Abstract
Malignant growth relies on rapid protein synthesis frequently leading to endoplasmic reticulum (ER) overload and accumulation of unfolded or misfolded protein in this cellular compartment. In the ER, protein homeostasis is finely regulated by a mechanism called the unfolded protein response (UPR), involving the activation of signalization pathways mediated by three transmembrane proteins, namely PERK, IRE1 and ATF6. IRE1 endoribonuclease activation leads in particular to the splicing of the cytosolic mRNA encoding the key UPR-specific transcription factor XBP1s. Our study shows that sustained activation of XBP1s expression in acute myeloid leukemia (AML) cells induces apoptosis in vitro and in vivo, whereas a moderate XBP1s expression sensitizes cells to chemotherapeutic treatments. ChIP-seq experiments identified specific XBP1s target genes including the MIR22HG lncRNA, the precursor transcript of microRNA-22-3p. miR-22-3p upregulation by XBP1s or forced expression of miR-22 significantly decreases cell’s viability and sensitizes leukemic cells to chemotherapy. We found that miR-22-3p intracellular effects result at least partially from the targeting of the mRNA encoding the deacetylase sirtuin-1 (SIRT1), a well-established pro-survival factor. Therefore, this novel XBP1s/miR-22/SIRT1 axis identified could play a pivotal role in the proliferation and chemotherapeutic response of leukemic cells.

Similar content being viewed by others
Introduction
Cancer development relies on a sustained protein synthesis machinery and therefore cancer cells adapt protein quality controls to meet the high demand for protein synthesis [1]. Most cellular stresses (hypoxia, nutrient deprivation, oxidative stress, acidosis, etc.) disrupt endoplasmic reticulum (ER) functions, causing misfolded protein accumulation, which activates the unfolded protein response (UPR) intended to restore ER homeostasis [2,3,4,5]. However, if the stress is not resolved, UPR can also lead to cell death [3]. UPR signaling involves the activation of three ER transmembrane proteins (i) the kinase PERK (protein kinase R (PKR)-like ER kinase) (ii) the bifunctional kinase/endoribonuclease IRE1 (inositol requiring enzyme 1), and (iii) the transcription factor ATF6 (activating transcription factor 6) [2, 6, 7].
IRE1, the most conserved arm of UPR, possesses two cytoplasmic catalytic domains (i) a protein kinase domain and (ii) an endoribonuclease domain [2, 8, 9]. Upon activation, the RNAse domain targets specific cytoplasmic mRNA and microRNAs, leading to their decay by the regulated IRE1-dependent decay (RIDD) pathway [10, 11]. In parallel, IRE1α endoribonuclease activity removes a 26 nucleotides sequence from X-box binding protein 1 (XBP1) mRNA coding sequence, inducing a translational frameshift prior re-ligation by the tRNA ligase RTCB [12,13,14]. This “spliced” mRNA is translated into a 376-amino-acids-long isoform XBP1s (s for “spliced”), coding for a potent transcription factor, which regulates stress response genes [15,16,17], including genes involved in protein folding, trafficking, and also some components of the ER-associated protein degradation machinery.
XBP1s expression in solid tumors is frequently associated with a poor prognosis, while in lymphoma and leukemia its role is controversial [18]. For instance, impaired XBP1 activation is a hallmark of one major germinal center B cell-like diffuse large B cell lymphoma subtype and contributes to tumor growth [19]. In acute myeloid leukemia (AML), XBP1s expression correlates with a favorable outcome upon cytarabine and etoposide therapy [20, 21]. Conversely, IRE1 has also been described to drive pro-survival signals in AML and pre-leukemic stem cells [22, 23].
In order to decipher the role of XBP1s in AML, we generated a model enabling conditional expression of the XBP1s isoform in six different cell line models. Our results demonstrate that sustained XBP1s expression can activate the apoptotic-signaling pathway both in vivo and in vitro. Interestingly, a lower and non-toxic XBP1s expression level sensitizes chemoresistant-AML-cell line OCI-AML3 to cytarabine (aracytine) treatment. By integrating RNA and ChIP-sequencing data, we revealed an XBP1s-dependent upregulation of the long noncoding RNA MIR22HG, precursor of miR-22. Interestingly, mature miR-22 is downregulated in AML patient cells [24] and represses genes involved in the DNA damage response [25, 26]. Here we demonstrate that miR-22 mediates apoptosis and enhances the efficacy of aracytine treatment in AML cells via sirtuin-1 (SIRT1) translational inhibition. Taken together these results identify a novel ER stress-induced axis: XBP1/miR-22/SIRT1, as an effector of apoptosis and chemosensitivity in AML.
Materials and methods
Generation of inducible cell lines and shRNA-mediated gene knockdown
The construction of lentiviral vectors expressing the XBP1s, XBP1 shRNAs and miR-22 constructs as well as transductions and establishment of stable and inducible cell lines are described in the Supplementary Information.
Cell culture and treatments
All leukemic cell lines were obtained from the Leibniz Institute DSMZ or ATCC and grown as recommended by the provider. The clinical and mutational features of our 6 AML cell lines are described in Supplementary Table 1. All drugs used were purchased from Sigma-Aldrich.
Reverse transcription and quantitative PCR
RNA was extracted as previously described [27]. For mRNA and long noncoding RNA, 500 ng of total RNA were reverse transcribed using the PrimeScriptTM RT-PCR Kit (Clontech) according to the manufacturer’s protocol. Reverse transcription (RT) reactions were diluted 20-fold and amplification was performed in a total volume of 10 µL containing 5 µL of the SYBR TB Green® Premix Ex TaqII™ from TakaraBio, 1 µL of both primers (final concentration of 300 nM each), and 2 µL of diluted cDNA. qPCR was performed on the StepOnePlus real-time PCR system (Applied Biosystems) and results were analyzed with the StepOne software. For mature microRNA quantification, RT was performed using the miRCURY™ LNA Universal RT kit and qPCR by the miRCURYTM LNA miRNA PCR system (both kits from Qiagen). hsa-let-7a-5p expression was used for normalization. All primers used in this study are listed in Supplementary Table 2.
Western blotting
Western blots were performed as previously described [28]. The list of the commercial antibodies is given in the Supplementary Methods together with a description of sample preparation and immunodetection processes.
siRNA and microRNA transfections
XBP1s siRNAs (Smart Pool, Dharmacon), siSIRT1 #1 (s23771, Thermo Fisher) siSIRT1 #2 (s23770, Thermo Fisher) as well as negative control siRNA were transfected in OCI-AML3 at a final concentration of 10 nM or 50 nM using the Lipofectamine RNAiMAX transfection reagent (Thermo Fisher) according to the manufacturer’s instructions. mirVanaTM miRNA mimic hsa-miR-22-3p and negative control (Thermo Fisher) were transfected using the same protocol. Cells were harvested 24 or 48 h later for protein and RNA analyses.
Flow cytometry measurements of apoptosis and cell viability assays
Analysis of apoptosis was done using Annexin V (Annexin-Pacific BlueTM) and propidium iodide (PI) (Biolegend # 640928) staining, according to manufacturer’s recommendations and followed by flow cytometry analysis using a MACSQuant® VYB from Miltenyi Biotec. Results were analyzed using the FlowJoTM software.
Murine xenograft models
All animal procedures were conducted in compliance with the Ethics Committee pursuant to European legislation translated into French Law as Décret 2013‐118 dated 1st February 2013 (APAFIS 3601‐2015121622062840). Animals were used in accordance to a protocol reviewed and approved by the Institutional Animal Care and Use Committee of Région Occitanie (France). All the experiments were carried out in accordance with relevant guidelines and regulations. NSG and nude mice were obtained from Charles River. Mice treatments, tumor engraftment and tumor growth monitoring are detailed in the Supplementary Methods section.
Chromatin immunoprecipitation (ChIP)
XBP1s expression was induced with 10 ng/mL of doxycycline for 48 h. ChIP was then performed using the ChIP-IT® Express kit (Active Motif) with minor modifications. Additional reagents used and procedures are given in the Supplementary Methods section.
AML patient samples
AML and normal bone marrow samples were obtained from patients at the Department of Hematology (CHU, Toulouse, France) after consent, in accordance with the Declaration of Helsinki.
Biotin pull-down assay
Biotinylated mimic miR-22 (purchased from Exiqon) was transfected in 5.106 OCI-AML3 cells using RNAiMAX (Thermo Fisher) at 10 nM final concentration for 24 h. Cells were then harvested and processed for pull-down assay as detailed in the Supplementary Methods section.
Statistical analyses
Results are presented as mean values ± standard deviations (SD). Differences between two groups were examined using a two-tailed Student’s t-test. Survival analyses were performed using the log-rank test. For the subcutaneous xenografts, determination of statistical significance was performed using two-way ANOVA followed by the Bonferroni test. All analyses were performed using GraphPad Prism version 8.4.
Results
XBP1s-inducible expression elicits an ER stress-like response and induces apoptosis in vitro
To identify the role of IRE1/XBP1 pathway activation in AML, we established an inducible OCI-AML3 cell line with doxycycline-controlled XBP1s expression. Tet-On cells expressing the rtTA transactivator were transduced with a bicistronic lentivector enabling simultaneous expression of XBP1s and the reporter eGFP (Fig. 1A). To validate our model, control cells (Tet-On) or XBP1s cells were treated with increasing amounts of doxycycline, resulting in increased XBP1s expression both at the protein (Fig. 1B) and mRNA level (Fig. 1C).
A Schematic of XBP1s-inducible model generation: OCI-AML3 cells were first transduced by a lentivirus expressing the rtTA doxycycline-inducible transactivator. These cells were used as “Tet-On” control cells in the following experiments. These Tet-On cells were then transduced with a lentivector expressing the XBP1s transgene followed by an IRES-eGFP cassette, under the control of a TET-inducible promoter. GFP-positive cells were sorted using flow cytometry after 24 h of doxycycline treatment. OCI-AML3 cells were treated with increasing amounts of doxycycline during 48 h. B XBP1s protein level was evaluated by western blot; GAPDH was used as loading control. C XBP1s expression was evaluated by RT-qPCR. Expression values were normalized to the housekeeping genes HPRT, MLN51 and ABL, and are depicted as a ratio of mRNA expression in doxycycline-induced cells relative to untreated cells. D Venn diagrams showing the overlap between RNA seq and ChIP seq experiments in OCI-AML3 XBP1s cells treated with 10 ng/ml of doxycycline for 48 h. The 911 genes selected in RNA seq are the ones with expression increased or inhibited by more than twofold in XBP1s cells compared to control cells. The 458 genes, shortlisted from the ChIP seq analysis, display an XBP1s peak at least at 750 bp upstream or downstream of the characterized transcription start site. E Variations in the expression level of the 390 genes showing modulated expression in OCI-AML3-XBP1s cells upon treatment with doxycycline and overlapping between RNA seq and ChIP seq experiments. F Gene ontology analysis was performed with the 390 genes up- and downregulated overlapping between RNA seq and ChIP seq experiments in OCI-AML3-XBP1s cells treated with doxycycline. G Gene set enrichment analysis (GSEA) was performed on the RNA seq full data of OCI-AML3-XBP1s cells treated with doxycycline. NES normalized enrichment score, FDR false discovery rate. H Apoptosis levels of OCI-AML3 Tet-On or OCI-AML3-XBP1s cells treated with increasing amounts of doxycycline during 48 h were measured by flow cytometry using Annexin V/PI staining. Data represent mean ± SD (n = 3).
We next confirmed the functionality of the exogenously expressed XBP1s protein after induction with low doses of doxycycline by analyzing the expression of DNAJB9 (ERdj4), and HSPA5 (BiP), which are highly induced in response to IRE1/XBP1 signaling activation [29] (Supplementary Fig. 1A, B). Moreover, the expression of DNAJB9 at the mRNA level increased proportionally to XBP1s (Supplementary Fig. 1C). In addition, XBP1s expression has no effect on eIF2α phosphorylation (Supplementary Fig. 1D) indicating that the PERK pathway is not activated upon XBP1s overexpression.
Post-UPR activation, XBP1s mainly regulates stress response genes [30,31,32]. To address the role of XBP1s, independently of the two other UPR pathways, we performed several analyses comparing XBP1s OCI-AML3 to Tet-On OCI-AML3 treated with 10 ng/mL of doxycycline: (i) ChIP followed by sequencing to determine the XBP1s genome-wide binding pattern, (ii) global RNA sequencing and (iii) micro-RNA sequencing. We cross-referenced RNA sequencing and ChIP sequencing data to identify XBP1s direct targets. We captured 911 mRNAs that were up or downregulated at least by twofold following XBP1s expression (Supplementary Table 3) along with 458 Bio-ChIP-seq peaks located at <750 bp from a characterized +1 transcription start site. We observed that more than 85% of XBP1s peaks (390 genes) overlapped with genes deregulated after XBP1s induction (Fig. 1D and Supplementary Table 4). Among these 390 genes, only 6 were downregulated (≤3.5-fold), while the 384 others were upregulated, including some of them more than 100-fold compared to the control condition (Fig. 1E). These results demonstrate that XBP1s, when activated independently of other UPR pathways, acts essentially as an activating factor. As expected, the 390 differentially expressed genes are mainly involved in response to unfolded protein accumulation, vesicle-mediated transport and are closely linked to the ER- and IRE1-mediated UPR (Fig. 1F). Notably, XBP1s directly upregulated numerous genes associated with protein processing in the ER (Supplementary Fig. 2A). In agreement, gene set enrichment analysis (GSEA) showed that genes associated with response to ER stress and unfolded protein were enriched in our model (Fig. 1G top and Supplementary Fig. 2B). Moreover, hallmark gene set enrichment analysis also revealed an upregulation of genes involved in apoptosis (Fig. 1G, bottom). Thus, we examined whether XBP1s expression could affect OCI-AML3 cell viability. Flow cytometry analysis demonstrated that increased expression of XBP1s triggers apoptosis in a doxycycline dose-dependent way while doxycycline had no effect on apoptosis in Tet-On control cells (Fig. 1H).
To further validate apoptosis mediated by XBP1s expression observed in OCI-AML3 in a wider panel of AML cell lines, we established five additional tetracycline-inducible cell lines expressing the XBP1 spliced isoform: OCI-AML2, MOLM-14, MV4-11, THP1 and HL60 (Supplementary Fig. 3A). Inducible XBP1s transgene expression was validated both at the mRNA and protein level (Supplementary Fig. 3B–F) and by analyzing the expression levels of downstream targets, i.e., DNAJB9 and HSPA5 (Supplementary Fig. 4). Flow cytometry analyses confirmed that apoptosis correlates XBP1s’ expression in a doxycycline dose-dependent manner in all cell lines while doxycycline had no effect on apoptosis in Tet-On control cells (Supplementary Fig. 3B–F). These results were further validated with cleaved-PARP detection in cells expressing XBP1s (Supplementary Fig. 5).
Increased XBP1s expression impairs tumoral progression and induces apoptosis in vivo
To assess the impact of chronic XBP1s expression on leukemic progression, we transduced OCI-AML3 cells expressing XBP1s with a lentivector constitutively expressing a firefly luciferase. These cells were then injected intravenously into NSG mice. After 18 days, when the luciferase signal was detectable by bioluminescence imaging in all the animals, drinking water was supplemented (or not) with 1 mg/ml of doxycycline (Fig. 2A). Upon treatment mice exhibited a profound delay in leukemia progression compared to untreated mice (Fig. 2B). Furthermore, overall survival of doxycycline-treated mice was significantly extended compared to untreated control mice (Fig. 2C). In order to evaluate whether a transient activation of XBP1s expression could also inhibit leukemic progression, OCI-AML3-, OCI-AML2- or HL60-XBP1s cells or control Tet-On cells were intravenously injected into NSG mice and, after 9 days of engraftment, drinking water was supplemented with 1 mg/ml of doxycycline for 8 days (Fig. 2D). Transient XBP1s expression significantly increased the overall survival of mice for all tested AML cell lines (Fig. 2E–G). Mice injected with the control Tet-On cells displayed splenomegaly, a feature that was absent in mice injected with the XBP1s-expressing cells (Fig. 2H).
A NSG mice (n = 6 for each group) were injected intravenously with OCI-AML3-XBP1s LucF/GFP-expressing cells. Eighteen days after engraftment, when the luciferase signal was detectable in all the animals, doxycycline was added (or not) at 1 mg/ml in the drinking water. B Leukemia development in mice was monitored by bioluminescence imaging with the IVIS Spectrum CT System (PerkinElmer) at the indicated time points. C Survival analysis of each group of mice was performed using a log-rank test (**p ≤ 0.01). D NSG mice were intravenously injected with either XBP1s-inducible AML cells or their control (Tet-On) counterparts. After a 9 days engraftment, doxycycline was added at the concentration of 1 mg/mL in drinking water during 8 days. E–G Survival analyses were then performed using the log-rank test (*p ≤ 0.05; ****p ≤ 0.0001) E injection of OCI-AML3 cells F injection of OCI-AML2 cells G injection of HL60 cells. H Analysis of spleen size after euthanasia of mice in each of the control (TET-ON) or XBP1s-expressing cell models.
In order to confirm that chronic activation of XBP1s expression impairs leukemic progression, we used an orthogonal approach where OCI-AML3 cells expressing XBP1s were subcutaneously xenografted in athymic mice (Fig. 3A). While efficient engraftment was observed in mice without doxycycline treatment, supplementation of drinking water with 2 or 0.2 mg/ml of doxycycline after injection completely prevented engraftment. Even after 3 weeks, and at a very low dose of doxycycline (0.02 mg/ml), engraftment was very inefficient (Fig. 3B). To further validate XBP1s anti-tumoral properties, doxycycline was added to the mice’s drinking water 17 days post injection when all tumors have reached an average size of 300 mm3 (Fig. 3C). XBP1s induction caused dramatic tumor regression compared to untreated mice (Fig. 3D). At day 23, experiment was stopped and any remaining tumors were collected for analysis of their protein content. Western blot analyses of individual mouse tumor extracts revealed that XBP1s expression was accompanied by the cleavage of both PARP and Caspase 3 (samples 8–14), thus confirming apoptosis-signaling induction in vivo (Fig. 3E). These data further demonstrated XBP1s anti-leukemic activity in vivo.
A Nude mice were grafted subcutaneously with OCI-AML3 XBP1s-expressing cells and were treated without (Ni) or with doxycycline at 2 mg/ml, 0.2 mg/mL or 0.02 mg/ml. B The mean tumor volume in mm3 was calculated over a 22-day period. Data represent mean ± SD (n = 8 for each group). Statistical analyses were performed by two-way ANOVA with Bonferroni correction (**p ≤ 0.01, ****p ≤ 0.0001). C Nude mice were injected subcutaneously with OCI-AML3 XBP1s cells. Seventeen days post injection, mice were exposed (or not), to doxycycline dissolved in their drinking water at a concentration of 2 mg/ml or 0.2 mg/mL. D The mean tumor volume in mm3 was calculated every 2 days after doxycycline treatment. Data represent mean ± SD (n = 7 for each group). Statistical analysis was performed by two-way ANOVA with Bonferroni correction (**p ≤ 0.01, ***p ≤ 0.001). E XBP1s expression, PARP and Caspase 3 cleavage were analyzed by western blotting on protein extracts from tumor samples collected at day 23 (end point) and using GAPDH as loading control. Samples 1–7 correspond to individual mice untreated with doxycycline (red curve). Samples 8–14 correspond to individual mice treated with a dose of 0.2 mg/mL of doxycycline in drinking water (black curve).
Increased XBP1s expression potentiates chemotherapeutic treatment in OCI-AML3 resistant cells
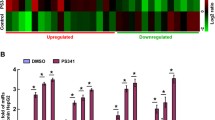
Considering that chronic low XBP1s level in cancer cells supports cell survival, we selected a doxycycline dose that does not trigger cell death (i.e., 4 ng/mL) to investigate the effects of XBP1s’ expression during chemotherapeutic treatment. Annexin V analysis of XBP1s-inducible OCI-AML3 cells by flow cytometry revealed that XBP1s expression sensitized cells to aracytine (Fig. 4A), but not to bortezomib, vinblastine or staurosporine (Fig. 4B) compared to Tet-On control cells. For completeness, we investigated whether ER stress induction by tunicamycin would impact cell survival upon aracytine treatment. Indeed, the combination potentiated apoptosis (Fig. 4C).
A OCI-AML3 Tet-On (control) and XBP1s received a 24 h treatment of doxycycline at 4 ng/mL, followed by a 24 h treatment with aracytine (10 µM). Percentage of apoptotic cells was measured by flow cytometry using Annexin V/PI staining. B OCI-AML3 Tet-On and XBP1s-inducible cells were treated with doxycycline (4 ng/mL) for 24 h and then with bortezomib (BTZ), vinblastine or staurosporine (Stauro), respectively, at 5 nM, 0.5 μM and 0.2 μM for 24 h or left untreated (NT). Data represent mean ± SD (n = 3). Statistical analyses were performed using unpaired t-tests; *p ≤ 0.05. C Apoptosis was assessed by Annexin V/propidium iodide (PI) staining in OCI-AML3 cells. Data represent mean ± SD (n = 3). The left side represents the apoptosis induced by tunicamycin and aracytine treatment alone. The right side represents the apoptotic response induced by a combinatorial treatment of the ER stress inducer tunicamycin and aracytine which appears significantly higher than the sum of the separate effects of each individual treatment, indicating that ER stress potentiates aracytine treatment. D Schematic of the in vivo procedure. NSG mice were injected (day 0) with 2 million of OCI-AML3 Tet-On (control) and XBP1s cells. After a 9 days engraftment, doxycycline was added at 1 mg/mL in drinking water during 8 days. At day 12, mice were daily injected intraperitoneally with aracytine at 30 mg/kg during 5 days. E Survival analyses of the treated mice were performed using a log-rank test (*p ≤ 0.05).
To further confirm in vivo the effect of XBP1s on chemosensitization, OCI-AML3 expressing XBP1s and control Tet-On cells were intravenously injected into NSG mice. After 9 days of engraftment, drinking water was supplemented with 1 mg/ml of doxycycline in order to induce XBP1s expression during 8 days. Three days after the beginning of doxycycline treatment, mice were daily injected with aracytine for 5 days (Fig. 4D). With the Tet-On control cells, no survival improvement was observed between untreated and aracytine-treated mice (Fig. 4E) while dox-induced XBP1s expression increased mice survival. Strikingly, XBP1s expression in combination with aracytine treatment further improved mice survival, thus confirming XBP1s-dependent chemosensitization in vivo (Fig. 4E).
MIR22HG is a direct target of XBP1s and is upregulated by XBP1s upon ER stress
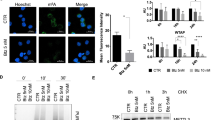
Most reported XBP1s targets in the literature are coding genes, with limited information on direct noncoding targets. Among the noncoding targets characterized here, miRnome analysis revealed a substantial upregulation, over 18-fold, of miR-22-3p following XBP1s expression activation (Supplementary Fig. 6A). Upregulation of the mature miR-22-3p microRNA upon XBP1s induction was validated by RT-qPCR in three different AML cell lines (Fig. 5A–C). Interestingly, we also identified MIR22HG (miR-22 precursor), as one of the most enriched direct XBP1s long noncoding RNA targets (Supplementary Fig. 6B). The initial ChIP-Seq results (Fig. 5D) were confirmed by ChIP-qPCR experiments which revealed an efficient and specific enrichment of miR22HG promoter region following XBP1 pull-down (Fig. 5E). To uncover potential binding sites for XBP1s in an unbiased manner, we performed de novo searches for over-represented sequences among the promoters bound in our ChIP-on-chip experiments and identified a consensus sequence also found in the MIR22HG promoter (Fig. 5F). Gene expression studies using a cohort of 55 primary AML patient samples demonstrated in parallel a positive correlation between XBP1s, MIR22HG and XBP1s known target DNAJB9 (Fig. 5G–I). The findings from our discovery cohort were confirmed by the validation cohort obtained from the BEAT AML data, showing that there is also a positive correlation between MIR22HG and DNAJB9 expression (Fig. 5J). Altogether these data demonstrate that XBP1s can directly activate transcription of MIR22HG and, as a consequence, induce miR-22 overexpression in AML cells.
Mature mir-22-3p expression levels were assessed by RT-qPCR in OCI-AML3- (A), THP1- (B) and MOLM-14-XBP1s (C) cells treated with doxycycline. Expression values were normalized to the control let-7a microRNA, and are depicted as the relative mir-22 expression ratio in doxycycline-treated cells compared to untreated cells. Data represent mean ± SD (n = 3). D Snapshots of ChIP-Seq signals (peaks) representing XBP1s-bound genomic regions in OCI-AML3 cells treated with 10 ng/mL of doxycycline for 48H compared to the input. The MIR22 Host Gene (MIR22HG) promoter region and exons are shown. E RT-qPCR analysis of MIR22HG and DNAJB9 (positive binding control) promoter regions immunoprecipitated following ChIP assay on OCI-AML3 XBP1s cells treated with 10 ng/mL of Dox and performed using anti-XBP1 or IgG isotype control antibodies. The enrichment of target gene promoter regions is expressed in % of input. The actin B (ACT B) promoter region is used as a negative control. F Sequence logos of the consensus cis-regulatory elements discovered in XBP1s target promoter, and localization of this consensus sequence in the MIR22HG promoter region. Pearson’s correlation analyses showing gene expression levels (quantified by RT-qPCR) of XBP1s versus DNAJB9 (G), XBP1s versus MIR22HG (H) and DNAJB9 versus MIR22HG (I) in a 55-AML-patient cohort. Expression values are expressed in Delta Ct (D Ct), calculated using housekeeping genes Actin, MLN51, GAPDH and TBP. J Correlation of MIR22HG and DNAJB9 expression in 483 AML patient samples from the BEAT AML database. Expression levels are shown on Log2 scales. K Survival curves are shown for MIR22HG expression in AML using BEAT AML data. To avoid potential bias in data interpretation, we removed here the samples that were not collected at diagnosis but later after the first line of therapy, those presenting with a myelodysplastic syndrome (MDS) or a myeloproliferative neoplasm (MPN) and the samples from patients who could benefit from a transplantation (either bone marrow or cord blood cells engraftment). The Kaplan–Meier curves were finally plotted with such a homogeneous set of n = 236 patients. Low and high expression levels of MIR22HG are drawn in blue and red, respectively. The p value represents the equality of survival curves based on a log-rank test.
Analysis of mir22HG level in samples categorized according to the French–American–British (FAB) classification enabled us to highlight a noticeable progressive increase in miR-22HG expression from M2 to M5 groups (Supplementary Fig. 7). Furthermore, to explore whether MIR22HG expression could affect the clinical outcomes, we used the Vizome data analysis tool, which contains data from the BEAT AML cohort. Interestingly, high miR22HG RNA expression was associated with improved overall survival in AML patients (Fig. 5K).
miR-22 has an anti-leukemic effect in vitro and participates in the XBP1s-induced phenotype
To explore the impact of miR-22 expression in cellulo, we transfected OCI-AML3 cells with a miR-22 mimic and observed a significant increase in the apoptotic response compared to the control mimic RNA, as evidenced by the increased number of Annexin V-positive cells as well as increased PARP cleavage (Fig. 6A, B).
A OCI-AML3 cells were transfected with a mir-22 mimic (22) or a non-relevant miRNA (Neg) for 48 h. Apoptosis was then assessed by cytometry using Annexin V/PI staining. Lower panel: mean ± SD of three independent experiments. B Western blot analysis of cleaved-PARP levels following mir-22 transfection in OCI-AML3. C OCI-AML3 Tet-On cells were transduced with a lentivector expressing a dox-inducible miR-22. Transduced cells and Tet-On control cells were then treated with increasing amounts of doxycycline during 48 h. Apoptosis was measured by flow cytometry using Annexin V/PI staining. Data represent mean ± SD (n = 3). D OCI-AML3 XBP1s cells were transfected with antisense oligonucleotides against mir-22 (anti-miR-22) or non-relevant oligonucleotides (anti-miR-neg) as control. After transfection, cells were treated with increasing amounts of doxycycline. After 24 h, apoptosis was quantified. Data represent mean ± SD (n = 3). E OCI-AML3 XBP1s cells were transfected with antisense oligonucleotides against mir-22-3p (anti-miR-22) or non-relevant oligonucleotide (anti-mir-neg) as control. After transfection, cells were treated with 4 ng/mL of doxycycline to induce XBP1s expression and with aracytine (AraC), at 10 μM. Apoptosis was quantified after 24 h. Data represent mean ± SD (n = 3). Statistical analyses were performed using unpaired t-tests (**p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0,0001).
To confirm this effect, we generated OCI-AML2 and OCI-AML3 models expressing miR-22-3p under the control of a doxycycline-inducible promoter. In both cell lines, the addition of increasing doses of doxycycline progressively elicited an apoptotic response in miR-22-expressing cells, but not in Tet-On control cells (Fig. 6C and Supplementary Fig. 8). Moreover, while transfection of an anti-miR-22-3p oligonucleotide had only a slight effect on OCI-AML3 basal apoptosis, it partially rescued XBP1s-induced apoptosis (Fig. 6D) demonstrating that the pro-apoptotic effect of XBP1s is driven at least partially by miR-22-3p expression. Additional experiments revealed that miR-22-3p inhibition through anti-miR-22-3p transfection also significantly decreased aracytine-induced apoptosis when compared to transfection with the negative control (anti-miR-Neg ; Fig. 6E), thus confirming that miR-22 participates in the apoptotic response to aracytine treatment.
XBP1s elicits apoptotic responses in AML cells in part through miR-22-3p-mediated downregulation of sirtuin-1 deacetylase (SIRT1) expression
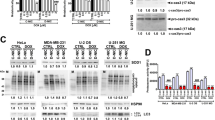
Some previously described miR-22-3p direct targets, MDC1, P21 and SIRT1 [33, 34], have been reported to promote cell survival. Using a biotin-labeling-based pull-down procedure we found that SIRT1 mRNA was efficiently enriched after miR-22 pull-down in OCI-AML3 cells. (Fig. 7A). Interestingly, the deacetylase SIRT1 was previously found to be frequently overexpressed and considered as an oncogene in AML [35,36,37]. Whereas no significant reduction of SIRT1 mRNA was observed after miR-22 mimic transfection (Fig. 7B) SIRT1 protein expression was lowered (Fig. 7C), indicating that miR-22 likely affects SIRT1 expression at the translational level. We confirmed the downregulation of SIRT1 protein expression by miR-22-3p using our two miR-22-inducible cell lines (Fig. 7D, E). We also observed a drop of SIRT1 protein levels following dose-dependent induction of XBP1s expression in OCI-AML3 (Fig. 7F) and XBP1s-inducible OCI-AML2, MOLM-14, MV4-11 and THP1 cells (Supplementary Fig. 9). Finally, we showed that anti-miR-22-3p treatment of doxycycline-induced OCI-AML3 XBP1s cells led to the re-expression of SIRT1 protein, both in untreated and aracytine-treated conditions (Fig. 7G). This supports the role of the XBP1s/miR-22-3p axis in downregulating SIRT1.
A RT-qPCR analysis of MDC1, P21, SIRT1, GAPDH and ABL mRNA levels following an RNA pull-down with a biotinylated mir-22 mimic transfected in OCI-AML3 wild-type cells. miR-neg: non-relevant pull-down with control miRNA. GAPDH and ABL: non-target mRNA controls. Data represent mean ± SD (n = 3). OCI-AML3 cells were either transfected with a miR-22 mimic (miR-22) or a non-relevant mimic (miR-neg). SIRT1 mRNA and protein levels were determined by RT-qPCR analysis (B) and western blotting (C), respectively. For RT-qPCR, values were normalized to the expression level of the housekeeping gene ABL and are shown as relative SIRT1 mRNA expression compared to non-relevant mimic transfection. SIRT1 protein levels were determined by western blotting after miR-22 induction in OCI-AML3-miR-22 (D) and OCI-AML2-miR-22 (E) inducible cells. F SIRT1 protein levels were analyzed by western blotting in OCI-AML3 XBP1s-expressing cells treated with 4 ng/mL or 10 ng/mL of doxycycline. G SIRT1 protein level was assessed by western blotting in OCI-AML3 XBP1s treated with 4 ng/mL of doxycycline and with (or not) 10 µM aracytine treatment and transfected with an antisense oligonucleotide against miR-22-3P (anti-miR-22) or non-relevant oligonucleotide (anti-miR-neg). GAPDH was used as a loading control. H–K OCI-AML3 and THP1 were transfected by two independent siRNAs directed against SIRT1 (si-1; si-2) or a non-relevant siRNA (siSCR) during 48 h. H, J siRNA efficiencies were evaluated by western blot using GAPDH or Actin as loading controls. I, K Apoptosis was measured by flow cytometry using Annexin V/PI staining. Data represent mean ± SD (n = 3). L OCI-AML3 cells were treated for 96 h with 30 µM of the SIRT1 inhibitor EX-527 or its vehicle DMSO, either with (AraC) or without (NT) aracytine treatment at 10 µM during the last 24 h. Apoptosis was measured by flow cytometry using Annexin V/PI staining. Data represent mean ± SD (n = 3). For all the presented experiments statistical analyses were performed using unpaired t-tests (*p ≤ 0.05; **p ≤ 0.01, ***p ≤ 0.001).
We then directly downregulated SIRT1 expression in OCI-AML3 and THP1 cells by RNAi. Two independent siRNAs directed against SIRT1 efficiently repressed its expression (Fig. 7H, J) and increased apoptosis (Fig. 7I, K) when compared to controls. Finally, SIRT1 pharmacological inhibition using the highly specific SIRT1 inhibitor EX-527 specifically enhanced the apoptotic response to treatment with aracytine (Fig. 7L). Altogether, these results demonstrated the implication of SIRT1 in the XBP1s-dependent chemosensitization to aracytine.
Discussion
In tumors, cancer cells are prone to many stresses which can disturb protein synthesis and folding accuracy and overwhelm protein quality control processes thus leading to ER stress and activation of the PERK, ATF6 and IRE1-XBP1s UPR signaling pathways. UPR induction acts initially as a cell protective process aimed to restore protein homeostasis. However, if the damage is excessive, UPR moves from protective-to-cytotoxic in order to eliminate dysfunctional cells [38]. Chemotherapeutic drugs, as strong inducers of stress in cancer cells, could exert their cytotoxic effect at least in part through boosting the transition from protective-to-cytotoxic UPR. Interestingly our recent bibliographic survey on the role played by the UPR in leukemia and the response to treatment with candidate therapeutic molecules revealed that the observed increases in cell death were extremely frequently associated with cytotoxic UPR induction [18], but mechanisms of action have not been explored so far in the vast majority of these studies.
In a large number of cancers, including hematological malignancies [22, 23, 39, 40], activation of IRE1-XBP1s branch was found to initially play an essential safeguard role against apoptosis during cellular stresses. In addition, some previously published functional studies of XBP1s in leukemic cells clearly pointed out a cell protective function [41, 42]. The increased expression of XBP1s, observed in certain cancers is therefore widely considered to primarily act by restoring cell homeostasis in conditions of moderate cellular stress, thus contributing both to promote precancerous development and to increase the resistance of cancer cells to the various stresses they undergo in vivo during tumor development. Blocking the protective IRE1-XBP1s signaling pathway appears therefore as an attractive anti-cancer therapy and the search for pharmacological inhibitors of IRE1 is under active development [43]. However, strong IRE1 activation can also contribute to apoptosis induction [42, 44, 45] by different molecular mechanisms such as hyperactivation of endonuclease activity boosting the RIDD process [42, 44] or activation of the apoptotic JNK signaling pathway [45, 46]. During IRE1-induced pro-apoptotic response, XBP1s expression was found, when tested [41, 42], to retain a cell protective effect which appeared outweighed by the other actions taken by IRE1. However, in lung cancer cells, XBP1s could also switch from a negative to a positive mediator of cytotoxic UPR once cell stress surpasses a specific threshold [47]. This effect was associated with a modification of XBP1’s transcriptional target selectivity allowing the activation of pro-apoptotic genes such as the transcription factor KLF9 [47]. The implication of XBP1s in the induction of pro-apoptotic programs leading either to slowed tumor progression in vivo [19] or to increased sensitivity to Bruton’s kinase tyrosine kinase inhibitor ibrutinib has also been reported in the case of diffuse large B cell lymphomas [48]. Therefore, XBP1s may as well directly contribute to apoptosis induction in certain conditions.
Interestingly, in AML, XBP1 activation has been previously correlated with a favorable outcome upon chemotherapy [21] but the underlying molecular mechanisms remained unknown.
In this study, we aimed to clarify the functions played by XBP1s in leukemic cells. We showed that sustained XBP1s expression induces apoptosis whereas a moderate XBP1s expression, compatible with cell viability, enhances response to aracytine treatment in AML-resistant cells. This last effect is achieved by increasing XBP1s expression at RNA level by about tenfold. Interestingly, this increase is of the same order of magnitude as that obtained on endogenous XBP1 in isolated patient cells treated with tunicamycin, for which activation of XBP1s expression ranges from 5- to 25-fold increase (Supplementary Fig. 10). In addition, highly deregulated XBP1s expression, able on its own to induce cytotoxic responses, could presumably also be achieved during treatment with anti-leukemic drugs, most of them having been reported to induce strong UPR response in leukemic cells [18].
We characterized direct XBP1s target genes using ChIP-sequencing, RNA sequencing and microRNA sequencing analyses. From the list of identified targets, we decided to focus our analysis mainly on XBP1s noncoding targets, still poorly characterized so far. We identified 13 annotated long noncoding RNAs as specific XBP1 target genes, including MIR22HG (C17orf91), the precursor transcript of the microRNA miR-22-3p (Supplementary Fig. 6). Interestingly, previous studies demonstrated that MIR22HG is expressed in response to hypoxic or chemical stress, two stimuli known to induce ER stress [49, 50]. Here, we showed that induction of MIR22HG expression upon ER stress is XBP1-dependent.
Mir-22 function and regulation appears very complex [51] and, in leukemia, it has been reported to behave either as oncogenic or anti-tumoral depending on the model system [24, 52, 53].
Here we found that in OCI-AML3 cells miR-22 interacts efficiently with the mRNA of the deacetylase SIRT1, a member of the mammalian sirtuin family, and inhibits SIRT1 expression at the translational level, as previously described [34]. SIRT1 was previously reported to be upregulated in AML and identified as an oncogene in this pathology [35, 36]. Consistently, SIRT1 repression in OCI-AML3 and THP1 cells using siRNAs triggered apoptosis (Fig. 7H–K). This clearly demonstrated that this protein acts as a pro-survival factor in AML cells.
SIRT1 is involved in many cellular processes including DNA repair [54] and protects against radiation-induced apoptosis in different diseases such as brain, lung and breast cancers [26, 55, 56]. Moreover, in the latter model, miR-22 overexpression inhibited SIRT1 expression and improved cells response to radiotherapy [26]. In AML, the SIRT1 pharmacological inhibitor Tenovin-6 (Tv-6) was previously shown to potentiate aracytine treatment [35]. Moreover, Tv-6 has also been reported to inhibit SIRT2 and activate p53. Our experiments performed with EX-527, a highly specific inhibitor of SIRT1 [57], confirmed the protective role of SIRT1 against aracytine treatment in AML cells. Conversely, we showed that inhibition of endogenous miR-22 in XBP1s-expressing cells restores SIRT1 expression (Fig. 7G) and partially reverses aracytine-induced apoptosis (Fig. 6E).
Our studies revealed that overexpression of XBP1s contributes to lower the expression level of SIRT1 at least in part through miR-22-3p upregulation and that this process may affect both cell viability and response to cytotoxic drugs. Interestingly, previous studies have shown that SIRT1 may play as well an inhibitory role on the UPR response. Indeed, the activation of SIRT1 in the liver of mice leads to reduced activation of both PERK and IRE1-XBP1s signaling [58]. Moreover, concomitant studies showed that SIRT1 can act as a negative regulator of XBP1s by repressing its transcriptional activity through direct deacetylation [59]. Therefore, a negative feedback loop between SIRT1 and XBP1 may take place in cells to balance their relative activities and finely modulate cell viability.
To conclude, the overall presented data suggest that IRE1/XBP1s signaling may in some conditions act as a tumor suppressor pathway in AML, in particular through activation of the XBP1s/miR-22/SIRT1 axis. Activating this branch of the UPR could therefore represent an attractive additional therapeutic strategy for AML treatment.
Data availability
The datasets generated during this study are available upon reasonable request to the corresponding author.
References
Moon HW, Han HG, Jeon YJ. Protein quality control in the endoplasmic reticulum and cancer. Int J Mol Sci. 2018;19:3020.
Hetz C. The unfolded protein response: controlling cell fate decisions under ER stress and beyond. Nat Rev Mol Cell Biol. 2012;13:89–102.
Hetz C, Papa FR. The unfolded protein response and cell fate control. Mol cell. 2018;69:169–81.
Hetz C, Zhang K, Kaufman RJ. Mechanisms, regulation and functions of the unfolded protein response. Nat Rev Mol Cell Biol. 2020;21:421–38.
Ron D, Walter P. Signal integration in the endoplasmic reticulum unfolded protein response. Nat Rev Mol Cell Biol. 2007;8:519–29.
Hetz C, Chevet E. Theme Series-UPR in cancer. Semin Cancer Biol. 2015;33:1–2.
Kaufman RJ. Stress signaling from the lumen of the endoplasmic reticulum: coordination of gene transcriptional and translational controls. Genes Dev. 1999;13:1211–33.
Lin JH, Li H, Yasumura D, Cohen HR, Zhang C, Panning B, et al. IRE1 signaling affects cell fate during the unfolded protein response. Science. 2007;318:944–9.
Shamu CE, Walter P. Oligomerization and phosphorylation of the Ire1p kinase during intracellular signaling from the endoplasmic reticulum to the nucleus. EMBO J. 1996;15:3028–39.
Maurel M, Chevet E, Tavernier J, Gerlo S. Getting RIDD of RNA: IRE1 in cell fate regulation. Trends Biochem Sci. 2014;39:245–54.
Hollien J, Weissman JS. Decay of endoplasmic reticulum-localized mRNAs during the unfolded protein response. Science. 2006;313:104–7.
Glimcher LH. XBP1: the last two decades. Ann Rheum Dis. 2010;69:i67–71.
Lu Y, Liang FX, Wang X. A synthetic biology approach identifies the mammalian UPR RNA ligase RtcB. Mol Cell. 2014;55:758–70.
Peschek J, Acosta-Alvear D, Mendez AS, Walter P. A conformational RNA zipper promotes intron ejection during non-conventional XBP1 mRNA splicing. EMBO Rep. 2015;16:1688–98.
Yamamoto K, Sato T, Matsui T, Sato M, Okada T, Yoshida H, et al. Transcriptional induction of mammalian ER quality control proteins is mediated by single or combined action of ATF6alpha and XBP1. Dev Cell. 2007;13:365–76.
Yamamoto K, Yoshida H, Kokame K, Kaufman RJ, Mori K. Differential contributions of ATF6 and XBP1 to the activation of endoplasmic reticulum stress-responsive cis-acting elements ERSE, UPRE and ERSE-II. J Biochem. 2004;136:343–50.
Acosta-Alvear D, Karagoz GE, Frohlich F, Li H, Walther TC, Walter P. The unfolded protein response and endoplasmic reticulum protein targeting machineries converge on the stress sensor IRE1. Elife. 2018;7:e43036.
Feral K, Jaud M, Philippe C, Di Bella D, Pyronnet S, Rouault-Pierre K, et al. ER stress and unfolded protein response in leukemia: friend, foe, or both? Biomolecules. 2021;11:199.
Bujisic B, De Gassart A. Impairment of both IRE1 expression and XBP1 activation is a hallmark of GCB DLBCL and contributes to tumor growth. Blood. 2017;129:2420–8.
Schardt JA, Mueller BU, Pabst T. Activation of the unfolded protein response in human acute myeloid leukemia. Methods Enzymol. 2011;489:227–43.
Schardt JA, Weber D, Eyholzer M, Mueller BU, Pabst T. Activation of the unfolded protein response is associated with favorable prognosis in acute myeloid leukemia. Clin Cancer Res Off J Am Assoc Cancer Res. 2009;15:3834–41.
Sun H, Lin DC, Guo X, Kharabi Masouleh B, Gery S, Cao Q, et al. Inhibition of IRE1α-driven pro-survival pathways is a promising therapeutic application in acute myeloid leukemia. Oncotarget. 2016;7:18736–49.
Liu L, Zhao M, Jin X, Ney G, Yang KB, Peng F, et al. Adaptive endoplasmic reticulum stress signalling via IRE1α-XBP1 preserves self-renewal of haematopoietic and pre-leukaemic stem cells. Nat Cell Biol. 2019;21:328–37.
Jiang X, Hu C, Arnovitz S, Bugno J, Yu M, Zuo Z, et al. miR-22 has a potent anti-tumour role with therapeutic potential in acute myeloid leukaemia. Nat Commun. 2016;7:11452.
Lee JH, Park SJ, Jeong SY, Kim MJ, Jun S, Lee HS, et al. MicroRNA-22 suppresses DNA repair and promotes genomic instability through targeting of MDC1. Cancer Res. 2015;75:1298–310.
Zhang X, Li Y, Wang D, Wei X. miR-22 suppresses tumorigenesis and improves radiosensitivity of breast cancer cells by targeting Sirt1. Biol Res. 2017;50:27.
Touriol C, Roussigne M, Gensac MC, Prats H, Prats AC. Alternative translation initiation of human fibroblast growth factor 2 mRNA controlled by its 3′-untranslated region involves a Poly(A) switch and a translational enhancer. J Biol Chem. 2000;275:19361–7.
Philippe C, Dubrac A, Quelen C, Desquesnes A, Van Den Berghe L, Ségura C, et al. PERK mediates the IRES-dependent translational activation of mRNAs encoding angiogenic growth factors after ischemic stress. Sci Signal. 2016;9:ra44.
Lee AH, Iwakoshi NN, Glimcher LH. XBP-1 regulates a subset of endoplasmic reticulum resident chaperone genes in the unfolded protein response. Mol Cell Biol. 2003;23:7448–59.
Shen T, Li Y, Liang S, Chen Z. XBP1 negatively regulates CENPF expression via recruiting ATF6α to the promoter during ER stress. Cancer Cell Int. 2020;20:459.
Acosta-Alvear D, Zhou Y, Blais A, Tsikitis M, Lents NH, Arias C, et al. XBP1 controls diverse cell type- and condition-specific transcriptional regulatory networks. Mol Cell. 2007;27:53–66.
Pramanik J, Chen X, Kar G, Henriksson J, Gomes T, Park JE, et al. Genome-wide analyses reveal the IRE1a-XBP1 pathway promotes T helper cell differentiation by resolving secretory stress and accelerating proliferation. Genome Med. 2018;10:76.
Huang ZP, Chen J, Seok HY, Zhang Z, Kataoka M, Hu X, et al. MicroRNA-22 regulates cardiac hypertrophy and remodeling in response to stress. Circ Res. 2013;112:1234–43.
Xu D, Takeshita F, Hino Y, Fukunaga S, Kudo Y, Tamaki A, et al. miR-22 represses cancer progression by inducing cellular senescence. J Cell Biol. 2011;193:409–24.
Sasca D, Hähnel PS, Szybinski J, Khawaja K, Kriege O, Pante SV, et al. SIRT1 prevents genotoxic stress-induced p53 activation in acute myeloid leukemia. Blood. 2014;124:121–33.
Li L, Osdal T, Ho Y, Chun S, McDonald T, Agarwal P, et al. SIRT1 activation by a c-MYC oncogenic network promotes the maintenance and drug resistance of human FLT3-ITD acute myeloid leukemia stem cells. Cell Stem Cell. 2014;15:431–46.
Li L, Bhatia R. Role of SIRT1 in the growth and regulation of normal hematopoietic and leukemia stem cells. Curr Opin Hematol. 2015;22:324–9.
Rufo N, Yang Y, De Vleeschouwer S, Agostinis P. The “Yin and Yang” of unfolded protein response in cancer and immunogenic cell death. Cells. 2022;11:2899.
Kharabi Masouleh B, Geng H, Hurtz C, Chan LN, Logan AC, Chang MS, et al. Mechanistic rationale for targeting the unfolded protein response in pre-B acute lymphoblastic leukemia. Proc Natl Acad Sci USA. 2014;111:E2219–2228.
Xie H, Tang CH, Song JH, Mancuso A, Del Valle JR, Cao J, et al. IRE1α RNase-dependent lipid homeostasis promotes survival in Myc-transformed cancers. J Clin Investig. 2018;128:1300–16.
Gebert M, Sobolewska A, Bartoszewska S, Cabaj A, Crossman DK, Kroliczewski J, et al. Genome-wide mRNA profiling identifies X-box-binding protein 1 (XBP1) as an IRE1 and PUMA repressor. Cell Mol Life Sci. 2021;78:7061–80.
Ghosh R, Wang L, Wang ES, Perera BG, Igbaria A, Morita S, et al. Allosteric inhibition of the IRE1α RNase preserves cell viability and function during endoplasmic reticulum stress. Cell. 2014;158:534–48.
Bartoszewska S, Slawski J, Collawn JF, Bartoszewski R. Dual RNase activity of IRE1 as a target for anticancer therapies. J Cell Commun Signal. 2023;17:1145–61.
Liu X, Wu B, Chen H, Sun H, Guo X, Sun T, et al. Intense endoplasmic reticulum stress (ERS) / IRE1α enhanced Oxaliplatin efficacy by decreased ABCC10 in colorectal cancer cells. BMC Cancer. 2022;22:1369.
Zeng T, Peng L, Chao H, Xi H, Fu B, Wang Y, et al. IRE1α-TRAF2-ASK1 complex-mediated endoplasmic reticulum stress and mitochondrial dysfunction contribute to CXC195-induced apoptosis in human bladder carcinoma T24 cells. Biochem Biophys Res Commun. 2015;460:530–6.
Urano F, Wang X, Bertolotti A, Zhang Y, Chung P, Harding HP, et al. Coupling of stress in the ER to activation of JNK protein kinases by transmembrane protein kinase IRE1. Science. 2000;287:664–6.
Fink EE, Moparthy S, Bagati A, Bianchi-Smiraglia A, Lipchick BC, Wolff DW, et al. XBP1-KLF9 axis acts as a molecular rheostat to control the transition from adaptive to cytotoxic unfolded protein response. Cell Rep. 2018;25:212–223 e214.
Zhang XT, Hu XB, Wang HL, Kan WJ, Xu L, Wang ZJ, et al. Activation of unfolded protein response overcomes Ibrutinib resistance in diffuse large B-cell lymphoma. Acta Pharmacol Sin. 2021;42:814–23.
Voellenkle C, Garcia-Manteiga JM, Pedrotti S, Perfetti A, De Toma I, Da Silva D, et al. Implication of Long noncoding RNAs in the endothelial cell response to hypoxia revealed by RNA-sequencing. Sci Rep. 2016;6:24141.
Tani H, Onuma Y, Ito Y, Torimura M. Long non-coding RNAs as surrogate indicators for chemical stress responses in human-induced pluripotent stem cells. PLoS ONE. 2014;9:e106282.
Wang J, Li Y, Ding M, Zhang H, Xu X, Tang J. Molecular mechanisms and clinical applications of miR-22 in regulating malignant progression in human cancer (review). Int J Oncol. 2017;50:345–55.
Song SJ, Ito K, Ala U, Kats L, Webster K, Sun SM, et al. The oncogenic microRNA miR-22 targets the TET2 tumor suppressor to promote hematopoietic stem cell self-renewal and transformation. Cell Stem Cell. 2013;13:87–101.
Song SJ, Pandolfi PP. miR-22 in tumorigenesis. Cell Cycle. 2014;13:11–12.
Chalkiadaki A, Guarente L. The multifaceted functions of sirtuins in cancer. Nat Rev Cancer. 2015;15:608–24.
Chang CJ, Hsu CC, Yung MC, Chen KY, Tzao C, Wu WF, et al. Enhanced radiosensitivity and radiation-induced apoptosis in glioma CD133-positive cells by knockdown of SirT1 expression. Biochem Biophys Res Commun. 2009;380:236–42.
Sun Y, Sun D, Li F, Tian L, Li C, Li L, et al. Downregulation of Sirt1 by antisense oligonucleotides induces apoptosis and enhances radiation sensitization in A549 lung cancer cells. Lung Cancer. 2007;58:21–29.
Solomon JM, Pasupuleti R, Xu L, McDonagh T, Curtis R, DiStefano PS, et al. Inhibition of SIRT1 catalytic activity increases p53 acetylation but does not alter cell survival following DNA damage. Mol Cell Biol. 2006;26:28–38.
Li Y, Xu S, Giles A, Nakamura K, Lee JW, Hou X, et al. Hepatic overexpression of SIRT1 in mice attenuates endoplasmic reticulum stress and insulin resistance in the liver. FASEB J Off Publ Fed Am Soc Exp Biol. 2011;25:1664–79.
Wang FM, Chen YJ, Ouyang HJ. Regulation of unfolded protein response modulator XBP1s by acetylation and deacetylation. Biochem J. 2011;433:245–52.
Acknowledgements
This work was supported by the Institut National de la Santé et de la Recherche Médicale (INSERM), Université Toulouse III (Paul Sabatier), by research grants from Association Laurette Fugain (ALF2018/03 awarded to CT), la Ligue Contre le Cancer, Fondation ARC pour la recherche sur le cancer, and by a generous donation of the “Club de la concorde de Genebrières”. CP and MJ were supported by a fellowship from the French Ministry of Higher Education and Research and MJ by a fellowship from la Ligue Contre le Cancer. KRP was supported by funding from the Barts Charity (G-002167), the Kay Kendall Leukemia Fund (KKL1149), and the Academy of Medical Sciences (SBF004\1099). We are thankful to Sandra Dailhau and Loelia Babin for their help in sequencing data analyses. We thank the Anexplo-Génotoul platform (UMS US006/CREFRE Toulouse, France) for animal facilities.
Author information
Authors and Affiliations
Contributions
CP, KRP, LM and CT wrote the manuscript and prepared the manuscript with contributions from all co-authors. CP, MJ, KRP, LM and CT designed the experiments and CP, MJ, KF, AG, LVDB, MF, LM and CT carried them out. AG, CP, MJ, LM and CT contributed to the sequencing study’s design and the data analysis. CP, MJ, LM, SP, MB, KRP and CT analyzed and interpreted the data. MB and SP commented on and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Philippe, C., Jaud, M., Féral, K. et al. Pivotal role of the endoplasmic reticulum stress-related XBP1s/miR-22/SIRT1 axis in acute myeloid leukemia apoptosis and response to chemotherapy. Leukemia 38, 1764–1776 (2024). https://doi.org/10.1038/s41375-024-02321-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-024-02321-8
- Springer Nature Limited