Abstract
Medulloblastoma (MB), the most common malignant paediatric brain tumor, is currently treated using a combination of surgery, craniospinal radiotherapy and chemotherapy. Owing to MB stem cells (MBSCs), a subset of MB patients remains untreatable despite standard therapy. CD133 is used to identify MBSCs although its functional role in tumorigenesis has yet to be determined. In this work, we showed enrichment of CD133 in Group 3 MB is associated with increased rate of metastasis and poor clinical outcome. The signal transducers and activators of transcription-3 (STAT3) pathway are selectively activated in CD133+ MBSCs and promote tumorigenesis through regulation of c-MYC, a key genetic driver of Group 3 MB. We screened compound libraries for STAT3 inhibitors and treatment with the selected STAT3 inhibitors resulted in tumor size reduction in vivo. We propose that inhibition of STAT3 signaling in MBSCs may represent a potential therapeutic strategy to treat patients with recurrent MB.
Similar content being viewed by others
Introduction
Brain tumors represent the leading cause of childhood cancer mortality, of which medulloblastoma (MB) is the most common malignant pediatric brain tumor. MB most often arises in the cerebellum of infants and young children, and was originally described as a primitive neuroectodermal tumor.1, 2 Although tremendous improvements in therapy regimens using a combination of surgical resection, craniospinal irradiation and chemotherapy with vincristine, cisplatin and cyclophosphamide have yielded 5-year overall survival rates of 70–80%, survivors are left with substantial neurocognitive impairments and treatment-related morbidity.3, 4
MB has been classified into four principal subgroups using integrative genomic platforms.1, 2 Groups 1 and 2, referred to as WNT and sonic hedgehog (SHH), derive from dysregulation of these key developmental signaling pathways and Groups 3 and 4 are associated with the strong upregulation of c-MYC and N-MYC, respectively.2, 5, 6 This subgroup stratification has generated four genetically distinct tumor identities, each with a characteristic clinical presentation and prognosis.7 Group 3 tumors have shown the worst prognosis of all, emphasizing the need for the development of specialized therapeutics in this subgroup.8 Recent reports suggest the presence of somatic copy-number aberrations and single-nucleotide variants that may present subgroup-specific therapeutic targets9, 10 but to date, druggable targets for Group 3 MB have not been identified.
As in many other solid tumors11 and hematopoietic cancers,12 MB is characterized by the presence of cancer stem cells (CSCs) or MB stem cells (MBSCs) that possess the ability to sustain tumor growth. These cells were first isolated and characterized in 2004.13 CD133 (Prom1), a pentaspan membrane glycoprotein, is a surface marker for MBSCs, although its functional role and signaling mechanisms are not well understood.13, 14 Although some studies debate the role of CD133 in initiating tumors,15 CD133-positive cells continue to retain a higher self-renewal capacity and multipotency.16
The signal transducers and activators of transcription (STAT) family of transcription factors are present in the cytoplasm as inactive monomeric/dimeric forms and upon phosphorylation at key tyrosine residues (Tyr705), STATs translocate to the nucleus. This movement from cytoplasm to nucleus is due to reciprocal SH2 domain–pTyr binding interactions between two STAT molecules, leading to the formation of activated homodimers.17 Upon nuclear translocation, these activated homodimers promote expression of genes that regulate cell growth, cell survival and cell cycle regulation. In cancer cells, continual activation of STATs, more specifically STAT3, leads to modulation of several cell cycle regulators to promote a state of tumorigenesis.
Numerous cancers such as leukemia, lymphoma,18 breast,19 ovarian19 and prostate20 cancer display aberrant activation of STAT3 signaling. In brain tumors, especially glioblastoma, recent signaling network profiling identified STAT3 as one of the key transcription factors modulating poor outcome in the mesenchymal subtype.21 In patient-derived glioblastoma brain tumor-initiating cells, STAT3 signaling is prognostically significant and can be targeted in orthotopic mouse models.22
Here, we describe a method for treatment of Group 3 MBSCs by targeting STAT3 transcription factor signaling. We sorted human Group 3 MBSCs based on CD133 expression and showed upregulation of p-STAT3 and c-MYC in these cells compared with CD133-negative subpopulations. Thereafter using structure-based drug design, we developed novel STAT3 inhibitors that have the potential to cross the blood-brain-barrier, to treat MB in vivo. Although CD133 has been used primarily as a marker to identify CSCs in multiple human cancers including brain tumors, its functional role remains obscure and cannot be currently targeted using small molecules or inhibitors. Similarly, c-MYC is difficult to target due to our poor understanding of binding ligands and inability to specify context-dependent MYC-associated transcriptional gene expression across different cancers.23 Our study presents a preclinical model applying STAT3 inhibitors to selectively target CD133+ MBSCs in Group 3 MB.
Results
CD133 expression is upregulated in Group 3 MB cells
CD133 is a marker used to enrich for human MBSCs,6, 13, 14, 24 but its role and expression within subgroups remains unclear. To evaluate CD133 protein expression in MBSCs, we propagated in neural stem cell (NSC) media the MB Group 3 lines Med8A, D425, D458, CHLA-01R-MED, and measured CD133 expression by flow cytometery (Figure 1a). It is noteworthy that D425 was derived from a primary cerebellar MB in a 6-year-old boy, whereas D458 was established from cerebrospinal metastasis after treatment failure from the same patient; thus D425 and D458 represent a matched primary and recurrent pair.25 Interestingly, CD133 expression was significantly higher in recurrent MB lines (D458, CHLA-01R-MED) compared with primary (D425) lines. Phase-contrast pictures of MB cell lines are shown in Figure 1b.
CD133 characterizes group 3 MB and CD133+ cells show increased tumorigenicty and proliferation as compared with CD133− populations: MB cell lines are grown in their respective media conditions as described in the ‘Materials and methods’ section. (a) Representative flow cytometry plots exhibiting percentage of CD133 protein levels in Med8A (4.75%), D425 (39.13%), D458 (68.62%) and CHLA-01R-MED (98.27%). (b) Phase-contrast pictures of the above cell lines showing difference in their morphology and growth patterns. (c) CD133 expression identifies Group 3 MB (indicated as Group C) that was probed using affymetrix exon array data and expression levels represented by log2–transformed intensity signal. Four independent databases were probed to measure CD133 expression levels. (d) The CD133 signature was originally identified and described by Venupogal et al.(55) and CD133 signature genes are included as Supplementary Table 2. CD133-high signature reflects a reduced overall survival of MB patients compared with CD133-low tumors (hazard ratio=2.4, P=0.01). (e and f) Group 3 MB cells D425 and D458 were flow cytometrically sorted into CD133+ and CD133− populations and Presto Blue Assay assessed their proliferative potential. CD133+ cells are more proliferative when compared with CD133− cells. Bar represents mean fluorescence intensity (arbitrary units (a.u.)) of three technical replicates, mean±s.d., two-tailed t-test, P⩽0.0007 (g) 500 000 cells of each of CD133+ and CD133− populations were xenografted intracranially in mice brains. CD133+ cells generated larger and infiltrative tumors than CD133− cells (scale bar=4 mm).
When we probed multiple published MB transcriptional databases (Toronto,5 Boston,26 Amsterdam27 and Memphis28) for CD133 we found its expression was strongly associated with Group 3 MB (Figure 1c). Data mined from the Boston database showed lower survival in CD133high tumors (P=0.01, hazard ratio=2.4) compared with CD133low tumors (Figure 1d). To provide a functional context for the clinical utility of CD133 expression in Group 3 MB, we compared the proliferative potential between sorted CD133-positive and CD133-negative populations. When compared with CD133-negative cells, CD133-positive cells had significantly higher proliferation capacity in both primary and matched recurrent cells lines (D425 and D458, respectively, Figures 1e and f).
To further validate the functional role of CD133 in tumor formation, we xenografted mice intracranially with sorted CD133-positive or CD133-negative Group 3 MB cells. We found that CD133-positive cells formed very large tumors, whereas tumors obtained by xenograting CD133-negative cells were much smaller (Figure 1g). Our data suggests that CD133 expression characterizes and may contribute to poor outcome in Group 3 MB through perpetuation of stem cell populations that drive tumorigenesis.
Activated STAT3 is enriched in CD133-high Group 3 MB
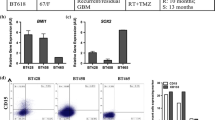
Activated STAT3 signaling has been described in other CSC populations, including CD133-positive human colon cancer cell lines29 and CD133high hepatocellular carcinoma cell lines.30 As high CD133 expression is a poor prognostic marker in many cancers31 and we show that it correlates with poor patient outcome in Group 3 MB (Figure 1d), we hypothesized that hyperactive signaling pathways in CD133-positive cells could be ideal therapeutic targets. In order to determine transcript levels of STAT3 among MB subgroups, we probed existing MB genomic platforms for its expression levels in different subgroups, as described previously for CD133. STAT3 expression levels were enriched in poor outcome non-Shh/Wnt subgroups, namely Group 3 and Group 4 MB (Figure 2a). We then determined protein levels of CD133, STAT3 and p-STAT3 in Group 3 MB cells using Western blot, and again validated that recurrent lines, D458 and CHLA-01R-MED have comparatively higher expression of CD133 than D425 line (Figure 2b). Although STAT3 expression is not significantly altered, activated STAT3 (p-STAT3) correlates with CD133 expression, and is especially elevated in recurrent Group 3 MB (Figure 2b). To further probe the relationship between CD133 and activated STAT3 signaling, we ectopically overexpressed CD133 in the CD133low Med8A Group 3 line. We validated overexpression of CD133 by RTPCR (Figure 2c) and Flow analysis (Figure 2d). Upon overexpression of CD133, MED8a cells showed a significant increase in p-STAT3 (Figure 2e). Med8A cells overexpressing CD133 also showed increased proliferation capacity (Figure 2f). Our data demonstrates that STAT3 signaling is activated in CD133+ Group 3 recurrent MB cells, and CD133 overexpression activates STAT3 to promote cell proliferation and growth of Group 3 MB.
STAT3 marks non-Shh/Wnt MB and its expression parallels CD133 expression. (a) affymetrix exon data from four independent databases shows STAT3 expression enriched in non-Shh/Wnt MB, that is, enrichment in Group 3 and Group 4 tumors (indicated as Group C and Group D respectively). Expression units are in log2 scale. (b) Immunoblot analysis for CD133, STAT3, activated STAT3 (p-STAT3) was performed in Group 3 and Group 2 MB lines. GAPDH was used as a loading control. To verify correlation between CD133 and STAT3, CD133 is overexpressed in Med8A. CD133 (c) transcript by qRT-PCR and (d) protein levels as shown by flow cytometric analysis (percentage increase from 5.03% to 82.18%, representative flow) are significantly increased following overexpression of CD133 (oe CD133). (e) Med8A oe CD133 showed increased p-STAT3, although no change in STAT3 was seen. (f) Proliferative capacity measured using Presto Blue assay shows OE CD133 cells have a significant increase in proliferation when compared with OE con (control cells). Bar represents mean fluorescence intensity (arbitrary units (a.u.)) of three technical replicates, mean±s.d., two-tailed t-test. *P<0.05.
p-STAT3 contributes to tumorigenesis in Group 3 MB
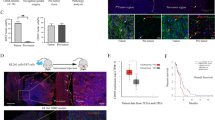
To investigate the role of STAT3 signaling in MB pathogenesis, we undertook lentiviral short-hairpin RNA-mediated knockdown of STAT3 in Group 3 patient-matched MB cell lines D425 (primary) and D458 (recurrent). Out of two vectors, shSTAT3-1 showed efficient knockdown in both cell lines D425 (Figure 3a) and D458 (Figure 3b) with prominent reduction in activated p-STAT3 levels. This knockdown correlated with reduction in proliferation capacity in both primary and recurrent MB (Figures 3c and d). To further elucidate the role of STAT3 in tumor initiation/maintenance, we performed intracranial injections of STAT3-knockdown MB cells into NOD-SCID mice and found a significant reduction in tumor size when compared with control (shControl-transduced) tumors (Figures 3e–g). Moreover, mice xenografted with shSTAT3-transduced cells treated survived longer than control mice (Figures 3h and i). These data demonstrate the prominent role of STAT3 in regulating proliferation and tumorigenesis of Group 3 MBSCs.
STAT3 knockdown using lentiviral approach reveals its role in proliferation and tumor formation. shSTAT3 lentiviral vector-mediated transduction shows reduction in both STAT3 and phosphorylated p-STAT3 protein levels as assessed by immunoblotting with shSTAT3-1 vector compared with shControl vector in both (a) D425 and (b) D458 cell lines. GAPDH was used as a loading control. Proliferative capacity as determined by Presto Blue assay shows decreased proliferation in shSTAT3-1 (c and d) when compared with shControl-transduced cells. Bar represents mean fluorescence intensity (arbitrary units (a.u.)) of three technical replicates, mean±s.d., two-tailed t-test. *P⩽0.01. (e) Xenografts were generated using 100 000 cells of shControl or shSTAT3-1 vectors in D425 (upper panel) and D458 (lower panel) into the frontal lobes of NOD-SCID mice (n=6 in each group). shSTAT3-1-treated cells displayed significantly small and less invasive tumors in both lines (arrows indicates tumor area). Mice were killed in pairs of shControl and shSTAT3-1 when one member of the pair reached end point and H&E sections of the brains are shown. (f and g) Total area of tumor was calculated using ImageJ64 software (NIH, Bethesda, MA, USA). Bar represents tumor area (mm2) of n=6 mice in each group, mean±s.d. and two-tailed t-test. *P⩽0.005. (h and i) shSTAT3 cells xenograted mice maintain a significant survival over shControl-treated mice (n=3 in each group). The median survival in case of D425 shSTAT3 mice increased to 38 days compared with 20 days in shControl mice (P=0.0072) and median survival in D458 shSTAT3 mice augmented to 33 days from 24 days in shControl mice (P=0.04).
Interplay between CD133, STAT3 and c-MYC may drive Group 3 MB
c-MYC drives a regulatory network in embryonic stem cells that is also active in many cancers and drives poor outcomes.32 c-MYC is further implicated in regulating Group 3 murine MBs that resemble their human counterpart.6 CD133 and c-MYC have clearly been associated with relapse and poor prognosis in MB patients.6 To explore how CD133, c-MYC and STAT3 may collectively regulate MBSCs,30, 33, 34 we first probed MB transcriptome data to validate that c-MYC is highly enriched in Group 3 MB tumors (Supplementary Figure 1). Although Group 3 MB lines used (D425, D458, Med8A and CHLA-01R-MED) showed varied mRNA expression levels of c-MYC (Figure 4a), protein levels of c-MYC were higher in Group 3 recurrent lines D458 and CHLA-01R-MED (Figure 4b).
CD133, STAT3 and c-MYC constitute a signaling axis in MBSC regulation. (a) transcript by qRT-PCR and (b) protein levels as shown by western blotting. Bar represents expression levels relative to GAPDH of three technical replicates, mean±s.d., one-way analysis of variance, P⩽0.004. D458 cells were flow cytometry sorted for CD133+ and CD133− populations. c-MYC (c) transcript by qRT-PCR and (d) protein levels in D458 as shown by western blotting are significantly increased in CD133-positive cells when compared with CD133-negative cells. CD133+ cells also showed increased protein expression of CD133 and p-STAT3 relative to GAPDH control as analyzed by western blotting. Bar represent expression levels relative to GAPDH of three technical replicates, mean±s.d., two-way analysis of variance, P=0.005. (e) D425 and D458 cells were treated with c-MYC inhibitor 10058-F4 and IC50 curves were generated from inhibition of cell proliferation using Presto Blue assay with four technical replicates. IC50 values are: D425: 189.148 μM and D458: 133.277 μM.
c-MYC is a known downstream target of STAT3 signaling in ovarian cancer and in glioblastoma stem cells, where STAT3 inhibitors have also reduced expression of stem cell-associated genes such as CD133. We show that p-STAT3 is upregulated in sorted CD133-positive populations from Group 3 MB lines, which also show enhanced c-MYC levels compared with CD133-negative populations (Figure 4c). Moreover, as a proof of principle we looked at the levels of STAT3 and c-MYC after knockdown of STAT3 in both D425 and D458 cell lines and found significant downregulation of c-MYC after STAT3 knockdown (Supplementary Figure 2). Expression levels of c-MYC are significantly higher in sorted CD133-positive cells compared with CD133-negative cells in D458 MB (Figure 4d). Given the high levels of c-MYC in Group 3 MB, we performed proliferation assays using a cell permeable c-MYC inhibitor, 10058-F4,35 to investigate whether targeting c-MYC could reduce tumor cell growth. Our results (Figure 4e) showed that Group 3 MB cell proliferation is attenuated by this c-MYC inhibitor, but only with an 50% inhibitory concentration values (IC50; concentration at which cell proliferation is inhibited by 50%) in the high micromolar range (D425: 189.148 μM and D458: 133.277 μM). However, pharmacodynamic studies and screening of other c-Myc inhibitors are necessary to draw conclusions about the utility of c-Myc inhibitors as therapeutics for Group 3 MB. Nevertheless, c-MYC is a highly pleiotropic transcription factor that controls expression of hundreds of genes and may prove challenging to target. This prompted us to seek alternate means to target the CD133/STAT3/c-MYC signaling axis in Group 3 MB.
Ex vivo treatment with STAT3 inhibitors reduces MB tumor burden in vivo
We tested various STAT3 inhibitors from our library (Supplementary Figures 3 and 4) and found PG-S3-002 to have selective activity towards CD133-positive MBSCs with IC50 in the low micromolar range, in both primary and recurrent MB cells (Figures 5a and b). A reduction in p-STAT3 protein levels was observed following treatment with PG-S3-002 at IC80 (Figures 5c and d). Intracranial xenograft injections of 1.0 × 105 viable STAT3 inhibitor-treated cells resulted in significantly smaller tumors (Figures 5e–g) compared with control cells. When we treated normal human NSCs with PG-S3-002, we found an IC50 value also in the low micromolar range (Supplementary Figure 5). This data shows that STAT3 inhibitor treatment at doses that would be tolerable in patients can decrease tumor burden of Group 3 MB xenografts, but with equal toxicity towards NSCs; implicating neurotoxicity as a potential side effect.
STAT3 inhibitors targeting MBSCs. (a) D425 and (b) D458 cells were flow cytometry sorted into CD133+/− populations and treated with PG-S3-002 STAT3 inhibitor. IC50 curves were generated from inhibition of cell proliferation using presto blue assay with four technical replicates. For D425, IC50 values are; CD133+: 2917 nM and CD133−: 13 794 nM. IC50 values for D458 are CD133+: 7807 nM and CD133−: 11 424 nM. (c and d) Western blot showing protein levels of STAT3 and phosphorylated STAT3 (p-STAT3) in DMSO-treated and PG-S3-002-treated (IC80) D425 and D458, relative to GAPDH control. (e) D425 (upper panel) and D458 (lower panel) cells were treated ex vivo with their respective IC80 of PG-S3-002 for 4 days and 100 000 viable cells were injected into frontal lobes of NOD-SCID mice (n=3 in each group). Cells treated with DMSO were used as control. Mice were killed in pairs as they fell sick. H&E sections of the brains are shown. Average tumor size reduction is 85% in (f) D425 and 70% in (g) D458. Tumor area was calculated using ImageJ64 software (NIH). Bar represents tumor area (mm2) of n=3 mice in each group, mean±s.d. and two-tailed t-test. *P⩽0.005.
Systemic treatment with STAT3 inhibitors PG-S3-009 and PG-S3-010 reduces tumor formation in vivo
As STAT3 signaling is active in MBSCs, we then investigated the role of STAT3 compounds in reducing tumor formation in our human-mouse xenograft model of Group 3 MB. We chose to xenograft recurrent MB lines that are refractory to conventional chemoradiotherapy, and first tested whether STAT3 inhibitors would target these cells in vitro. Both PG-S3-009 and PG-S3-010 compounds showed IC50 values in the low micromolar range and selectivity for recurrent MBSCs (Figure 6a, Supplementary Figures 7a and b). These two inhibitors are highly selective for recurrent (D458) MB, when compared with matched primary MB (Supplementary Figure 6). Our 2-week treatment protocol comprised PG-S3-009 and PG-S3-010 administration by intraperitoneal injections (20 mg/kg dose, twice a week), following tumor engraftment. Mice were culled and brains harvested at treatment completion for both control (DMSO) and inhibitor-treated mice (Figure 6b). Mice treated with PG-S3-009 or PG-S3-010 maintained a reduction in tumor burden (Figures 6c and d), with PG-S3-009 showing the greatest efficacy and potency. These data demonstrates selected STAT3 inhibitors to be efficacious in targeting recurrent, treatment-refractory Group 3 MBSCs.
In vivo treatment with STAT3 inhibitors in recurrent MB cells. (a) unsorted D458 cells were treated with STAT3 inhibitors PG-S3-009 and PG-S3-010 and IC50 curves generated from inhibition of cell proliferation using Presto Blue assay with four technical replicates. For middle and right panels, D458 cell are sorted by flow cytometry into CD133+ and CD133− populations and then IC50 curves were generated. Left panel: IC50 values for PG-S3-009 and PG-S3-010 are 3943 nM and 2747 nM for total population, respectively. Middle panel: after treatment with PG-S3-009, sorted cells have IC50 values 3254 nM and 44 232 nM for CD133+ and CD133− population, respectively. Right panel: after PG-S3-010 treatment, IC50 values were 2517 nM and 39 511 nM for CD133+ and CD133− and populations, respectively. (b) Timeline of the in vivo treatment protocol. Day 0 is the first day of tumor implantation in mice (n=15 mice). D458 recurrent MB cells are used. After a week of implantation, treatment protocol of 20 mg/kg body weight of DMSO, PG-S3-009 or PG-S3-010 (n=5 mice in each set) started with intraperitoneal injections twice a week for 2 weeks. At the end of 2 weeks, mice were killed and their brains were harvested. (c) Representative H&E sections of the brains are shown (n=5). (d) Tumor area was calculated using ImageJ64 software (NIH). Bar represents tumor area (mm2) of n=5 mice in each group, mean±s.d. and two-tailed t-test. *P⩽0.005. Average tumor size reduction is 80% in treatment with PG-S3-009 and 35% in PG-S3-010.
PG-S3-009 treatment significantly decreases tumor size of recurrent Group 3 MB in vivo, but does not increase survival
D458 cells were treated with PG-S3-009 in varying doses in vitro and found that cellular levels of STAT3, p-STAT3 and c-MYC gradually decreases with increasing concentration of PG-S3-009 (Figure 7a). These data implicate the direct targeting of STAT3 by PG-S3-009 and subsequent downregulation of STAT3 leading to the reduction of the downstream target c-MYC in MBSCs (Supplementary Figure 7). In order to evaluate the effect of the drug in vivo, NODSCID mice were intracranially xenografted with D458 cells and treated intraperitoneal with escalating doses of PG-S3-009 (DMSO, 5, 10 or 20 mg/kg) (Figure 7b). Cohorts treated at 20 mg/kg with PG-S3-009 showed significant reduction in tumor burden compared with DMSO or lower-dose treated mice (Figures 7c and d). However, no survival benefit was observed in treated mice compared with DMSO controls (Figure 7e). Treated mice became anorexic and withdrew from feeding and activity, suggestive of treatment-induced neurotoxicity, although no toxicity was seen in liver, kidneys and lungs (data not shown). These data implicate the direct targeting of STAT3 by PG-S3-009 and subsequent downregulation of STAT3 leading to the reduction of the downstream target c-MYC in MBSCs (Supplementary Figure 8).
Dose-dependence study of PG-S3-009. (a) Western blot showing c-MYC, STAT3 and p-STAT3 levels compared to GAPDH after treatment of D458 with serially increasing concentration of PG-S3-009. (b) Timeline of the in vivo protocol starting with 48 mice intracranial injections (frontal lobe) of D458 at day 0, day of tumor cell implantation. After a week, (n=12 mice in each set) were given 5, 10 or 20 mg/kg body weight of either DMSO or PG-S3-009 for 2 weeks, twice a week. At the end of third week, mice were killed (n=6 mice in each set) for H&E staining and remaining 6 mice per set were monitored for their survival. (c) Representative H&E sections of the brains are shown after each treatment regimens (n=6). (d) There is a significant reduction in tumor area (calculated using ImageJ software). Bar represents tumor area (mm2) of n=6 mice in each group, mean±s.d. and two-tailed t-test. *P=0.03. (e) Percentage survival for mice treated with 5 mg (upper panel), 10 mg (middle panel) and 20 mg/kg body weight of PG-S3-009 (lower panel).
Discussion
The recent molecular classification of MB into four distinct subgroups implies that different subrgoup-specific therapeutic strategies need to be devised for individualized patient therapy. This therapeutic roadmap may only be realized by elucidating the biological and genetic drivers of tumorigenesis in each subgroup, in order to discover tractable and druggable therapeutic targets. In this study, we link the high expression of CD133 with Group 3 MB and further correlate its expression with c-MYC, via STAT3. High c-Myc expression often reflects a classic Group 3 MB with metastasis at diagnosis, large cell histology and overall poor prognosis.2 Novel mouse models have been developed using genetics to recapitulate human Group 3 MB that have the potential to be used for preclinical testing.6, 36
Our observation of enrichment of CD133 in Group 3 MB is in accordance with the expression of this cell-surface marker in multiple CSC populations, which are thought to evade current chemoradiotherapy only to seed recurrent disease and drive patient relapse and poor outcome.14 In light of the fact that Group 3 MB is the most aggressive among all subgroups, our understanding of the functional role of CD133 in Group 3 MB can provide a strategic platform to identify novel signaling inhibitors in CD133-positive cells for the effective management of these tumors. c-MYC expression can be regulated by numerous signaling pathways including the JAK/STAT, rendering STAT3 inhibitors attractive therapeutic options. As the Wnt subtype of MB (Group 1) also demonstrates elevated levels of c-Myc, STAT3 inhibitors could be considered as a therapeutic option for patients with this positive prognostic MB subtype.
Activated STAT3 signaling has been shown to be essential for cell survival and tumorsphere-forming capacity in human colon cancer cell lines sorted for both ALDH (aldehyde dehydrogenase) and CD133 (ALDH+/CD133+).29 Previous studies have shown that downregulation and inhibition of STAT3 signaling suppresses proliferation and induces apoptosis in human glioma cells37,38,39 as well as various cancers.40, 41 Xiao et al., have already shown that disruption of STAT3 signaling leads to inhibition of cell growth and apoptosis of medulloblastoma cells.42 Using bioinformatics and microarray platforms, another study revealed STAT3/p-STAT3 signaling upregulated in CD133-positive MB cells compared with CD133-negative cells.43 STAT3 has a key role in maintaining both tumor cells and its microenvironment, although little is known about both upstream and downstream regulators of JAK–STAT3 signaling. Despite advances in studying the role of STAT3 as an ideal target for CSC-driven tumors,30, 33, 44 challenges have arisen in developing these drugs for clinical use. In general STAT3 inhibitors targeting the SH2 domain have suffered from limited clinical utility due to inclusion of polar groups designed to mimic SH2 domain binding Tyr705 residue that has resulted in metabolic instability and poor cell permeability.42, 45 Non-phosphorylated small molecule inhibitors such as STA-201,46 BP-1-102,47 BP-5-87,48 SH-4-5444 and LY542 have, to a certain extent circumvented this problem.
To enhance in vivo efficacy, SH-4-54 was optimized specifically to yield inhibitors with improved bioavailability. PG-S3-009 and PG-S3-010 were two of the most potent anti-cancer compounds to emerge from this program. These inhibitors have shown great promise in treating other treatment-refractory and aggressive cancers such as brain metastases.49
Previous studies have suggested that c-MYC is a critical regulator of MB cell growth and proliferation.32 The activation of STAT3 in MB50 and its translocation to the nucleus permits activation of oncogenes and cell cycle regulatory genes such as c-MYC, cyclin-D1 and Cox-2, leading to cancer progression.51 In another study, DNA microarray analysis in CD133-positive MB cells showed an upregulation of c-MYC.52 Although both CD133 and c-MYC upregulation have clear implications in Group 3 MB pathogenesis, both are difficult to target owing to insufficient understanding of molecular mechanisms and their role in normal development and cancer. Currently there are no available drugs targeting CD133, and the c-MYC targeting compound 10058-F435 has a very high IC50 value in MBSCs that may preclude clinical use. There are currently no clinically approved therapies for directly targeting c-MYC, as MYC is a pleiotropic transcription factor that controls expression of hundreds of genes. Moreover, 10058-F4 inhibits c-MYC/Max dimerization,35 implying multiple off-target cellular activities through modulation of various transcription factors in other tissues and organs. Within the CD133/STAT3/c-MYC signaling axis, the only attractive target for drug development is STAT3. This concept prompted us to search for novel STAT3 inhibitors to treat Group 3 MB with the potential to translate into clinical settings.
In this study, we demonstrate the enrichment of CD133 in Group 3 MBSCs and discover potential signaling mechanisms involving activated STAT3 (p-STAT3) and c-MYC in CD133-positive cells. Treatment with STAT3 dimerization inhibitor PG-S3-002 ex vivo significantly reduces tumor burden in mice. Our preclinical trial of another STAT3 inhibitor, PG-S3-009, showed reduced tumor burden in mice, but no significant survival benefit due to presumed neurotoxicity of the compound. However, if neurotoxicity could be mitigated with chemical modifications to the compound, these inhibitors show great promise in effectively treating recurrent MB (D458), an important result as Group 3 recurrence is currently treated with palliation alone. Our study describes the first preclinical evidence that novel STAT3 inhibitors cross the blood-brain-barrier and effectively treat recurrent Group 3 MB.
Materials and methods
CD133/STAT3/c-MYC gene profiling in subgroups of MB
MB microarray data for 103 MBs were downloaded from GEO (GSE21140). These data consisted of processed Affymetrix CEL files (Affymetrix Human Exon 1.0 ST Array (transcript (gene) version)) that had undergone gene-level analysis (CORE content), quantile normalization (sketch) and summarization using PLIER with PM-GCBG background correction for 103 MBs. Clinical annotations for each MB tumor, including subgroup (Wnt, Shh, Group 3, Group 4) were also downloaded from this location. The expression of each gene was plotted as the normalized fluorescence intensity of the corresponding affymetrix probes. Similarly, raw Affymetrix datasets for 62 primary human MBs, 40 primary human MBs, 15 Daoy MB cell line samples and 46 primary human MBs were respectively downloaded and processed from GEO (GSE10327, GSE12992 and GSE7578) or http://www.stjuderesearch.org/site/data/medulloblastoma28 and normalized using RMA.53 When available, clinical annotations for each tumor were also downloaded from these locations.
CD133 signature for survival using Pomeroy database
Survival analysis (for CD133) was completed in (R core team, Vienna, Austria), and Kaplan–Meier survival curves were drawn using GraphPad prism (GraphPad software, SanDiego, CA, USA).
MB cell culture
MB cell lines Med8A, D425 and D458 were cultured in DMEM high glucose (Life Technologies, Carlsbad, CA, USA; #11965-118) supplemented with 1% penicillin–streptomycin, 1% glutamine (Gibco) and 20% serum for Med8A, D425 and D548. CHLA-01R-MED (ATCC, Manassas, VA, USA; CRL-3034) was obtained the American Type Culture Collection (ATCC) and maintained in tumor sphere media as described previously.54
Virus preparation and transduction
Lentiviral vectors shSTAT3-1 and shSTAT3-2 expressing short-hairpin RNA-targeting human STAT3 (target sequence: 5′-TGCATGTCTCCTTGACTCT-3′, 5′-TACCTAAGGCCATGAACTT-3′, respectively) and control scrambled short-hairpin RNA vector were purchased from Thermo Scientific (Waltham, MA, USA). CD133 overexpression vector was purchased from Genecopoeia (Rockville, MD, USA). Replication-incompetent lentivirus was produced. Briefly, after co-transfection of expression vector and packaging plasmids, viral supernatants were harvested 48 h after transfection. Thereafter, supernatant was filtered through a 0.45 μm cellulose acetate filter and precipitated with PEGit (System biosciences, Palo Alto, CA, USA). After precipitation, viral pellets were re-suspended in 1.0 ml of DMEM F-12 media and stored at −80 °C until further used. To generate stable shSTAT3 lines, MB cells were transduced with lentivirus prepared as stated above. After 48 h of transduction, cells were grown in media containing 1 μg/ml of puromycin to generate stable cell lines.
Western immunoblotting
Proteins from cell lysates were extracted and quantified as described previously.24 Briefly, 20 μg of total protein was run using 10% sodium dodecyl sulphate–polyacrylamide gel electrophoresis and proteins were then transferred to polyvinylidene fluoride membrane. Blots were probed for the following antibodies: anti-CD133 (1:500; Millipore, Etobicoke, ON, Canada; #MAB4399 clone 17A6.1), anti-c-MYC (1:1000; Cell signaling, Danvers, MA, USA; #5605), anti-STAT3 (1:1000; Cell Signaling #9139), anti-phospho STAT3 (1:1000; Cell Signaling; #4113), anti-GAPDH (1:40,000; Abcam, Toronto, ON, Canada; #ab8245). Secondary antibodies were conjugated with horseradish peroxidase and procured from Bio-Rad (Mississauga, ON, Canada; Goat anti-mouse IgG) or Sigma (Oakville, ON, Canada; goat anti-rabbit IgG) and chemidoc was used for visualization of protein bands.
Cell proliferation assay
Using 96-well plates, single cells were plated at a density of 1000 cells/200μl per well in quadruplicate for each sample and incubated for 4 days. After 4 days, 20μl of Presto Blue (Life technologies), a cell viability indicator, were added to each well ~4 h prior to the readout time point. Fluorescence was measured using a FLUOstar Omega Fluorescence 556 Microplate reader (BMG LABTECH) at excitation and emission wavelengths of 540–570 nm and 580–610 nm, respectively. Readings were analyzed using MARS data analysis software (Guelph, ON, Canada).
Cell sorting and analysis using flow cytometry
Tumorspheres/cell aggregates were dissociated using liberase (Roche, Laval, QC, Canada) and cells at a concentration of 1 million single cells/ml in phosphate-buffered saline+2 mM EDTA were stained with APC-conjugated anti-CD133 or a matched isotype control (Miltenyi, Auburn, CA, USA). The MoFlo XDP Cell Sorter was used to analyze/sort samples and Kaluza software (Beckman Coulter, Mississauga, ON, Canada) was used for analysis. Mouse IgG CompBeads (BD, San Jose, CA, USA) were used for compensation and viability dye 7-AAD dye (1:10; Beckman Coulter) was added in samples to exclude dead cells. Purity check for CD133 expression levels after sorting was also performed.
Quantitative real-time–polymerase chain
Norgen Total RNA isolation kit was used for RNA extraction. RNA was quantified using NanoDrop Spectrophotometer ND-1000 (Thermoscientific, Wilmington, DE, USA). Using 1 μg RNA, complementary DNA was synthesized using qScript cDNA Super Mix (Quanta Biosciences, Beverly, MA, USA) and a C1000 Thermo Cycler (Bio-Rad, Mississauga, ON, Canada) with the following cycle parameters: 4 min at 25 °C, 30 min at 42 °C, 5 min at 85 °C, hold at 4 °C. qRT-PCR was performed using Perfecta SybrGreen (Quanta Biosciences) and an 4CFX96 instrument (Bio-Rad) as described previously.55 Gene expression was quantified by using CFX Manager 3.0 (Bio Rad) software and expression levels were normalized to GAPDH/Actin expression. Primers are listed in Supplementary Table 1.
Screening of MBSCs with STAT3 inhibitors
Dr Patrick Gunning provided a library of proprietary direct-binding STAT3 inhibitors. Cell proliferation in MBSCs (grown in DMEM supplemented with 20% fetal bovine serum) was determined by plotting percentage cell viability versus log dilutions of the inhibitors, giving IC50. To assess tumor inhibition in mice, compound PG-S3-002 was chosen for ex vivo treatment and modification of compound PG-S3-002 resulted in synthesis of compound PG-S3-009, which has increased bioavailability and was used for in vivo treatment. See Supplementary Figure 8 for details about PG-S3-009.
In vivo intracranial injections of MB stem cells and H&E staining of xenograft tumors
The McMaster University Animal Research Ethics Board (AREB) approved all experimental procedures and intracranial injections were performed as described previously.13 Briefly, live cells (numbers mentioned in figure legends) re-suspended in 10 μl of phosphate-buffered saline were injected into the frontal lobe of NOD-SCID mice (both males and females of 8-10 weeks of age). Mice injected with CD133 sorted cells (positive or negative) were killed when the CD133-positive group reached end point. Treated mice (with either shSTAT3 or ex vivo STAT3 inhibitor or in vivo STAT3 inhibitor) were killed when control Group/untreated mice reached end point as shown by signs of illness, for tumor size quantification. Mice were randomized into control or treatment groups. Group allocation was not blinded. For survival studies, treated or control mice were killed when they reached end point. The treatment did not induce adverse weight loss that would be contraindicated by our animal utilization protocol (AUP) endpoints policy.
Mouse brains were harvested, formalin-fixed and thereafter brain sections were paraffin-embedded and also stained for hematoxylin and eosin (Cell Signaling). Aperio Slide Scanner (Leica Biosystems, Concord, ON, Canada) was used to scan images and images were analyzed using ImageScope v11.1.2.760 software (Aperio, Concord, ON, Canada). Survival curves were plotted using Graphpad Prism 5 (SanDiego, CA, USA).
Statistical analysis
Technical/experimental replicates from at least three samples were compiled for each experiment, unless otherwise stated in figure legends. Respective data represent mean±s.d. with n values listed in figure legends. Student’s t-test analyses and two-way analysis of variance were performed using GraphPad Prism 5. P<0.05 was considered significant. For in silico analyses, statistical tests were complied and completed in R.
References
Northcott PA, Jones DT, Kool M, Robinson GW, Gilbertson RJ, Cho YJ et al. Medulloblastomics: the end of the beginning. Nat Rev Cancer 2012; 12: 818–834.
Taylor MD, Northcott PA, Korshunov A, Remke M, Cho YJ, Clifford SC et al. Molecular subgroups of medulloblastoma: the current consensus. Acta Neuropathol 2012; 123: 465–472.
Gajjar A, Chintagumpala M, Ashley D, Kellie S, Kun LE, Merchant TE et al. Risk-adapted craniospinal radiotherapy followed by high-dose chemotherapy and stem-cell rescue in children with newly diagnosed medulloblastoma (St Jude Medulloblastoma-96): long-term results from a prospective, multicentre trial. Lancet Oncol 2006; 7: 813–820.
Mabbott DJ, Spiegler BJ, Greenberg ML, Rutka JT, Hyder DJ, Bouffet E . Serial evaluation of academic and behavioral outcome after treatment with cranial radiation in childhood. J Clin Oncol 2005; 23: 2256–2263.
Northcott PA, Korshunov A, Witt H, Hielscher T, Eberhart CG, Mack S et al. Medulloblastoma comprises four distinct molecular variants. J Clin Oncol 2011; 29: 1408–1414.
Pei Y, Moore CE, Wang J, Tewari AK, Eroshkin A, Cho YJ et al. An animal model of MYC-driven medulloblastoma. Cancer Cell 2012; 21: 155–167.
Robinson G, Parker M, Kranenburg TA, Lu C, Chen X, Ding L et al. Novel mutations target distinct subgroups of medulloblastoma. Nature 2012; 488: 43–48.
Kool M, Korshunov A, Remke M, Jones DT, Schlanstein M, Northcott PA et al. Molecular subgroups of medulloblastoma: an international meta-analysis of transcriptome, genetic aberrations, and clinical data of WNT, SHH, Group 3, and Group 4 medulloblastomas. Acta Neuropathol 2012; 123: 473–484.
Northcott PA, Shih DJ, Peacock J, Garzia L, Morrissy AS, Zichner T et al. Subgroup-specific structural variation across 1,000 medulloblastoma genomes. Nature 2012; 488: 49–56.
Parsons DW, Li M, Zhang X, Jones S, Leary RJ, Lin JC et al. The genetic landscape of the childhood cancer medulloblastoma. Science 2011; 331: 435–439.
Grosse-Gehling P, Fargeas CA, Dittfeld C, Garbe Y, Alison MR, Corbeil D et al. CD133 as a biomarker for putative cancer stem cells in solid tumours: limitations, problems and challenges. J Pathol 2013; 229: 355–378.
Cox CV, Diamanti P, Evely RS, Kearns PR, Blair A . Expression of CD133 on leukemia-initiating cells in childhood ALL. Blood 2009; 113: 3287–3296.
Singh SK, Hawkins C, Clarke ID, Squire JA, Bayani J, Hide T et al. Identification of human brain tumour initiating cells. Nature 2004; 432: 396–401.
Singh SK, Clarke ID, Terasaki M, Bonn VE, Hawkins C, Squire J et al. Identification of a cancer stem cell in human brain tumors. Cancer Res. 2003; 63: 5821–5828.
Beier D, Hau P, Proescholdt M, Lohmeier A, Wischhusen J, Oefner PJ et al. CD133(+) and CD133(−) glioblastoma-derived cancer stem cells show differential growth characteristics and molecular profiles. Cancer Res 2007; 67: 4010–4015.
Shmelkov SV, Butler JM, Hooper AT, Hormigo A, Kushner J, Milde T et al. CD133 expression is not restricted to stem cells, and both CD133+ and CD133- metastatic colon cancer cells initiate tumors. J Clin Invest 2008; 118: 2111–2120.
Haftchenary S, Avadisian M, Gunning PT . Inhibiting aberrant Stat3 function with molecular therapeutics: a progress report. Anti-Cancer Drugs 2011; 22: 115–127.
Holtick U, Vockerodt M, Pinkert D, Schoof N, Sturzenhofecker B, Kussebi N et al. STAT3 is essential for Hodgkin lymphoma cell proliferation and is a target of tyrphostin AG17 which confers sensitization for apoptosis. Leukemia 2005; 19: 936–944.
Burke WM, Jin X, Lin HJ, Huang M, Liu R, Reynolds RK et al. Inhibition of constitutively active Stat3 suppresses growth of human ovarian and breast cancer cells. Oncogene 2001; 20: 7925–7934.
Abdulghani J, Gu L, Dagvadorj A, Lutz J, Leiby B, Bonuccelli G et al. Stat3 promotes metastatic progression of prostate cancer. Am J Pathol 2008; 172: 1717–1728.
Carro MS, Lim WK, Alvarez MJ, Bollo RJ, Zhao X, Snyder EY et al. The transcriptional network for mesenchymal transformation of brain tumours. Nature 2010; 463: 318–325.
Stechishin OD, Luchman HA, Ruan Y, Blough MD, Nguyen SA, Kelly JJ et al. On-target JAK2/STAT3 inhibition slows disease progression in orthotopic xenografts of human glioblastoma brain tumor stem cells. Neuro-oncology 2013; 15: 198–207.
Bandopadhayay P, Bergthold G, Nguyen B, Schubert S, Gholamin S, Tang Y et al. BET bromodomain inhibition of MYC-amplified medulloblastoma. Clin Cancer Res 2014; 20: 912–925.
Wang X, Venugopal C, Manoranjan B, McFarlane N, O'Farrell E, Nolte S et al. Sonic hedgehog regulates Bmi1 in human medulloblastoma brain tumor-initiating cells. Oncogene 2012; 31: 187–199.
Friedman HS, Colvin OM, Kaufmann SH, Ludeman SM, Bullock N, Bigner DD et al. Cyclophosphamide resistance in medulloblastoma. Cancer Res 1992; 52: 5373–5378.
Cho YJ, Tsherniak A, Tamayo P, Santagata S, Ligon A, Greulich H et al. Integrative genomic analysis of medulloblastoma identifies a molecular subgroup that drives poor clinical outcome. J Clin Oncol 2011; 29: 1424–1430.
Kool M, Koster J, Bunt J, Hasselt NE, Lakeman A, van Sluis P et al. Integrated genomics identifies five medulloblastoma subtypes with distinct genetic profiles, pathway signatures and clinicopathological features. PloS One 2008; 3: e3088.
Thompson MC, Fuller C, Hogg TL, Dalton J, Finkelstein D, Lau CC et al. Genomics identifies medulloblastoma subgroups that are enriched for specific genetic alterations. J Clin Oncol 2006; 24: 1924–1931.
Lin L, Fuchs J, Li C, Olson V, Bekaii-Saab T, Lin J . STAT3 signaling pathway is necessary for cell survival and tumorsphere forming capacity in ALDH(+)/CD133(+) stem cell-like human colon cancer cells. Biochem Biophys Res Commun 2011; 416: 246–251.
Won C, Kim BH, Hee YiE, Choi KJ, Kim EK, Jeong JM et al. STAT3-mediated CD133 upregulation contributes to promotion of hepatocellular carcinoma. Hepatology 2015;; 62: 1160–1173.
Zeppernick F, Ahmadi R, Campos B, Dictus C, Helmke BM, Becker N et al. Stem cell marker CD133 affects clinical outcome in glioma patients. Clin Cancer Res 2008; 14: 123–129.
Kim J, Woo AJ, Chu J, Snow JW, Fujiwara Y, Kim CG et al. A Myc network accounts for similarities between embryonic stem and cancer cell transcription programs. Cell 2010; 143: 313–324.
Ashizawa T, Miyata H, Iizuka A, Komiyama M, Oshita C, Kume A et al. Effect of the STAT3 inhibitor STX-0119 on the proliferation of cancer stem-like cells derived from recurrent glioblastoma. Int J Oncol 2013; 43: 219–227.
Sai K, Wang S, Balasubramaniyan V, Conrad C, Lang FF, Aldape K et al. Induction of cell-cycle arrest and apoptosis in glioblastoma stem-like cells by WP1193, a novel small molecule inhibitor of the JAK2/STAT3 pathway. J Neuro-oncology 2012; 107: 487–501.
Huang MJ, Cheng YC, Liu CR, Lin S, Liu HE . A small-molecule c-Myc inhibitor, 10058-F4, induces cell-cycle arrest, apoptosis, and myeloid differentiation of human acute myeloid leukemia. Exp Hematol 2006; 34: 1480–1489.
Kawauchi D, Robinson G, Uziel T, Gibson P, Rehg J, Gao C et al. A mouse model of the most aggressive subgroup of human medulloblastoma. Cancer Cell 2012; 21: 168–180.
Chen F, Xu Y, Luo Y, Zheng D, Song Y, Yu K et alDown-regulation of Stat3 decreases invasion activity and induces apoptosis of human glioma cells Journal of molecular neuroscience: MN, 2010; 40: 353–359.
Li GH, Wei H, Lv SQ, Ji H, Wang DL . Knockdown of STAT3 expression by RNAi suppresses growth and induces apoptosis and differentiation in glioblastoma stem cells. International journal of oncology 2010; 37: 103–110.
Rahaman SO, Harbor PC, Chernova O, Barnett GH, Vogelbaum MA, Haque SJ . Inhibition of constitutively active Stat3 suppresses proliferation and induces apoptosis in glioblastoma multiforme cells. Oncogene 2002; 21: 8404–8413.
Aoki Y, Feldman GM, Tosato G . Inhibition of STAT3 signaling induces apoptosis and decreases survivin expression in primary effusion lymphoma. Blood 2003; 101: 1535–1542.
Ding BB, Yu JJ, Yu RY, Mendez LM, Shaknovich R, Zhang Y et al. Constitutively activated STAT3 promotes cell proliferation and survival in the activated B-cell subtype of diffuse large B-cell lymphomas. Blood 2008; 111: 1515–1523.
Xiao H, Bid HK, Jou D, Wu X, Yu W, Li C et al. A novel small molecular STAT3 inhibitor, LY5, inhibits cell viability, cell migration, and angiogenesis in medulloblastoma cells. The Journal of biological chemistry 2015; 290: 3418–3429.
Chang CJ, Chiang CH, Song WS, Tsai SK, Woung LC, Chang CH et al. Inhibition of phosphorylated STAT3 by cucurbitacin I enhances chemoradiosensitivity in medulloblastoma-derived cancer stem cells. Child Nervous Syst 2012; 28: 363–373.
Haftchenary S, Luchman HA, Jouk AO, Veloso AJ, Page BD, Cheng XR et al. Potent targeting of the STAT3 protein in brain cancer stem cells: a promising route for treating glioblastoma. ACS Med Chem Lett 2013; 4: 1102–1107.
Turkson J, Ryan D, Kim JS, Zhang Y, Chen Z, Haura E et al. Phosphotyrosyl peptides block Stat3-mediated DNA binding activity, gene regulation, and cell transformation. J Biol Chem 2001; 276: 45443–45455.
Song H, Wang R, Wang S, Lin J . A low-molecular-weight compound discovered through virtual database screening inhibits Stat3 function in breast cancer cells. Proceedings Natl Acad Sci USA 2005; 102: 4700–4705.
Zhang X, Yue P, Page BD, Li T, Zhao W, Namanja AT et al. Orally bioavailable small-molecule inhibitor of transcription factor Stat3 regresses human breast and lung cancer xenografts. Proceedings Natl Acad Sci USA 2012; 109: 9623–9628.
Eiring AM, Page BD, Kraft IL, Mason CC, Vellore NA, Resetca D et al. Combined STAT3 and BCR-ABL1 inhibition induces synthetic lethality in therapy-resistant chronic myeloid leukemia. Leukemia 2015; 29: 586–597.
Singh M, Garg N, Venugopal C, Hallett R, Tokar T, McFarlane N et al. STAT3 pathway regulates lung-derived brain metastasis initiating cell capacity through miR-21 activation. Oncotarget 2015; 6: 27461–27477.
Schaefer LK, Ren Z, Fuller GN, Schaefer TS . Constitutive activation of Stat3alpha in brain tumors: localization to tumor endothelial cells and activation by the endothelial tyrosine kinase receptor (VEGFR-2). Oncogene 2002; 21: 2058–2065.
Zhao D, Pan C, Sun J, Gilbert C, Drews-Elger K, Azzam DJ et al. VEGF drives cancer-initiating stem cells through VEGFR-2/Stat3 signaling to upregulate Myc and Sox2. Oncogene 2015; 34: 3107–3119.
Gu C, Yokota N, Gao Y, Yamamoto J, Tokuyama T, Namba H . Gene expression of growth signaling pathways is up-regulated in CD133-positive medulloblastoma cells. Oncology Lett 2011; 2: 357–361.
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 2003; 4: 249–264.
Venugopal C, McFarlane NM, Nolte S, Manoranjan B, Singh SK . Processing of primary brain tumor tissue for stem cell assays and flow sorting. J Vis Exp 2012.
Singh SK, Venugopal C, Hallett R, Vora P, Manoranjan B, Mahendram S et al. Pyrvinium targets CD133 in human glioblastoma brain tumor-initiating cells. Clin Cancer Res 2015; 21: 5324–5337.
Acknowledgements
Dr Sheila K Singh is supported by funds from the Department of Surgery at McMaster University, Canada Research Chair award, Canadian Institutes of Health Research (CIHR) Operating Grant #RN193483–299211, Stem Cell Network Drug Discovery Grant, Ontario Institute for Cancer Research Cancer Stem Cell Program, the Canadian Cancer Society Research Institute and Terry Fox Research Institute New Investigator Award. The Alberta Innovates, Stem Cell Network, CIHR, ORF and CFI fund Dr Patrick T. Gunning’s research. Dr John A. Hassell is supported by the funds from CIHR Operating Grant # MOP-142353
Author contributions
NG and SKS: conception and design, development of methodology, acquisition of data, analysis and interpretation of data, writing, review and/or revision of the manuscript, and study supervision. DB: acquisition of data, writing, review and/or revision of the manuscript. CV: conception and design, analysis and interpretation of data, writing, review and/or revision of the manuscript. SM and NM: acquisition of data, analysis and interpretation of data. DAR, CCA, PSL, RFGB AMA and DE: technical or material support. TV: acquisition of data. BM: writing, review and/or revision of the manuscript. RH: acquisition of data, analysis and interpretation of data. KHD, JMK: analysis and interpretation of data. OAA, JAH: administrative, technical or material support. PTG: conception and design, development of methodology, analysis and interpretation of data, writing, review and/or revision of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Garg, N., Bakhshinyan, D., Venugopal, C. et al. CD133+ brain tumor-initiating cells are dependent on STAT3 signaling to drive medulloblastoma recurrence. Oncogene 36, 606–617 (2017). https://doi.org/10.1038/onc.2016.235
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2016.235
- Springer Nature Limited
This article is cited by
-
Marinopyrrole derivative MP1 as a novel anti-cancer agent in group 3 MYC-amplified Medulloblastoma
Journal of Experimental & Clinical Cancer Research (2024)
-
PFKFB3 works on the FAK-STAT3-SOX2 axis to regulate the stemness in MPM
British Journal of Cancer (2022)
-
STAT3 in medulloblastoma: a key transcriptional regulator and potential therapeutic target
Molecular Biology Reports (2022)
-
RBM5-AS1 promotes radioresistance in medulloblastoma through stabilization of SIRT6 protein
Acta Neuropathologica Communications (2021)
-
MiR-212-3p functions as a tumor suppressor gene in group 3 medulloblastoma via targeting nuclear factor I/B (NFIB)
Acta Neuropathologica Communications (2021)