Abstract
Little is known about the impact of DNA methylation on the evolution/progression of Ph+ chronic myeloid leukemia (CML). We investigated the methylome of CML patients in chronic phase (CP-CML), accelerated phase (AP-CML) and blast crisis (BC-CML) as well as in controls by reduced representation bisulfite sequencing. Although only ~600 differentially methylated CpG sites were identified in samples obtained from CP-CML patients compared with controls, ~6500 differentially methylated CpG sites were found in samples from BC-CML patients. In the majority of affected CpG sites, methylation was increased. In CP-CML patients who progressed to AP-CML/BC-CML, we identified up to 897 genes that were methylated at the time of progression but not at the time of diagnosis. Using RNA-sequencing, we observed downregulated expression of many of these genes in BC-CML compared with CP-CML samples. Several of them are well-known tumor-suppressor genes or regulators of cell proliferation, and gene re-expression was observed by the use of epigenetic active drugs. Together, our results demonstrate that CpG site methylation clearly increases during CML progression and that it may provide a useful basis for revealing new targets of therapy in advanced CML.
Similar content being viewed by others
Introduction
Chronic myeloid leukemia (CML) is a stem cell-derived malignancy characterized by the expansion and accumulation of myeloid cells in the blood and bone marrow (BM), the chromosomal translocation t(9;22) and the associated oncoprotein BCR-ABL1.1 The disease can be divided into a chronic phase (CP), accelerated phase (AP) and blast crisis (BC). In untreated and drug-resistant patients, evolution of CP to AP and finally to BC reflects the natural course of disease.1 The prognosis of patients in AP and BC is poor unless these patients can undergo stem cell transplantation.
In CP-CML, BCR-ABL1 is a key driver of oncogenesis, cellular proliferation and survival. Until now, only few BCR-ABL1-independent genetic lesions (for example, c-Myc copy number gain, RB1 mutations) are known to be involved in CML progression.2, 3, 4 The potential contribution of epigenetic changes, especially of DNA methylation (referred to as methylation) to CML progression, has not been investigated in detail so far.5, 6, 7
Methylation is an epigenetic modification involved in the regulation of many biological processes, including embryogenesis, genomic imprinting or X chromosome inactivation.8, 9, 10, 11 In the mammalian genome, cytosines within CpG sites are the main targets of methylation. CpG sites are spread throughout the genome but are mainly located in small regions of about 0.5–2 kb in length, called CpG islands.12 These regions are found in ~60% of human gene promoters and less frequently in gene bodies or in intergenic regions.13, 14 However, in total CpG sites are under-represented in the mammalian genome. Transcriptional gene silencing is associated with methylation of CpG sites and CpG islands located near the transcription start sites (TSS) of genes.13 In cancer cells, these regions may be frequently methylated resulting in transcriptional gene silencing of many cancer-associated genes.15, 16, 17 Although methylation of CpG sites in gene bodies may contribute to cancer-causing somatic and germline mutations, the function of intergenic CpG site methylation is barely understood so far.18 As methylation is reversible, methylated genes may be re-expressed by the use of DNA methyltransferase inhibitors (DNMTi; for example, 5-aza-2′-deoxycytidine (Aza-dC); 5-azacytidine).19 Moreover, DNMTi and histone deacetylase inhibitors (for example, trichostatin A (TSA)) act synergistically on gene re-expression.20
Methylation is considered biologically important in the pathogenesis of many malignant diseases. Therefore, we were interested to study the methylome in CML patients at diagnosis as well as at disease progression. Using reduced representation bisulfite sequencing (RRBS), we analyzed samples from CP, AP and BC of CML patients and from control individuals for differences in CpG site methylation. Moreover, we performed RNA-sequencing (RNA-seq) to study gene expression in these samples.
Overall, we were able (1) to demonstrate that CpG site methylation increases dramatically during the progression from CP-CML to AP-CML/BC-CML and (2) to identify genes that are transcriptionally regulated by methylation in BC-CML samples. Some of these genes are known to be involved in the pathogenesis of various malignancies; however, the role of many other genes in the pathogenesis and progression of malignant diseases, particularly in CML, remains to be determined.
Materials and methods
CML samples and cell lines
Mononuclear cells were isolated from peripheral blood (PB) or BM of 31 samples (17 CP-CML, 5 AP-CML, 9 BC-CML) from 23 CML patients using Ficoll (Sigma, St Louis, MO, USA). PB and BM samples were collected from all patients at diagnosis and from a subset of four patients during their follow-up. All four patients progressed to AP or BC during therapy with BCR-ABL1 tyrosine kinase inhibitors. Clinico-pathological characteristics of the patients are shown in Supplementary Table S1. Mononuclear cells obtained from five individuals were used as controls. All donors gave written informed consent. The study was approved by the ethics committee of the Medical University of Vienna, Austria (no. 1594/2015).
Authenticated cell lines K562, KCL22, NALM-1, TOM-1 and BV-173 were purchased from the Leibniz Institute DSMZ (Braunschweig, Germany). KU812 cells were kindly provided by Dr K Kishi (Niigata University, Niigata, Japan), and K562 R cells (imatinib-resistant) were kindly provided by James D Griffin (Dana Farber Cancer Institute, Boston, MA, USA). Cells were cultured in RPMI1640 medium with 10% fetal calf serum at 37 °C. Mycoplasma contamination was tested using the Venor GeM Classic Mycoplasma Detection Kit (Minerva Biolabs, Berlin, Germany). Aza-dC and TSA treatment was performed as reported.21 Untreated control cells were cultured in parallel.
Reduced representation bisulfite sequencing
Genomic DNA was MspI (New England Biolabs, NEB, Ipswich, MA, USA) digested, end-repaired and A-tailed using Klenow Polymerase (NEB). Illumina adapters were ligated using Quick Ligase (NEB) followed by AMPure XP size selection (Beckman Coulter, Fullerton, CA, USA). RRBS libraries were bisulfite treated using the EZ-DNA Methylation‐Direct kit (Zymo Research Corp, Orange, CA, USA) followed by quantitative PCR quantification. Enrichment PCR was performed using the PfuTurboCx Hotstart Kit (Agilent Technologies, Santa Clara, CA, USA) followed by AMPure XP clean up. Quality of final libraries was checked by Experion analysis (Bio-Rad, Hercules, CA, USA). Sequencing was performed on a HiSeq2000 sequencer (Illumina Inc, San Diego, CA, USA). A detailed description of the RRBS assay is provided as Supplementary Information.
RNA-sequencing
Total RNA was extracted using the RNeasy Kit (Qiagen, Hilden, Germany) and processed for sequencing using the TruSeq RNA Sample Preparation Kit (Illumina Inc) as described.22 mRNAs were purified using poly(T)-oligo-attached magnetic beads, fragmented and applied to first-strand complementary DNA (cDNA) synthesis. Second-strand cDNA synthesis was performed using DNA polymerase I and RNase H. cDNAs were end-repaired, A-tailed, ligated to adapters and amplified to create the final cDNA library for sequencing (HiSeq2000, Illumina Inc).
Methylation-sensitive high-resolution melting (MS-HRM) analyses
Genomic DNA was modified by treatment with sodium bisulfite using the EpiTect Bisulfite Kit (Qiagen) as recommended.23 The following primer sequences for methylation-insensitive amplification of a part of the CYP1B1 5′ region (ENSEMBL database, release 69) were designed using the Methyl Primer Express v1.0 software (Applied Biosystems, Carlsbad, CA, USA): fwd, 5′-GGTTAAAGYGGTTTGGTGT-3′ and rev, 5′-CTCCCACTCCAAAATCAAAA-3′. MS-HRM analyses were performed using the EpiTect HRM PCR Kit in a RotorGeneQ cycler (Qiagen).23 Methylation standards were constructed by diluting 100% methylated and unmethylated control DNA (Qiagen) at 100, 75, 50, 25, 10 and 0% ratios. Melting curves were normalized by calculation of two normalization regions before and after the major fluorescence decrease using the RotorGeneQ software. Normalized fluorescence values were plotted against the percentage of methylation for each of the methylation standards to generate a standard curve for the calculation of methylation levels of genes in leukemia cells. Water blanks were used as negative controls.
Real-time reverse transcription-PCR (RT-PCR)
Total RNA was reverse-transcribed using the OmniScript Reverse Transcriptase Kit (Qiagen), and cDNA was used for standard RT-PCR analyses. Detailed information about used RT-PCR assays is provided as Supplementary Information.
Statistical analyses
RRBS reads were processed using the -RRBS option of the Trim Galore! software (http://www.bioinformatics.babraham.ac.uk/projects/trim_galore/). Bismark software was used to align reads to hg19 and for methylation calling.24 Codes are available from Trim Galore! and bismark documentation. Bam files from RRBS analyses were deposited in ArrayExpress database (E-MTAB-4341). Methylation was quantified using the 'Bisulfite methylation over features' pipeline of the SeqMonk v0.27.0 software (Babraham Institute, Cambridge, UK). Differential methylation (increased or decreased) was calculated using a windowed replicate filter over a window of 1000 bp using the following cutoff levels: at least 25% difference in methylation and false discovery rate (FDR)<0.05. Hierarchical clustering was performed using the hclust function of R-2.15.0. Circos software (Canada’s Michael Smith Genome Sciences Center, Vancouver, BC, Canada) was used to visualize RRBS data.25 The R packages MethylSig and MethylKit were used for quality control, calculation of correlation coefficients to describe similarities of CpG site methylation between samples, for CpG island/Refseq annotation and for transcription factor (TF)-binding site enrichment analyses. Pathway enrichment analyses were performed using the GREAT software (Stanford University, Stanford, CA, USA; default settings; binomial test and hypergeometric test cutoff: FDR<0.05).26
Galaxy platform and TopHat2 algorithm were used to align raw RNA-seq data to hg19.22, 27, 28, 29, 30 Bam files from RNA-seq analyses were deposited in ArrayExpress database (E-MTAB-4333). Aligned .bam files were processed using RNA-seq quantitation pipeline of the SeqMonk v0.27.0 software (Babraham Institute). An adjusted P<0.05 was defined as cutoff for differentially expressed genes. Single-nucleotide variants (SNVs) were identified using RNAseqmut (https://github.com/davidliwei/rnaseqmut) and verified using IGV.31
R software (https://cran.r-project.org/) and GraphPad Prism v.6.03 (GraphPad Software, Inc., San Diego, CA, USA) were used to calculate Bartlett tests, t-tests and Mann–Whitney U-tests. P-values of <0.05 (two-sided) were considered as significant.
Results
Comprehensive CpG site methylation analyses in CML and in control samples
We investigated the methylome of 17 CP-CML, 5 AP-CML, 9 BC-CML and 5 control samples using RRBS. The bisulfite conversion rate of mapped RRBS reads was >99% for all samples analyzed. A mean of 1.4 × 107 cytosines were analyzed per sample and about 20% (2.6 × 106) of these cytosines were located in CpG sites. Examples of RRBS library quality and RRBS data quality are shown in Supplementary Figures S1 and S2.
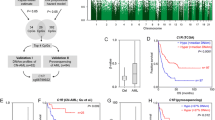
When we compared the content of methylated CpG sites in CML samples with that in control samples, we observed a higher content in CML samples (Figure 1). Moreover, a higher content of methylated CpG sites was also seen in BC-CML samples compared with CP-CML samples (P=0.02, Figure 1). However, no statistically significant difference in the content of methylated CpG sites between AP-CML and CP-CML samples was found.
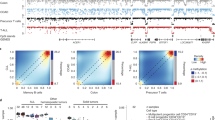
Methylome profiling in CML samples and in control samples by RRBS. The percentage of methylated CpG sites in control samples and in different CML stages is demonstrated. Mean values±s.e.m. are shown. P-values were calculated using the Mann–Whitney U-test. Ctrl, controls; ns, not statistically significant.
Hierarchical clustering of CML samples and of control samples based on CpG site methylation clearly separated CP-CML, AP-CML and BC-CML samples from control samples (Figure 2a and Supplementary Figure S3A).
Cluster analyses and identification of differentially methylated CpG sites. (a) Hierarchical clustering of CML samples and of control samples based on CpG site methylation is shown. The samples were obtained either from PB or from BM. Green dots, CP-CML; blue dots, BC-CML; red dots, controls. (b) Bland–Altman MA-plots summarizing differentially methylated CpG sites between control samples (normal peripheral blood (nPB) and normal bone marrow (nBM)) and CP-CML or BC-CML samples are shown. Each dot represents a single CpG site. The areas above the gray line indicate increased CpG site methylation and the areas below the gray line indicate decreased CpG site methylation.
Detailed comparison of CpG site methylation between CML and control samples
Analyses were performed separately for the PB (CP-CML, n=6; BC-CML, n=4; controls, n=3) and BM (CP-CML, n=11; AP-CML, n=4; BC-CML, n=5; controls, n=2) sample groups. In the PB cohort, 666 CpG sites (0.4% of the CpG sites analyzed) were differentially methylated when we compared CP-CML and control samples. While methylation of 292 CpG sites was increased, methylation of 374 CpG sites was decreased (Figure 2b). Six thousand nine hundred and twelve CpG sites (2.3% of the CpG sites analyzed) were differentially methylated when we compared BC-CML and control samples. While methylation of 6081 CpG sites was increased, methylation of 831 CpG sites was decreased (Figure 2b). All these differences were statistically significant.
Similar results were observed in the BM cohort. Two hundred and seventy-six CpG sites (0.2% of the CpG sites analyzed) were differentially methylated between CP-CML and control samples. Increased methylation was observed in 237 CpG sites and decreased methylation in 39 CpG sites. Six thousand one hundred and one CpG sites (1.8% of the CpG sites analyzed) were differentially methylated between BC-CML and control samples and increased methylation was observed in 5729 CpG sites, whereas decreased methylation was found in 372 CpG sites (Figure 2b). Moreover, different CpG site methylation was observed in AP-CML samples compared with control samples. Four thousand one hundred and thirty CpG sites (0.9% of the CpG sites analyzed) were differentially methylated and increased methylation was observed in 3575 CpG sites, whereas decreased methylation was found in 555 CpG sites (Supplementary Figure S3B). Again, all these differences were statistically significant.
From the CpG sites found to be differentially methylated in AP-CML/BC-CML samples, we also determined their location within, around and between CpG islands. The majority of CpG sites with increased methylation (88%; range 87–89%) were located in or close to CpG islands and only 12% were located in between CpG islands (P<0.0001). In contrast, about half of the CpG sites with decreased methylation (49%; range 45–53%) were located in between CpG islands mainly in introns and intergenic regions.
Additionally, we assigned CpG sites with increased methylation in AP-CML/BC-CML samples to genomic regions located around TSS of genes (±2000 bp), and we identified 348 methylated genes in BC-CML samples from the PB cohort and 141 and 292 methylated genes in AP-CML and BC-CML samples from the BM cohort, respectively (Figure 3). A detailed description of these genes is listed in Supplementary Tables S2–S4.
Heatmaps summarizing methylation values of genes with increased methylation in AP-CML and BC-CML samples compared with control samples. (a) 348 genes with increased methylation in BC-CML samples compared with control samples (PB cohort), (b) 141 genes with increased methylation in AP-CML samples compared with control samples (BM cohort) and (c) 292 genes with increased methylation in BC-CML samples compared with control samples (BM cohort) are shown. The colors range from blue (low methylation) to red (high methylation). Mean methylation values are shown. Ctrl, controls.
CpG site methylation, gene expression and SNV analyses of individual CML patients
Comparison of the methylome of CML patients at different stages of disease
To determine differences between CP-CML and disease progression, we compared CpG site methylation in samples of four patients who were initially diagnosed with CP-CML and who progressed several months/years later (Supplementary Table S1, Figure 4a). From patient 1, PB samples from three different time points were available (CP-CML, CP-CML ~5 years later and AP-CML). Overall, we identified 20 172 CpG sites that were differentially methylated in the AP-CML sample compared with the CP-CML samples of this patient. In 72% of these CpG sites, methylation was increased. Interestingly, only 1966 differentially methylated CpG sites were identified when we compared the 2 CP-CML samples of this patient. This finding suggests that CpG site methylation does not differ significantly between various time points during CP-CML even when they are several years apart.
CpG site methylation in four CML patients at diagnosis and at disease progression. (a) The time from first diagnosis of CP-CML to disease progression, BCR-ABL1 count (Ph+), blast count and therapies are shown. (b) The circos plots demonstrate the mean percentage of CpG site methylation of genomic regions located ±2000 bp from TSS of genes. Normal peripheral blood (nPB) and normal bone marrow (nBM) samples from control individuals were used as references. na, not available.
Patients 2–4 were initially diagnosed with CP-CML and developed BC-CML after several months or years. PB samples from patients 2 and 3 and BM samples from patient 4 from two different time points were available for analyses, respectively (Figure 4a). We identified 29 280, 58 691 and 28 268 differentially methylated CpG sites in patients 2–4 in the BC-CML samples compared with the CP-CML samples, respectively. In 96, 89 and 67% of these CpG sites, methylation was increased.
In patients 1–3, increased CpG site methylation was accompanied with increased BCR-ABL1 expression determined by RT-PCR analyses using GUSB as well as ABL1 as reference genes (Figure 4a). In patient 4, material for BCR-ABL1 testing by RT-PCR analyses using GUSB as reference gene was used up; however, BCR-ABL1 expression in both samples of this patient was observed in previous RT-PCR analyses using ABL1 as reference gene.
Genomic annotation of differentially methylated CpG sites
We additionally determined the location of differentially methylated CpG sites within, around and between CpG islands in the samples from these four patients. The majority of CpG sites with increased methylation (87%; range 82–92%) were located in or close to CpG islands. Only 13% of them were located in between CpG islands (P<0.0001; Supplementary Figure S4A) and were evenly distributed in promoter, gene body and intergenic regions. In contrast, about half of the CpG sites with decreased methylation (54%; range 45–63%) were located outside of CpG islands mainly in introns and intergenic regions (Supplementary Figure S4B).
Moreover, we were interested to identify TFs whose activity might be affected by methylation of their binding sites. By TF-binding site enrichment analyses in the samples of these four patients, we observed an overrepresentation of CpG sites with increased methylation in several TF-binding sites. These TF-binding sites include C-myc (P=5 × 10−16), nuclear factor-κB (P=1 × 10−14), C-fos (P=5 × 10−13) and YY1 (P=2 × 10−10). Detailed results are shown in Supplementary Table S5.
To investigate whether CpG sites with increased methylation in the samples of these four patients are associated with specific biological processes or molecular functions, we performed Gene Ontology (GO) and molecular signature (MSigDB) enrichment analyses. Highly enriched GO/MSigDB terms were mainly associated with cell differentiation, developmental processes, transcriptional gene regulation, targets of polycomb proteins (EED, SUZ12, PRC2), H3K27 trimethylation and gene hypermethylation in cells of different malignant diseases (Supplementary Table S6).
In addition, we assigned CpG sites with increased methylation in AP-CML/BC-CML samples to genomic regions located around TSS of genes (±2000 bp), and we identified 344 (patient 1), 381 (patient 2), 897 (patient 3) and 597 (patient 4) methylated genes in these samples, respectively (Figure 4b). A detailed description of the genes is listed in Supplementary Tables S7–S10. Three hundred and sixty-three of these genes were methylated in samples of at least two of the patients (Figure 5).
Overlap of genes found to be methylated in BC-CML samples from patients 2–4 who were initially diagnosed with CP-CML and who progressed to BC-CML. The circos plot represents methylation values in CP-CML (outer ring) and BC-CML (inner ring) determined by RRBS. Patients 2 (blue), 3 (green) and 4 (pink) are shown. Blue lines indicate genes that are methylated in two of the patients and red lines indicate genes that are methylated in all three patients.
Transcriptional gene expression of methylated genes
From patients 3 and 4 also total RNA from CP-CML and BC-CML samples was available and was used together with total RNA samples from four control individuals for RNA-sequencing analyses to investigate gene expression. Overall, the median expression of methylated genes was lower in BC-CML samples compared with both CP-CML and control samples (Figure 6a). Expression of 193 out of 897 (patient 3) and of 141 out of 597 (patient 4) methylated genes in BC-CML samples was downregulated less than twofold in BC-CML samples compared with CP-CML samples of these patients, respectively (Figure 6a). Compared with control samples, expression of a large majority of them was downregulated in the BC-CML samples. Some of these genes are known tumor-suppressor genes (EPB41L3, PRDX2), putative tumor-suppressor genes (PLCL1, TUSC1), regulators of cell proliferation (BCL11B, NDRG2, PID1) or regulators of drug metabolism (CYP1B1).21, 32, 33, 34, 35 A description of the downregulated genes is shown in Supplementary Tables S11 and S12.
Methylation-associated downregulation of transcriptional gene expression. (a) The boxplots summarize mRNA expression of genes found to be methylated in BC-CML samples, and the heatmaps demonstrate mRNA expression values of genes found to be downregulated at least twofold in expression. Colors range from red (low expression) to blue (high expression). (b) The results from RNA-seq analyses regarding CYP1B1 expression in the BC-CML sample and the CP-CML sample of patient 3 and in the control samples were confirmed by RT-PCR. The expression values of CYP1B1 were normalized against 18 S rRNA or GAPDH. Each experiment was performed in triplicate. Mean fold changes (FCs)+s.e.m. are shown. (c) The FCs (low to high) of CYP1B1 expression in seven leukemia cell lines compared with controls and the percentages (low to high) of methylation are demonstrated. (d) The FC in CYP1B1 expression after treatment of K562 and K562 R cells with Aza-dC and TSA is shown. Mean FCs+s.e.m. are shown. nPB, normal peripheral blood; nBM, normal bone marrow; Exp, gene expression; Meth, methylation.
We confirmed CYP1B1, EPB41L3 and PID1 expression data obtained by RNA-seq using RT-PCR in CP-CML and BC-CML samples of patient 3 and in control samples (Figure 6b and Supplementary Figure S5). In addition, we determined both CYP1B1 expression and methylation by MS-HRM in seven leukemia cell lines. In five of them, CYP1B1 was methylated and was not transcribed (Figure 6c). To investigate whether transcription of CYP1B1 may be re-expressed by epigenetically active drugs, we treated K562 and K562 R cells with Aza-dC and TSA and performed RT-PCR. CYP1B1 expression was found to be upregulated 218-fold in K562 cells and 9647-fold in K562 R cells (Figure 6d).
SNV analyses
When we compared the mutational landscapes of CP-CML and BC-CML samples from patients 3 and 4, we found 73 (patient 3) and 248 (patient 4) SNVs to be present only in BC-CML samples, respectively (Supplementary Tables S13 and S14). Examples for non-synonymous SNVs are ABL1 (p.L248V), NR3C1 (p.G239X), PLK4 (p.S200T) and VRK3 (p.V290A) (Supplementary Figures S6 and S7). Interestingly, BC-CML-specific SNVs of certain epigenetic modifiers (DNMT1, DNMT3A, TET2, TET3, EZH1, EZH2, RUNX1, IKZF1, IDH1 and IDH2) and of certain oncogenes (MYC, RUNX1, MDM2, BCL2, NFKB1, NFKB2, KRAS, NRAS, HRAS, RAF1, MEK/MAP2K1, ERK/MAPK1) were not found. Moreover, no BC-CML-specific karyotypic changes were observed in patients 1, 3 and 4.
Discussion
Although changes in methylation patterns were widely analyzed in various malignant diseases, data generated on the impact of methylation in CML are limited and mainly restricted to analyses of single genes or small numbers of genes.5, 6, 16, 36, 37, 38 However, as recently developed techniques for detecting methylation are much more advanced, we were able to perform extensive methylation analyses in CML samples using the next-generation sequencing approach RRBS. The main advantages of this method compared with microarray-based techniques are the high sensitivity, the high CpG coverage, the single-nucleotide resolution and the low input DNA requirement.39, 40, 41
On average, we analyzed 2.6 × 106 CpG sites in each CML and each control sample for methylation. Overall, the frequency of methylated CpG sites was significantly higher in CML samples compared with control samples, indicating that CpG site methylation is involved in CML. Interestingly, we observed differences in CpG site methylation between CP-, AP- and BC-CML samples. In CP-CML samples, the frequency of CpG site methylation was not significantly different from control samples, suggesting that CpG site methylation does not have a central role in CP-CML. These findings support the assumption that CP-CML is mainly driven by the BCR-ABL1 translocation.42 However, we observed a high increase of methylated CpG sites in AP and BC, indicating that CpG site methylation may be important for disease progression.
To investigate differences in CpG site methylation between CP-CML and CML progression in detail, we compared the methylome of four CML patients at the time of diagnosis (CP-CML) and at the time of disease progression (AP-CML or BC-CML). Although we found similar frequencies of differentially methylated CpG sites between both CP-CML samples of patient 1, we observed a significant increase of differentially methylated CpG sites in the AP-CML sample of this patient. However, the highest increase of differentially methylated CpG sites (up to 58 691) was found in BC-CML samples. In the majority of CpG sites, methylation was increased, suggesting that increased but not decreased CpG site methylation has an important role in the progression of this disease.
Genomic annotation of CpG sites with increased methylation in AP-CML/BC-CML revealed that they are mainly located in or around CpG islands. Our results suggest that these CpG sites may be involved in regulation of transcriptional gene expression as CpG island methylation is strongly associated with gene silencing.13 Moreover, many methylated CpG sites were found in the binding sites of certain TFs. This finding is of interest as it was reported that methylation of TF-binding sites may deregulate transcriptional gene expression by changing the structure of DNA or by altering the affinity of TFs to DNA.43, 44, 45 In addition, GO/MSigDB enrichment analyses revealed that increased CpG site methylation frequently happens at genomic regions prone for epigenetic modifications associated with gene silencing. Similar findings were reported by Viré et al.46 who demonstrated that polycomb group proteins, H3K27 methylation and CpG site methylation are closely linked in epigenetic gene silencing.
Further assignment of CpG sites with increased methylation in BC-CML samples to genomic regions ±2000 bp from TSS of genes resulted in the identification of up to 897 genes. Many of them were methylated in the BC-CML samples of at least two of the three patients, suggesting that changes in methylation during disease progression are not random events. Although methylation of some of these genes (for example, TFAP2A, EBF2) was previously reported in CML, methylation of the majority of them was unknown so far.6 Interestingly, some of them were described to be methylated in other hematological and/or solid malignancies, for example, BNIP3 (multiple myeloma, gastric cancer), HIC1 (myelodysplastic syndrome), CADM1 and EPB41L3 (lung and breast cancers), PDLIM4 and CIDEB (neoplastic mast cells) and CDH1 (myelodysplastic syndrome).15, 21, 47, 48, 49, 50
We also performed genome-wide gene expression analyses in CP-CML and BC-CML samples of two patients who were initially diagnosed with CP-CML and who progressed to BC-CML. These samples were also analyzed by RRBS. Expression of many genes identified to be methylated in BC-CML samples was found to be downregulated in these samples, suggesting that expression of them is affected by methylation. Examples are the known/putative tumor-suppressor genes CYP1B1, EPB41L3 and PID1, which downregulated expression observed by RNA-seq was confirmed by RT-PCR. CYP1B1 belongs to the cytochrome P450 family, is involved in drug metabolism and steroid synthesis and was found to be associated with a shorter survival of acute lymphocytic leukemia patients.51, 52 EPB41L3 is involved in cell adhesion, regulation of cell motility, cell proliferation and apoptosis and is frequently inactivated by methylation in various cancer types.21, 49, 53 PID1 is a tumor cell growth suppressor and a low PID1 expression was found to be associated with shorter overall survival of patients with brain tumors.32 Additional experiments to define the role of these genes during CML progression will be performed.
Because we demonstrated that treatment of CML cells (both sensitive and resistant to imatinib) with Aza-dC lead to gene re-expression and that CpG site methylation is significantly increased in AP-CML/BC-CML samples, we hypothesize that besides BCR-ABL1 tyrosine kinase inhibitor also DNMTi (for example, decitabine) might be useful for the treatment of advanced-stage CML patients. Indeed, Issa et al.54 reported a complete and partial hematological response rate in 34% and 20% of imatinib-resistant patients treated with low-dose decitabine, respectively. Another study described a complete hematological response rate in 32% and a partial hematological response rate in 4% of AP-CML/BC-CML patients treated with decitabine and imatinib.55 The efficacy of the combination decitabine and the BCR-ABL1 tyrosine kinase inhibitor dasatinib is currently being investigated in AP-CML/BC-CML patients in a clinical trial (ClinicalTrials.gov, NCT01498445).
It was reported that SNVs may occur in epigenetic modifiers in CML patients.56, 57, 58 While Schmidt et al.56 found SNVs in the epigenetic modifiers DNMT3A, EZH2, RUNX1 and TET2 in CP-CML samples, Makishima et al.58 observed BC-CML-specific SNVs in TET2, ASXL1 and IDH family genes when they compared BC-CML and CP-CML samples. However, in our sample collection we neither detected BC-CML-specific SNVs of any of these genes nor of any other epigenetic modifiers investigated which suggests that their role in CML, and particularly in disease progression, is still unclear. Additional studies are necessary to clarify the occurrence of molecular changes of epigenetic modifiers and to investigate a potential role of them in methylation changes in CML patients.
In conclusion, using a novel next-generation sequencing approach, we were able to demonstrate that CpG site methylation is heavily increased in AP-CML/BC-CML compared with CP-CML. Moreover, a large number of genes transcriptionally regulated by methylation were identified in BC-CML samples. From the majority of these genes, methylation in CML was unknown so far. Finally, our findings suggest that investigating the efficacy of DNMTi in patients with progressed CML should be considered.
References
Melo JV, Barnes DJ . Chronic myeloid leukaemia as a model of disease evolution in human cancer. Nat Rev Cancer 2007; 7: 441–453.
Greulich-Bode KM, Heinze B . On the power of additional and complex chromosomal aberrations in CML. Curr Genomics 2012; 13: 471–476.
Perrotti D, Jamieson C, Goldman J, Skorski T . Chronic myeloid leukemia: mechanisms of blastic transformation. J Clin Invest 2010; 120: 2254–2264.
Di Bacco A, Keeshan K, McKenna SL, Cotter TG . Molecular abnormalities in chronic myeloid leukemia: deregulation of cell growth and apoptosis. Oncologist 2000; 5: 405–415.
Jelinek J, Gharibyan V, Estecio MR, Kondo K, He R, Chung W et al. Aberrant DNA methylation is associated with disease progression, resistance to imatinib and shortened survival in chronic myelogenous leukemia. PLoS One 2011; 6: e22110.
Dunwell T, Hesson L, Rauch TA, Wang L, Clark RE, Dallol A et al. A genome-wide screen identifies frequently methylated genes in haematological and epithelial cancers. Mol Cancer 2010; 9: 44.
Amabile G, Di Ruscio A, Muller F, Welner RS, Yang H, Ebralidze AK et al. Dissecting the role of aberrant DNA methylation in human leukaemia. Nat Commun 2015; 6: 7091.
Straussman R, Nejman D, Roberts D, Steinfeld I, Blum B, Benvenisty N et al. Developmental programming of CpG island methylation profiles in the human genome. Nat Struct Mol Biol 2009; 16: 564–571.
Reik W . Stability and flexibility of epigenetic gene regulation in mammalian development. Nature 2007; 447: 425–432.
Okano M, Bell DW, Haber DA, Li E . DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 1999; 99: 247–257.
Weber M, Schubeler D . Genomic patterns of DNA methylation: targets and function of an epigenetic mark. Curr Opin Cell Biol 2007; 19: 273–280.
Takai D, Jones PA . Comprehensive analysis of CpG islands in human chromosomes 21 and 22. Proc Natl Acad Sci USA 2002; 99: 3740–3745.
Jones PA . Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet 2012; 13: 484–492.
Deaton AM, Bird A . CpG islands and the regulation of transcription. Genes Dev 2011; 25: 1010–1022.
Ghanim V, Herrmann H, Heller G, Peter B, Hadzijusufovic E, Blatt K et al. 5-azacytidine and decitabine exert proapoptotic effects on neoplastic mast cells: role of FAS-demethylation and FAS re-expression, and synergism with FAS-ligand. Blood 2012; 119: 4242–4252.
Heller G, Babinsky VN, Ziegler B, Weinzierl M, Noll C, Altenberger C et al. Genome-wide CpG island methylation analyses in non-small cell lung cancer patients. Carcinogenesis 2013; 34: 513–521.
Esteller M . Epigenetics in cancer. N Engl J Med 2008; 358: 1148–1159.
Rideout WM 3rd, Coetzee GA, Olumi AF, Jones PA . 5-Methylcytosine as an endogenous mutagen in the human LDL receptor and p53 genes. Science 1990; 249: 1288–1290.
Navada SC, Steinmann J, Lubbert M, Silverman LR . Clinical development of demethylating agents in hematology. J Clin Invest 2014; 124: 40–46.
Cameron EE, Bachman KE, Myohanen S, Herman JG, Baylin SB . Synergy of demethylation and histone deacetylase inhibition in the re-expression of genes silenced in cancer. Nat Genet 1999; 21: 103–107.
Heller G, Fong KM, Girard L, Seidl S, End-Pfützenreuter A, Lang G et al. Expression and methylation pattern of TSLC1 cascade genes in lung carcinomas. Oncogene 2006; 25: 959–968.
Heller G, Altenberger C, Schmid B, Marhold M, Tomasich E, Ziegler B et al. DNA methylation transcriptionally regulates the putative tumor cell growth suppressor ZNF677 in non-small cell lung cancers. Oncotarget 2015; 6: 394–408.
Heller G, Weinzierl M, Noll C, Babinsky V, Ziegler B, Altenberger C et al. Genome-wide miRNA expression profiling identifies miR-9-3 and miR-193a as targets for DNA methylation in non-small cell lung cancers. Clin Cancer Res 2012; 18: 1619–1629.
Krueger F, Andrews SR . Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 2011; 27: 1571–1572.
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D et al. Circos: an information aesthetic for comparative genomics. Genome Res 2009; 19: 1639–1645.
McLean CY, Bristor D, Hiller M, Clarke SL, Schaar BT, Lowe CB et al. GREAT improves functional interpretation of cis-regulatory regions. Nat Biotechnol 2010; 28: 495–501.
Goecks J, Nekrutenko A, Taylor J . Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome Biol 2010; 11: R86.
Blankenberg D, Von Kuster G, Coraor N, Ananda G, Lazarus R, Mangan M et al. Galaxy: a web-based genome analysis tool for experimentalists. Curr Protoc Mol Biol Chapter 19 (Unit): 191–21.
Giardine B, Riemer C, Hardison RC, Burhans R, Elnitski L, Shah P et al. Galaxy: a platform for interactive large-scale genome analysis. Genome Res 2005; 15: 1451–1455.
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL . TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 2013; 14: R36.
Robinson JT, Thorvaldsdottir H, Winckler W, Guttman M, Lander ES, Getz G et al. Integrative genomics viewer. Nat Biotechnol 2011; 29: 24–26.
Erdreich-Epstein A, Robison N, Ren X, Zhou H, Xu J, Davidson TB et al. PID1 (NYGGF4), a new growth-inhibitory gene in embryonal brain tumors and gliomas. Clin Cancer Res 2014; 20: 827–836.
Agrawal-Singh S, Isken F, Agelopoulos K, Klein HU, Thoennissen NH, Koehler G et al. Genome-wide analysis of histone H3 acetylation patterns in AML identifies PRDX2 as an epigenetically silenced tumor suppressor gene. Blood 2012; 119: 2346–2357.
Kohno T, Otsuka T, Takano H, Yamamoto T, Hamaguchi M, Terada M et al. Identification of a novel phospholipase C family gene at chromosome 2q33 that is homozygously deleted in human small cell lung carcinoma. Hum Mol Genet 1995; 4: 667–674.
Shan Z, Shakoori A, Bodaghi S, Goldsmith P, Jin J, Wiest JS . TUSC1, a putative tumor suppressor gene, reduces tumor cell growth in vitro and tumor growth in vivo. PLoS One 2013; 8: e66114.
Heller G, Zielinski CC, Zöchbauer-Müller S . Lung cancer: from single-gene methylation to methylome profiling. Cancer Metastasis Rev 2010; 29: 95–107.
Uehara E, Takeuchi S, Yang Y, Fukumoto T, Matsuhashi Y, Tamura T et al. Aberrant methylation in promoter-associated CpG islands of multiple genes in chronic myelogenous leukemia blast crisis. Oncol Lett 2012; 3: 190–192.
Janssen JJ, Denkers F, Valk P, Cornelissen JJ, Schuurhuis GJ, Ossenkoppele GJ . Methylation patterns in CD34 positive chronic myeloid leukemia blast crisis cells. Haematologica 2010; 95: 1036–1037.
Michels KB, Binder AM, Dedeurwaerder S, Epstein CB, Greally JM, Gut I et al. Recommendations for the design and analysis of epigenome-wide association studies. Nat Methods 2013; 10: 949–955.
Gu H, Bock C, Mikkelsen TS, Jager N, Smith ZD, Tomazou E et al. Genome-scale DNA methylation mapping of clinical samples at single-nucleotide resolution. Nat Methods 2010; 7: 133–136.
Bock C . Analysing and interpreting DNA methylation data. Nat Rev Genet 2012; 13: 705–719.
Cea M, Cagnetta A, Nencioni A, Gobbi M, Patrone F . New insights into biology of chronic myeloid leukemia: implications in therapy. Curr Cancer Drug Targets 2013; 13: 711–723.
Perini G, Diolaiti D, Porro A, Della Valle G . In vivo transcriptional regulation of N-Myc target genes is controlled by E-box methylation. Proc Natl Acad Sci USA 2005; 102: 12117–12122.
Kim J, Kollhoff A, Bergmann A, Stubbs L . Methylation-sensitive binding of transcription factor YY1 to an insulator sequence within the paternally expressed imprinted gene, Peg3. Hum Mol Genet 2003; 12: 233–245.
Wang H, Maurano MT, Qu H, Varley KE, Gertz J, Pauli F et al. Widespread plasticity in CTCF occupancy linked to DNA methylation. Genome Res 2012; 22: 1680–1688.
Vire E, Brenner C, Deplus R, Blanchon L, Fraga M, Didelot C et al. The Polycomb group protein EZH2 directly controls DNA methylation. Nature 2006; 439: 871–874.
Heller G, Schmidt WM, Ziegler B, Holzer S, Müllauer L, Bilban M et al. Genome-wide transcriptional response to 5-aza-2'-deoxycytidine and trichostatin a in multiple myeloma cells. Cancer Res 2008; 68: 44–54.
Liu ZJ, Zhang J, Gao YH, Pei LR, Zhou J, Gu LK et al. Large-scale characterization of DNA methylation changes in human gastric carcinomas with and without metastasis. Clin Cancer Res 2014; 20: 4598–4612.
Heller G, Geradts J, Ziegler B, Newsham I, Filipits M, Markis-Ritzinger EM et al. Downregulation of TSLC1 and DAL-1 expression occurs frequently in breast cancer. Breast Cancer Res Treat 2007; 103: 283–291.
Aggerholm A, Holm MS, Guldberg P, Olesen LH, Hokland P . Promoter hypermethylation of p15INK4B, HIC1, CDH1, and ER is frequent in myelodysplastic syndrome and predicts poor prognosis in early-stage patients. Eur J Haematol 2006; 76: 23–32.
Tang YM, Wo YY, Stewart J, Hawkins AL, Griffin CA, Sutter TR et al. Isolation and characterization of the human cytochrome P450 CYP1B1 gene. J Biol Chem 1996; 271: 28324–28330.
DiNardo CD, Gharibyan V, Yang H, Wei Y, Pierce S, Kantarjian HM et al. Impact of aberrant DNA methylation patterns including CYP1B1 methylation in adolescents and young adults with acute lymphocytic leukemia. Am J Hematol 2013; 88: 784–789.
Wang Z, Zhang J, Ye M, Zhu M, Zhang B, Roy M et al. Tumor suppressor role of protein 4.1B/DAL-1. Cell Mol Life Sci 2014; 71: 4815–4830.
Issa JP, Gharibyan V, Cortes J, Jelinek J, Morris G, Verstovsek S et al. Phase II study of low-dose decitabine in patients with chronic myelogenous leukemia resistant to imatinib mesylate. J Clin Oncol 2005; 23: 3948–3956.
Oki Y, Kantarjian HM, Gharibyan V, Jones D, O'Brien S, Verstovsek S et al. Phase II study of low-dose decitabine in combination with imatinib mesylate in patients with accelerated or myeloid blastic phase of chronic myelogenous leukemia. Cancer 2007; 109: 899–906.
Schmidt M, Rinke J, Schafer V, Schnittger S, Kohlmann A, Obstfelder E et al. Molecular-defined clonal evolution in patients with chronic myeloid leukemia independent of the BCR-ABL status. Leukemia 2014; 28: 2292–2299.
Soverini S, de Benedittis C, Mancini M, Martinelli G . Mutations in the BCR-ABL1 kinase domain and elsewhere in chronic myeloid leukemia. Clin Lymphoma Myeloma Leuk 2015; 15 (Suppl): S120–S128.
Makishima H, Jankowska AM, McDevitt MA, O'Keefe C, Dujardin S, Cazzolli H et al. CBL, CBLB, TET2, ASXL1, and IDH1/2 mutations and additional chromosomal aberrations constitute molecular events in chronic myelogenous leukemia. Blood 2011; 117: e198–e206.
Acknowledgements
This study was supported by research funding from the Austrian Science Fund (FWF) through projects P24130 and SFB F4709-B20 to SZM, by a Clinical Research Grant of the Austrian Society of Hematology and Oncology to GH, SFB project F4704-B20 to PV and SFB project F4705-B20 to TL. We thank the Biomedical Sequencing Facility at CeMM for technical advice and for performing the next-generation sequencing experiments.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
PV received research grants from Novartis and Ariad. The other authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Heller, G., Topakian, T., Altenberger, C. et al. Next-generation sequencing identifies major DNA methylation changes during progression of Ph+ chronic myeloid leukemia. Leukemia 30, 1861–1868 (2016). https://doi.org/10.1038/leu.2016.143
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2016.143
- Springer Nature Limited
This article is cited by
-
SLIT2 promoter hypermethylation predicts disease progression in chronic myeloid leukemia
European Journal of Medical Research (2022)
-
DNA methylation: a saga of genome maintenance in hematological perspective
Human Cell (2022)
-
DNA methylation as predictive marker of response to immunotherapy?
memo - Magazine of European Medical Oncology (2021)
-
Imatinib independent aberrant methylation of NOV/CCN3 in chronic myelogenous leukemia patients: a mechanism upstream of BCR-ABL1 function?
Cell Communication and Signaling (2019)
-
Dnmt1 links BCR-ABLp210 to epigenetic tumor stem cell priming in myeloid leukemia
Leukemia (2019)