirmed the activation of DDR.4 The dose and duration of damage were chosen to avoid apoptotic induction, confirmed by the absence of PARP cleavage (Figure 1a and Supplementary Figure S1).
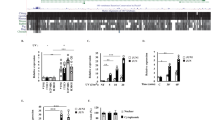
Identification of microRNAs upregulated following DNA damage. (a) DNA damage was induced in MCF10A cells with doxorubicin (Dox.) at a concentration of 400 nM for 8 h, and was confirmed by western blots for phospho-p53 (Ser15) and γ-H2AX. Absence of cleaved PARP (89 kDa) confirmed cells were not undergoing apoptosis. Representative western blots shown (n=3). (b) Small RNA deep sequencing (Illumina) was carried out on total RNA from control and DNA damaged MCF10A cells. MicroRNAs with a significant fold-change (P<0.05) (n=3) are shown, with normalised reads displayed as reads per million (RPM). (c) Validation of miR-486-5p upregulation following DNA damage in MCF10A cells that had been exposed to doxorubicin (400 nM) for 0–48 h. MiR-486-5p levels were quantified via qPCR, as well as miR-34 family members as a comparison. Values are mean±S.D. (t-test, n=3, compared with 0 h control), *P<0.05, **P<0.01 and ***P<0.001; NS, not significant. Corresponding western blots are shown in Figure 2c
Analysis revealed six miRNAs that showed significant upregulation following damage (P<0.05), all of which showed >2-fold change (Figure 1b). Two miRNAs were downregulated ~3-fold (P<0.05) (miR-92a-1-5p and miR-509-3p). Among the upregulated miRNAs were members of the miR-34 family, including miR-34c and miR-34a which are known to be induced following DNA damage;19, 20, 21 miR-143-3p which has been previously implicated in DDR;22 and miR-200a which has a well-known role in cancer metastasis23 and DNA damage checkpoint control.24 Importantly, we also found that miR-486-5p increased by 3.8-fold following DNA damage and was relatively abundant, suggesting a possible functional role in DDR. MiR-486 is a muscle-enriched miRNA, but relatively little is known about its involvement in DDR.25
To validate these results and determine the temporal nature of miR-486-5p induction, we performed a doxorubicin time-course using MCF10A cells. MiR-34a and miR-34c were used as positive controls (Figure 1c). As expected, miR-34a increased 5.4-fold after 24 h of doxorubicin treatment and 8.0-fold after 48 h. MiR-34c increased 21.5-fold after 24 h of doxorubicin treatment and remained high at 48 h. A significant 1.5-fold increase in miR-486-5p was detectable as early as 4 h following doxorubicin exposure, and increased to 8.3-fold after 48 h. We therefore selected miR-486 for further investigation.
The miR-486 host gene ANK1 is also induced following DNA damage
Approximately half of the 2588 miRNAs in the human genome are intragenic,26 and there is often a functional relationship between the miRNA and its host gene.27 Intragenic miRNAs can be regulated either by the host gene’s promoter or an independent promoter.28 MiR-486 is located within the last intron of the cytoskeleton adaptor gene ANK1 (Figure 2a). Therefore we asked if the primary host gene transcript, ANK1, was also induced following DNA damage. It was previously shown that miR-486 is controlled by a p53 promoter located 21.4 kb upstream of ANK1,29 but to our knowledge no one had previously examined the role of ANK1 in relation to DNA damage, miR-486 or activation of the p53 pathway.
ANK1 is upregulated following exposure to different inducers of DNA damage and in a variety of cell types. (a) Diagram showing the location of miR-486 in relation to the open reading frame of the cytoskeleton adaptor gene ANK1, as well as the small ankyrin-1 isoform (sAnk1). MiR-486 is located in the intronic region between the last two exons of the ANK1 gene. (b) MCF10A cells were treated with doxorubicin (dox.) as in Figure 1c, and ANK1 mRNA levels were quantified by qPCR. The p53-regulated p21(Waf1/ Cip1) transcript served as a positive control (n=3). (c) Representative western analysis of protein samples harvested from cells undergoing DNA damage (n=3). The ankyrin-1 protein (ANK1) (246 kDa) was quantified (right panel) and normalised to β-actin levels. (d) Ankyrin-1 was induced following ionising radiation (IR, 10 Gy) exposure in MCF10A cells but not by ultraviolet light-C (UVC, 100 J/m2). Cells were harvested 24 h after IR/UVC exposure. Cisplatin (10 μg/ml) and mitomycin C (300 nM) treatment for 24 h also induced ankyrin-1. Representative western blots are shown (n=3). (e) Ankyrin-1 was upregulated following 25 μM etoposide (etop.) treatment for 24 h in retinal pigment epithelium (RPE) and MCF7 cells, and in A549 cells following IR (5 Gy) exposure where cells were harvested after 6 h. Representative western blots are shown (n=3). For all bar charts, values are mean±S.D. (t-test, n=3, compared with 0 h control), *P<0.05, **P<0.01 and ***P<0.001; NS, not significant
Strikingly, we found that ANK1 mRNA was upregulated 16-fold after 8 h of doxorubicin-induced DNA damage (Figure 2b) and 110-fold after 16 h, which was markedly higher than the increase in miR-486-5p expression (Figure 1c). To compare this with a well-known DNA damage-induced transcript, we measured mRNA levels of the p53-regulated gene p21Waf1/Cip1, which showed a 25-fold increase following 16 h of doxorubicin treatment (Figure 2b). The dramatic induction of ANK1 mRNA expression levels led us to investigate whether the ankyrin-1 protein was similarly induced. Indeed, we observed that the 246 kDa ankyrin-1 protein was induced ~31-fold after 16 h of doxorubicin treatment (Figure 2c), and ~76-fold after 48 h. Importantly, this was restricted to the canonical ankyrin-1 protein, as the shorter, muscle-enriched isoform of ankyrin-1 (sAnk1) only increased by ~2-fold after 24 h of treatment (Figure 2c).
Next we examined if ankyrin-1 levels increased following exposure to other DNA damage inducers. We found that ionising radiation (IR) exposure induced ankyrin-1 expression in MCF10A cells (Figure 2d), despite low levels of the DNA damage marker γ-H2AX. Ankyrin-1 levels were also induced by the DNA crosslinking reagents cisplatin and mitomycin C (Figure 2d), and also by a different DNA topoisomerase II inhibitor, etoposide (Figure 3c). Ultraviolet-C irradiation did not markedly induce p53 phosphorylation or ANK1 expression, although high levels of γ-H2AX were observed (Figure 2d).
ANK1 and miR-486 expression are p53 dependent. (a) MCF10A cells were subjected to DNA damage using 400 nM doxorubicin (dox.) for 24 h following 48 h siRNA-mediated depletion of p53 or a control siRNA. ANK1 and miR-486-5p RNA levels were determined by qPCR and representative western blots show ankyrin-1 (ANK1), γ-H2AX and phospho-p53 (Ser15) levels following DNA damage and p53 depletion (n=3). (b) Doxycycline inducible wild-type p53 or mutant p53R175H H1299 cells were either treated (induced) or not treated with doxycycline. Wild type or mutant p53 expression was confirmed via western blot of p53 and p21(Waf1/ Cip1), and ANK1 and miR-486-5p RNA levels were determined by qPCR (n=3). (c) Cells were treated with the p53 activator, nutlin. Total p53 and ankyrin-1 levels were analysed by western blot and compared with DNA damage-induced cells treated with the topoisomerase II inhibitor etoposide (etop.) (25 μM) for 24 h. Representative western blots are shown (n=3). For bar charts, values are mean±S.D. (t-test, n=3), *P<0.05, **P<0.01 and ***P<0.001
To examine if the upregulation of ankyrin-1 was specific to MCF10A cells, we treated another non-tumorigenic cell line, retinal pigment epithelium (RPE) cells, and also breast cancer-derived MCF7 cells with etoposide18 (Figure 2e). Both RPE and MCF7 cells showed a marked increase of ankyrin-1 protein levels following DNA damage. Ankyrin-1 was also induced by IR exposure in A549 lung carcinoma cells (Figure 2e).
Taken together, these data demonstrated that ANK1 was induced in response to various DNA damaging agents in different cell lines, suggesting an important role for ANK1 in DDR activation.
ANK1 and miR-486 expression are p53 dependent
To investigate if the transcription factor p53 was co-regulating both miR-486 and ANK1 following DNA damage, we depleted p53 with siRNA prior to doxorubicin treatment (Figure 3a). In control cells, DNA damage induced miR-486-5p expression by 11.6-fold and ANK1 expression by 26.5-fold. However, knockdown of p53 impaired the induction of both miR-486-5p and ANK1 (Figure 3a).
We then used p53-null cells that harbour either a doxycycline-inducible wild-type p53 or dominant-negative mutant p53R175H construct (Figure 3b).30 Upon induction of wild-type p53 expression with doxycycline, a 3.7-fold induction of miR-486-5p and a 4.9-fold increase in ANK1 mRNA were observed. Importantly, ankyrin-1 was also induced at the protein level (Figure 3b). As expected, expression of mutant p53R175H did not induce p21Waf1/Cip1. Likewise, both miR-486-5p and ANK1 were not induced by mutant p53, demonstrating that only wild-type p53 could induce their expression.
Finally, we asked whether MCF10A cells treated with the p53 activator nutlin31 could induce ANK1 expression independently of DNA damage. In control cells ankyrin-1 was barely detectable (Figure 3c); however, its expression was induced following treatment with etoposide or nutlin. The stabilisation of p53 by nutlin was confirmed by MDM2 and p53 western analysis (Figure 3c).
Taken together, these data strongly suggest that both miR-486 and ANK1 are induced following DNA damage in a p53-dependent manner.
miR-486-5p inhibition reverses DNA damage-induced cell-cycle arrest
Previously it has been reported that miR-486 expression can inhibit cell growth resulting in cell-cycle arrest in G1.25, 29, 32, 33 However, these publications did not examine the role of miR-486 following DNA damage in conjunction with the role of the host gene ANK1 in cell-cycle control. To clarify this, we first set out to establish the individual contributions that increased ankyrin-1 or miR-486-5p levels had on cells following DNA damage. We used siRNA to deplete ANK1 and anti-miRs to inhibit miR-486-5p (Figures 4a and b). We confirmed that siRNA depletion of ANK1 resulted in the loss of ankyrin-1 induction following DNA damage (Figure 4a), but did not affect the induction of miR-486-5p (Figure 4b). Likewise, the inhibition of miR-486-5p resulted in ablation of the miRNA induction post-DNA damage (Figure 4b), but did not significantly affect the induction of ankyrin-1 protein (Figure 4a). Now that we could separate the contributions of the mRNA and miRNA components, we examined the individual effects of ANK1 and miR-486-5p on cell-cycle arrest following DNA damage (Figure 4c). We used etoposide treatment for the fluorescence-activated cell sorting (FACS) experiments because doxorubicin and propidium iodide have overlapping fluorescence emission wavelengths. In control siRNA-transfected cells treated with etoposide, we observed a significant decrease in the G1 population, with a corresponding increase in G2, implying G2/M arrest. Activation of DDR was confirmed by detection of phospho-ATM, phospho-p53 and γ-H2A.X (Figure 4a). The depletion of ankyrin-1 following DNA damage (Figure 4a) did not have any effect on the proportion of cells arrested in G2 (Figure 4c). This suggests that DNA damage-induced G2 arrest was ANK1-independent. Importantly, the inhibition of miR-486-5p in cells treated with etoposide resulted in a reduction of cells in G2/M (Figure 4c).
miR-486-5p inhibition partly reverses the DNA damage-induced accumulation of cells in G2. (a) Cells were either transfected with a negative control siRNA, an siRNA against ANK1, a negative control 2′-O-methyl inhibitor or a 2′-O-methyl inhibitor against miR-486-5p. After 24 h, DNA damage was induced using 25 μM etoposide (etop.) for 24 h. DNA damage was assessed by p-ATM, p-p53 (Ser15) and γ-H2AX while a lack of cleaved caspase-3 and PARP showed that the treatment had not induced apoptosis. Representative western blots are shown (n=3). (b) miR-486-5p levels were quantified via qPCR (n=3). (c) Cell-cycle analysis was achieved via propidium iodide staining and subsequent flow cytometry. For all bar charts, values are mean±S.D. (t-test, n=3), **P<0.01; NS, not significant
M-phase arrest induced after nocodazole treatment was used to confirm that the increase in the G1/S population is not caused by cells escaping G2/M arrest and re-entering the cell cycle. Nocodazole-treated cells had a higher proportion of cells remaining in G1 following miR-486-5p inhibition, with or without DNA damage (Supplementary Figure S2). This suggests that miR-486 contributes to G1/S cell-cycle checkpoint control.
Ankyrin proteins are composed of three parts:34, 35 an N-terminal membrane-binding region containing multiple ankyrin repeat domains, a central spectrin-binding region and a C-terminal regulatory region. Examination of the ankyrin-1 protein sequence revealed a death domain within the C-terminus (Supplementary Figure S3). Death domains are known to interact with components of the apoptosis pathway.36 We therefore examined if the induction of ankyrin-1 protein could induce apoptosis. We did not observe any cleaved caspase-3 following either induction or siRNA depletion of ANK1 (Figure 4a). Similarly, miR-486 expression in the presence or absence of etoposide treatment did not induce caspase-3 activation despite previous work, implicating miR-486 in augmenting DDR.25 To investigate further, we conducted caspase-3/7 activity assays using cell lysates and annexin V-FITC staining of live cells, but failed to observe any increase in caspase activity or apoptosis (data not shown). Similarly, experiments to test if increased expression of the death domain-containing ankyrin-1 protein might change the ability of cells to engage the extrinsic apoptosis pathways37 did not show any effects on cell death (data not shown).
Taken together, these data suggest miR-486-5p is involved in maintaining cell-cycle arrest following DNA damage. However, increased ankyrin-1 levels do not appear to affect regulation of the cell-cycle or apoptosis in damaged cells. Owing to the marked induction of ankyrin-1 protein following DNA damage, we decided to further explore other functions of ankyrin-1 following DNA damage.
Ankyrin-1 depletion inhibits cell migration and affects the formation of actin-rich plasma membrane protrusions following DNA damage
Upon DNA damage, cell migration is decreased38, 39 but is actively maintained through ATM signalling.40 Ankyrin-1 functions as a membrane adaptor protein, connecting cell membrane proteins to the spectrin-actin cytoskeleton, thus implicating it in cell migration.41 Moreover, it is well-known that p53 can inhibit cell motility through regulation of cell morphology, cell spreading and Rho signalling.15 Given that ANK1 is controlled by the p53 pathway, we investigated whether ANK1 plays a role in modulating cell motility. To this end, we performed wound-healing assays on cells treated with either control or ANK1 siRNA, with or without etoposide (Figure 5a). Cells that had not been exposed to DNA damage underwent wound closure at the same rate in the presence or absence of ANK1 siRNA (Figures 5b–d). As expected, cells treated with etoposide did not achieve full wound closure (Figures 5b–d). Surprisingly, cells that lacked ankyrin-1 were even more severely affected upon etoposide treatment. For example, at 36 h after wound generation, ANK1-expressing cells treated with etoposide achieved 68.6% scratch wound closure compared with 38.3% in the absence of ankyrin-1 (Figure 5b). Ankyrin-1 depletion also impaired cell motility in a transwell migration assay (Supplementary Figure S4). These data suggested that ankyrin-1 has a role in sustaining cell movement following DNA damage. Importantly, videos of wound healing assays (Supplementary Movies) confirmed impaired movement of ANK1-depleted cells following DNA damage, showing lamellipodia ruffling which is indicative of inhibited actin filament turnover.42 Time-lapse videos also revealed that DNA damaged cells depleted of ankyrin-1 displayed rapid extension and retraction of membrane protrusions with very little movement of the cell body. The lack of directional movement of these cells was demonstrated by cell tracking assays, which depict the movement patterns of individual cells (Figure 5d).
Ankyrin-1 depletion inhibits cell migration following DNA damage. (a) siRNA was used to deplete ANK1, or cells were treated with a negative control siRNA. Following siRNA treatment, DNA damage was induced with etoposide (etop.) (25 μM) for 24 h. Representative western blots are shown (n=3). (b) Wound healing assays (scratch assay) was performed on confluent MCF10A cells and migration into the wound was monitored for 48 h using time-lapse microscopy. Values are mean±S.D. (t-test, n=3), *P<0.05. (c) Representative images of MCF10A scratch assays at the initial time of scratch (0 h) and after 24 and 48 h (n=3). Scale bar represents 200 μm. (d) Representative cell tracking assays taken from 10 randomly selected cells from each condition undergoing wound healing
Intrigued by these observations, we proceeded to examine the cytoskeleton, specifically looking at the morphology of actin filaments in response to DNA damage in the presence or absence of ankyrin-1. Interestingly, it was observed that control cells treated with etoposide had longer, more prevalent actin-rich plasma membrane protrusions compared with etoposide-treated cells following ankyrin-1 depletion (Figure 6a). Quantification revealed that the removal of ANK1 by siRNA decreased the proportion of cells with long actin protrusions following DNA damage from 27.1% to 8.8% (Figures 6b–d). These data suggested that ankyrin-1 may play a role in regulating actin cytoskeleton structure following DNA damage.
Expression of ANK1 following DNA damage affects actin-rich plasma membrane protrusions. (a) Representative immunofluorescent confocal images of cells treated with either control or ANK1-specific siRNAs, followed by etoposide (etop.) (25 μM) treatment for 24 h to induce DNA damage (n=3). Actin was stained using Alexa Fluor 488 phalloidin (green) and nuclei were stained using DAPI (blue). White arrows point to actin-rich plasma membrane protrusions. Scale bar represents 50 μm. (b) Long actin processes protruding from cells (examples indicated in (a) with white arrows) were measured on 80 randomly selected cells from each condition. Only cells that were not in colonies were recorded. Measurements were taken from three independent repeat (rep.) experiments and actin protrusion length from the cell membrane was plotted by size (longest to shortest) for each condition in each repeat experiment, with cells lacking any protrusions being represented by a blank space (a length of zero). (c) Percentage of cells with actin protrusions that were greater than 50 units (arbitrary) from each condition. Values are mean±S.D. (t-test, n=3), **P<0.01. (d) Representative western blots (n=3) of proteins from cells used in confocal microscopy, showing ankyrin-1 (ANK-1) levels following DNA damage, with or without siRNA depletion
Actin remodelling factors are modulated by ankyrin-1
As ankyrin-1 appeared to affect actin filament structure and cell motility, we set out to identify how upregulation of ankyrin-1 regulates actin remodelling. It is possible that increased ankyrin-1 levels alone could inhibit actin filament organisation through increased spectrin-actin binding or, alternatively, ankyrin-1 could act to modulate actin remodelling signalling pathways.
Since ankyrin repeats (ANK) have been shown to facilitate nuclear import for some proteins,43 we first investigated the localisation of ankyrin-1 protein following DNA damage. We found that the ankyrin-1 protein was present predominantly in the cytoplasmic compartment (Figure 7a). We then investigated the Rho GTPase signalling pathway, which controls the remodelling of the actin cytoskeleton.17 To ensure the changes in this pathway were specific to increased levels of ankyrin-1 and not due to other events triggered by DNA damage, we overexpressed ANK1 in MCF10A cells (Figure 7b). We observed a strong reduction in PAK1 levels and also a reduction in the phosphorylation of the actin-severing protein cofilin (Figures 7b and c). Both of these proteins are key regulators of actin remodelling.17 Examination of the tumour suppressor p27Kip1, which has recently been implicated in cell migration by regulating RhoA,44 revealed that it too was significantly reduced upon ANK1 overexpression (Figures 7b and c).
Actin remodelling factors are modulated by ankyrin-1 levels. (a) Western blot analysis of cells treated with control vehicle or 25 μM etoposide (etop.) for 24 h. MCF10A cells were fractionated into cytoplasmic and nuclear fractions to show ankyrin-1 (ANK1) is localised to the cytoplasm following DNA damage (n=3). (b) Representative western blot analysis showing levels of actin cytoskeleton regulators following ANK1 overexpression (n=3). MCF10A cells were nucleofected with an ANK1 expression construct or with an empty vector control. Proteins involved in the actin remodelling pathway were analysed following 48 h of expression. (c) Western blot quantification of p27Kip1, p-cofilin and PAK1 for experiments shown in (b) following normalisation to β-actin levels. Values are mean±S.D. (t-test, n=3), **P<0.01 and ***P<0.001
The impact of ANK1 expression in cancer patients
To understand the potential clinical relevance of ANK1 expression levels, we analysed gene expression data from cancer patients using a database tool.45 This analysis interrogates gene expression data and identifies correlations with patient outcome. Since the p53 pathway controls ANK1 expression, we divided the patient cohort into two groups: the first, where a positive correlation between TP53 and ANK1 expression is present, and the second, where it is absent or negative. This analysis allowed us to ask if the link between TP53 and ANK1 was significant in a clinical setting. Importantly, we found that patients with a high positive correlation between TP53 and ANK1 showed increased survival (Supplementary Figure S5).
Having established the relationship between TP53 and ANK1 in the clinical setting, we next asked how ANK1 expression alone relates to patient survival. Interestingly, we found that high ANK1 expression correlated with decreased survival in lung cancer, breast cancer and CLL (Figure 8a). Since we have demonstrated ANK1 plays a role in cell motility following DNA damage (Figures 5 and 6), we hypothesised that high levels of ANK1 could accelerate the metastasis of cancer cells that are resistant to chemotherapy. To examine this we looked at the relationship between ANK1 and genes associated with DNA repair, such as ATM and CHEK1. Indeed, we identified that high levels of both ANK1 and either ATM or CHEK1 have a negative additive effect on survival (Figure 8b). These data suggest that ANK1 is connected with the DDR and its expression has clinical significance.
ANK1 expression in cancer patient survival. (a) Kaplan–Meier survival plots of patients with lung cancer (GEO data set: GSE31210), breast cancer (GEO data set: GSE25055) and chronic lymphocytic leukaemia (GEO data set: GSE22762) with either high (red) or low (green) ANK1 expression. Data obtained using the PPISURV database tool using different Gene Expression Omnibus (GEO) data sets. (b) Correlation between expression of DNA damage response regulators (ATM and CHECK1) and survival in breast cancer patients (GEO data set GSE25066), and the synergistic effect of high ANK1 levels. High expression of genes shown in red or low in blue. Data obtained using SynTarget bioinformatics analysis tool (www.bioprofiling.de)
Discussion
Previous work investigating the function of miR-486 demonstrated roles in skeletal muscle development,32 regulation of NF-κB signalling46 and the modulation of phosphoinositide-3-kinase (PI3K)/Akt signalling.47 In this study, we have shown that both miR-486 and ANK1 are co-regulated by p53 and are both induced following DNA damage. Previously, miR-486 had been shown to be DDR induced48 and p53 regulated.29 However, these studies overlooked the induction of its host gene, ANK1. Our data show that ANK1 is induced to a greater extent than the embedded miRNA following DNA damage. Genome-wide ChIP analysis for p53-binding sites has identified a p53-binding motif upstream of ANK1.49
MiR-486 has been shown to be controlled by the transcription factor MRTF-A at a promoter immediately upstream of exon 39a within the ANK1 gene, which also allows for the muscle-specific production of sAnk1 (ANK1.5).47, 50, 51 Concurrently, miR-486 has also been shown to be regulated by a p53 promoter upstream of the ANK1 gene in lung cancer.29 Our work shows that p53 controls both miR-486 and full-length ANK1 expression following DNA damage. It is known that the ANK1 gene is regulated by erythroid transcription factor GATA-1 and transcription factor NF-E2,52 but its expression in relation to p53 has never been examined. It is not unreasonable to assume that the same p53-binding site located 21.4 kb upstream of ANK1, previously shown to regulate miR-486,29 may control the expression of both transcripts. However, we cannot rule out the possible usage of other uncharacterised p53-binding sites.
Previous reports demonstrated that the overexpression of miR-486 results in inhibition of cell growth and blocks cell-cycle progression,25, 29 but the effect of miR-486 on cell-cycle arrest following DNA damage has not been examined. We investigated this using non-tumorigenic MCF10A cells that arrest at G2/M following DNA damage, and observed that inhibition of miR-486-5p resulted in a reduction of cells in G2/M. This suggests that, similar to the miR-34 family,14 miR-486-5p may play a role in the maintenance of cell-cycle arrest following DNA damage. Further investigation is required to identify mRNA targets of miR-486-5p, the repression of which would facilitate cell-cycle arrest during DDR.
Intriguingly, whereas miR-486-5p leads to the inhibition of cell-cycle progression, ankyrin-1 appears to play no role in cell-cycle maintenance, but rather has an effect on actin remodelling and facilitating cell motility. Indeed, the ankyrin protein has been implicated in cell migration.53 Ankyrins are highly conserved adaptor proteins that connect a variety of membrane-spanning proteins with the spectrin-actin cytoskeleton.41 As ankyrin-1 has been shown to bind to TIAM1, which in turn stimulates GDP/GTP exchange activity on Rac1,53 we looked at a panel of molecular components in the actin remodelling pathway. We found that overexpression of ANK1 severely hampered the expression of PAK1, p27Kip1 and phospho-cofilin levels, but had little effect on levels of ROCK1/ROCK2 or TIAM1. It is known that cofilin, along with ARP2/3 and profilin, is one of the main effectors of actin remodelling. Therefore, we speculate that the lack of actin protrusions and perturbed migratory behaviour in DNA damaged cells lacking ankyrin-1 may be attributed to ankyrin-1 levels modulating components of the Rho GTPase signalling pathway. This ultimately results in impaired cell motility. The physiological reason for the apparent facilitation of cell migration as a result of ANK1 upregulation following DNA damage is currently unclear; however, it is in line with other previously documented observations.40
In summary, we have shown how p53 can regulate a single transcript that produces two distinct molecular units with very different functions: a structural protein to facilitate motility and a co-expressed miRNA to attenuate cell-cycle progression following DNA damage. We also implicate a novel role for ankyrin-1 in the DDR. In line with our findings, a lack of TP53/ANK1 correlation predicts a poor patient outcome in clinical data sets. Importantly, we found that higher ANK1 levels alone correlated with lower tumour-free patient survival in the clinical setting. Moreover, high expression levels of ANK1 and DNA repair genes, such as ATM and CHEK1, in combination, are additive in predicting negative outcome. In light of the role ANK1 plays in cell motility post-DNA damage, we postulate that high levels of ANK1 could accelerate the metastasis of cancer cells that are resistant to chemotherapy. This suggests that ANK1 expression might potentially be used as a prognostic marker in patients with specific tumours.
Materials and Methods
Cell culture
MCF10A cells were cultured in DMEM/F12 (Gibco, Grand Island, NY, USA) with 5% horse serum, 1% glutamine, 20 ng/ml recombinant human EGF (Peprotech, Rocky Hill, NJ, USA), 10 μg/ml insulin, 500 ng/ml hydrocortisone and 10 ng/ml cholera toxin. MCF7 and A549 cells were cultured in DMEM with 10% fetal bovine serum (FBS) and 1% l-glutamine. The p53/R175H doxycycline-inducible H1299 cells were described previously16 and were cultured in DMEM using 10% tetracycline-free FBS and 1% glutamine. Gene expression of p53/R175H was induced by culturing cells in 2 μg/ml doxycycline for 48 h. RPE cells were cultured in DMEM/F12 with 10% FBS, 1% glutamine and 0.16% sodium bicarbonate. All cells were grown at 37 °C in a 5% CO2 humidified incubator.
DNA damage and p53 induction
DNA damage was achieved using either 25 μM etoposide (24 h) (Cayman, Ann Arbor, MI, USA) in dimethyl suphoxide (DMSO), 400 nM doxorubicin hydrochloride (0–48 h) (Sigma, St. Louis, MO, USA), 10 μg/ml cisplatin (24 h) (Sigma) or 300 nM mitomycin C (24 h) (Sigma). IR was produced by an Xstrahl X-Ray irradiator: Xstrahl (Camberley, UK) and ultraviolet light was produced using a UV crosslinker CX-2000 (UVP, Upland, CA, USA). The p53 activator nutlin-3 (Sigma) was prepared in DMSO and used at 10 μM for 24 h.
Western blotting and antibodies
Cells were harvested in RIPA buffer (150 mM NaCl) and lysates were sonicated for 2.5 min prior to clarification. Total protein (20–40 μg) was separated on polyacrylamide gels and transferred onto nitrocellulose membrane, 0.45 μm. All antibodies were used at dilutions recommended by the manufacturer. Antibodies used were: ANK1 (Novus Biologicals, Littleton, CO, USA, NBP1-71805) – for detection of endogenous protein, ANK1 (Atlas Antibodies, Stockholm, Sweden, HPA056953) – for detection of cloned protein, sAnk1 (Abcam, Cambridge, UK, ab58698) – for detection of the small isoform, Total p53 (Santa Cruz, Dallas, TX, USA, sc-126), Phospho-p53 (Ser15) (Cell Signalling, Danvers, MA, USA, #9284), MDM2 (Santa Cruz, sc-965), Total H2A.X (Cell Signalling, #2595), Phospho-histone (γ) H2A.X (Ser139) (Cell Signalling, #9718), PARP (Cell Signalling, #9542), B-actin (Sigma, A5441), p21Waf1/Cip1 (Santa Cruz, sc-397), Phospho-ATM (Ser1981) (Abcam, ab81292), Caspase-3 (Cell Signalling, #9662), GAPDH (Santa Cruz, sc-32233), eIF4G (Cell Signalling, #2498), Lamin A/C (Santa Cruz, sc-7292), p27Kip1 (Cell Signalling, #3686), PAK1 (Bethyl Laboratories, Montgomery, TX, USA, A301-259AT), Total cofilin (Cell Signalling, #5175), Phospho-cofilin (Ser3) (Cell Signalling, #3313), ROCK1 (Cell Signalling, #4035), ROCK2 (Bethyl Laboratories, A300-047A-T), TIAM1 (Bethyl Laboratories, A300-099AT) and Vinculin (Abcam, ab18058). IR-Dye-labelled secondary antibodies (LI-COR) were used to detect primary antibodies. Blots were scanned with a LI-COR laser-based image detector: LI-COR Biosciences, (Lincoln, NE, USA) and analysed using Image Studio software (v.2.1).
Small RNA library preparation
Total RNA was purified using Trizol reagent (Invitrogen, Carlsbad, CA, USA). Small RNA libraries were prepared using the TruSeq Small RNA kit (Illumina, San Diego, CA, USA) with 0.5 μg total RNA. Libraries were sent to The Genome Analysis Centre (TGAC), Norwich. The small RNA libraries were checked on a BioAnalyser (Agilent DNA 2100 chip, Agilent, Santa Clara, CA, USA) and LabChip GX, followed by quantification. Libraries were pooled and sequenced on the Illumina HiSeq 2500 (Rapid-Run mode, cBot clustering) platform using 50 bp single end reads.
Bioinformatics analysis of deep sequencing data and accession numbers
All sequencing files were transformed from FASTQ to FASTA format and the adaptors were removed by matching the first 8 nt of adaptor sequence (TGGAATTC). These reads were then then mapped to mature human miRNA sequences from miRBase version 2054 using PatMaN55 to obtain counts for all annotated miRNAs in each of the samples. Differentially expressed miRNAs were called using the edgeR package.56 A P-value of 0.05 was used as a cutoff to analyse microRNAs that were upregulated following DNA damage. Deep sequencing data are available from National Center for Biotechnology Information (NCBI) Sequence Read Archive (SRA) accession SRP064590.
Quantitative PCR and TaqMan assays
MicroRNAs were quantified using TaqMan assays (Applied Biosystems, Foster City, CA, USA) with 100 ng input total RNA. MicroRNA and sRNA-specific reverse transcription primer and probe reagents used were: hsa-miR-486-5p (#001278), hsa-miR-34a-5p (#000426), hsa-miR-34c-5p (#000428) and U6 snRNA (#001973). SYBR green (Applied Biosystems) based quantitative PCR (qPCR) was used for mRNA quantification. Superscript III reverse transcriptase (Invitrogen) was used to convert 1 μg total RNA to cDNA using random primers. cDNA was used as a template in qPCR with gene-specific primers: ANK1: forward 5′-TGAAGACGGGAGCCTCGAT-3′ and reverse 5′-GGGGTCTCCACTTTCACGTT-3′; p21Waf1/Cip1: forward 5′-GGAGACTCTCAGGGTCGAAA-3′ and reverse 5′-GGATTAGGGCTTCCTCTTGG-3′ and GAPDH: forward 5′-AGCCACATCGCTCAGACAC-3′ and reverse 5′-GCCCAATACGACCAAATCC-3′.
The ΔΔCT method was used to calculate fold-change between the control sample and the other conditions using U6 snRNA as an internal control gene for microRNAs and GAPDH for mRNAs.
Statistical analysis
Where stated, statistical significance was analysed using the Student’s t-test (two sample, two tailed, unpaired) and expressed as a P-value. In all bar charts and graphs, error bars represent the standard deviation of relative values/fold-changes from three experiments. Statistical significance for cancer patient data analysis was calculated using the relevant bioinformatics tool (see below).
siRNAs and transfection
SiRNA transfections were performed using Lipofectamine RNAiMAX (Invitrogen) and Opti-MEM+GlutaMAX (Gibco). The negative control siRNA used in all experiments was the ON-TARGETplus non-targeting siRNA #3 (Thermo Scientific, Waltham, MA, USA, D-001810-03-50), (+)5′-UCAGAAAACAUGUAAACCA(dTdT)-3′, (−)5′-UGGUUUACAUGUUUUCUGA-3′. The p53 targeting siRNA was custom ordered from Eurofins (Luxembourg City, Luxembourg), (+)5′-GACUCCAGUGGUAAUCUAC(dTdT)-3′, (−)5′-GUAGAUUACCACUGGAGUC(dTdT)-3′, and transfected at 50 nM twice over 2 days. The ANK1 targeting siRNA was a Silencer Select siRNA (Ambion, s1364), (+)5′-AAAUCAGUUUAACACCUUA(dTdT)-3′, (−)5′-UAAGGUGUUAAACUGAUUU(dCdA)-3′ and was transfected once at 20 nM.
FACS
For cell-cycle analysis, cells were fixed in ice cold 70% ethanol/30% PBS and left at −20 °C for 6–12 h. Cells were then washed in PBS and stained with 10 μg/ml propidium iodide and 0.1 mg/ml RNase A in PBS and left at 37 °C for 30 min, followed by overnight incubation at 4 °C. Propidium iodide incorporation into DNA was measured using the BD FACSCanto II (BD Biosciences, San Jose, CA, USA) with the 488 nm laser, measuring forward scatter, side scatter and red fluorescence. 10 000 cells were measured for each experimental condition.
Wound healing assay (scratch assay)
MCF10A cells were transfected with the appropriate siRNA and then treated the following day with either DMSO or 25 μM etoposide for 24 h. Cells were allowed to grow to full confluency in six-well plates. A linear scratch was made using a 10 μl pipette tip and cell migration into the wound was monitored for 48 h using a phase contrast time-lapse microscope at × 5 magnification. Images were taken at three different points along the scratch at 10 min intervals using MetaMorph Microscopy Automation & Image Analysis Software (v.7.8): Molecular Devices (Sunnyvale, CA, USA). Scratch wound closure was measured at specific time points along each scratch using the ImageJ software: National Institutes of Health, (Bethesda, MD, USA). Cell tracking was achieved using the Manual Tracking plugin on ImageJ.
Immunofluorescence confocal microscopy
Cells were grown on coverslips at low confluency and transfected with the appropriate siRNA. The following day cells were treated with either DMSO or 25 μM etoposide for 24 h. Cells were then fixed using 4% formaldehyde in PBS for 10 min and permbealised with 0.25% Triton X-100 in PBS for a further 10 min. Cells were blocked in 5% goat serum and 0.1% Triton X-100 in PBS for 1 h, followed by staining with Alexa Fluor 488 phalloidin (Life Technologies, Carlsbad, CA, USA) according to the manufacturer’s instructions. Coverslips were mounted using Vectashield hard set mounting medium with DAPI (Vector Laboratories, Burlingame, CA, USA). Confocal microscopy was performed using a Zeiss LSM 510 microscope at × 63 magnification, equipped with ZEN software (Zeiss, Oberkochen, Germany). Only isolated cells (not in colonies) had their protruding actin processes measured using ImageJ software.
Cytoplasmic and nuclear fractionation
Cells were washed twice in PBS and pelleted by centrifugation at 250 × g for 2 min, then resuspended in fractionation buffer (10 mM HEPES pH 7.9, 10 mM KCl, 1.5 mM MgCl2, 0.34 M sucrose, 10% glycerol, 1 mM DTT, 1 × protease inhibitor (Roche, Basel, Switzerland), 10 mM β-glycerophosphate, 50 mM NaF, 15 μM MG-132), followed by addition of 0.1% Triton X-100. Cells were incubated on ice for 8 min followed by centrifugation at 1300 × g, 4 °C for 5 min, followed by collection and clarification of the supernatant (cytoplasmic) fraction. The pellet (nucleus) was washed twice with fractionation buffer and lysed in RIPA buffer (same volume as cytoplasmic fraction), followed by sonication for 5 min and clarification. Equal volumes of nuclear and cytoplasmic lysate were analysed via western blot.
ANK1 cloning and nucleofection
The ANK1 gene was cloned from ORFeome Collaboration vector BC156401 (Thermo) into pcDNA3.1(+) (Invitrogen) using primers: Forward: 5′-CGCGGAATTCATGCCCTATTCTGTGGGCTTC-3′ (with EcoRI restriction site) and Reverse: 5′- CGGCCTCGAGTCACTTGTCGTCATCGTCTTTGTAGTCGGGGTTGGGTGTCGAGGTG-3′ (with XhoI site and FLAG tag). PCR was performed using Phusion DNA Polymerase (NEB) in HF buffer with DMSO. Temperature cycling parameters were: 98 °C for 1 min, followed by four cycles of: 98 °C – 10 s, 50 °C – 20 s, 72 °C – 5 m 30 s, followed by 25 cycles of: 98 °C – 10 s, 60 °C – 20 s, 72 °C – 5 m 30 s, followed by a final 10 min extension at 72 °C. The entire cloned ANK1 sequence was checked by Sanger sequencing following cloning. The pcDNA-ANK1-FLAG construct was nucleofected using programme T-020 (AMAXA nucleofector kit L) (Lonza, Basel, Switzerland) into 1 million MCF10A cells using 1.5 μg plasmid. Cells were harvested 48 h after nucleofection.
Datamining of gene expression data
The analyses of translational relevance of ANK1 have been performed with the BioProfiling.de online datamining tools45 using publically available clinical gene expression data from the GEO omnibus repository. The PPISURV57 tool was used to correlate gene expression data with cancer patient survival. To demonstrate translational relevance of potential association between ANK1 and DNA damage genes we used the SynTarget tool. SynTarget estimates the synergy of gene expression of two genes on survival of cancer patients. Patients in the clinical data sets are split into four groups with respect to the expression of the ANK1 and CHK1/ATM genes (high–high, high–low, low–high and low–low). Next, each group is tested versus other samples to find any statistical differences in survival outcome (i.e. high–high versus others, high–low versus others and so on).
Determining p53/ANK1 correlation and survival outcome in cancer patients
In contrast to the standard approach, where patients are grouped based on high/low expression of a gene, in this case, patients are grouped based on presence/absence of correlation between p53/ANK1 genes. Namely, in the first group we selected patients so as to maximise positive correlation of p53/ANK1, while all the other patients formed the second group. To identify statistical differences in survival outcome between the two groups of patients, the R statistical package was used to perform statistical tests and to derive the P-value.
Abbreviations
- ANK1:
-
ankyrin-1
- ATM:
-
ataxia telangiectasia mutated
- CHEK1:
-
checkpoint kinase 1
- CLL:
-
chronic lymphocytic leukaemia
- DDR:
-
DNA damage response pathway
- DMSO:
-
dimethyl sulphoxide
- Dox.:
-
doxorubicin
- DSB:
-
DNA double-strand break
- Etop.:
-
etoposide
- G1/G2:
-
gap phases of the cell-cycle
- γ-H2A.X:
-
phosphorylated H2A histone family member X
- IR:
-
ionising radiation
- miRNA:
-
microRNA
- MDM2:
-
mouse double minute 2 homologue
- p53:
-
tumour suppressor protein p53
- PAK1:
-
p21 protein (Cdc42/Rac)-activated kinase 1
- PARP:
-
poly (ADP-ribose) polymerase
- RPE:
-
retinal pigment epithelium cells
- S-phase:
-
synthesis phase of the cell cycle
- sAnk1/ANK1.5:
-
shorter muscle-enriched isoform of ankyrin-1
- siRNA:
-
small interfering RNA
- TIAM1:
-
T-lymphoma invasion and metastasis 1
- UVC:
-
ultraviolet-C irradiation
References
Bennett CB, Lewis AL, Baldwin KK, Resnick MA . Lethality induced by a single site-specific double-strand break in a dispensable yeast plasmid. Proc Natl Acad Sci USA 1993; 90: 5613–5617.
Shrivastav M, De Haro LP, Nickoloff JA . Regulation of DNA double-strand break repair pathway choice. Cell Res 2008; 18: 134–147.
Houtgraaf JH, Versmissen J, van der Giessen WJ . A concise review of DNA damage checkpoints and repair in mammalian cells. Cardiovasc Revasc Med 2006; 7: 165–172.
Liu C, Srihari S, Cao KL, Chenevix-Trench G, Simpson PT, Ragan MA et al. A fine-scale dissection of the DNA double-strand break repair machinery and its implications for breast cancer therapy. Nucleic Acids Res 2014; 42: 6106–6127.
Ciccia A, Elledge SJ . The DNA damage response: making it safe to play with knives. Mol Cell 2010; 40: 179–204.
Hu H, Gatti RA . MicroRNAs: new players in the DNA damage response. J Mol Cell Biol 2011; 3: 151–158.
Lakin ND, Jackson SP . Regulation of p53 in response to DNA damage. Oncogene 1999; 18: 7644–7655.
Kruiswijk F, Labuschagne CF, Vousden KH . P53 in survival, death and metabolic health: a lifeguard with a licence to kill. Nat Rev Mol Cell Biol 2015; 16: 393–405.
Kubbutat MHG, Jones SN, Vousden KH . Regulation of p53 stability by Mdm2. Nature 1997; 387: 299–303.
Liao J, Cao B, Zhou X, Lu H . New insights into p53 functions through its target microRNAs. J Mol Cell Biol 2014; 6: 206–213.
Wilczynska A, Bushell M . The complexity of miRNA-mediated repression. Cell Death Differ 2015; 22: 22–33.
Cannell IG, Bushell M . Regulation of Myc by miR-34c: a mechanism to prevent genomic instability? Cell Cycle 2010; 9: 2726–2730.
Adhikary S, Eilers M . Transcriptional regulation and transformation by MYC proteins. Nat Rev Mol Cell Biol 2005; 6: 635–645.
Cannell IG, Kong YW, Johnston SJ, Chen ML, Collins HM, Dobbyn HC et al. p38 MAPK/MK2-mediated induction of miR-34c following DNA damage prevents Myc-dependent DNA replication. Proc Natl Acad Sci USA 2010; 107: 5375–5380.
Roger L, Gadea G, Roux P . Control of cell migration: a tumour suppressor function for p53? Biol Cell 2006; 98: 141–152.
Muller PAJ, Caswell PT, Doyle B, Iwanicki MP, Tan EH, Karim S et al. Mutant p53 drives invasion by promoting integrin recycling. Cell 2009; 139: 1327–1341.
Schmitz AAP, Govek EE, Bottner B, Van Aelst L . Rho GTPases: signaling, migration, and invasion. Exp Cell Res 2000; 261: 1–12.
Burden DA, Osheroff N . Mechanism of action of eukaryotic topoisomerase II and drugs targeted to the enzyme. Biochim Biophys Acta 1998; 1400: 139–154.
Chang T, Wentzel EA, Kent OA, Ramachandran K, Mullendore M, Lee KH et al. Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol Cell 2007; 26: 745–752.
He L, He X, Lim LP, De Stanchina E, Xuan Z, Liang Y et al. A microRNA component of the p53 tumour suppressor network. Nature 2007; 447: 1130–1134.
Kumamoto K, Spillare EA, Fujita K, Horikawa I, Yamashita T, Appella E et al. Nutlin-3a activates p53 to both down-regulate inhibitor of growth 2 and up-regulate mir-34a, mir-34b, and mir-34c expression, and induce senescence. Cancer Res 2008; 68: 3193–3203.
Suzuki HI, Yamagata K, Sugimoto K, Iwamoto T, Kato S, Miyazono K . Modulation of microRNA processing by p53. Nature 2009; 460: 529–533.
Feng X, Wang Z, Fillmore R, Xi Y . MiR-200, a new star miRNA in human cancer. Cancer Lett 2014; 344: 166–173.
Bhatt S, Gupta MK, Khamaisi M, Martinez R, Gritsenko MA, Wagner BK et al. Preserved DNA damage checkpoint pathway protects against complications in long-standing type 1 diabetes. Cell Metab 2015; 22: 239–252.
Faraonio R, Salerno P, Passaro F, Sedia C, Iaccio A, Bellelli R et al. A set of miRNAs participates in the cellular senescence program in human diploid fibroblasts. Cell Death Differ 2012; 19: 713–721.
Hinske LCG, Galante PAF, Kuo WP, Ohno-Machado L . A potential role for intragenic miRNAs on their hosts' interactome. BMC Genomics 2010; 11: 533.
Hinske LC, Franca GS, Torres HAM, Ohara DT, Lopes-Ramos CM, Heyn J et al. miRIAD-integrating microRNA inter- and intragenic data. Database 2014; 2014: 1–9.
Ferland-McCollough D, Fernandez-Twinn DS, Cannell IG, David H, Warner M, Vaag AA et al. Programming of adipose tissue miR-483-3p and GDF-3 expression by maternal diet in type 2 diabetes. Cell Death Differ 2012; 19: 1003–1012.
Peng Y, Dai Y, Hitchcock C, Yang X, Kassis ES, Liu L et al. Insulin growth factor signaling is regulated by microRNA-486, an underexpressed microRNA in lung cancer. Proc Natl Acad Sci USA 2013; 110: 15043–15048.
Muller PAJ, Vousden KH . Mutant p53 in cancer: new functions and therapeutic opportunities. Cancer Cell 2014; 25: 304–317.
Vassilev LT, Vu BT, Graves B, Carvajal D, Podlaski F, Filipovic Z et al. In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science 2004; 303: 844–848.
Dey BK, Gagan J, Dutta A . miR-206 and -486 induce myoblast differentiation by downregulating Pax7. Mol Cell Biol 2011; 31: 203–214.
Ji X, Wu B, Fan J, Han R, Luo C, Wang T et al. The anti-fibrotic effects and mechanisms of microRNA-486-5p in pulmonary fibrosis. Sci Rep 2015; 5: 14131.
Lux SE, John KM, Bennett V . Analysis of Cdna for human erythrocyte Ankyrin indicates a repeated structure with homology to tissue-differentiation and cell-cycle control proteins. Nature 1990; 344: 36–42.
Lambert S, Yu H, Prchal JT, Lawler J, Ruff P, Speicher D et al. Cdna sequence for human erythrocyte Ankyrin. Proc Natl Acad Sci USA 1990; 87: 1730–1734.
Lahm A, Paradisi A, Green D, Melino G . Death fold domain interaction in apoptosis. Cell Death Differ 2003; 10: 10–12.
Morgan M, Thorburn J, Thomas L, Maxwell T, Brothman A, Thorburn A . An apoptosis signaling pathway induced by the death domain of FADD selectively kills normal but not cancerous prostate epithelial cells. Cell Death Differ 2001; 8: 696–705.
Herraiz C, Calvo F, Pandya P, Cantelli G, Rodriguez-Hernandez I, Orgaz JL et al. Reactivation of p53 by a cytoskeletal sensor to control the balance between DNA damage and tumor dissemination. J Natl Cancer Inst 2016; 108: 1–14.
Hsu F-, Lin T-, Chen J-, Shieh S- . p53-mediated transactivation of LIMK2b links actin dynamics to cell cycle checkpoint control. Oncogene 2010; 29: 2864–2876.
Chen W, Ebelt ND, Stracker TH, Xhemalce B, Van Den Berg CL, Miller KM . ATM regulation of IL-8 links oxidative stress to cancer cell migration and invasion. eLife 2015; 4: 1–21.
Bennett V, Baines AJ . Spectrin and ankyrin-based pathways: metazoan inventions for integrating cells into tissues. Physiol Rev 2001; 81: 1353–1392.
Borm B, Requardt R, Herzog V, Kirfel G . Membrane ruffles in cell migration: indicators of inefficient lamellipodia adhesion and compartments of actin filament reorganization. Exp Cell Res 2005; 302: 83–95.
Lu M, Zak J, Chen S, Sanchez-Pulido L, Severson DT, Endicott J et al. A code for RanGDP binding in Ankyrin repeats defines a nuclear import pathway. Cell 2014; 157: 1130–1145.
Besson A, Gurian-West M, Schmidt A, Hall A, Roberts JM . p27(Kip1) modulates cell migration through the regulation of RhoA activation. Genes Dev 2004; 18: 862–876.
Antonov AV . BioProfiling.de: analytical web portal for high-throughput cell biology. Nucleic Acids Res 2011; 39: W323–W327.
Song L, Lin C, Gong H, Wang C, Liu L, Wu J et al. miR-486 sustains NF-kappa B activity by disrupting multiple NF-kappa B-negative feedback loops. Cell Res 2013; 23: 274–289.
Small EM, O'Rourke JR, Moresi V, Sutherland LB, McAnally J, Gerard RD et al. Regulation of PI3-kinase/Akt signaling by muscle-enriched microRNA-486. Proc Natl Acad Sci USA 2010; 107: 4218–4223.
Hattori H, Janky R, Nietfeld W, Aerts S, Babu MM, Venkitaraman AR . p53 shapes genome-wide and cell type-specific changes in microRNA expression during the human DNA damage response. Cell Cycle 2014; 13: 2572–2586.
Wei CL, Wu Q, Vega VB, Chiu KP, Ng P, Zhang T et al. A global map of p53 transcription-factor binding sites in the human genome. Cell 2006; 124: 207–219.
Gallagher PG, Forget BG . An alternate promoter directs expression of a truncated, muscle-specific isoform of the human ankyrin 1 gene. J Biol Chem 1998; 273: 1339–1348.
Porter NC, Resneck WG, O'Neill A, Van Rossum DB, Stone MR, Bloch RJ . Association of small ankyrin 1 with the sarcoplasmic reticulum. Mol Membr Biol 2005; 22: 421–432.
Yocum AO, Steiner LA, Seidel NE, Cline AP, Rout ED, Lin JY et al. A tissue-specific chromatin loop activates the erythroid ankyrin-1 promoter. Blood 2012; 120: 3586–3593.
Bourguignon LYW, Zhu HB, Shao LJ, Chen YW . Ankyrin-Tiam1 interaction promotes Rac1 signaling and metastatic breast tumor cell invasion and migration. J Cell Biol 2000; 150: 177–191.
Kozomara A, Griffiths-Jones S . miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res 2014; 42: D68–D73.
Pruefer K, Stenzel U, Dannemann M, Green RE, Lachmann M, Kelso J . PatMaN: rapid alignment of short sequences to large databases. Bioinformatics 2008; 24: 1530–1531.
Robinson MD, McCarthy DJ, Smyth GK . edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010; 26: 139–140.
Antonov AV, Krestyaninova M, Knight RA, Rodchenkov I, Melino G, Barlev NA . PPISURV: a novel bioinformatics tool for uncovering the hidden role of specific genes in cancer survival outcome. Oncogene 2014; 33: 1621–1628.
Acknowledgements
This work was funded by the Medical Research Council (MRC), UK. AEH is an MRC post-doctoral research fellow and MB is an MRC Senior Fellow. W-TL was partially supported by an MRC Centenary Award. We thank David Read (microscopist), Lucia Pinon (flow cytometrist) and Debra Evens (laboratory technician) for providing technical support for some of the experiments.
Author contributions
The project was conceived and experiments were planned by AEH and MB. All experiments were conducted by AEH except the ionising radiation experiments in A549 cells, which were conducted by W-TL. Bioinformatics analysis of deep sequencing data was conducted by CP and SM. TD provided guidance on deep sequencing and subsequent analysis. JDG and PAJM provided expertise and advice on the p53 and cell motility aspects of the project. JDG also provided guidance on statistical analysis. AVA performed the clinical bioinformatics analyses. AW helped with the cloning of the ANK1 gene. All aspects of the study were supervised by MB. The manuscript was prepared by AEH, W-TL, AW and MB and all authors read and approved it.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by M Agostini
Supplementary Information accompanies this paper on Cell Death and Disease website
Supplementary information
Rights and permissions
Cell Death and Disease is an open-access journal published by Nature Publishing Group. This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Hall, A., Lu, WT., Godfrey, J. et al. The cytoskeleton adaptor protein ankyrin-1 is upregulated by p53 following DNA damage and alters cell migration. Cell Death Dis 7, e2184 (2016). https://doi.org/10.1038/cddis.2016.91
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2016.91
- Springer Nature Limited
This article is cited by
-
Differentially expressed genes in the femur cartilage transcriptome clarify the understanding of femoral head separation in chickens
Scientific Reports (2021)
-
The related miRNAs involved in doxorubicin resistance or sensitivity of various cancers: an update
Cancer Chemotherapy and Pharmacology (2021)
-
Could Alzheimer’s Disease Originate in the Periphery and If So How So?
Molecular Neurobiology (2019)
-
Drosha drives the formation of DNA:RNA hybrids around DNA break sites to facilitate DNA repair
Nature Communications (2018)
-
p73 Regulates Primary Cortical Neuron Metabolism: a Global Metabolic Profile
Molecular Neurobiology (2018)