Abstract
Children with P2RY8-CRLF2-positive acute lymphoblastic leukemia have an increased relapse risk. Their mutational and transcriptional landscape, as well as the respective patterns at relapse remain largely elusive. We, therefore, performed an integrated analysis of whole-exome and RNA sequencing in 41 major clone fusion-positive cases including 19 matched diagnosis/relapse pairs. We detected a variety of frequently subclonal and highly instable JAK/STAT but also RTK/Ras pathway-activating mutations in 76% of cases at diagnosis and virtually all relapses. Unlike P2RY8-CRLF2 that was lost in 32% of relapses, all other genomic alterations affecting lymphoid development (58%) and cell cycle (39%) remained stable. Only IKZF1 alterations predominated in relapsing cases (P=0.001) and increased from initially 36 to 58% in matched cases. IKZF1’s critical role is further corroborated by its specific transcriptional signature comprising stem cell features with signs of impaired lymphoid differentiation, enhanced focal adhesion, activated hypoxia pathway, deregulated cell cycle and increased drug resistance. Our findings support the notion that P2RY8-CRLF2 is dispensable for relapse development and instead highlight the prominent rank of IKZF1 for relapse development by mediating self-renewal and homing to the bone marrow niche. Consequently, reverting aberrant IKAROS signaling or its disparate programs emerges as an attractive potential treatment option in these leukemias.
Similar content being viewed by others

Introduction
Acute lymphoblastic leukemia (ALL) is the predominant malignancy in children and adolescents and one of the leading causes of death in these age groups. It comprises multiple entities with gene fusions, such as ETV6-RUNX1, BCR-ABL1, TCF3-PBX1 and those with KMT2A that are associated with distinct clinical features and prognosis.1 Recently, a novel subgroup has been described whose defining characteristic is the deregulated expression of the cytokine receptor-like factor 2 (CRLF2) gene, which is located in the pseudoautosomal region 1 on the short arm of the X and Y chromosome. The two most common causative genetic defects are a small interstitial deletion that fuses the first non-coding exon of the purinergic receptor P2Y, G-protein coupled, 8 (P2RY8) to the entire coding region of CRLF2 and occurs in 5–8% of childhood B-cell precursor (BCP) ALL cases and a more rare translocation that places CRLF2 under the control of the IGH enhancer.2, 3 Activating mutations of the CRLF2 or IL7R genes are far less common.4, 5, 6 In contrast to IGH-CRLF2 translocations, which occur in a hematopoietic stem/precursor cell and are initiating events, P2RY8-CRLF2 fusions are caused by illegitimate V(D)J-mediated recombination in a B precursor cell, and are, therefore, likely secondary events and define a more heterogeneous group of leukemias.7 As such, they accompany bona fide primary abnormalities, such as intrachromosomal amplification of chromosome 21 (iAMP21) and, albeit less frequently, hyperdiploidy, but are present only in small subclones in the vast majority of cases and these fusions are often lost at relapse.8, 9 Further, they show a strong affinity for constitutional as well as acquired numerical chromosome 21 abnormalities.3, 4, 8 P2RY8-CRLF2 fusions often carry additional alterations in JAK/STAT pathway genes and they may cooperatively activate downstream pathways.4, 8, 10, 11 They are associated with a significantly increased relapse risk in AIEOP/BFM protocols, which is independent of the size of the P2RY8-CRLF2-positive clone. Respective cases are primarily classified as non-high risk by clinical and molecular response criteria and relapses occur predominantly late.9, 12, 13, 14
IKZF1 encodes the lymphoid transcription factor IKAROS, which is a key regulator in early lymphocyte development.15 Alterations mainly comprise deletions and only rarely sequence alterations.16, 17, 18 Irrespective of the type of IKZF1 alteration, they prevail in poor responding cases in major treatment protocols.5, 11, 17, 19, 20, 21, 22, 23 They are a hallmark of high-risk BCP ALL, especially those, which carry a BCR-ABL1 and other cytokine- and kinase-activating fusions, including IGH-CRLF2 and P2RY8-CRLF2.11, 24 The common characteristic of such cases is a gene expression signature that resembles that of genuine BCR-ABL1-positive cases and which are, therefore, also referred to as either 'BCR-ABL1-like' or 'Ph-like'.11, 24 IKZF1 alterations still confer a dismal prognosis in such cases, even when treated on high risk or kinase inhibitor-containing protocols.1, 16, 17, 24, 25, 26, 27, 28
IKZF1 deletions were also reported in small cohorts of P2RY8-CRLF2-positive leukemia cases recruited to various treatment protocols. It, therefore, seems likely that their presence may contribute to relapse development in this particular subgroup, a notion that has so far not been systematically investigated in large and well-characterized cohorts. Notably, hardly any detailed information about concurring alterations, their subclonal or clonal nature as well as whether any or which of them are preserved at relapse, is available. To address these issues we performed whole-exome sequencing (WES) and determined the genome-wide mutational pattern in a quantitative manner in 41 relapsing and non-relapsing P2RY8-CRLF2-positive cases that were treated primarily according to BFM protocols. Special emphasis was put on defining the size, mutation patterns and distribution of clones at diagnosis and relapse, which enabled us to deduce the kinetics and indirectly also the in vivo selection of particular clones. Together with the transcriptional profiling of these cases, this approach allowed us not only to gain new insights into the mechanisms of relapse development but also to infer the biological impact of genomic alterations, their potential role in resistance mechanisms as well as their applicability as biomarkers and drug targets.
Materials and methods
Patients and samples
Selection of BCP ALL cases was based on the presence of a P2RY8-CRLF2 fusion in the major clone at initial diagnosis, the absence of a KMT2A, BCR-ABL or ETV6-RUNX1 rearrangement, and on the availability of DNA or frozen viable cells with a blast count of ⩾85% at diagnosis or ⩾70% at relapse. In total, 52 children (including 23 individuals with Down syndrome (DS)) with a median age of 3.8 years (range, 1.5–13.7) were included. Cases were initially treated on BFM ALL 2000/09 (n=32), CoALL-08-09 (n=9), FRALLE, EORTC58951 and EORTC58081 protocols (n=11; www.ClinicalTrials.gov: NCT00411541, NCT00005603, NCT00343369 and NCT0114348). Leukemic samples from 41 cases, including 19 with matched relapses, were available for WES. From the remaining 11 cases (from FRALLE and EORTC studies) only multiplex ligation-dependent probe amplification (MLPA)-derived copy number alterations (CNA) were available. Remission samples served as germline control in all instances. All non-relapsing cases were in long-term remission (median, 7 years; range, 4–14 years). Relapsing cases had higher white blood cell count at initial diagnosis (Table 1) but did not differ in any other clinical or response criteria from the non-relapsing cohort. The study cases are representative of an unselected cohort of 20 respective ALL cases consecutively recruited to the BFM ALL 2009 study in Austria (Table 1). The detailed clinical data of cases with matched diagnosis and relapse samples are summarized in Table 2. The median time to relapse was 35 months (range 4–97). At relapse, the majority of cases (9 of 16 with available data) had a poor molecular response.29
Samples were obtained from the respective study centers upon institutional review and ethics committees’ approval. Informed consent for tissue banking and research studies was obtained from patients, their parents or legal guardians in accordance with the Declaration of Helsinki.
Sample preparation and WES
WES was performed as described previously.30 Samples were prepared using Nextera Rapid Capture Exome Kit (Illumina, San Diego, CA, USA). Paired-end sequencing of 100 bp reads was performed on a HiSeq 2000 (Illumina) to obtain at least a 50 × coverage. More detailed coverage metrics computed by Picard (https://github.com/broadinstitute/picard) are provided in the Supplementary Methods, additional file 1. Mutations were validated with Sanger sequencing (Supplementary Results).
Determination of CNA by PCR, MLPA and single-nucleotide polymorphism array analysis
Genomic breakpoint amplification of P2RY8-CRLF2 was performed as described previously.9 The MLPA probe mixes used for detection of IKZF1 deletions were SALSA MLPA probe mix P202-B1 IKZF1 and P335 ALL-IKZF1 (MRC-Holland, Amsterdam, The Netherlands). Single-nucleotide polymorphism array analysis was performed using CytoScan HD arrays (Affymetrix, Santa Clara, CA, USA) and data analysis was done with the Chromosome Analysis Suite (ChAS) version 3.1 (Affymetrix). Summary of methods and analyzed samples can be found in Supplementary Methods, Supplementary Figure 1 and Supplementary Table 1.
Mutation and CNA detection from WES
Reads were aligned with BWA-MEM and mutations called with MuTect. SnpEff, dbNSFP and custom Perl scripts were used for variant annotation and filtering. WES-based CNA detection was performed with exomeCopy.
RNA sequencing and differential gene expression analysis
Total RNA was extracted from frozen primary leukemic samples (>90% blasts) upon Ficoll gradient separation using Quick-RNA MiniPrep kit (Zymo Research, Irvine, CA, USA). Selection of cases was based on the availability of material. Sequencing libraries were prepared using the NEBNext Poly(A) mRNA Magnetic Isolation Module and the NEBNext Ultra Directional RNA Library Prep Kit for Illumina (New England Biolabs, Ipswich, MA, USA) and sequenced on a HiSeq 2000 (Illumina) at the Genomics Core Facility, EMBL, Heidelberg. RNA-Seq data analysis was performed with a custom pipeline implemented in Anduril using GSNAP for read alignment, htseq-count for read counting and DESeq2 for differential gene expression analysis. Gene set enrichment analysis was performed with GSEA.
Statistical analyses
Associations between categorical variables were examined using Fisher's exact and Mann–Whitney tests. Event-free survival (EFS) and overall survival (OS) were analyzed according to the Kaplan–Meier method and compared by the log-rank test.
Data availability
WES and RNA-Seq data have been deposited at the European Genome-phenome Archive (accession number EGAS00001001847). WES data are available for academic purposes by contacting the corresponding author, as the patient/parent consent does not cover depositing data that can be used for large-scale determination of germline variants.
Results
Somatic sequence and CNAs of 41 P2RY8-CRLF2-positive ALL cases are summarized in Figure 1. Including mutations with an allelic frequency of ⩾10%, the median number of somatic, predicted deleterious sequence mutations per case was 12 (range, 4–44) at initial diagnosis and 14 (range, 4–474) at relapse (Supplementary Methods, additional file 2). CNAs were detected using a custom pipeline that predicts CNAs/microdeletions with high sensitivity and specificity from WES data. Single-nucleotide polymorphism array, MLPA and P2RY8-CRLF2 breakpoint analyses served for validation (Supplementary Figures 2 and 3, additional files 3 and 4). Integrated analysis of genomic sequence and CNAs showed that all leukemias had at least one aberration in putative leukemic driver genes that were grouped according to pathways or biological processes (Figure 1). In addition, copy number and sequence mutations are represented according to the constitutional status of cases (Supplementary Results and Supplementary Figure 4).
Genomic sequence and copy number alterations of 41 P2RY8-CRLF2-positive ALL cases. Recurrently altered genes in relapsing cases (left) and non-relapsing cases (right) in columns. Non-silent, predicted deleterious sequence and copy number alterations in genes (rows) are listed according to functional groups in diagnostic (top) and relapse (bottom) samples. Mutations are marked by color codes (as indicated) to show their clonal or subclonal nature based on adjusted allelic frequency (adj. AF), predicted functional effects and conservation from diagnosis to relapse. CN chr21>2, somatic gain of chromosome 21; DS, Down syndrome; OG, oncogene; Sex Chrs. abnorm, copy number aberration of sex chromosomes; TS, tumor suppressor; TTR, time to relapse; UPN, unique patient number (columns).
Clonal heterogeneity and instability of kinase-activating pathway alterations
Irrespective of the presence of a P2RY8-CRLF2 fusion, 31/60 leukemic samples harbored mutations in JAK/STAT pathway components: JAK2 (n=21), CRLF2 (n=3), JAK3 (n=3), JAK1 (n=2), IL7RA (n=1) and SYK (n=1; Supplementary Table 2). JAK1 and, more frequently, JAK2 mutations are known to occur in BCP ALL,31, 32 whereas the recurrent, activating JAK3 mutation in the pseudokinase domain (Supplementary Figure 5) has only been documented in JMML and T-ALL so far,33, 34 but drives B-cell leukemia in mice.35 All mutations in CRLF2 (F232C) and IL7RA (S185C) are activating mutations and have been reported previously.4, 6, 36
At initial diagnosis, the overall frequency of JAK/STAT pathway mutations was 51% and equally distributed between relapsing and non-relapsing cases. They were subclonal in 47% of cases, including two with two mutations each. Notwithstanding of their original extent, JAK/STAT pathway mutation carrying clones were lost at relapse in 60% of the cases. Two subclonal mutations developed into the dominant clone and four were newly acquired, resulting in an overall relapse frequency of 42% (Figures 1, 2a and 4a; Supplementary Table 2).
Clonal composition and stability of signaling mutations. (a) Clonal composition of JAK/STAT (blue) and RTK/Ras pathway (red) signaling gene mutations according to individual genes (symbol code). Dots represent the adjusted allelic frequency (adj. AF) of mutations. Black dots mark conserved mutations at diagnosis and relapse. (b) Representative examples of the clonal distribution of selected abnormalities at diagnosis and relapse with either a conserved (top three) or lost (bottom three) P2RY8-CRLF2 fusion. Blots show the adj. AF of deleterious sequence (green), copy number (blue) and P2RY8-CRLF2 (pseudoautosomal region 1 (PAR1); red) alterations at initial diagnosis (x axis), and at relapse (y axis). Dashed lines indicate the border between subclonal and clonal mutations. (c) Mutational patterns of genomic alterations of cases with multiple relapses. Adj. AF of sequence and copy number alterations (y axis) at indicated leukemia occasions (x axis). Dashed lines highlight the loss or gain of distinct mutations. A gray horizontal line separates subclonal from clonal (⩾30% adj. AF) mutations. D, initial diagnosis; R, first relapse; RR, second relapse.
Mutations in RTK/Ras pathway genes were seen in 29% of the cases at diagnosis, whereby the majority of them concurred with a somatic gain of chromosome 21, but, in accordance with a recent publication, they were only found in 1/23 DS ALL cases.37 In line with the high instability of these clones, the frequency of such mutations increased to 68% in relapses (P=0.06), including four with DS (Figures 1, 2a and 4a; Supplementary Table 3). Alterations preferably affected KRAS, followed by NRAS, FLT3, PTPN11 and CBL.
Taken together, JAK/STAT (excluding P2RY8-CRLF2) and RTK/Ras pathway mutations were found in 78% of all cases at diagnosis and increased to 95% at relapse, but the original mutations were nevertheless only conserved in less than half of the cases (Supplementary Figure 6). In accordance with this finding, P2RY8-CRLF2 was also lost in 32% of relapses. It is worth noting that RTK/RAS and JAK/STAT pathway alterations were found in three instances, in two of them they obviously even concurred within the same clone.
Recurrent alterations affecting lymphoid differentiation and tumor suppressor genes
The vast majority of alterations in genes that are critical for B-cell development and differentiation, tumor suppression and cell cycle regulation are microdeletions. Including CNA data from additional 11 non-relapsing ALL cases (Supplementary Figure 7), the total evaluable number of cases amounts to 52.
At diagnosis, 58% of them had lesions in lymphoid differentiation genes, which accords with previous observations.38 They comprise alterations in PAX5 (30.7%), IKZF1 (23%), ETV6 (11.5%), EBF1 (4%) and IKZF3 (2%). Deletions in the tumor suppressor/cell cycle regulator genes CDKN2A/B were found in 38.5%.
At relapse, respective sequence and CNA were always conserved and only few were additionally acquired in some instances, altogether adding up to 58% IKZF1, 42% CDKN2A/B and 26% PAX5 deletions, respectively (Figure 1 and 4a). Similarly, all four ETV6 alterations (two deletions and two mutations in the ETS domain (E425* and R369P39)) remained conserved. The E425* mutation is novel in BCP ALL.
Gains or losses of sex chromosomes were seen in 27% of relapsing and non-relapsing cases. Sex chromosome losses at relapse did not concord with that of P2RY8-CRLF2, which indicates that at least in the informative cases relapses must have evolved from a fusion-negative ancestral clone.
Relapse-predicting and -associated alterations in P2RY8-CRLF2-positive ALL
Except for IKZF1, the frequency of alterations in genes implicated in lymphoid development and in tumor suppression/cell cycle regulation did not significantly differ between relapsing and non-relapsing cases. IKZF1 alterations prevailed in the relapsing cohort with a frequency of 41% versus 10% in non-relapsing cases (P=0.001). Two of the latter had deletions and one a subclonal mutation (Figure 1; Supplementary Table 4).
At relapse, 58% of cases harbored an IKZF1 alteration (including three relapse-specific ones), whereby all initially detected IKZF1 alterations remained conserved. The only exception was one case (#92) with an originally biallelic alteration, in which the G158S mutation27, 40 was lost but the deletion preserved. Intragenic IKZF1 deletions result in the expression of the dominant-negative isoform (IK6) in 36% of relapses, whereas deletions of the entire gene presumably lead to haploinsufficiency.16, 17, 41 In two cases a dominant-negative acting IKZF1 mutation concurred with a deletion. In 63% of cases, either PAX5 and/or CDKN2A/B alterations were also present implying that these combinations might confer an evolutionary advantage.17, 27
In line with previous reports, we noted also a high number of alterations (32–38%) in epigenetic regulator encoding genes such as CREBBP, KMT2D and SETD2.30, 42, 43, 44 However, the frequency of these alterations was similar in relapsing and non-relapsing cases as well as in diagnosis and relapse samples, which indicates that they have little direct relevance in the disease recurrence process.
Size and kinetics of mutant clones in relapsing leukemias
Given the instability of P2RY8-CRLF2-positive clones,9 we next compared the mutation patterns in matched diagnosis and relapse samples from the 19 cases in different settings. Figure 2b shows six prototypic cases, three in which the P2RY8-CRLF2 fusion was conserved (top) and three in which it was lost at relapse (bottom). Other concurring kinase-activating alterations were either retained (n=6), replaced by another one (n=5), lost (n=1) or evolved from an initial subclonal to a clonal one at relapse (n=2). Moreover, five cases without an initial kinase-activating mutation acquired one at relapse (Supplementary Figure 8). The respective patterns were similar in cases with conserved or lost P2RY8-CLRF2 fusions.
In five cases we were also able to study subsequent relapses (Figure 2c). CDKN2A, PAX5 and IKZF1 were either retained or newly acquired in second relapses (#715, S23, AL9890), whereas kinase-activating alterations were exchanged in the second relapse in three cases.
IKZF1 alterations are associated with dismal outcome
Except for the genomic IKZF1 status and white blood cell count that were not associated with each other (P=0.66), we found no other biological or clinical parameters such as other genetic alterations, age at diagnosis, clinical risk group assignment or morphological and molecular response to treatment to be correlated with the occurrence of relapses. There was no difference between DS and non-DS cases. Of note, only one of eight IKZF1-altered relapsing cases had initially a poor MRD response and was, therefore, assigned to high-risk treatment, but none of them had initially a poor response to glucocorticoids in vivo, as one might have expected based on in vitro experiments.45 Yet, IKZF1-mutated cases had a significantly poorer outcome than their IKZF1 wild-type counterparts as evidenced by an adverse pEFS (P=0.026) and pOS (P=0.051; Supplementary Figure 9).
IKZF1 alterations display distinctive transcriptional signatures
As IKZF1 alterations are highly associated with relapses in P2RY8-CRLF2-positive ALL cases, we profiled 22 leukemia samples and analyzed the RNA-Seq data according to their IKZF1 status (Supplementary Table 4). We designated IK6 deletions and biallelic alterations as 'IKN', larger deletions of IKZF1 as 'IKD' and wtIKZF1 ones as 'IKC'. These analyses revealed specific regulations in the IKN group and, albeit to a much lesser degree, in the IKD group compared with the IKC one (Supplementary Methods, additional files 5 and 6). Unsupervised hierarchical clustering of the 200 most differentially expressed genes in the IKN and IKD cohort grouped them according to their IKZF1 status (Figure 3a; Supplementary Figure 10). This finding was confirmed in an independent RNA-Seq data set of 20 mainly B-other ALL cases with known IKZF1 status, where a significant correlation in the log2-fold expression of the top 50 up- and downregulated genes from the IKN cohort was observed (P=0.0039, Spearman correlation; Supplementary Methods, additional file 7).
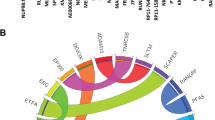
Transcriptional signature of leukemias according to IKZF1 status. (a) Cluster heatmap of the top 50 up and downregulated genes in both IKN (IKZF1 alterations leading to a dominant-negative effect and biallelic alterations) and IKD (IKZF1 deletions resulting in haploinsufficiency) samples according to fold-change (P⩽1E−8 for IKN versus IKC, P⩽2E−3 for IKD versus IKC); IKN cases are indicated in red, IKD ones in blue and controls in gray at the top of the map. (b) Selected gene set enrichment plots showing a significant concordant regulation with IKN and IKD signatures. P-values are indicated in each analysis. FDR, false discovery rate; NES, normalized enrichment score.
Consequently, we used the IKN and IKD cohorts for gene set enrichment analysis and found a highly significant correlation of differentially expressed genes with developmental and biological processes (Supplementary Methods, additional file 8). Representative examples are shown in Figure 3b and Table 3. Differentially expressed genes in the IKN group, and less so in the IKD cohort, were significantly enriched for various human hematopoietic and lymphoid stem cells sets, and concordantly for gene sets specifically expressed in immature B cells. This suggests that IKZF1 alterations lead to impaired B-cell differentiation and the acquisition of stem cell-like features, as described for murine and human BCR-ABL+ leukemias and 'Ph-like' human leukemias.17, 27 Consistent with the adhesive properties of Ikzf1−/− pro/pre-B cells in mice,46, 47 gene sets that are upregulated in the context of microenvironment, focal adhesion kinase and integrin pathways were enriched. We also found an enrichment of genes that are higher expressed in response to hypoxia, downstream VEGF/VEGFR signaling and to EPO signaling.
In line with normal murine pro-B cells lacking Ikzf1,46 genes critical for cell cycle regulation were negatively enriched in IKZF1-altered compared with wtIKZF1 samples and they had lower expression of genes involved in DNA replication and proliferation. Moreover, IKZF1-mutant samples were negatively enriched for major DNA damage and repair pathways. By contrast, JAK/STAT, PI3K and mTOR, as well as Ras and MAPK pathway signaling gene sets were positively enriched in the IKZF1-mutant samples, a finding that partly might reflect the proposed negative regulation of STAT5 signaling by IKZF1.48 In concordance with the enrichment of resistance gene sets derived from primary childhood ALL samples, the majority of our relapse cases with IKZF1 alterations had also a poor response to chemotherapy (Figure 1; Table 2).
Our IKZF1 signature was further corroborated by comparison with respective data from human leukemias and murine models by revealing a significant enrichment of differentially expressed genes in IKZF1-mutant samples (Supplementary Figure 11).
Discussion
CRLF2 overexpression is the common hallmark of two distinct genetic subtypes of childhood ALL with a high recurrence risk.2, 3, 13 Most of the cases with IGH-CRLF2 translocations belong to the high-risk group, whereas those with a P2RY8-CRLF2 fusion primarily fall into the intermediate-risk group.8, 9, 12, 13, 14 The genomic components of the latter and especially their role in the disease and particularly in the relapse process remain, to a large extent, elusive. We therefore analyzed the genomic and transcriptional profile of altogether 41 cases, including 21 cases with DS, that had a P2RY8-CRLF2 fusion in the dominant clone at initial diagnosis. They comprised 22 relapsing cases with 19 matched diagnosis and relapse pairs as well as 20 non-relapsing ones. This largest cohort of its kind reported so far allowed us to determine and examine those genetic lesions that were lost, retained or acquired in relapses.
The P2RY8-CRLF2 fusion was lost in one-third of the relapses. Together with previous observations-that, for instance, these fusions frequently affect subclones that never evolve into major relapse clones-indicate that it is primarily a secondary change that may potentially supply the respective cells with a certain proliferative but certainly not with an evolutionary advantage.9
In line with other types of childhood BCP ALL,49 B-cell differentiation genes are also commonly deleted in cases with a P2RY8-CRLF2 fusion. These abnormalities are usually preserved in the corresponding relapses. Consistent with their role in drug resistance, IKZF1 alterations prevail already in relapse-prone cases at diagnosis but become even more abundant in relapses. IKZF1-deleted cases also seem to profit from other associated B-cell development and cell cycle gene defects, particularly those of PAX5 and/or CDKN2A/B, which are found in half of the cases, but also from mutations in specific proliferation-promoting pathway genes.16, 17, 50
Aside from a high white blood cell count, IKZF1 deletions are also the only relapse-predicting molecular indicator of P2RY8-CRLF2-positive ALL cases. As IKZF1 deletions as well as the vast majority of other genomic abnormalities are similarly frequent in P2RY8-CRLF2-positive DS and non-DS ALL at diagnosis and relapse, one can expect that IKZF1 deletions have a similar role in both cohorts. This seems indeed to be the case, although confounding factors, such as the greater treatment toxicity in DS patients, may influence and bias such outcome results to some extent.23, 51, 52
As primarily reported for DS ALL, our P2RY8-CRLF2-positive cases frequently harbor JAK/STAT but also Ras pathway mutations, with those in the first pathway prevailing at diagnosis.3, 4, 5, 10, 37, 53, 54 These alterations were usually subclonal, even concurred in three cases, and were lost at relapse in half of the cases. Although almost all relapse cases had mutations in genes that are involved in either signaling pathway, those affecting the Ras pathway predominated. This switch from the JAK/STAT to the Ras pathway was neither related to the preexisting or acquired copy number state of chromosome 21 nor to the leukemias’ underlying primary abnormality as reported in DS ALL.37 As P2RY8-CRLF2-positive leukemias not only have JAK/STAT but also PI3K/mTOR signaling activated,55, 56 it is interesting to note that RAS alterations activate both the RAF/MEK/ERK and PI3K/mTOR pathways.57 Thus, this leukemia subgroup seems to be particularly promiscuous regarding its preference for proliferative driver mutations, which they likely require for the emergence of drug-resistant relapses. Although IKZF1 deletions might enhance STAT5 signaling in JAK-mutated cases,48 our findings provide some circumstantial evidence that Ras pathway alterations could interact in a similar fashion and substitute those in the JAK/STAT pathway.
Taken together, we consider the P2RY8-CRLF2 fusion as one of the secondary proliferative driver alterations that-in line with those activating JAK/STAT and RTK/Ras pathways-are highly instable at relapse, albeit to a lesser degree if initially present in the major clone,30 and not as a bona fide primary genetic alteration that is always stable at relapse and critical for the maintenance of the leukemia (Figure 4a). We, therefore, illustrate the various scenarios by showing prototypic cases where either the P2RY8-CRLF2 fusion and/or a JAK/STAT and/or RTK/Ras pathway mutation is lost at relapse, and instead another proliferative driver mutation in one of the two pathways occurs (Figures 4b–e). As chemotherapy primarily eliminates the bulk of rapidly proliferating cells, it spares those less active, resistant stem cell-like ones that eventually generate relapses.58 In this scenario, the P2RY8-CRLF2 fusion is only one of several proliferation-activating alterations and merely serves as a common marker for an otherwise genetically heterogeneous group, whose other and probably more relevant features are IKZF1 deletions. These deletions are well-known disease drivers in many types of drug-resistant leukemias, such as BCR-ABL1-positive ones, which share a similar gene expression signature.11, 24 The transcriptional profile of IKZF1-deleted cases reflects their strong homing preference to the bone marrow niche as well as their high repopulation capacity, attributes that also become apparent in mouse models in which Ikzf1−/− pro/pre-B cells acquire stem cell and adhesion properties including activation of the focal adhesion kinase pathway.27, 46, 47, 59
Clonal evolution of leukemia and selection of relapse clones. On the basis of the frequency and stability of genomic alterations, depicted in (a) for the most frequently altered genes and pathways, we envision that RAG-mediated microdeletions affecting B-cell differentiation and tumor suppressor genes follow the occurrence of a leukemia-initiating (founder) alteration. Given their stability and increased frequency at relapse they apparently foster resistance and emergence of relapse. The ensuing continuous emergence of JAK/STAT and/or RTK/Ras pathway mutations eventually leads to the clinical manifestation of leukemia, whereby the type of proliferation-promoting alterations greatly varies. (b) Shown is case 108, which harbors, in addition to alterations in IKZF1, CDKN2A/B and P2RY8-CRLF2 (pseudoautosomal region 1 (PAR1) deletion), also a JAK2 mutation in 50% of the population. The latter clone predominates at relapse. (c) Similarly, case DL2 carries a JAK2 mutation in 70% of the P2RY8-CRLF2+-, IKZF1-mutant population at diagnosis. The initial JAK2 mutation is, however, replaced by another one at relapse, where also a PAX5 deletion is present. (d) Case HV57, harboring an IKZF1 alteration, illustrates the loss of an initial predominant P2RY8-CRLF2+- and JAK2-mutant clone (the latter affecting ~60% of cells) and the emergence of a fusion negative, FLT3-mutant one. (e) In case B36, the P2RY8-CRLF2 fusion occurs in a CDKN2A/B- and JAK3-mutant clone and even harbors a KRAS mutation in 36% of cells. At relapse, P2RY8-CRLF2- and KRAS-mutant clones are replaced by an IKZF1-altered population carrying an NRAS mutation. Color code shown as inset of the graph.
Besides these biological insights, our findings also provide some clues that may become relevant in future treatment decisions. Apart from their prognostic implications, IKZF1 alterations may eventually serve as markers for specific therapeutic interventions. It appears reasonable to try to restore IKAROS signaling especially in those IKZF1-deleted cases that still have retained a functional wild-type allele. For the other 20% of cases with biallelic IKZF1 alterations,60 inhibition of the activated focal adhesion kinase pathway may become a viable treatment option. Such approaches might perhaps be combined with a cocktail of other signaling inhibitors given the availability of various JAK/STAT, Ras/MEK/ERK and PI3K/mTOR pathway inhibitors.55
References
Hunger SP, Mullighan CG . Redefining ALL classification: toward detecting high-risk ALL and implementing precision medicine. Blood 2015; 125: 3977–3987.
Russell LJ, Capasso M, Vater I, Akasaka T, Bernard OA, Calasanz MJ et al. Deregulated expression of cytokine receptor gene, CRLF2, is involved in lymphoid transformation in B-cell precursor acute lymphoblastic leukemia. Blood 2009; 114: 2688–2698.
Mullighan CG, Collins-Underwood JR, Phillips LA, Loudin MG, Liu W, Zhang J et al. Rearrangement of CRLF2 in B-progenitor- and down syndrome-associated acute lymphoblastic leukemia. Nat Genet 2009; 41: 1243–1246.
Hertzberg L, Vendramini E, Ganmore I, Cazzaniga G, Schmitz M, Chalker J et al. Down syndrome acute lymphoblastic leukemia, a highly heterogeneous disease in which aberrant expression of CRLF2 is associated with mutated JAK2: a report from the International BFM Study Group. Blood 2010; 115: 1006–1017.
Chen IM, Harvey RC, Mullighan CG, Gastier-Foster J, Wharton W, Kang H et al. Outcome modeling with CRLF2, IKZF1, JAK, and minimal residual disease in pediatric acute lymphoblastic leukemia: a Children's Oncology Group study. Blood 2012; 119: 3512–3522.
Yoda A, Yoda Y, Chiaretti S, Bar-Natan M, Mani K, Rodig SJ et al. Functional screening identifies CRLF2 in precursor B-cell acute lymphoblastic leukemia. Proc Natl Acad Sci USA 2010; 107: 252–257.
Tsai AG, Yoda A, Weinstock DM, Lieber MR . t(X;14)(p22;q32)/t(Y;14)(p11;q32) CRLF2-IGH translocations from human B-lineage ALLs involve CpG-type breaks at CRLF2, but CRLF2/P2RY8 intrachromosomal deletions do not. Blood 2010; 116: 1993–1994.
Ensor HM, Schwab C, Russell LJ, Richards SM, Morrison H, Masic D et al. Demographic, clinical, and outcome features of children with acute lymphoblastic leukemia and CRLF2 deregulation: results from the MRC ALL97 clinical trial. Blood 2011; 117: 2129–2136.
Morak M, Attarbaschi A, Fischer S, Nassimbeni C, Grausenburger R, Bastelberger S et al. Small sizes and indolent evolutionary dynamics challenge the potential role of P2RY8-CRLF2-harboring clones as main relapse-driving force in childhood ALL. Blood 2012; 120: 5134–5142.
Harvey RC, Mullighan CG, Chen IM, Wharton W, Mikhail FM, Carroll AJ et al. Rearrangement of CRLF2 is associated with mutation of JAK kinases, alteration of IKZF1, Hispanic/Latino ethnicity, and a poor outcome in pediatric B-progenitor acute lymphoblastic leukemia. Blood 2010; 115: 5312–5321.
Harvey RC, Mullighan CG, Wang X, Dobbin KK, Davidson GS, Bedrick EJ et al. Identification of novel cluster groups in pediatric high-risk B-precursor acute lymphoblastic leukemia with gene expression profiling: correlation with genome-wide DNA copy number alterations, clinical characteristics, and outcome. Blood 2010; 116: 4874–4884.
Palmi C, Vendramini E, Silvestri D, Longinotti G, Frison D, Cario G et al. Poor prognosis for P2RY8-CRLF2 fusion but not for CRLF2 over-expression in children with intermediate risk B-cell precursor acute lymphoblastic leukemia. Leukemia 2012; 26: 2245–2253.
Cario G, Zimmermann M, Romey R, Gesk S, Vater I, Harbott J et al. Presence of the P2RY8-CRLF2 rearrangement is associated with a poor prognosis in non-high-risk precursor B-cell acute lymphoblastic leukemia in children treated according to the ALL-BFM 2000 protocol. Blood 2010; 115: 5393–5397.
Attarbaschi A, Morak M, Cario G, Cazzaniga G, Ensor HM, te Kronnie T et al. Treatment outcome of CRLF2-rearranged childhood acute lymphoblastic leukaemia: a comparative analysis of the AIEOP-BFM and UK NCRI-CCLG study groups. Br J Haematol 2012; 158: 772–777.
Georgopoulos K . Haematopoietic cell-fate decisions, chromatin regulation and ikaros. Nat Rev Immunol 2002; 2: 162–174.
Mullighan CG, Miller CB, Radtke I, Phillips LA, Dalton J, Ma J et al. BCR-ABL1 lymphoblastic leukaemia is characterized by the deletion of Ikaros. Nature 2008; 453: 110–114.
Mullighan CG, Su X, Zhang J, Radtke I, Phillips LA, Miller CB et al. Deletion of IKZF1 and prognosis in acute lymphoblastic leukemia. N Engl J Med 2009; 360: 470–480.
Boer JM, van der Veer A, Rizopoulos D, Fiocco M, Sonneveld E, de Groot-Kruseman HA et al. Prognostic value of rare IKZF1 deletion in childhood B-cell precursor acute lymphoblastic leukemia: an international collaborative study. Leukemia 2016; 30: 32–38.
Clappier E, Grardel N, Bakkus M, Rapion J, De Moerloose B, Kastner P et al. IKZF1 deletion is an independent prognostic marker in childhood B-cell precursor acute lymphoblastic leukemia, and distinguishes patients benefiting from pulses during maintenance therapy: results of the EORTC Children's Leukemia Group study 58951. Leukemia 2015; 29: 2154–2161.
Dorge P, Meissner B, Zimmermann M, Moricke A, Schrauder A, Bouquin JP et al. IKZF1 deletion is an independent predictor of outcome in pediatric acute lymphoblastic leukemia treated according to the ALL-BFM 2000 protocol. Haematologica 2013; 98: 428–432.
Palmi C, Lana T, Silvestri D, Savino A, Kronnie GT, Conter V et al. Impact of IKZF1 deletions on IKZF1 expression and outcome in Philadelphia chromosome negative childhood BCP-ALL. Reply to 'incidence and biological significance of IKZF1/Ikaros gene deletions in pediatric Philadelphia chromosome negative and Philadelphia chromosome positive B-cell precursor acute lymphoblastic leukemia'. Haematologica 2013; 98: e164–e165.
Asai D, Imamura T, Suenobu S, Saito A, Hasegawa D, Deguchi T et al. IKZF1 deletion is associated with a poor outcome in pediatric B-cell precursor acute lymphoblastic leukemia in Japan. Cancer Med 2013; 2: 412–419.
Buitenkamp TD, Pieters R, Gallimore NE, van der Veer A, Meijerink JP, Beverloo HB et al. Outcome in children with Down's syndrome and acute lymphoblastic leukemia: role of IKZF1 deletions and CRLF2 aberrations. Leukemia 2012; 26: 2204–2211.
Den Boer ML, van Slegtenhorst M, De Menezes RX, Cheok MH, Buijs-Gladdines JG, Peters ST et al. A subtype of childhood acute lymphoblastic leukaemia with poor treatment outcome: a genome-wide classification study. Lancet Oncol 2009; 10: 125–134.
van der Veer A, Zaliova M, Mottadelli F, De Lorenzo P, Te Kronnie G, Harrison CJ et al. IKZF1 status as a prognostic feature in BCR-ABL1-positive childhood ALL. Blood 2014; 123: 1691–1698.
Roberts KG, Morin RD, Zhang J, Hirst M, Zhao Y, Su X et al. Genetic alterations activating kinase and cytokine receptor signaling in high-risk acute lymphoblastic leukemia. Cancer Cell 2012; 22: 153–166.
Churchman ML, Low J, Qu C, Paietta EM, Kasper LH, Chang Y et al. Efficacy of retinoids in IKZF1-mutated BCR-ABL1 acute lymphoblastic leukemia. Cancer Cell 2015; 28: 343–356.
Iacobucci I, Iraci N, Messina M, Lonetti A, Chiaretti S, Valli E et al. IKAROS deletions dictate a unique gene expression signature in patients with adult B-cell acute lymphoblastic leukemia. PLoS One 2012; 7: e40934.
von Stackelberg A, Seeger K, Henze G, Eckert C . Clinical significance of minimal residual disease in childhood acute lymphoblastic leukemia after first relapse. Leukemia 2004; 18: 1727–1728.
Malinowska-Ozdowy K, Frech C, Schonegger A, Eckert C, Cazzaniga G, Stanulla M et al. KRAS and CREBBP mutations: a relapse-linked malicious liaison in childhood high hyperdiploid acute lymphoblastic leukemia. Leukemia 2015; 29: 1656–1667.
Mullighan CG, Zhang J, Harvey RC, Collins-Underwood JR, Schulman BA, Phillips LA et al. JAK mutations in high-risk childhood acute lymphoblastic leukemia. Proc Natl Acad Sci USA 2009; 106: 9414–9418.
Izraeli S, Shochat C, Tal N, Geron I . Towards precision medicine in childhood leukemia—insights from mutationally activated cytokine receptor pathways in acute lymphoblastic leukemia. Cancer Lett 2014; 352: 15–20.
Sakaguchi H, Okuno Y, Muramatsu H, Yoshida K, Shiraishi Y, Takahashi M et al. Exome sequencing identifies secondary mutations of SETBP1 and JAK3 in juvenile myelomonocytic leukemia. Nat Genet 2013; 45: 937–941.
Bains T, Heinrich MC, Loriaux MM, Beadling C, Nelson D, Warrick A et al. Newly described activating JAK3 mutations in T-cell acute lymphoblastic leukemia. Leukemia 2012; 26: 2144–2146.
Degryse S, de Bock CE, Cox L, Demeyer S, Gielen O, Mentens N et al. JAK3 mutants transform hematopoietic cells through JAK1 activation, causing T-cell acute lymphoblastic leukemia in a mouse model. Blood 2014; 124: 3092–3100.
Shochat C, Tal N, Bandapalli OR, Palmi C, Ganmore I, te Kronnie G et al. Gain-of-function mutations in interleukin-7 receptor-alpha (IL7R) in childhood acute lymphoblastic leukemias. J Exp Med 2011; 208: 901–908.
Nikolaev SI, Garieri M, Santoni F, Falconnet E, Ribaux P, Guipponi M et al. Frequent cases of RAS-mutated Down syndrome acute lymphoblastic leukaemia lack JAK2 mutations. Nat Commun 2014; 5: 4654.
Mullighan CG, Goorha S, Radtke I, Miller CB, Coustan-Smith E, Dalton JD et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature 2007; 446: 758–764.
Topka S, Vijai J, Walsh MF, Jacobs L, Maria A, Villano D et al. Germline ETV6 mutations confer susceptibility to acute lymphoblastic leukemia and thrombocytopenia. PLoS Genet 2015; 11: e1005262.
Olsson L, Albitar F, Castor A, Behrendtz M, Biloglav A, Paulsson K et al. Cooperative genetic changes in pediatric B-cell precursor acute lymphoblastic leukemia with deletions or mutations of IKZF1. Genes Chromosomes Cancer 2015; 54: 315–325.
Olsson L, Johansson B . Ikaros and leukaemia. Br J Haematol 2015; 169: 479–491.
Mullighan CG, Zhang J, Kasper LH, Lerach S, Payne-Turner D, Phillips LA et al. CREBBP mutations in relapsed acute lymphoblastic leukaemia. Nature 2011; 471: 235–239.
Mar BG, Bullinger LB, McLean KM, Grauman PV, Harris MH, Stevenson K et al. Mutations in epigenetic regulators including SETD2 are gained during relapse in paediatric acute lymphoblastic leukaemia. Nat Commun 2014; 5: 3469.
Zhu X, He F, Zeng H, Ling S, Chen A, Wang Y et al. Identification of functional cooperative mutations of SETD2 in human acute leukemia. Nat Genet 2014; 46: 287–293.
Marke R, Havinga J, Cloos J, Demkes M, Poelmans G, Yuniati L et al. Tumor suppressor IKZF1 mediates glucocorticoid resistance in B-cell precursor acute lymphoblastic leukemia. Leukemia 2015; 30: 1599–1603.
Schwickert TA, Tagoh H, Gultekin S, Dakic A, Axelsson E, Minnich M et al. Stage-specific control of early B cell development by the transcription factor Ikaros. Nat Immunol 2014; 15: 283–293.
Joshi I, Yoshida T, Jena N, Qi X, Zhang J, Van Etten RA et al. Loss of Ikaros DNA-binding function confers integrin-dependent survival on pre-B cells and progression to acute lymphoblastic leukemia. Nat Immunol 2014; 15: 294–304.
Heizmann B, Kastner P, Chan S . Ikaros is absolutely required for pre-B cell differentiation by attenuating IL-7 signals. J Exp Med 2013; 210: 2823–2832.
Mullighan CG, Phillips LA, Su X, Ma J, Miller CB, Shurtleff SA et al. Genomic analysis of the clonal origins of relapsed acute lymphoblastic leukemia. Science 2008; 322: 1377–1380.
Kuiper RP, Waanders E, van der Velden VH, van Reijmersdal SV, Venkatachalam R, Scheijen B et al. IKZF1 deletions predict relapse in uniformly treated pediatric precursor B-ALL. Leukemia 2010; 24: 1258–1264.
Xavier AC, Ge Y, Taub J . Unique clinical and biological features of leukemia in Down syndrome children. Expert Rev Hematol 2010; 3: 175–186.
Buitenkamp TD, Izraeli S, Zimmermann M, Forestier E, Heerema NA, van den Heuvel-Eibrink MM et al. Acute lymphoblastic leukemia in children with Down syndrome: a retrospective analysis from the Ponte di Legno study group. Blood 2014; 123: 70–77.
van Bodegom D, Zhong J, Kopp N, Dutta C, Kim MS, Bird L et al. Differences in signaling through the B-cell leukemia oncoprotein CRLF2 in response to TSLP and through mutant JAK2. Blood 2012; 120: 2853–2863.
Roll JD, Reuther GW . CRLF2 and JAK2 in B-progenitor acute lymphoblastic leukemia: a novel association in oncogenesis. Cancer Res 2010; 70: 7347–7352.
Tasian SK, Doral MY, Borowitz MJ, Wood BL, Chen IM, Harvey RC et al. Aberrant STAT5 and PI3K/mTOR pathway signaling occurs in human CRLF2-rearranged B-precursor acute lymphoblastic leukemia. Blood 2012; 120: 833–842.
Maude SL, Tasian SK, Vincent T, Hall JW, Sheen C, Roberts KG et al. Targeting JAK1/2 and mTOR in murine xenograft models of Ph-like acute lymphoblastic leukemia. Blood 2012; 120: 3510–3518.
Steelman LS, Abrams SL, Whelan J, Bertrand FE, Ludwig DE, Basecke J et al. Contributions of the Raf/MEK/ERK, PI3K/PTEN/Akt/mTOR and Jak/STAT pathways to leukemia. Leukemia 2008; 22: 686–707.
Greaves M, Maley CC . Clonal evolution in cancer. Nature 2012; 481: 306–313.
Park SY, Wolfram P, Canty K, Harley B, Nombela-Arrieta C, Pivarnik G et al. Focal adhesion kinase regulates the localization and retention of pro-B cells in bone marrow microenvironments. J Immunol 2013; 190: 1094–1102.
Dupuis A, Gaub MP, Legrain M, Drenou B, Mauvieux L, Lutz P et al. Biclonal and biallelic deletions occur in 20% of B-ALL cases with IKZF1 mutations. Leukemia 2013; 27: 503–507.
Acknowledgements
We like to acknowledge the technical support of Vladimir Benes (EMBL, GeneCore, Heidelberg) for RNA-Seq, Maria Morak, Christina Satke, Andrea Inthal and Sabrina Haslinger (CCRI) and Aurélie CAYE (Hôpitaux de Paris, Hôpital Robert Debré, Département de Génétique, Paris) for technical support and Fikret Rifatbegovic for help with Figure 4. This work was supported in part by research funding from The Austrian Science Fund, FWF I-1226-B19 to RP-G, the FP7-ERA-NET grant TRANSCALL and by charitable donations of the Kapsch group (http://www.kapsch.net/kapschgroup) to CV and RP-G.
Author contributions
CV performed wet lab work, analyzed and compiled data, and contributed to manuscript writing; CF performed bioinformatics and statistical analyses, and contributed to manuscript writing; CE, GC, UzS, HC, AA, AvS, MS, MAH, GM provided patient samples and clinical information; AM, FK and SF performed experiments; KN performed SNP array analysis; MS and CB performed WES and primary analysis; OAH is responsible for cytogenetics, MLPA and SNP array analysis, and contributed to manuscript writing; RP-G conceived the study, supervised research, interpreted results and wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Vesely, C., Frech, C., Eckert, C. et al. Genomic and transcriptional landscape of P2RY8-CRLF2-positive childhood acute lymphoblastic leukemia. Leukemia 31, 1491–1501 (2017). https://doi.org/10.1038/leu.2016.365
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2016.365
- Springer Nature Limited
This article is cited by
-
Philadelphia chromosome-like acute lymphoblastic leukemia with concomitant rearrangements of CRLF2 and ABL1: a pediatric case report
BMC Pediatrics (2024)
-
BCR::ABL1-like acute lymphoblastic leukaemia: a single institution experience on identification of potentially therapeutic targetable cases
Molecular Cytogenetics (2023)
-
Prospective Identification of Prognostic Hot-Spot Mutant Gene Signatures for Leukemia: A Computational Study Based on Integrative Analysis of TCGA and cBioPortal Data
Molecular Biotechnology (2023)
-
HMGN1 plays a significant role in CRLF2 driven Down Syndrome leukemia and provides a potential therapeutic target in this high-risk cohort
Oncogene (2022)
-
Prognostic Impact of Somatic Copy Number Alterations in Childhood B-Lineage Acute Lymphoblastic Leukemia
Current Oncology Reports (2021)